The Transcription Factor Sfp1 Regulates the Oxidative Stress Response in Candida albicans
Abstract
1. Introduction
2. Materials and Methods
2.1. Strains and Growth Conditions
2.2. DNA Microarray Analysis
2.3. Reverse Transcription (RT) Real-Time Quantitative PCR (qPCR)
2.4. Extraction and Quantification of Total Protein
2.5. Determination of Total Antioxidant Capacity
2.6. Cell Susceptibility to Oxidants and Measurement of Intracellular ROS
2.7. Examination of Cell Morphology by Scanning Electron Microscopy (SEM)
2.8. Measurement of Sod and Catalase Enzyme Activity
2.9. Measurement of Total Glutathione, Oxidized-Glutathione (GSSG), and Reduced-Glutathione (GSH) Content
2.10. Measurement of Glutathione Peroxidase Enzyme Activity
2.11. Western Blotting
2.12. Macrophage Killing and Phagocytosis Assay
2.13. Statistical Analysis
3. Results
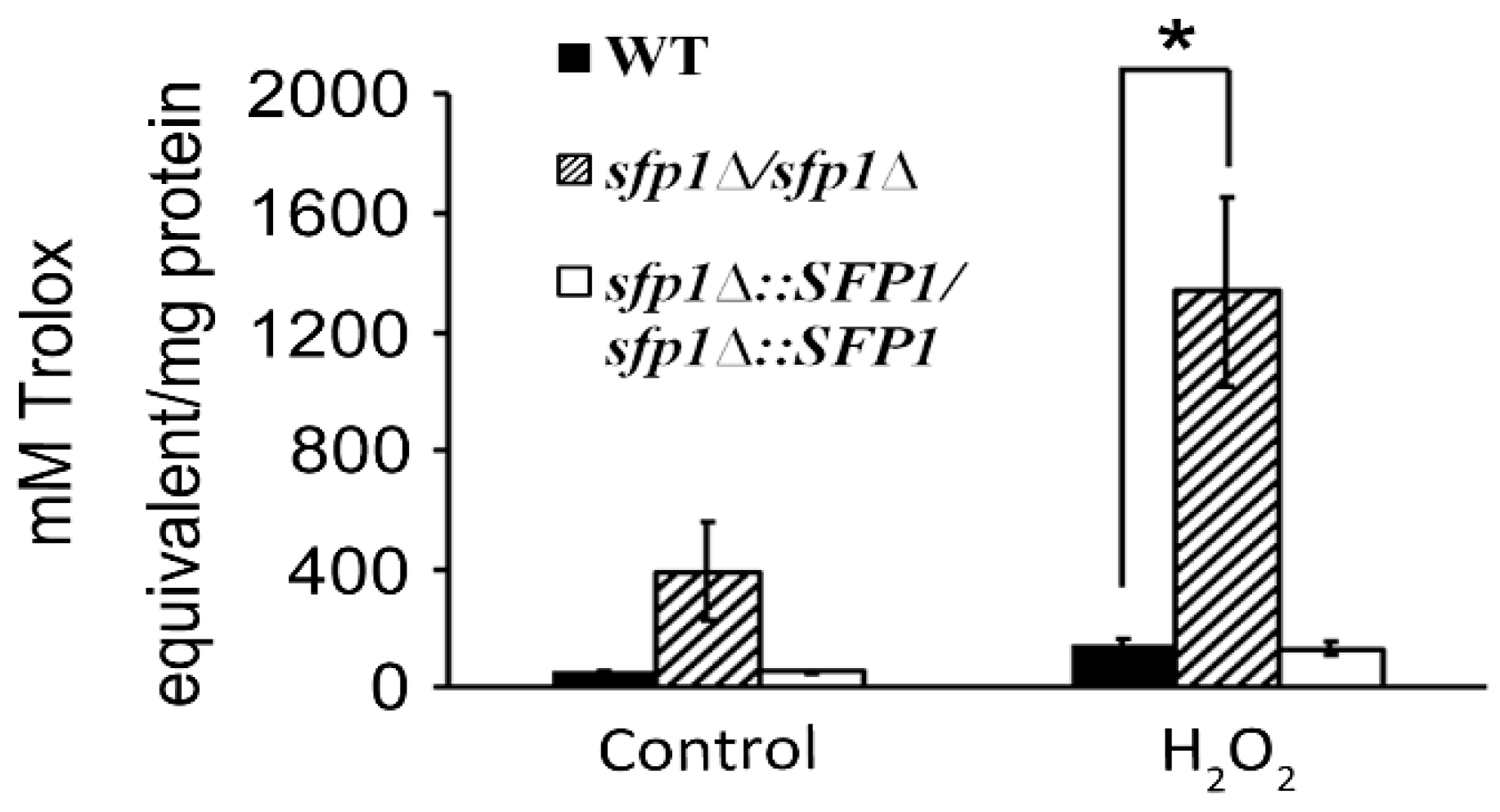
3.1. Sfp1 is Involved in the C. albicans Response to Oxidative Stress
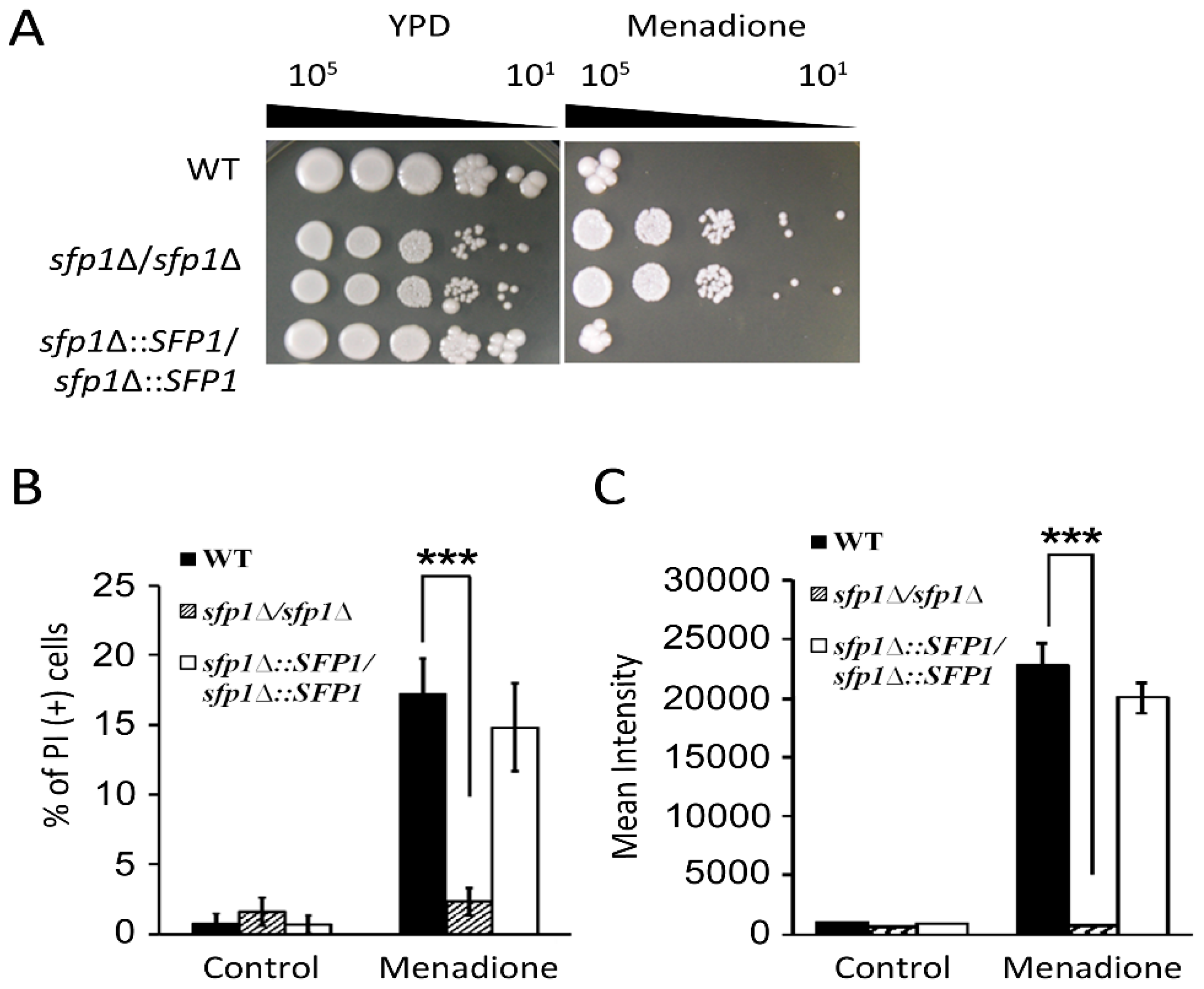
3.2. Sfp1 is Related to Cellular Susceptibility to Menadione/Superoxide
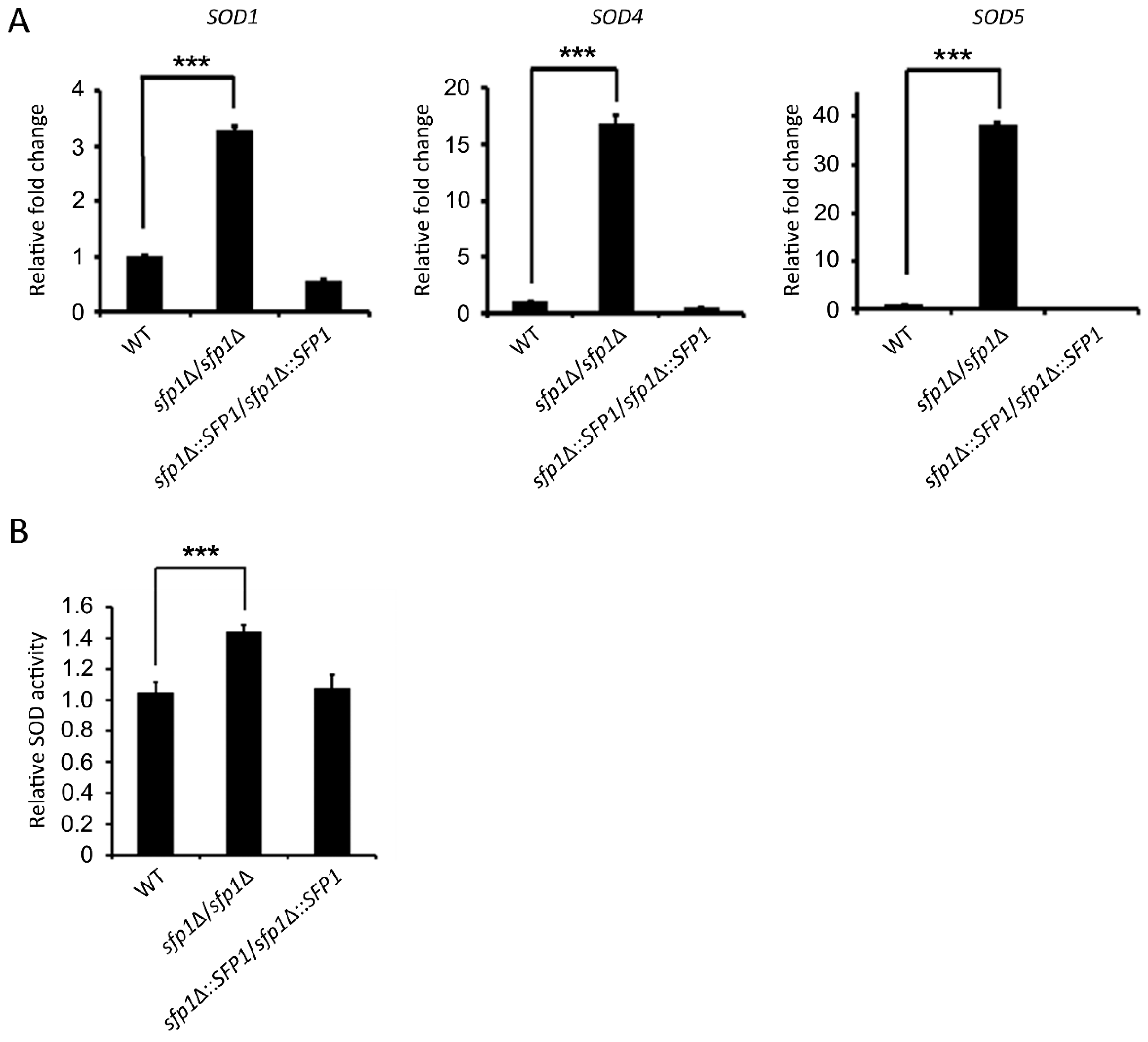
3.3. Sfp1 Affects SOD Gene Expression and Enzyme Activity
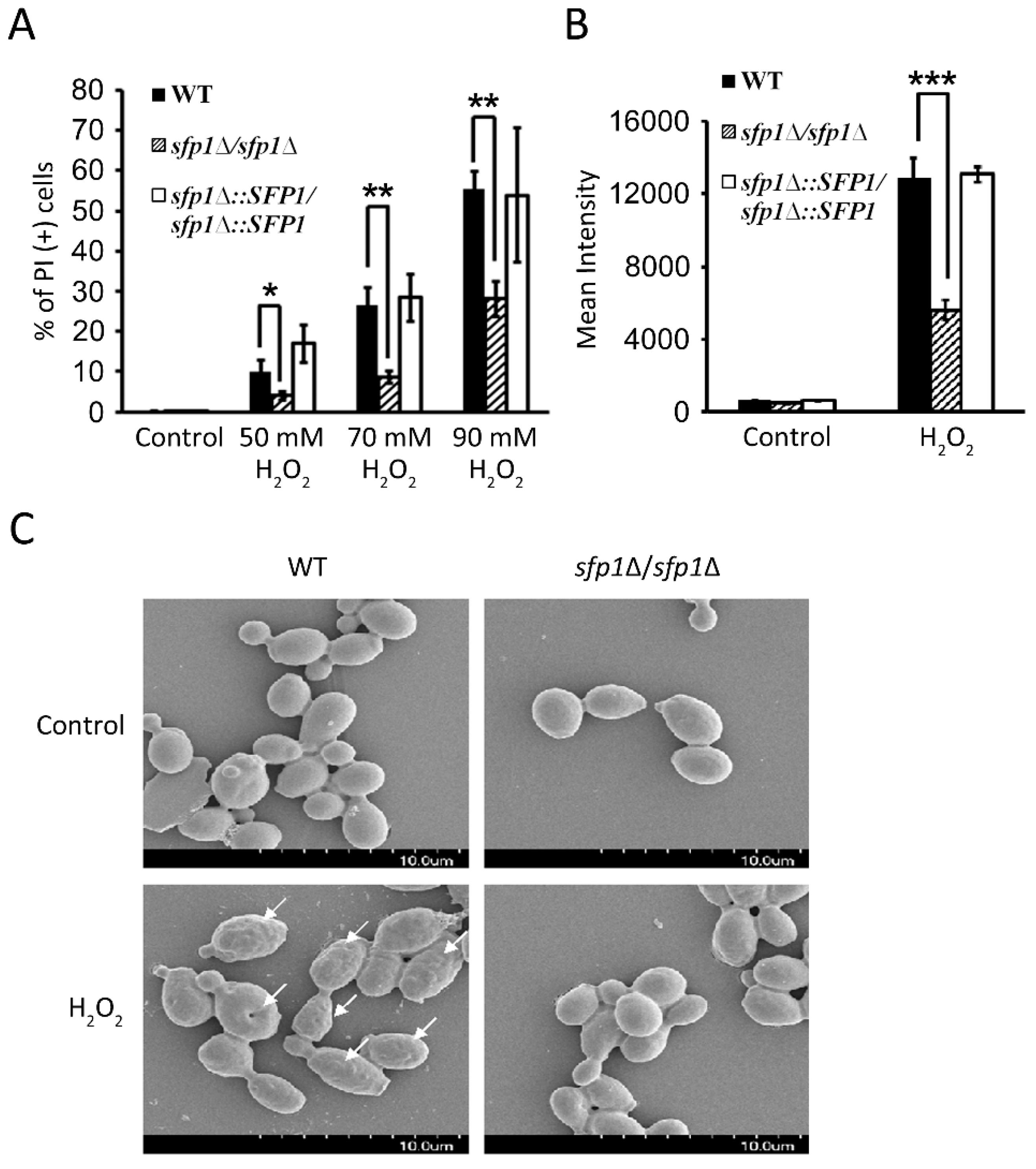
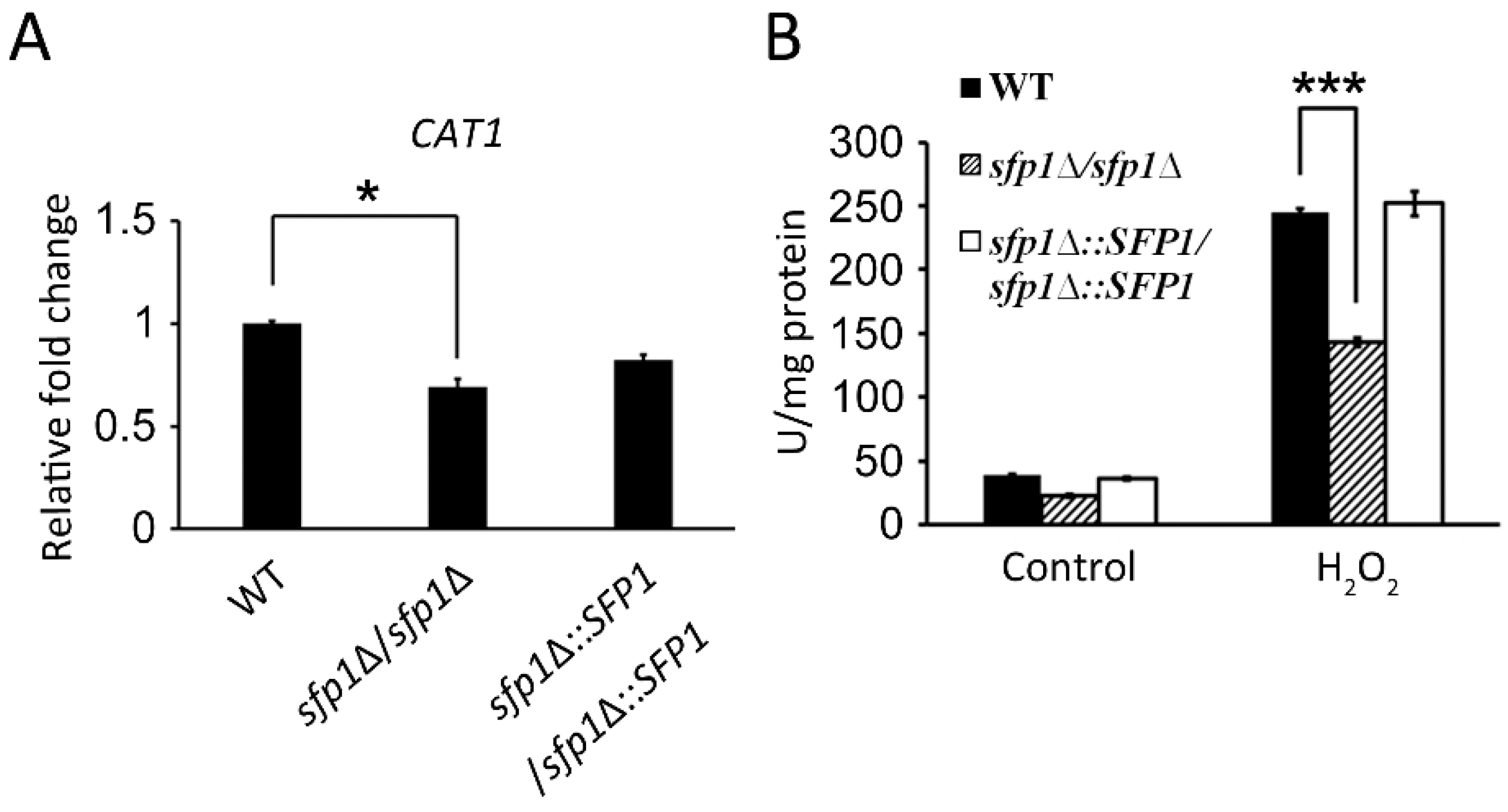
3.4. Sfp1 Is Also Related to Cellular Susceptibility to Hydrogen Peroxide
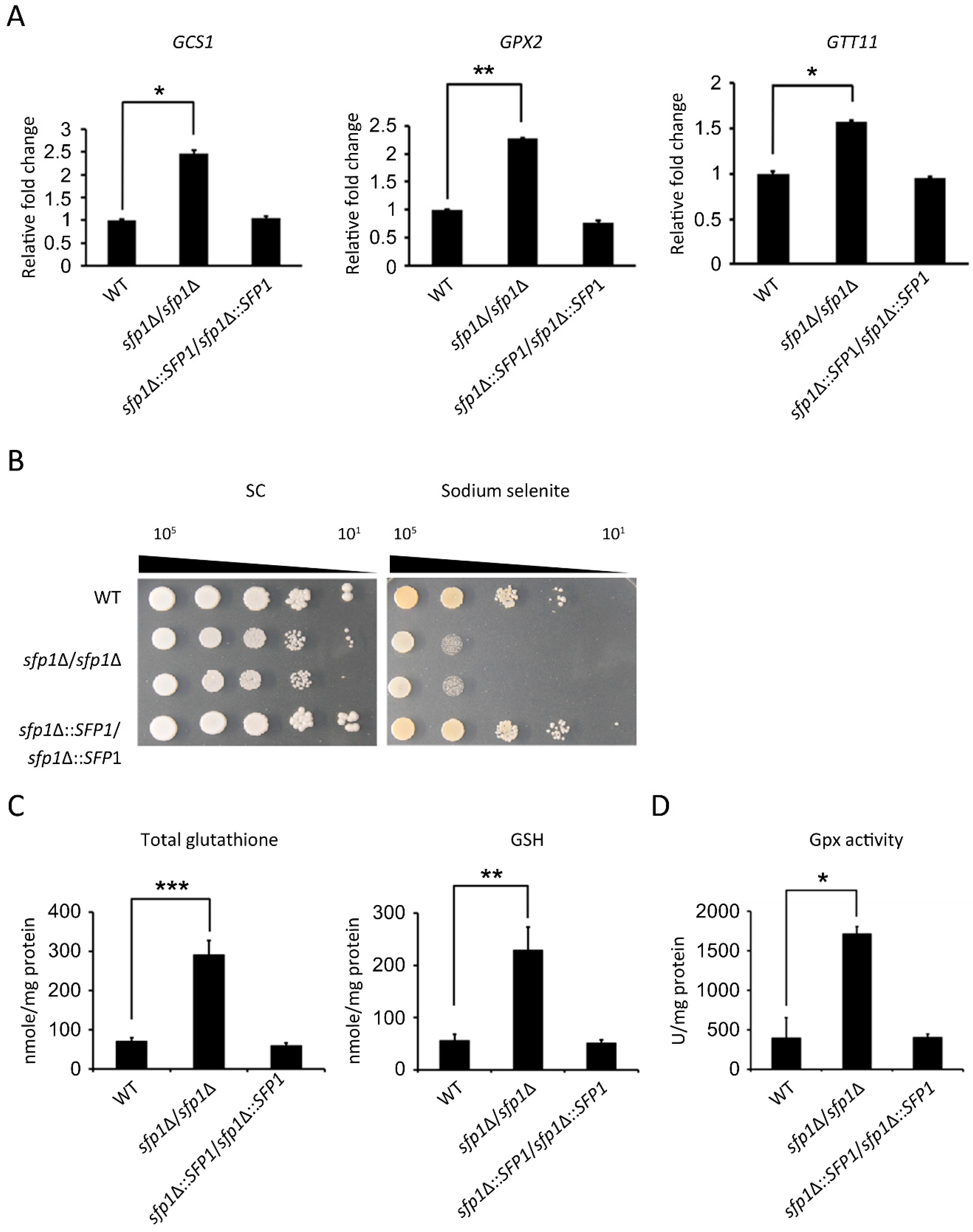
3.5. Sfp1 Regulates the Glutathione System to Detoxify Hydrogen Peroxide
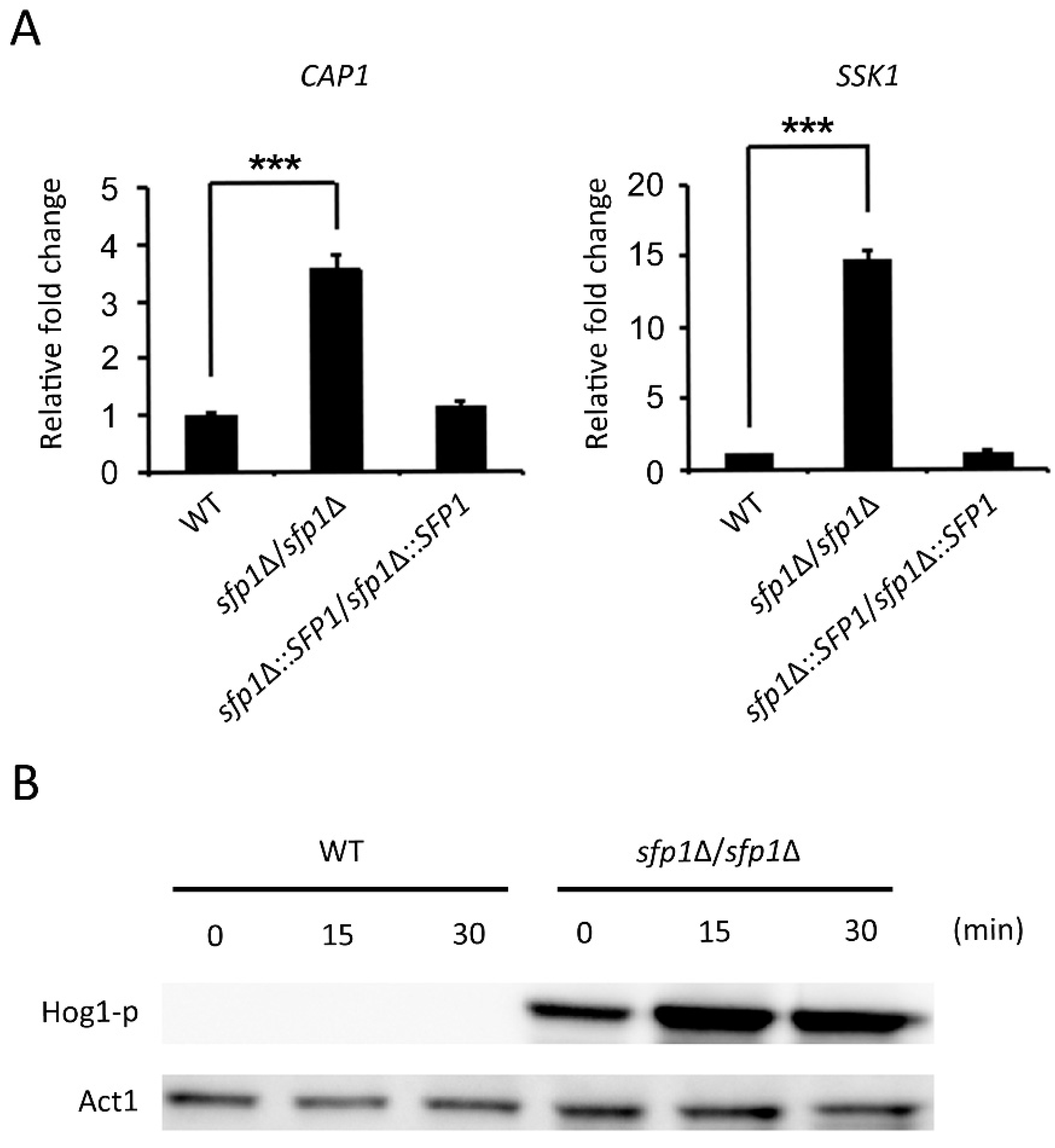
3.6. The Hog1 Signaling Pathway and the Transcription Factor Cap1 Are Related to the Sfp1-Mediated Oxidative Stress Response
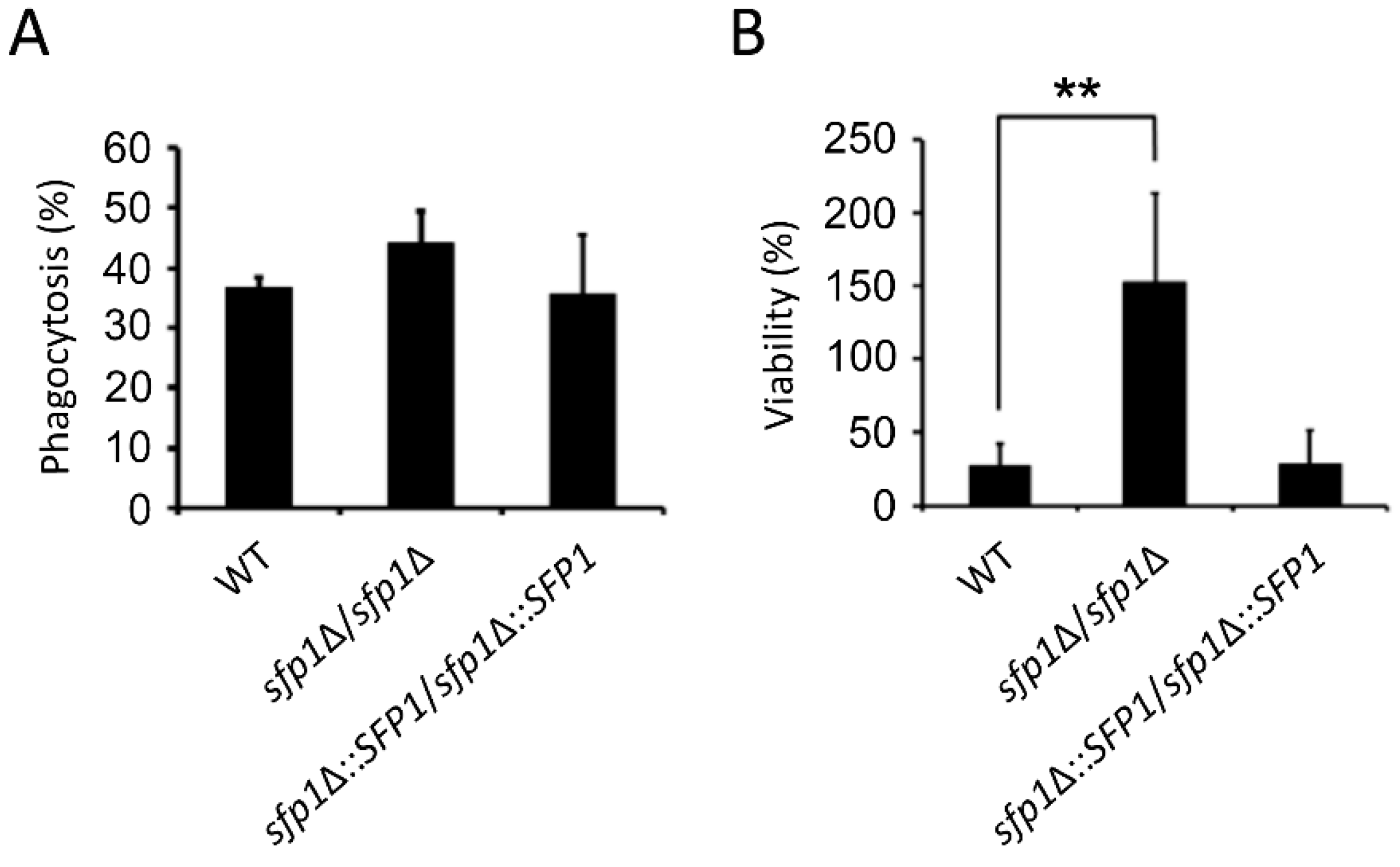
3.7. The sfp1Δ/sfp1Δ Mutant is Resistant to Macrophage Killing
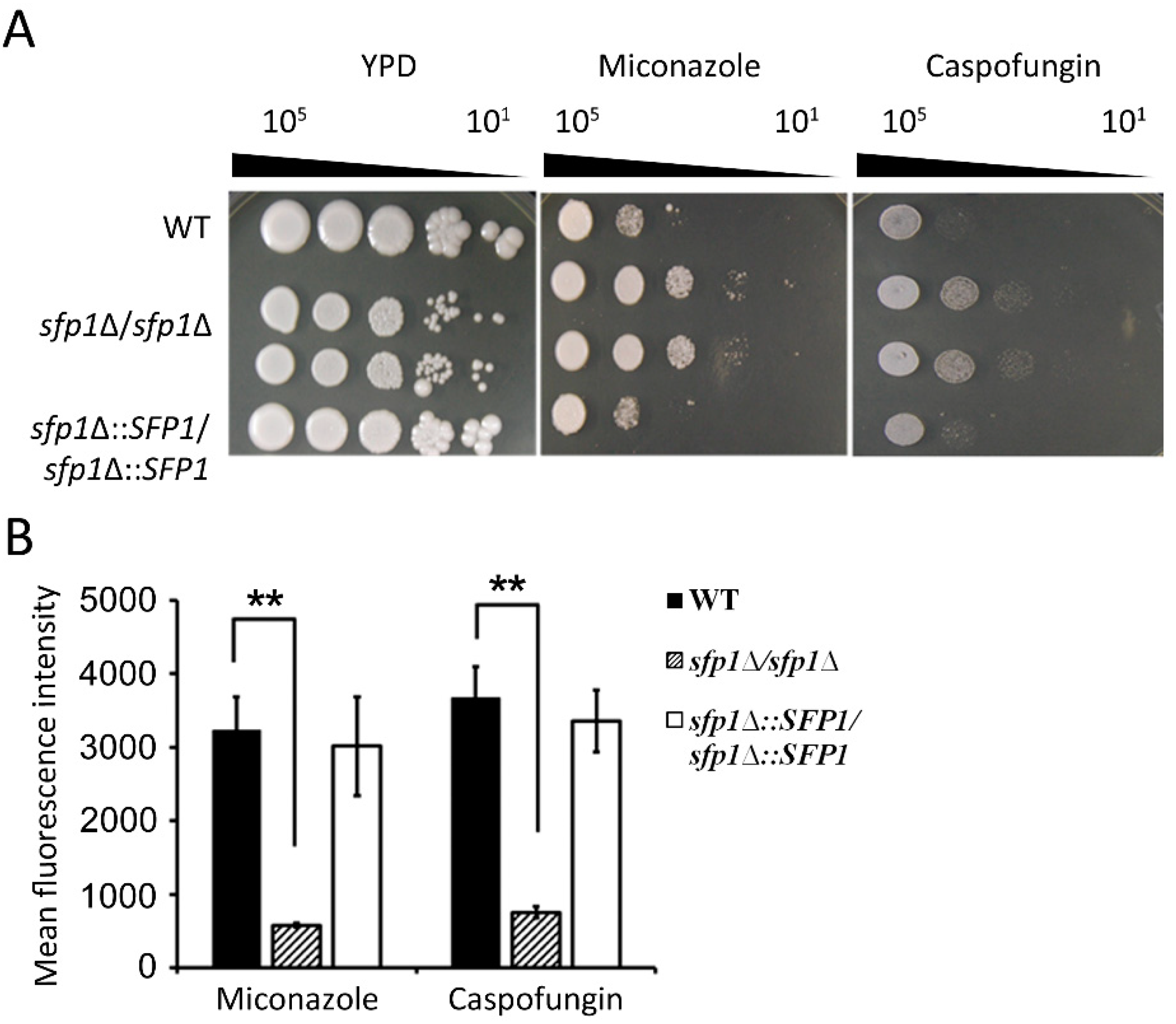
3.8. The sfp1Δ/sfp1Δ Mutant is Resistant to ROS-Generating Antifungals
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Odd, F. Candida and Candidiasis: A Review and Bibliography; Bailliere Tindal: London, UK, 1988. [Google Scholar]
- Arendrup, M.C.; Patterson, T.F. Multidrug-resistant Candida: Epidemiology, molecular mechanisms, and treatment. J. Infect. Dis. 2017, 216, S445–S451. [Google Scholar] [CrossRef]
- Dantas Ada, S.; Day, A.; Ikeh, M.; Kos, I.; Achan, B.; Quinn, J. Oxidative stress responses in the human fungal pathogen, Candida albicans. Biomolecules 2015, 5, 142–165. [Google Scholar] [CrossRef] [PubMed]
- Delattin, N.; Cammue, B.P.; Thevissen, K. Reactive oxygen species-inducing antifungal agents and their activity against fungal biofilms. Future Med. Chem. 2014, 6, 77–90. [Google Scholar] [CrossRef]
- Kaloriti, D.; Tillmann, A.; Cook, E.; Jacobsen, M.; You, T.; Lenardon, M.; Ames, L.; Barahona, M.; Chandrasekaran, K.; Coghill, G.; et al. Combinatorial stresses kill pathogenic Candida species. Med. Mycol. 2012, 50, 699–709. [Google Scholar] [CrossRef] [PubMed]
- Mesa-Arango, A.C.; Trevijano-Contador, N.; Roman, E.; Sanchez-Fresneda, R.; Casas, C.; Herrero, E.; Arguelles, J.C.; Pla, J.; Cuenca-Estrella, M.; Zaragoza, O. The production of reactive oxygen species is a universal action mechanism of amphotericin B against pathogenic yeasts and contributes to the fungicidal effect of this drug. Antimicrob. Agents Chemother. 2014, 58, 6627–6638. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, D.; Kondo, K.; Uehara, N.; Otokozawa, S.; Tsuji, N.; Yagihashi, A.; Watanabe, N. Endogenous reactive oxygen species Is an important mediator of miconazole antifungal effect. Antimicrob. Agents Chemother. 2002, 46, 3113–3117. [Google Scholar] [CrossRef]
- Hao, B.; Cheng, S.; Clancy, C.J.; Nguyen, H.M. Caspofungin kills Candida albicans by causing both cellular apoptosis and necrosis. Antimicrob. Agents Chemother. 2012, 57, 326–332. [Google Scholar] [CrossRef]
- Bink, A.; Vandenbosch, D.; Coenye, T.; Nelis, H.; Cammue, B.P.A.; Thevissen, K. Superoxide dismutases are involved in Candida albicans biofilm persistence against miconazole. Antimicrob. Agents Chemother. 2011, 55, 4033–4037. [Google Scholar] [CrossRef]
- Brucker, K.; Bink, A.; Meert, E.; Cammue, B.P.A.; Thevissen, K. Potentiation of antibiofilm activity of amphotericin B by superoxide dismutase inhibition. Oxid. Med. Cell. Longev. 2013, 2013, 1–7. [Google Scholar] [CrossRef]
- Komalapriya, C.; Kaloriti, D.; Tillmann, A.T.; Yin, Z.; Herrero-de-Dios, C.; Jacobsen, M.D.; Belmonte, R.C.; Cameron, G.; Haynes, K.; Grebogi, C.; et al. Integrative model of oxidative stress adaptation in the fungal pathogen Candida albicans. PLoS ONE 2015, 10, e0137750. [Google Scholar] [CrossRef] [PubMed]
- Broxton, C.N.; Culotta, V.C. SOD enzymes and microbial pathogens: Surviving the oxidative storm of infection. PLoS Pathog. 2016, 12, e1005295. [Google Scholar] [CrossRef]
- Wysong, D.R.; Christin, L.; Sugar, A.M.; Robbins, P.W.; Diamond, R.D. Cloning and sequencing of a Candida albicans catalase gene and effects of disruption of this gene. Infect. Immun. 1998, 66, 1953–1961. [Google Scholar]
- Smith, D.A.; Nicholls, S.; Morgan, B.A.; Brown, A.J.; Quinn, J. A conserved stress-activated protein kinase regulates a core stress response in the human pathogen Candida albicans. Mol. Biol. Cell 2004, 15, 4179–4190. [Google Scholar] [CrossRef]
- Arana, D.M.; Alonso-Monge, R.; Du, C.; Calderone, R.; Pla, J. Differential susceptibility of mitogen-activated protein kinase pathway mutants to oxidative-mediated killing by phagocytes in the fungal pathogen Candida albicans. Cell. Microbiol. 2007, 9, 1647–1659. [Google Scholar] [CrossRef] [PubMed]
- Alonso-Monge, R.; Navarro-Garcia, F.; Molero, G.; Diez-Orejas, R.; Gustin, M.; Pla, J.; Sanchez, M.; Nombela, C. Role of the mitogen-activated protein kinase Hog1p in morphogenesis and virulence of Candida albicans. J. Bacteriol. 1999, 181, 3058–3068. [Google Scholar]
- Patterson, M.J.; McKenzie, C.G.; Smith, D.A.; da Silva Dantas, A.; Sherston, S.; Veal, E.A.; Morgan, B.A.; MacCallum, D.M.; Erwig, L.-P.P.; Quinn, J. Ybp1 and Gpx3 signaling in Candida albicans govern hydrogen peroxide-induced oxidation of the Cap1 transcription factor and macrophage escape. Antioxid. Redox Signal. 2013, 19, 2244–2260. [Google Scholar] [CrossRef]
- Hwang, C.-S.S.; Rhie, G.-e.E.; Oh, J.-H.H.; Huh, W.-K.K.; Yim, H.-S.S.; Kang, S.-O.O. Copper- and zinc-containing superoxide dismutase (Cu/ZnSOD) is required for the protection of Candida albicans against oxidative stresses and the expression of its full virulence. Microbiology 2002, 148, 3705–3713. [Google Scholar] [CrossRef]
- Frohner, I.E.; Bourgeois, C.; Yatsyk, K.; Majer, O.; Kuchler, K. Candida albicans cell surface superoxide dismutases degrade host-derived reactive oxygen species to escape innate immune surveillance. Mol. Microbiol. 2009, 71, 240–252. [Google Scholar] [CrossRef] [PubMed]
- Miramón, P.; Dunker, C.; Kasper, L.; Jacobsen, I.D.; Barz, D.; Kurzai, O.; Hube, B. A family of glutathione peroxidases contributes to oxidative stress resistance in Candida albicans. Med. Mycol. 2014, 52, 223–239. [Google Scholar] [CrossRef] [PubMed]
- Roman, E.; Prieto, D.; Martin, R.; Correia, I.; Mesa Arango, A.C.; Alonso-Monge, R.; Zaragoza, O.; Pla, J. Role of catalase overproduction in drug resistance and virulence in Candida albicans. Future Microbiol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.-F.; Lan, C.-Y. Role of SFP1 in the regulation of Candida albicans biofilm formation. PLoS ONE 2015, 10, e0129903. [Google Scholar] [CrossRef]
- Hsu, P.C.; Chao, C.C.; Yang, C.Y.; Ye, Y.L.; Liu, F.C.; Chuang, Y.J.; Lan, C.Y. Diverse Hap43-independent functions of the Candida albicans CCAAT-binding complex. Eukaryot. Cell 2013, 12, 804–815. [Google Scholar] [CrossRef] [PubMed]
- Nailis, H.; Coenye, T.; Van Nieuwerburgh, F.; Deforce, D.; Nelis, H.J. Development and evaluation of different normalization strategies for gene expression studies in Candida albicans biofilms by real-time PCR. BMC Mol. Biol. 2006, 7, 25. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Yu, Q.; Zhang, B.; Li, J.; Zhang, B.; Wang, H.; Li, M. Endoplasmic reticulum-derived reactive oxygen species (ROS) is involved in toxicity of cell wall stress to Candida albicans. Free Radic. Biol. Med. 2016, 99, 572–583. [Google Scholar] [CrossRef]
- Dai, B.-D.D.; Wang, Y.; Zhao, L.-X.X.; Li, D.-D.D.; Li, M.-B.B.; Cao, Y.-B.B.; Jiang, Y.-Y.Y. Cap1p attenuates the apoptosis of Candida albicans. FEBS J. 2013, 280, 2633–2643. [Google Scholar] [CrossRef]
- Beers, R.F.; Sizer, I.W. A spectrophotometric method for measuring the breakdown of hydrogen peroxide by catalase. J. Biol. Chem. 1952, 195, 133–140. [Google Scholar] [PubMed]
- Rahman, I.; Kode, A.; Biswas, S.K. Assay for quantitative determination of glutathione and glutathione disulfide levels using enzymatic recycling method. Nat. Protoc. 2006, 1, 3159–3165. [Google Scholar] [CrossRef]
- Maras, B.; Angiolella, L.; Mignogna, G.; Vavala, E.; Macone, A.; Colone, M.; Pitari, G.; Stringaro, A.; Dupré, S.; Palamara, A.T. Glutathione metabolism in Candida albicans resistant strains to fluconazole and micafungin. PLoS ONE 2014, 9, e98387. [Google Scholar] [CrossRef]
- Tsao, C.C.; Chen, Y.T.; Lan, C.Y. A small G protein Rhb1 and a GTPase-activating protein Tsc2 involved in nitrogen starvation-induced morphogenesis and cell wall integrity of Candida albicans. Fungal Genet. Biol. 2009, 46, 126–136. [Google Scholar] [CrossRef]
- Chauhan, N.; Inglis, D.; Roman, E.; Pla, J.; Li, D.; Calera, J.A.; Calderone, R. Candida albicans response regulator gene SSK1 regulates a subset of genes whose functions are associated with cell wall biosynthesis and adaptation to oxidative stress. Eukaryot. Cell 2003, 2, 1018–1024. [Google Scholar] [CrossRef]
- Wang, Y.; Cao, Y.Y.; Jia, X.M.; Cao, Y.B.; Gao, P.H.; Fu, X.P.; Ying, K.; Chen, W.S.; Jiang, Y.Y. Cap1p is involved in multiple pathways of oxidative stress response in Candida albicans. Free Radic. Biol. Med. 2006, 40, 1201–1209. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; De Micheli, M.; Coleman, S.T.; Sanglard, D.; Moye-Rowley, W.S. Analysis of the oxidative stress regulation of the Candida albicans transcription factor, Cap1p. Mol. Microbiol. 2000, 36, 618–629. [Google Scholar] [CrossRef] [PubMed]
- Missall, T.A.; Lodge, J.K.; McEwen, J.E. Mechanisms of resistance to oxidative and nitrosative stress: Implications for fungal survival in mammalian hosts. Eukaryot. Cell 2004, 3, 835–846. [Google Scholar] [CrossRef] [PubMed]
- Lin, G.-Y.; Chen, H.-F.; Xue, Y.-P.; Yeh, Y.-C.; Chen, C.-L.; Liu, M.-S.; Cheng, W.-C.; Lan, C.-Y. The Antimicrobial Peptides P-113Du and P-113Tri Function against Candida albicans. Antimicrob. Agents Chemother. 2016, 60, 6369–6373. [Google Scholar] [CrossRef]
- Ramírez-Quijas, M.D.; Zazueta-Sandoval, R. Effect of oxidative stress on cell wall morphology in four pathogenic Candida species. Mycol. Prog. 2015, 14, 8. [Google Scholar] [CrossRef]
- Nakagawa, Y.; Koide, K.; Watanabe, K.; Morita, Y.; Mizuguchi, I.; Akashi, T. The expression of the pathogenic yeast Candida albicans catalase gene in response to hydrogen peroxide. Microbiol. Immunol. 1999, 43, 645–651. [Google Scholar] [CrossRef] [PubMed]
- Quintana-Cabrera, R.; Bolanos, J.P. Glutathione and gamma-glutamylcysteine in hydrogen peroxide detoxification. Methods Enzymol. 2013, 527, 129–144. [Google Scholar] [CrossRef]
- Grant, C.M.; Perrone, G.; Dawes, I.W. Glutathione and catalase provide overlapping defenses for protection against hydrogen peroxide in the yeast Saccharomyces cerevisiae. Biochem. Biophys. Res. Commun. 1998, 253, 893–898. [Google Scholar] [CrossRef]
- Garcerá, A.; Casas, C.; Herrero, E. Expression of Candida albicans glutathione transferases is induced inside phagocytes and upon diverse environmental stresses. FEMS Yeast Res. 2010, 10, 422–431. [Google Scholar] [CrossRef][Green Version]
- Yadav, A.K.; Desai, P.R.; Rai, M.N.; Kaur, R.; Ganesan, K.; Bachhawat, A.K. Glutathione biosynthesis in the yeast pathogens Candida glabrata and Candida albicans: Essential in C. glabrata, and essential for virulence in C. albicans. Microbiology 2011, 157, 484–495. [Google Scholar] [CrossRef]
- Chen, J.J.; Boylan, L.M.; Wu, C.K.; Spallholz, J.E. Oxidation of glutathione and superoxide generation by inorganic and organic selenium compounds. Biofactors 2007, 31, 55–66. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez-Escobedo, G.; Orta-Zavalza, E.; Castano, I.; De Las Penas, A. Role of glutathione in the oxidative stress response in the fungal pathogen Candida glabrata. Curr. Genet. 2013, 59, 91–106. [Google Scholar] [CrossRef]
- Alonso-Monge, R.; Navarro-García, F.; Román, E.; Negredo, A.I.; Eisman, B.; Nombela, C.; Pla, J. The Hog1 mitogen-activated protein kinase is essential in the oxidative stress response and chlamydospore formation in Candida albicans. Eukaryot. Cell 2003, 2, 351–361. [Google Scholar] [CrossRef] [PubMed]
- Znaidi, S.; Barker, K.S.; Weber, S.; Alarco, A.M.; Liu, T.T.; Boucher, G.; Rogers, P.D.; Raymond, M. Identification of the Candida albicans Cap1p regulon. Eukaryot. Cell 2009, 8, 806–820. [Google Scholar] [CrossRef]
- Moye-Rowley, W.S. Regulation of the transcriptional response to oxidative stress in fungi: Similarities and differences. Eukaryot. Cell 2003, 2, 381–389. [Google Scholar] [CrossRef] [PubMed]
- Kos, I.; Patterson, M.J.; Znaidi, S.; Kaloriti, D.; da Silva Dantas, A.; Herrero-de-Dios, C.M.; d’Enfert, C.; Brown, A.J.; Quinn, J. Mechanisms underlying the delayed activation of the Cap1 transcription factor in Candida albicans following combinatorial oxidative and cationic stress important for phagocytic potency. mBio 2016, 7, e00331. [Google Scholar] [CrossRef]
- Jimenez-Lopez, C.; Lorenz, M.C. Fungal immune evasion in a model host-pathogen interaction: Candida albicans versus macrophages. PLoS Pathog. 2013, 9, e1003741. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.A.; Morgan, B.A.; Quinn, J. Stress signalling to fungal stress-activated protein kinase pathways. FEMS Microbiol. Lett. 2010, 306, 1–8. [Google Scholar] [CrossRef]
- Enjalbert, B.; Smith, D.A.; Cornell, M.J.; Alam, I.; Nicholls, S.; Brown, A.J.; Quinn, J. Role of the Hog1 stress-activated protein kinase in the global transcriptional response to stress in the fungal pathogen Candida albicans. Mol. Biol. Cell 2006, 17, 1018–1032. [Google Scholar] [CrossRef]
- Kastora, S.L.; Herrero-de-Dios, C.; Avelar, G.M.; Munro, C.A.; Brown, A.J.P. Sfp1 and Rtg3 reciprocally modulate carbon source-conditional stress adaptation in the pathogenic yeast Candida albicans. Mol. Microbiol. 2017, 105, 620–636. [Google Scholar] [CrossRef] [PubMed]
- Monge, R.A.; Roman, E.; Nombela, C.; Pla, J. The MAP kinase signal transduction network in Candida albicans. Microbiology 2006, 152, 905–912. [Google Scholar] [CrossRef]
- Menon, V.; De Bernardis, F.; Calderone, R.; Chauhan, N. Transcriptional profiling of the Candida albicans Ssk1p receiver domain point mutants and their virulence. FEMS Yeast Res. 2008, 8, 756–763. [Google Scholar] [CrossRef][Green Version]
- Calera, J.A.; Zhao, X.J.; Calderone, R. Defective hyphal development and avirulence caused by a deletion of the SSK1 response regulator gene in Candida albicans. Infect. Immun. 2000, 68, 518–525. [Google Scholar] [CrossRef] [PubMed]
- Du, C.; Calderone, R.; Richert, J.; Li, D. Deletion of the SSK1 response regulator gene in Candida albicans contributes to enhanced killing by human polymorphonuclear neutrophils. Infect. Immun. 2005, 73, 865–871. [Google Scholar] [CrossRef] [PubMed]
- Grant, C.M.; Maclver, F.H.; Dawes, I.W. Glutathione is an essential metabolite required for resistance to oxidative stress in the yeast Saccharomyces cerevisiae. Curr. Genet. 1996, 29, 511–515. [Google Scholar] [CrossRef]
- Meister, A.; Anderson, M.E. Glutathione. Annu. Rev. Biochem. 1983, 52, 711–760. [Google Scholar] [CrossRef]
- Tillmann, A.T.; Strijbis, S.; Cameron, G.; Radmaneshfar, E.; Thiel, M.; Munro, C.A.; MacCallum, D.M.; Distel, B.; Gow, N.A.R.; Brown, A.J.P. Contribution of Fdh3 and Glr1 to glutathione redox state, stress adaptation and virulence in Candida albicans. PLoS ONE 2015, 10, e0126940. [Google Scholar] [CrossRef]
- Sellam, A.; Charillot, J.; Mallick, J.; Tebbji, F.; Richard, J.; Michael, A.; Cook, M.; Tyers, M. The p38/HOG stress-activated protein kinase network couples growth to division in Candida albicans. PLoS Genet. 2019, 15, e1008052. [Google Scholar] [CrossRef]
- Brown, A.J.P.; Cowen, L.E.; di Pietro, A.; Quinn, J. Stress adaptation. Microbiol. Spectr. 2017, 5, 66–75. [Google Scholar] [CrossRef]
- Morano, K.A.; Grant, C.M.; Moye-Rowley, W.S. The response to heat shock and oxidative stress in Saccharomyces cerevisiae. Genetics 2012, 190, 1157–1195. [Google Scholar] [CrossRef] [PubMed]
- Roetzer, A.; Gregori, C.; Jennings, A.M.; Quintin, J.; Ferrandon, D.; Butler, G.; Kuchler, K.; Ammerer, G.; Schuller, C. Candida glabrata environmental stress response involves Saccharomyces cerevisiae Msn2/4 orthologous transcription factors. Mol. Microbiol. 2008, 69, 603–620. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Chen, J.; Liua, J.; Han, B. A transcriptome analysis reveals the oxidative stress response in Saccharomyces cerevisiae. RSC. Adv. 2015, 5, 22923–22934. [Google Scholar] [CrossRef]
- Charizanis, C.; Juhnke, H.; Krems, B.; Entian, K.D. The oxidative stress response mediated via Pos9/Skn7 is negatively regulated by the Ras/PKA pathway in Saccharomyces cerevisiae. Mol. Gen. Genet. 1999, 261, 740–752. [Google Scholar] [CrossRef]
- Gomez-Pastor, R.; Garre, E.; Perez-Torrado, R.; Matallana, E. Trx2p-dependent regulation of Saccharomyces cerevisiae oxidative stress response by the Skn7p transcription factor under respiring conditions. PLoS ONE 2013, 8, e85404. [Google Scholar] [CrossRef]
- Lee, J.; Godon, C.; Lagniel, G.; Spector, D.; Garin, J.; Labarre, J.; Toledano, M.B. Yap1 and Skn7 control two specialized oxidative stress response regulons in yeast. J. Biol. Chem. 1999, 274, 16040–16046. [Google Scholar] [CrossRef]
- Roetzer, A.; Klopf, E.; Gratz, N.; Marcet-Houben, M.; Hiller, E.; Rupp, S.; Gabaldon, T.; Kovarik, P.; Schuller, C. Regulation of Candida glabrata oxidative stress resistance is adapted to host environment. FEBS Lett. 2011, 585, 319–327. [Google Scholar] [CrossRef] [PubMed]
- Kounatidis, I.; Ames, L.; Mistry, R.; Ho, H.L.; Haynes, K.; Ligoxygakis, P. A gost-pathogen interaction screen identifies ada2 as a mediator of Candida glabrata defenses against reactive oxygen species. G3 (Bethesda) 2018, 8, 1637–1647. [Google Scholar] [CrossRef]
- Fassler, J.S.; West, A.H. Fungal Skn7 stress responses and their relationship to virulence. Eukaryot. Cell 2011, 10, 156–167. [Google Scholar] [CrossRef]
| ORF | Gene | Function | Relative fold change (sfp1∆/∆: WT) |
|---|---|---|---|
| Transcription factor | |||
| orf19.1623 | CAP1 | bZIP transcription factor, responding to oxidative stress | 1.58 |
| Hog MAPK pathway | |||
| orf19.5031 | SSK1 | Response regulator | 2.01 |
| Glutathione system | |||
| orf19.5059 | GCS1 | Gamma-glutamylcysteine synthetase | 2.32 |
| orf19.85 | GPX2 | Glutathione peroxidase | 2.27 |
| orf19.6947 | GTT11 | Glutathione S-transferase | 1.65 |
| orf19.359 | GTT12 | Glutathione S-transferase | 1.70 |
| orf19.356 | GTT13 | Glutathione S-transferase | 1.71 |
| orf19.6478 | YCF1 | Glutathione S-conjugate transporter | 1.79 |
| orf19.5673 | OPT7 | Glutathione transmembrane transporter | 4.04 |
| orf19.6402 | CYS3 | Cystathionine gamma-lyase | 2.18 |
| orf19.5811 | MET1 | Uroporphyrin-3 C-methyltransferase | 3.59 |
| Thioredoxin system | |||
| orf19.5180 | PRX1 | Thioredoxin peroxidase | 3.43 |
| Oxidoreductase | |||
| orf19.113 | CIP1 | Oxidoreductase, induced by oxidative stress | 2.11 |
| orf19.125 | EBP1 | NADPH oxidoreductase | 2.26 |
| orf19.3131 | OYE32 | NAD(P)H oxidoreductase | 1.88 |
| orf19.1048 | IFD6 | Aldo-keto reductase | 2.04 |
| Others | |||
| orf19.5843 | SRR1 | Two-component system response regulator involving in multiple stress responses | 3.84 |
| orf19.7293 | MPS1 | Monopolar spindle protein | 2.13 |
| orf19.4772 | SHO1 | Adaptor protein | 1.87 |
| orf19.2028 | MXR1 | Methionine sulfoxide reductase | 1.83 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, S.-Y.; Chen, H.-F.; Yeh, Y.-C.; Xue, Y.-P.; Lan, C.-Y. The Transcription Factor Sfp1 Regulates the Oxidative Stress Response in Candida albicans. Microorganisms 2019, 7, 131. https://doi.org/10.3390/microorganisms7050131
Lee S-Y, Chen H-F, Yeh Y-C, Xue Y-P, Lan C-Y. The Transcription Factor Sfp1 Regulates the Oxidative Stress Response in Candida albicans. Microorganisms. 2019; 7(5):131. https://doi.org/10.3390/microorganisms7050131
Chicago/Turabian StyleLee, Shao-Yu, Hsueh-Fen Chen, Ying-Chieh Yeh, Yao-Peng Xue, and Chung-Yu Lan. 2019. "The Transcription Factor Sfp1 Regulates the Oxidative Stress Response in Candida albicans" Microorganisms 7, no. 5: 131. https://doi.org/10.3390/microorganisms7050131
APA StyleLee, S.-Y., Chen, H.-F., Yeh, Y.-C., Xue, Y.-P., & Lan, C.-Y. (2019). The Transcription Factor Sfp1 Regulates the Oxidative Stress Response in Candida albicans. Microorganisms, 7(5), 131. https://doi.org/10.3390/microorganisms7050131





