Probiotics: If It Does Not Help It Does Not Do Any Harm. Really?
Abstract
1. Introduction
1.1. The Probiotic Market is Booming Sky High
1.2. Probiotic: Medical and Non-Medical Uses
1.3. Can Probiotic Intake Change Microbiotic Composition and Restore Eubiosis?
1.4. Probiotics as a Supplemental Therapy in Autoimmune Diseases
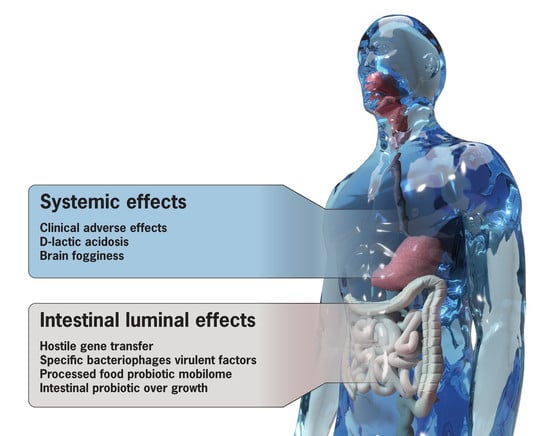
2. The Dark Side of Probiotics
2.1. Horizontal Gene Transfer (HGT)
2.2. Bacteriophages of Probiotics Transfer Mobile Virulent Genes
2.3. Processed Food and the Probiotic Mobilome
2.4. d-lactate, Metabolic Acidosis, and Brain Fogginess
2.5. Intestinal Bacterial Overgrowth, Gas, and Bloating
2.6. Additional Clinical Probiotic Side Effects
3. Problematic Inadequate Design, Incomplete Reporting, and Lack of Transparency
4. Lack of Effective Regulation of Probiotics
5. Probiotic Safety is Under-Reported
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Doré, J.; Multon, M.C.; Béhier, J.M.; Affagard, H.; Andremont, A.; Barthélémy, P.; Batitsa, R.; Bonneville, M.; Bonny, C.; Boyaval, G.; et al. The human gut microbiome as source of innovation for health: Which physiological and therapeutic outcomes could we expect? Therapie 2017, 72, 21–38. [Google Scholar] [CrossRef]
- Glendinning, L.; Free, A. Supra-organismal interactions in the human intestine. Front. Cell. Infect. Microbiol. 2014, 4, 47. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, J.K.; Holmes, E.; Wilson, I.D. Gut microorganisms, mammalian metabolism and personalized health care. Nat. Rev. Microbiol. 2005, 3, 431–438. [Google Scholar] [CrossRef] [PubMed]
- Lerner, A.; Matthias, T. GUT-the Trojan horse in remote organs’ autoimmunity. J. Clin. Cell. Immunol. 2016, 7, 401. [Google Scholar]
- Lerner, A.; Neidhöfer, S.; Matthias, T. The gut microbiome feelings of the brain: Perspective for Non-Microbiologists. Microorganisms 2017, 5, 66. [Google Scholar] [CrossRef]
- Harari, Y.N. Sapiens: A Brief History of Humankind; Vintage Books: London, UK, 2011; pp. 1–466. [Google Scholar]
- Lerner, A. The last two millennias eco-catastrophes are the driving forces for the potential genetic advantage mechanisms in celiac disease. Med. Hypotheses 2011, 77, 773–776. [Google Scholar]
- Lerner, A.; Aminov, R.; Matthias, T. Potential effects of horizontal gene exchange in the human gut. Front. Microbiol. 2017, 8, 1630. [Google Scholar] [CrossRef]
- Lerner, A.; Aminov, R.; Matthias, T. Dysbiosis may trigger autoimmune diseases via inappropriate posttranslational modification of host proteins. Front. Microbiol. 2016, 7, 84. [Google Scholar] [CrossRef]
- Ozen, M.; Dinleyici, E.C. The history of probiotics: The untold story. Benef. Microbes 2015, 6, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Gasbarrini, G.; Bonvicini, F.; Gramenzi, A. Probiotics History. In Proceedings of the 8th Probiotics, Prebiotics & New Foods for Microbiota and Human Health Meeting, Rome, Italy, 13–15 September 2015; pp. S116–S119. [Google Scholar]
- Villena, J.; Kitazawa, H. Probiotic Microorganisms: A Closer Look. Microorganisms 2017, 5, 17. [Google Scholar] [CrossRef]
- Liu, Y.; Alookaran, J.J.; Rhoads, J.M. Probiotics in Autoimmune and Inflammatory Disorders. Nutrients 2018, 10, 1537. [Google Scholar] [CrossRef]
- Mathipa, M.G.; Thantsha, M.S. Probiotic engineering: Towards development of robust probiotic strains with enhanced functional properties and for targeted control of enteric pathogens. Gut Pathog. 2017, 9, 28. [Google Scholar] [CrossRef]
- Liu, Y.; Tran, D.Q.; Rhoads, J.M. Probiotics in Disease Prevention and Treatment. J. Clin. Pharmacol. 2018, 58 (Suppl. 10), S164–S179. [Google Scholar] [CrossRef]
- Pirbaglou, M.; Katz, J.; de Souza, R.J.; Stearns, J.C.; Motamed, M.; Ritvo, P. Probiotic supplementation can positively affect anxiety and depressive symptoms: A systematic review of randomized controlled trials. Nutr. Res. 2016, 36, 889–898. [Google Scholar] [CrossRef]
- Saxelin, M. Probiotic formulations and applications, the current probiotics market, and changes in the marketplace: A European perspective. Clin. Infect. Dis. 2008, 46 (Suppl. 2), S76–S79; discussion S144–S151. [Google Scholar] [CrossRef]
- Di Cerbo, A.; Palmieri, B. Review: The market of probiotics. Pak. J. Pharm. Sci. 2015, 28, 2199–2206. [Google Scholar]
- Reid, G.; Kort, R.; Alvarez, S.; Bourdet-Sicard, R.; Benoit, V.; Cunningham, M.; Saulnier, D.; Vlieg, J.V.H.; Verstraelen, H.; Sybesma, W. Expanding the reach of probiotics through social enterprises. Benef. Microbes 2018, 9, 707–715. [Google Scholar] [CrossRef]
- Global Market Insights, Inc. Available online: https://www.gminsights.com/pressrelease/probiotics-market (accessed on 23 November 2018).
- Global Market Insights, Inc. Available online: https://www.gminsights.com/industry-analysis/probiotics-market (accessed on 23 November 2018).
- de Vrese, M.; Schrezenmeir, J. Probiotics, prebiotics, and synbiotics. Adv. Biochem. Eng. Biotechnol. 2008, 111, 1–66. [Google Scholar] [CrossRef]
- Goldin, B.R.; Gorbach, S.L. Clinical indications for probiotics: An overview. Clin. Infect. Dis. 2008, 46 (Suppl. 2), S96–S100; discussion S144–S151. [Google Scholar] [CrossRef]
- Minocha, A. Probiotics for preventive health. Nutr. Clin. Pract. 2009, 24, 227–241. [Google Scholar] [CrossRef]
- D’Angelo, C.; Reale, M.; Costantini, E. Microbiota and Probiotics in Health and HIV Infection. Nutrients 2017, 9, 615. [Google Scholar] [CrossRef]
- Westfall, E.; Brandenburg, D. Probiotics for Symptoms of Depression and Anxiety. Am. Fam. Phys. 2018, 97. online. [Google Scholar]
- Lieske, J.C. Probiotics for prevention of urinary stones. Ann. Transl. Med. 2017, 5, 29. [Google Scholar] [CrossRef]
- Misra, S.; Medhi, B. Role of probiotics as memory enhancer. Indian J. Pharmacol. 2013, 45, 311–312. [Google Scholar] [CrossRef]
- Chong, H.X.; Yusoff, N.A.A.; Hor, Y.Y.; Lew, L.C.; Jaafar, M.H.; Choi, S.B.; Yusoff, M.; Wahid, N.; Abdullah, M.; Zakaria, N.; et al. Lactobacillus plantarum DR7 alleviates stress and anxiety in adults: A randomised, double-blind, placebo-controlled study. Benef. Microbes 2019, 18, 1–20. [Google Scholar] [CrossRef]
- Gorbach, S.L. Probiotics in the third millennium. Dig. Liver Dis. 2002, 34 (Suppl. 2), S2–S7. [Google Scholar] [CrossRef]
- Mahmoodpoor, A.; Hamishehkar, H.; Asghari, R.; Abri, R.; Shadvar, K.; Sanaie, S. Effect of a Probiotic Preparation on Ventilator-Associated Pneumonia in Critically Ill Patients Admitted to the Intensive Care Unit: A Prospective Double-Blind Randomized Controlled Trial. Nutr. Clin. Pract. 2019, 34, 156–162. [Google Scholar] [CrossRef]
- Sohn, K.; Underwood, M.A. Prenatal and postnatal administration of prebiotics and probiotics. Semin. Fetal Neonatal Med. 2017, 22, 284–289. [Google Scholar] [CrossRef]
- Ayala, F.R.; Bauman, C.; Cogliati, S.; Leñini, C.; Bartolini, M.; Grau, R. Microbial flora, probiotics, Bacillus subtilis and the search for a long and healthy human longevity. Microb. Cell 2017, 4, 133–136. [Google Scholar] [CrossRef]
- Guslandi, M. Role of Probiotics in Crohn’s Disease and in Pouchitis. J. Clin. Gastroenterol. 2015, 49 (Suppl. 1), S46–S49. [Google Scholar] [CrossRef]
- Tirandaz, H.; Ebrahim-Habibi, M.B.; Moradveisi, B.; Raoofi, S.; Salehi-Najafabadi, A.; Mohammadi, E. Microbiota potential for the treatment of sexual dysfunction. Med. Hypotheses 2018, 115, 46–49. [Google Scholar] [CrossRef]
- Ahtesh, F.B.; Stojanovska, L.; Apostolopoulos, V. Anti-hypertensive peptides released from milk proteins by probiotics. Maturitas 2018, 115, 103–109. [Google Scholar] [CrossRef]
- Santocchi, E.; Guiducci, L.; Fulceri, F.; Billeci, L.; Buzzigoli, E.; Apicella, F.; Calderoni, S.; Grossi, E.; Morales, M.A.; Muratori, F. Gut to brain interaction in Autism Spectrum Disorders: A randomized controlled trial on the role of probiotics on clinical, biochemical and neurophysiological parameters. BMC Psychiatry 2016, 16, 183. [Google Scholar] [CrossRef]
- Welcome, M.O. Current Perspectives and Mechanisms of Relationship between Intestinal Microbiota Dysfunction and Dementia: A Review. Dement. Geriatr. Cogn. Dis. Extra 2018, 8, 360–381. [Google Scholar] [CrossRef]
- Bezirtzoglou, E.; Stavropoulou, E. Immunology and probiotic impact of the newborn and young children intestinal microflora. Anaerobe 2011, 17, 369–374. [Google Scholar] [CrossRef]
- Khoruts, A. Targeting the microbiome: From probiotics to fecal microbiota transplantation. Genome Med. 2018, 10, 80. [Google Scholar] [CrossRef]
- Rao, S.S.C.; Rehman, A.; Yu, S.; Andino, N.M. Brain fogginess, gas and bloating: A link between SIBO, probiotics and metabolic acidosis. Clin. Transl. Gastroenterol. 2018, 9, 162. [Google Scholar] [CrossRef]
- Zmora, N.; Zilberman-Schapira, G.; Suez, J.; Mor, U.; Dori-Bachash, M.; Bashiardes, S.; Kotler, E.; Zur, M.; Regev-Lehavi, D.; Brik, R.B.-Z.; et al. Personalized Gut Mucosal Colonization Resistance to Empiric Probiotics Is Associated with Unique Host and Microbiome Features. Cell 2018, 174, 1388–1405. [Google Scholar] [CrossRef]
- Suez, J.; Zmora, N.; Zilberman-Schapira, G.; Mor, U.; Dori-Bachash, M.; Bashiardes, S.; Zur, M.; Regev-Lehavi, D.; Brik, R.B.-Z.; Federici, S.; et al. Post-Antibiotic Gut Mucosal Microbiome Reconstitution Is Impaired by Probiotics and Improved by Autologous FMT. Cell 2018, 174, 1406–1423. [Google Scholar] [CrossRef]
- Singh, A.; Sarangi, A.N.; Goel, A.; Srivastava, R.; Bhargava, R.; Gaur, P.; Aggarwal, A.; Aggarwal, R. Effect of administration of a probiotic preparation on gut microbiota and immune response in healthy women in India: An open-label, single-arm pilot study. BMC Gastroenterol. 2018, 18, 85. [Google Scholar] [CrossRef]
- Derrien, M.; van Hylckama Vlieg, J.E. Fate, activity, and impact of ingested bacteria within the human gut microbiota. Trends Microbiol. 2015, 23, 354–366. [Google Scholar] [CrossRef]
- Maldonado-Gómez, M.X.; Martínez, I.; Bottacini, F.; O’Callaghan, A.; Ventura, M.; van Sinderen, D.; Hillmann, B.; Vangay, P.; Knights, D.; Hutkins, R.W.; et al. Stable Engraftment of Bifidobacterium longum AH1206 in the Human Gut Depends on Individualized Features of the Resident Microbiome. Cell Host Microbe 2016, 20, 515–526. [Google Scholar] [CrossRef]
- Laursen, M.F.; Laursen, R.P.; Larnkjær, A.; Michaelsen, K.F.; Bahl, M.I.; Licht, T.R. Administration of two probiotic strains during early childhood does not affect the endogenous gut microbiota composition despite probiotic proliferation. BMC Microbiol. 2017, 17, 175. [Google Scholar] [CrossRef]
- Kristensen, N.B.; Bryrup, T.; Allin, K.H.; Nielsen, T.; Hansen, T.H.; Pedersen, O. Alterations in fecal microbiota composition by probiotic supplementation in healthy adults: A systematic review of randomized controlled trials. Genome Med. 2016, 8, 52. [Google Scholar] [CrossRef]
- Selmi, C.; Gao, B.; Gershwin, M.E. The long and latent road to autoimmunity. Cell. Mol. Immunol. 2018, 15, 543–546. [Google Scholar] [CrossRef]
- De Luca, F.; Shoenfeld, Y. The microbiome in autoimmune diseases. Clin. Exp. Immunol. 2019, 195, 74–85. [Google Scholar] [CrossRef]
- Haase, S.; Haghikia, A.; Wilck, N.; Müller, D.N.; Linker, R.A. Impacts of microbiome metabolites on immune regulation and autoimmunity. Immunology 2018, 154, 230–238. [Google Scholar] [CrossRef]
- Chen, B.; Sun, L.; Zhang, X. Integration of microbiome and epigenome to decipher the pathogenesis of autoimmune diseases. J. Autoimmun. 2017, 83, 31–42. [Google Scholar] [CrossRef]
- de Oliveira, G.L.V.; Leite, A.Z.; Higuchi, B.S.; Gonzaga, M.I.; Mariano, V.S. Intestinal Dysbiosis and pRobiotic Applications in Autoimmune Diseases. Immunology 2017, 152, 1–12. [Google Scholar] [CrossRef]
- de Oliveira, G.L.V. Probiotic applications in autoimmune diseases In Probiotics-Current Knowledge and Future Prospects; IntechOpen: London, UK, 2018; pp. 69–89. [Google Scholar]
- Marietta, E.; Horwath, I.; Balakrishnan, B.; Taneja, V. Role of the intestinal microbiome in autoimmune diseases and its use in treatments. Cell Immunol. 2018. [Google Scholar] [CrossRef]
- Balakrishnan, B.; Taneja, V. Microbial modulation of the gut microbiome for treating autoimmune diseases. Expert Rev Gastroenterol. Hepatol. 2018, 12, 985–996. [Google Scholar] [CrossRef]
- Dwivedi, M.; Kumar, P.; Laddha, N.C.; Kemp, E.H. Induction of regulatory T cells: A role for probiotics and prebiotics to suppress autoimmunity. Autoimmun. Rev. 2016, 15, 379–392. [Google Scholar] [CrossRef]
- Tankou, S.K.; Regev, K.; Healy, B.C.; Cox, L.M.; Tjon, E.; Kivisakk, P.; Vanande, I.P.; Cook, S.; Gandhi, R.; Glanz, B.; et al. Investigation of probiotics in multiple sclerosis. Mult. Scler. 2018, 24, 58–63. [Google Scholar] [CrossRef]
- Kota, R.K.; Ambati, R.R.; Yalakurthi, A.K.; Srirama, K.; Reddy, P.N. Recent advances in probiotics as live biotherapeutics against gastrointestinal diseases. Curr. Pharm. Des. 2018, 24, 3162–3171. [Google Scholar] [CrossRef]
- Eom, T.; Kim, Y.S.; Choi, C.H.; Sadowsky, M.J.; Unno, T. Current understanding of microbiota- and dietary-therapies for treating inflammatory bowel disease. J. Microbiol. 2018, 56, 189–198. [Google Scholar] [CrossRef]
- Rinaldi, E.; Consonni, A.; Guidesi, E.; Elli, M.; Mantegazza, R.; Baggi, F. Gut microbiota and probiotics: Novel immune system modulators in myasthenia gravis? Ann. N. Y. Acad. Sci. 2018, 1413, 49–58. [Google Scholar] [CrossRef]
- Aqaeinezhad Rudbane, S.M.; Rahmdel, S.; Abdollahzadeh, S.M.; Zare, M.; Bazrafshan, A.; Mazloomi, S.M. The efficacy of probiotic supplementation in rheumatoid arthritis: A meta-analysis of randomized, controlled trials. Inflammopharmacology 2018, 26, 67–76. [Google Scholar] [CrossRef]
- Aminov, R.I. The extent and regulation of lateral gene transfer in natural microbial ecosystems. In Horizontal Gene Transfer in Microorganisms; Francino, M.P., Ed.; Horizon Scientific Press: Norwich, UK, 2012; Chapter 6; pp. 93–100. [Google Scholar]
- Wong, A.; Ngu, D.Y.; Dan, L.A.; Ooi, A.; Lim, R.L. Detection of antibiotic resistance in probiotics of dietary supplements. Nutr. J. 2015, 14, 95. [Google Scholar] [CrossRef]
- Zheng, M.; Zhang, R.; Tian, X.; Zhou, X.; Pan, X.; Wong, A. Assessing the Risk of Probiotic Dietary Supplements in the Context of Antibiotic Resistance. Front. Microbiol. 2017, 8, 908. [Google Scholar] [CrossRef]
- van Reenen, C.A.; Dicks, L.M. Horizontal gene transfer amongst probiotic lactic acid bacteria and other intestinal microbiota: What are the possibilities? A review. Arch. Microbiol. 2011, 193, 157–168. [Google Scholar] [CrossRef]
- Mercanti, D.J.; Rousseau, G.M.; Capra, M.L.; Quiberoni, A.; Tremblay, D.M.; Labrie, S.J.; Moineau, S. Genomic Diversity of Phages Infecting Probiotic Strains of Lactobacillus paracasei. Appl. Environ. Microbiol. 2015, 82, 95–105. [Google Scholar] [CrossRef]
- Ceapa, C.; Davids, M.; Ritari, J.; Lambert, J.; Wels, M.; Douillard, F.P.; Smokvina, T.; De Vos, W.M.; Knol, J.; Kleerebezem, M. The Variable Regions of Lactobacillus rhamnosus Genomes Reveal the Dynamic Evolution of Metabolic and Host-Adaptation Repertoires. Genome Biol. Evol. 2016, 8, 1889–1905. [Google Scholar] [CrossRef]
- Tannock, G.W.; Luchansky, J.B.; Miller, L.; Connell, H.; Thode-Andersen, S.; Mercer, A.A.; Klaenhammer, T.R. Molecular characterization of a plasmid-borne (pGT633) erythromycin resistance determinant (ermGT) from Lactobacillus reuteri 100-63. Plasmid 1994, 31, 60–71. [Google Scholar] [CrossRef]
- Lin, C.F.; Fung, Z.F.; Wu, C.L.; Chung, T.C. Molecular characterization of a plasmid-borne (pTC82) chloramphenicol resistance determinant (cat-TC) from Lactobacillus reuteri G4. Plasmid 1996, 36, 116–124. [Google Scholar] [CrossRef]
- Egervärn, M.; Roos, S.; Lindmark, H. Identification and characterization of antibiotic resistance genes in Lactobacillus reuteri and Lactobacillus plantarum. J. Appl. Microbiol. 2009, 107, 1658–1668. [Google Scholar] [CrossRef]
- Baugher, J.L.; Durmaz, E.; Klaenhammer, T.R. Spontaneously induced prophages in Lactobacillus gasseri contribute to horizontal gene transfer. Appl. Environ. Microbiol. 2014, 80, 3508–3517. [Google Scholar] [CrossRef]
- Courvalin, P. Predictable and unpredictable evolution of antibiotic resistance. J. Intern. Med. 2008, 264, 4–16. [Google Scholar] [CrossRef]
- Doron, S.; Snydman, D.R. Risk and safety of probiotics. Clin. Infect. Dis. 2015, 60 (Suppl. 2), S129–S134. [Google Scholar] [CrossRef]
- Egervärn, M.; Lindmark, H.; Olsson, J.; Roos, S. Transferability of a tetracycline resistance gene from probiotic Lactobacillus reuteri to bacteria in the gastrointestinal tract of humans. Antonie Van Leeuwenhoek 2010, 97, 189–200. [Google Scholar] [CrossRef]
- Imperial, I.C.; Ibana, J.A. Addressing the Antibiotic Resistance Problem with Probiotics: Reducing the Risk of Its Double-Edged Sword Effect. Front. Microbiol. 2016, 7, 1983. [Google Scholar] [CrossRef]
- Rosander, A.; Connolly, E.; Roos, S. Removal of antibiotic resistance gene-carrying plasmids from Lactobacillus reuteri ATCC 55730 and characterization of the resulting daughter strain, L. reuteri DSM 17938. Appl. Environ. Microbiol. 2008, 74, 6032–6040. [Google Scholar] [CrossRef]
- Muñoz-Atienza, E.; Gómez-Sala, B.; Araújo, C.; Campanero, C.; del Campo, R.; Hernández, P.E.; Herranz, C.; Cintas, L.M. Antimicrobial activity, antibiotic susceptibility and virulence factors of Lactic Acid Bacteria of aquatic origin intended for use as probiotics in aquaculture. BMC Microbiol. 2013, 13, 15. [Google Scholar] [CrossRef]
- Fittipaldi, N.; Segura, M.; Grenier, D.; Gottschalk, M. Virulence factors involved in the pathogenesis of the infection caused by the swine pathogen and zoonotic agent Streptococcus suis. Future Microbiol. 2012, 7, 259–279. [Google Scholar] [CrossRef]
- Yu, J.; Pian, Y.; Ge, J.; Guo, J.; Zheng, Y.; Jiang, H.; Hao, H.; Yuan, Y.; Jiang, Y.; Yang, M. Functional and Structural Characterization of the Antiphagocytic Properties of a Novel Transglutaminase from Streptococcus suis. J. Biol. Chem. 2015, 290, 19081–19092. [Google Scholar] [CrossRef]
- Lerner, A.; Matthias, T. Changes in intestinal tight junction permeability associated with industrial food additives explain the rising incidence of autoimmune disease. Autoimmun. Rev. 2015, 14, 479–489. [Google Scholar] [CrossRef]
- Lerner, A.; Matthias, T. Possible association between celiac disease and bacterial transglutaminase in food processing: A hypothesis. Nutr. Rev. 2015, 73, 544–552. [Google Scholar] [CrossRef]
- Lerner, A.; Aminov, R.; Matthias, T. Intestinal dysbiotic transglutaminases are potential environmental drivers of systemic autoimmunogenesis. Front. Microbiol. 2017, 8, 66. [Google Scholar] [CrossRef]
- Matthias, T.; Jeremias, P.; Neidhöfer, S.; Lerner, A. The industrial food additive microbial transglutaminase, mimics the tissue transglutaminase and is immunogenic in celiac disease patients. Autoimmun. Rev. 2016, 15, 1111–1119. [Google Scholar] [CrossRef]
- Matthias, T.; Lerner, A. Microbial transglutaminase is immunogenic and potentially pathogenic in pediatric celiac disease. Front. Pediatr. 2018, 6, 389. [Google Scholar] [CrossRef]
- Zhang, D.; de Souza, R.F.; Anantharaman, V.; Lyer, L.M.; Aravind, L. Polymorphic toxin systems: Comprehensive characterization of trafficking modes, processing, mechanisms of action, immunity and ecology using comparative genomics. Biol. Direct 2012, 7, 18. [Google Scholar] [CrossRef]
- Masuda, M.; Betancourt, L.; Matsuzawa, T.; Kashimoto, T.; Takao, T.; Shimonishi, Y.; Horiguchi, Y. Activation of Rho through a cross-link with polyamines catalyzed by Bordetella dermonecrotizing toxin. EMBO J. 2000, 19, 521–530. [Google Scholar] [CrossRef]
- Jubelin, G.; Chavez, C.V.; Taieb, F.; Banfield, M.J.; Samba-Louaka, A.; Nobe, R.; Nougayrède, J.-P.; Zumbihl, R.; Givaudan, A.; Escoubas, J.-M.; et al. Cycle Inhibiting Factors (CIFs) Are a Growing Family of Functional Cyclomodulins Present in Invertebrate and Mammal Bacterial Pathogens. PLoS ONE 2009, 4, e4855. [Google Scholar] [CrossRef]
- Schmidt, G.; Selzer, J.; Lerm, M.; Aktories, K. The Rho-deamidating cytotoxic necrotizing factor 1 from Escherichia coli possesses transglutaminase activity. Cysteine 866 and histidine 881 are essential for enzyme activity. J. Biol. Chem. 1998, 273, 13669–13674. [Google Scholar] [CrossRef]
- Makarova, K.S.; Aravind, L.; Koonin, E.V. A superfamily of archaeal, bacterial, and eukaryotic proteins homologous to animal transglutaminases. Protein Sci. 1999, 8, 1714–1719. [Google Scholar] [CrossRef]
- Lerner, A.; Matthias, T. Microbial transglutaminase is beneficial to food industries but a caveat to public health. Med One 2019, 4, e190001. [Google Scholar] [CrossRef]
- Sitaraman, R. Prokaryotic horizontal gene transfer within the human holobiont: Ecological-evolutionary inferences, implications and possibilities. Microbiome 2018, 6, 163. [Google Scholar] [CrossRef]
- Xu, F.; Wang, J.; Guo, Y.; Fu, P.; Zeng, H.; Li, Z.; Pei, X.; Liu, X.; Wang, S. Antibiotic resistance, biochemical typing, and PFGE typing of Bifidobacterium strains commonly used in probiotic health foods. Food Sci. Biotechnol. 2018, 27, 467–477. [Google Scholar] [CrossRef]
- Gueimonde, M.; Sánchez, B.; de Los Reyes-Gavilán, C.; Margolles, A. Antibiotic resistance in probiotic bacteria. Front. Microbiol. 2013, 4, 202. [Google Scholar] [CrossRef]
- Comunian, R.; Daga, E.; Dupré, I.; Paba, A.; Devirgiliis, C.; Piccioni, V.; Perozzi, G.; Zonenschain, D.; Rebecchi, A.; Morelli, L.; et al. Susceptibility to tetracycline and erythromycin of Lactobacillus paracasei strains isolated from traditional Italian fermented foods. Int. J. Food Microbiol. 2010, 138, 151–156. [Google Scholar] [CrossRef]
- Guo, H.; Pan, L.; Li, L.; Lu, J.; Kwok, L.; Menghe, B.; Zhang, H. Characterization of Antibiotic Resistance Genes from Lactobacillus Isolated from Traditional Dairy Products. J. Food Sci. 2017, 82, 724–730. [Google Scholar] [CrossRef]
- Dec, M.; Nowaczek, A.; Stępień-Pyśniak, D.; Wawrzykowski, J.; Urban-Chmiel, R. Identification and antibiotic susceptibility of lactobacilli isolated from turkeys. BMC Microbiol. 2018, 18, 168. [Google Scholar] [CrossRef]
- Agersø, Y.; Stuer-Lauridsen, B.; Bjerre, K.; Jensen, M.G.; Johansen, E.; Bennedsen, M.; Brockmann, E.; Nielsen, B. Antimicrobial Susceptibility Testing and Tentative Epidemiological Cutoff Values for Five Bacillus Species Relevant for Use as Animal Feed Additives or for Plant Protection. Appl. Environ. Microbiol. 2018, 84, e01108-18. [Google Scholar] [CrossRef]
- Abriouel, H.; Casado Muñoz, M.D.C.; Lavilla Lerma, L.; Pérez Montoro, B.; Bockelmann, W.; Pichner, R.; Kabisch, J.; Cho, G.S.; Franz, C.M.A.P.; Gálvez, A.; et al. New insights in antibiotic resistance of Lactobacillus species from fermented foods. Food Res. Int. 2015, 78, 465–481. [Google Scholar] [CrossRef]
- Vitetta, L.; Vitetta, G.; Hall, S. Immunological Tolerance and Function: Associations Between Intestinal Bacteria, Probiotics, Prebiotics, and Phages. Front. Immunol. 2018, 9, 2240. [Google Scholar] [CrossRef]
- Bakhshinejad, B.; Ghiasvand, S. Bacteriophages in the human gut: Our fellow travelers throughout life and potential biomarkers of heath or disease. Virus Res. 2017, 240, 47–55. [Google Scholar] [CrossRef]
- Tetz, G.; Tetz, V. Bacteriophages as New Human Viral Pathogens. Microorganisms 2018, 6, 54. [Google Scholar] [CrossRef]
- Pfeiffer, J.K.; Virgin, H.W. Viral immunity. Transkingdom control of viral infection and immunity in the mammalian intestine. Science 2016, 351, aad5872. [Google Scholar] [CrossRef]
- Proal, A.; Marshall, T. Myalgic Encephalomyelitis/Chronic Fatigue Syndrome in the Era of the Human Microbiome: Persistent Pathogens Drive Chronic Symptoms by Interfering With Host Metabolism, Gene Expression, and Immunity. Front. Pediatr. 2018. [Google Scholar] [CrossRef]
- Tetz, G.; Tetz, V. Prion-Like Domains in Phagobiota. Front. Microbiol. 2017, 8, 2239. [Google Scholar] [CrossRef]
- Muniesa, M.; Colomer-Lluch, M.; Jofre, J. Potential impact of environmental bacteriophages in spreading antibiotic resistance genes. Future Microbiol. 2013, 8, 739–751. [Google Scholar] [CrossRef]
- Muniesa, M.; Colomer-Lluch, M.; Jofre, J. Could bacteriophages transfer antibiotic resistance genes from environmental bacteria to human-body associated bacterial populations? Mob. Genet. Elem. 2013, 3, e25847. [Google Scholar] [CrossRef]
- Lood, R.; Ertürk, G.; Mattiasson, B. Revisiting Antibiotic Resistance Spreading in Wastewater Treatment Plants—Bacteriophages as a Much Neglected Potential Transmission Vehicle. Front. Microbiol. 2017, 8, 2298. [Google Scholar] [CrossRef]
- Quirós, P.; Colomer-Lluch, M.; Martínez-Castillo, A.; Miró, E.; Argente, M.; Jofre, J.; Navarro, F.; Muniesa, M. Antibiotic resistance genes in the bacteriophage DNA fraction of human fecal samples. Antimicrob. Agents Chemother. 2014, 58, 606–609. [Google Scholar] [CrossRef]
- Eaton, T.J.; Gasson, M.J. Molecular screening of Enterococcus virulence determinants and potential for genetic exchange between food and medical isolates. Appl. Environ. Microbiol. 2001, 67, 1628–1635. [Google Scholar] [CrossRef]
- Marcó, M.B.; Moineau, S.; Quiberoni, A. Bacteriophages and dairy fermentations. Bacteriophage 2012, 2, 149–158. [Google Scholar] [CrossRef]
- Domingo-Calap, P.; Delgado-Martínez, J. Bacteriophages: Protagonists of a Post-Antibiotic Era. Antibiotics 2018, 7, 66. [Google Scholar] [CrossRef]
- Li, J.; Cao, J.; Zhu, Y.G.; Chen, Q.L.; Shen, F.; Wu, Y.; Xu, S.; Fan, H.; Da, G.; Huang, R.-J.; et al. Global Survey of Antibiotic Resistance Genes in Air. Environ. Sci. Technol. 2018, 52, 10975–10984. [Google Scholar] [CrossRef]
- Villion, M.; Moineau, S. Bacteriophages of lactobacillus. Front. Biosci. 2009, 14, 1661–1683. [Google Scholar] [CrossRef]
- Capra, M.L.; Del L Quiberoni, A.; Ackermann, H.W.; Moineau, S.; Reinheimer, J.A. Characterization of a new virulent phage (MLC-A) of Lactobacillus paracasei. J. Dairy Sci. 2006, 89, 2414–2423. [Google Scholar] [CrossRef]
- Aucouturier, A.; Chain, F.; Langella, P.; Bidnenko, E. Characterization of a Prophage-Free Derivative Strain of Lactococcus lactis ssp. lactis IL1403 Reveals the Importance of Prophages for Phenotypic Plasticity of the Host. Front. Microbiol. 2018, 9, 2032. [Google Scholar] [CrossRef]
- Waller, A.S.; Yamada, T.; Kristensen, D.M.; Kultima, J.R.; Sunagawa, S.; Koonin, E.V.; Bork, P. Classification and quantification of bacteriophage taxa in human gut metagenomes. ISME J. 2014, 8, 1391–1402. [Google Scholar] [CrossRef]
- Visweswaran, G.R.; Kurek, D.; Szeliga, M.; Pastrana, F.R.; Kuipers, O.P.; Kok, J.; Buist, G. Expression of prophage-encoded endolysins contributes to autolysis of Lactococcus lactis. Appl. Microbiol. Biotechnol. 2017, 101, 1099–1110. [Google Scholar] [CrossRef]
- Labrie, S.J.; Moineau, S. Abortive infection mechanisms and prophage sequences significantly influence the genetic makeup of emerging lytic lactococcal phages. J. Bacteriol. 2007, 189, 1482–1487. [Google Scholar] [CrossRef]
- Chopin, M.C.; Chopin, A.; Bidnenko, E. Phage abortive infection in lactococci: Variations on a theme. Curr. Opin. Microbiol. 2005, 8, 473–479. [Google Scholar] [CrossRef]
- Devirgiliis, C.; Zinno, P.; Perozzi, G. Update on antibiotic resistance in foodborne Lactobacillus and Lactococcus species. Front. Microbiol. 2013, 4, 301. [Google Scholar] [CrossRef]
- Zhang, D.; Aravind, L. Novel transglutaminase-like peptidase and C2 domains elucidate the structure, biogenesis and evolution of the ciliary compartment. Cell Cycle 2012, 11, 3861–3875. [Google Scholar] [CrossRef]
- Khan, S.H.; Ansari, F.A. Probiotics—The friendly bacteria with market potential in global market. Pak. J. Pharm. Sci. 2007, 20, 76–82. [Google Scholar]
- Kopp-Hoolihan, L. Prophylactic and therapeutic uses of probiotics: A review. J. Am. Diet. Assoc. 2001, 101, 229–238. [Google Scholar] [CrossRef]
- Kaur, I.P.; Chopra, K.; Saini, A. Probiotics: Potential pharmaceutical applications. Eur. J. Pharm. Sci. 2002, 15, 1–9. [Google Scholar] [CrossRef]
- O’Toole, P.W.; Marchesi, J.R.; Hill, C. Next-generation probiotics: The spectrum from probiotics to live biotherapeutics. Nat. Microbiol. 2017, 2, 17057. [Google Scholar] [CrossRef]
- Boesmans, L.; Valles-Colomer, M.; Wang, J.; Eeckhaut, V.; Falony, G.; Ducatelle, R.; van Immerseel, F.; Raes, J.; Verbeke, K. Butyrate Producers as Potential Next-Generation Probiotics: Safety Assessment of the Administration of Butyricicoccus pullicaecorum to Healthy Volunteers. mSystems 2018, e00094-18. [Google Scholar] [CrossRef]
- Roman, P.; Estévez, A.F.; Miras, A.; Sánchez-Labraca, N.; Cañadas, F.; Vivas, A.B.; Cardona, D. A Pilot Randomized Controlled Trial to Explore Cognitive and Emotional Effects of Probiotics in Fibromyalgia. Sci. Rep. 2018, 8, 10965. [Google Scholar] [CrossRef]
- Rezac, S.; Kok, C.R.; Heermann, M.; Hutkins, R. Fermented Foods as a Dietary Source of Live Organisms. Front. Microbiol. 2018, 9, 1785. [Google Scholar] [CrossRef]
- Oniciuc, E.A.; Likotrafiti, E.; Alvarez-Molina, A.; Prieto, M.; Santos, J.A.; Alvarez-Ordóñez, A. The Present and Future of Whole Genome Sequencing (WGS) and Whole Metagenome Sequencing (WMS) for Surveillance of Antimicrobial Resistant Microorganisms and Antimicrobial Resistance Genes across the Food Chain. Genes 2018, 9, 268. [Google Scholar] [CrossRef]
- Flórez, A.B.; Vázquez, L.; Mayo, B. A Functional Metagenomic Analysis of Tetracycline Resistance in Cheese Bacteria. Front. Microbiol. 2017, 8, 907. [Google Scholar] [CrossRef]
- de Paula, A.C.L.; Medeiros, J.D.; de Azevedo, A.C.; de Assis Chagas, J.M.; da Silva, V.L.; Diniz, C.G. Antibiotic Resistance Genetic Markers and Integrons in White Soft Cheese: Aspects of Clinical Resistome and Potentiality of Horizontal Gene Transfer. Genes 2018, 9, 106. [Google Scholar] [CrossRef]
- Devirgiliis, C.; Zinno, P.; Stirpe, M.; Barile, S.; Perozzi, G. Functional screening of antibiotic resistance genes from a representative metagenomic library of food fermenting microbiota. Biomed. Res. Int. 2014, 2014, 290967. [Google Scholar] [CrossRef]
- Cole, M.L.; Singh, O.V. Microbial occurrence and antibiotic resistance in ready-to-go food items. J. Food Sci. Technol. 2018, 55, 2600–2609. [Google Scholar] [CrossRef]
- Papagaroufalis, K.; Fotiou, A.; Egli, D.; Tran, L.A.; Steenhout, P. A Randomized Double Blind Controlled Safety Trial Evaluating d-Lactic Acid Production in Healthy Infants Fed a Lactobacillus reuteri-containing Formula. Nutr. Metab. Insights 2014, 7, 19–27. [Google Scholar] [CrossRef]
- Haschke-Becher, E.; Brunser, O.; Cruchet, S.; Gotteland, M.; Haschke, F.; Bachmann, C. Urinary D-lactate excretion in infants receiving Lactobacillus johnsonii with formula. Ann. Nutr. Metab. 2008, 53, 240–244. [Google Scholar] [CrossRef]
- Vitetta, L.; Coulson, S.; Thomsen, M.; Nguyen, T.; Hall, S. Probiotics, D-Lactic acidosis, oxidative stress and strain specificity. Gut Microbes 2017, 8, 311–322. [Google Scholar] [CrossRef]
- Munakata, S.; Arakawa, C.; Kohira, R.; Fujita, Y.; Fuchigami, T.; Mugishima, H. A case of D-lactic acid encephalopathy associated with use of probiotics. Brain Dev. 2010, 32, 691–694. [Google Scholar] [CrossRef]
- Rao, S.S.C.; Yu, S.; Tetangco, E.P.; Yan, Y. Probiotics can Cause D-Lactic Acidosis and Brain Fogginess: Reply to Quigley et al. Clin. Transl. Gastroenterol. 2018, 9, 207. [Google Scholar] [CrossRef]
- Muñoz, J.A.; Chenoll, E.; Casinos, B.; Bataller, E.; Ramón, D.; Genovés, S.; Montava, R.; Ribes, J.M.; Buesa, J.; Fàbrega, J.; et al. Novel probiotic Bifidobacterium longum subsp. infantis CECT 7210 strain active against rotavirus infections. Appl. Environ. Microbiol. 2011, 77, 8775–8783. [Google Scholar] [CrossRef]
- Ewaschuk, J.B.; Naylor, J.M.; Zello, G.A. D-lactate in human and ruminant metabolism. J. Nutr. 2005, 135, 1619–1625. [Google Scholar] [CrossRef]
- Ku, W.H.; Lau, D.C.Y.; Huen, K.F. Probiotics Provoked D-lactic Acidosis in Short Bowel Syndrome: Case Report and Literature. HK J. Paediatr. 2006, 11, 246–254. [Google Scholar]
- Petersen, C. D-lactic acidosis. Nutr. Clin. Pract. 2005, 20, 634–645. [Google Scholar] [CrossRef]
- Uribarri, J.; Oh, M.S.; Carroll, H.J. D-lactic acidosis. A review of clinical presentation, biochemical features, and pathophysiologic mechanisms. Medicine 1998, 77, 73–82. [Google Scholar] [CrossRef]
- Oh, M.S.; Phelps, K.R.; Traube, M.; Barbosa-Saldivar, J.L.; Boxhill, C.; Carroll, H.J. D-lactic acidosis in a man with the short-bowel syndrome. N. Engl. J. Med. 1979, 301, 249–252. [Google Scholar] [CrossRef]
- Quigley, E.M.M.; Pot, B.; Sanders, M.E. ‘Brain Fogginess’ and D-Lactic Acidosis: Probiotics Are Not the Cause. Clin. Transl. Gastroenterol. 2018, 9, 187. [Google Scholar] [CrossRef]
- Sachdeva, S.; Puri, A.S.; Kumar, A.; Dalal, A.; Dahale, A.S. Brain Fogginess and SIBO: A Link or Just a Mirage? Clin. Transl. Gastroenterol. 2018, 9, 184. [Google Scholar] [CrossRef]
- Van den Nieuwboer, M.; Brummer, R.J.; Guarner, F.; Morelli, L.; Cabana, M.; Claasen, E. The administration of probiotics and synbiotics in immune compromised adults: Is it safe? Benef. Microbes 2015, 6, 3–17. [Google Scholar] [CrossRef]
- Sanders, M.E.; Akkermans, L.M.; Haller, D.; Hammerman, C.; Heimbach, J.; Hörmannsperger, G.; Huys, G. Safety assessment of probiotics for human use. Gut Microbes 2010, 1, 164–185. [Google Scholar] [CrossRef]
- Sherid, M.; Samo, S.; Sulaiman, S.; Husein, H.; Sifuentes, H.; Sridhar, S. Liver abscess and bacteremia caused by lactobacillus: Role of probiotics? Case report and review of the literature. BMC Gastroenterol. 2016, 16, 138. [Google Scholar] [CrossRef]
- Kopp, M.V.; Hennemuth, I.; Heinzmann, A.; Urbanek, R. Randomized, double-blind, placebo-controlled trial of probiotics for primary prevention: no clinical effects of Lactobacillus GG supplementation. Pediatrics 2008, 121, e850-6. [Google Scholar] [CrossRef]
- Franz, C.M.; Muscholl-Silberhorn, A.B.; Yousif, N.M.; Vancanneyt, M.; Swings, J.; Holzapfel, W.H. Incidence of virulence factors and antibiotic resistance among Enterococci isolated from food. Appl. Environ. Microbiol. 2001, 67, 4385–4389. [Google Scholar] [CrossRef]
- Rathnayake, I.U.; Hargreaves, M.; Huygens, F. Antibiotic resistance and virulence traits in clinical and environmental Enterococcus faecalis and Enterococcus faecium isolates. Syst. Appl. Microbiol. 2012, 35, 326–333. [Google Scholar] [CrossRef]
- Kho, Z.Y.; Lal, S.K. The Human Gut Microbiome—A Potential Controller of Wellness and Disease. Front. Microbiol. 2018, 9, 1835. [Google Scholar] [CrossRef]
- Fatal gastrointestinal mucormycosis in an infant following use of contaminated ABC Dophilus powder from Solgar Inc. Available online: www.cdc.gov/fungal/outbreaks/rhizopus-investigation.html (accessed on 20 February 2015).
- Naqvi, S.S.B.; Nagendra, V.; Hofmeyr, A. Probiotic related Lactobacillus rhamnosus endocarditis in a patient with liver cirrhosis. IDCases 2018, 13, e00439. [Google Scholar] [CrossRef]
- Boumis, E.; Capone, A.; Galati, V.; Venditti, C.; Petrosillo, N. Probiotics and infective endocarditis in patients with hereditary hemorrhagic telangiectasia: A clinical case and a review of the literature. BMC Infect. Dis. 2018, 18, 65. [Google Scholar] [CrossRef]
- Jäsberg, H.; Tervahartiala, T.; Sorsa, T.; Söderling, E.; Haukioja, A. Probiotic intervention influences the salivary levels of Matrix Metalloproteinase (MMP)-9 and Tissue Inhibitor of metalloproteinases (TIMP)-1 in healthy adults. Arch. Oral Biol. 2018, 85, 58–63. [Google Scholar] [CrossRef]
- Falcinelli, S.; Rodiles, A.; Hatef, A.; Picchietti, S.; Cossignani, L.; Merrifield, D.L.; Unniappan, S.; Carnevali, O. Influence of Probiotics Administration on Gut Microbiota Core: A Review on the Effects on Appetite Control, Glucose, and Lipid Metabolism. J. Clin. Gastroenterol. 2018, 52 (Suppl. 1), S50–S56. [Google Scholar] [CrossRef]
- Dufour, A.; Hindré, T.; Haras, D.; Le Pennec, J.P. The biology of lantibiotics from the lacticin 481 group is coming of age. FEMS Microbiol. Rev. 2007, 31, 134–167. [Google Scholar] [CrossRef]
- Muthu Selvam, R.; Vinothini, G.; Palliyarai Thaiyammal, S.; Latha, S.; Chinnathambi, A.; Dhanasekaran, D.; Padmanabhan, P.; Alharbi, S.A.; Archunan, G. The cell aggregating propensity of probiotic actinobacterial isolates: Isolation and characterization of the aggregation inducing peptide pheromone. Biofouling 2016, 32, 71–79. [Google Scholar] [CrossRef]
- Kitazawa, H.; Ino, T.; Kawai, Y.; Itoh, T.; Saito, T. A novel immunostimulating aspect of Lactobacillus gasseri: Induction of “Gasserokine” as chemoattractants for macrophages. Int. J. Food Microbiol. 2002, 77, 29–38. [Google Scholar] [CrossRef]
- Nagy, G.; Pinczes, G.; Pinter, G.; Pocsi, I.; Prokisch, J.; Banfalvi, G. In Situ Electron Microscopy of Lactomicroselenium Particles in Probiotic Bacteria. Int. J. Mol. Sci. 2016, 17, 1047. [Google Scholar] [CrossRef]
- Washington, E.J.; Banfield, M.J.; Dangl, J.L. What a difference a Dalton makes: Bacterial virulence factors modulate eukaryotic host cell signaling systems via deamidation. Microbiol. Mol. Biol. Rev. 2013, 77, 527–539. [Google Scholar] [CrossRef]
- Rondanelli, M.; Faliva, M.A.; Perna, S.; Giacosa, A.; Peroni, G.; Castellazzi, A.M. Using probiotics in clinical practice: Where are we now? A review of existing meta-analyses. Gut Microbes 2017, 8, 521–543. [Google Scholar] [CrossRef]
- Hempel, S.; Newberry, S.; Ruelaz, A.; Wang, Z.; Miles, J.N.; Suttorp, M.J.; Johnsen, B.; Shanman, R.; Slusser, W.; Fu, N.; et al. Safety of probiotics used to reduce risk and prevent or treat disease. Evid. Rep. Technol. Assess. 2011, 200, 1–645. [Google Scholar]
- van den Nieuwboer, M.; Brummer, R.J.; Guarner, F.; Morelli, L.; Cabana, M.; Claassen, E. Safety of probiotics and synbiotics in children under 18 years of age. Benef. Microbes 2015, 6, 615–630. [Google Scholar] [CrossRef]
- Doherty, G.A.; Bennett, G.C.; Cheifetz, A.S.; Moss, A.C. Meta-analysis: Targeting the intestinal microbiota in prophylaxis for post-operative Crohn’s disease. Aliment. Pharmacol. Ther. 2010, 31, 802–809. [Google Scholar] [CrossRef]
- Tan-Lim, C.S.C.; Esteban-Ipac, N.A.R. Probiotics as treatment for food allergies among pediatric patients: A meta-analysis. World Allergy Organ. J. 2018, 11, 25. [Google Scholar] [CrossRef]
- Osborn, D.A.; Sinn, J.K. Probiotics in infants for prevention of allergic disease and food hypersensitivity. Cochrane Database Syst. Rev. 2007, 4, CD006475. [Google Scholar] [CrossRef]
- Reis, D.J.; Ilardi, S.S.; Punt, S.E.W. The anxiolytic effect of probiotics: A systematic review and meta-analysis of the clinical and preclinical literature. PLoS ONE 2018, 13, e0199041. [Google Scholar] [CrossRef]
- Borgeraas, H.; Johnson, L.K.; Skattebu, J.; Hertel, J.K.; Hjelmesaeth, J. Effects of probiotics on body weight, body mass index, fat mass and fat percentage in subjects with overweight or obesity: A systematic review and meta-analysis of randomized controlled trials. Obes. Rev. 2018, 19, 219–232. [Google Scholar] [CrossRef]
- Wardill, H.R.; Van Sebille, Y.Z.A.; Ciorba, M.A.; Bowen, J.M. Prophylactic probiotics for cancer therapy-induced diarrhoea: A meta-analysis. Curr. Opin. Support Palliat. Care 2018, 12, 187–197. [Google Scholar] [CrossRef]
- Canales, J.; Rada, G. Are probiotics effective in preventing urinary tract infection? Medwave 2018, 18, e7186. [Google Scholar] [CrossRef]
- Hassan, H.; Rompola, M.; Glaser, A.W.; Kinsey, S.E.; Phillips, R.S. Systematic review and meta-analysis investigating the efficacy and safety of probiotics in people with cancer. Support Care Cancer 2018, 26, 2503–2509. [Google Scholar] [CrossRef]
- Mohammed, A.T.; Khattab, M.; Ahmed, A.M.; Turk, T.; Sakr, N.; Khalil, A.M.; Abdelhalim, M.; Sawaf, B.; Hirayama, K.; Huy, N.T. The therapeutic effect of probiotics on rheumatoid arthritis: A systematic review and meta-analysis of randomized control trials. Clin. Rheumatol. 2017, 36, 2697–2707. [Google Scholar] [CrossRef]
- Liu, B.; He, Y.; Wang, M.; Liu, J.; Ju, Y.; Zhang, Y.; Liu, T.; Li, L.; Li, Q. Efficacy of probiotics on anxiety-A meta-analysis of randomized controlled trials. Depress Anxiety 2018, 35, 935–945. [Google Scholar] [CrossRef]
- Lin, J.; Zhang, Y.; He, C.; Dai, J. Probiotics supplementation in children with asthma: A systematic review and meta-analysis. J. Paediatr. Child Health 2018, 54, 953–961. [Google Scholar] [CrossRef]
- McFarland, L.V.; Goh, S. Are probiotics and prebiotics effective in the prevention of travellers’ diarrhea: A systematic review and meta-analysis. Travel Med. Infect Dis. 2018. [Google Scholar] [CrossRef]
- Jin, L.; Deng, L.; Wu, W.; Wang, Z.; Shao, W.; Liu, J. Systematic review and meta-analysis of the effect of probiotic supplementation on functional constipation in children. Medicine 2018, 97, e12174. [Google Scholar] [CrossRef]
- Rouhani, M.H.; Hadi, A.; Ghaedi, E.; Salehi, M.; Mahdavi, A.; Mohammadi, H. Do probiotics, prebiotics and synbiotics affect adiponectin and leptin in adults? A systematic review and meta-analysis of clinical trials. Clin. Nutr. 2018. [Google Scholar] [CrossRef]
- Upadhyay, R.P.; Taneja, S.; Chowdhury, R.; Strand, T.A.; Bhandari, N. Effect of prebiotic and probiotic supplementation on neurodevelopment in preterm very low birth weight infants: Findings from a meta-analysis. Pediatr. Res. 2018. [Google Scholar] [CrossRef]
- Dalal, R.; McGee, R.G.; Riordan, S.M.; Webster, A.C. Probiotics for people with hepatic encephalopathy. Cochrane Database Syst. Rev. 2017, 2, CD008716. [Google Scholar] [CrossRef]
- Goldenberg, J.Z.; Lytvyn, L.; Steurich, J.; Parkin, P.; Mahant, S.; Johnston, B.C. Probiotics for the prevention of pediatric antibiotic-associated diarrhea. Cochrane Database Syst. Rev. 2015, 12, CD004827. [Google Scholar] [CrossRef]
- Szajewska, H.; Horvath, A. Lactobacillus rhamnosus GG in the Primary Prevention of Eczema in Children: A Systematic Review and Meta-Analysis. Nutrients 2018, 10, 1319. [Google Scholar] [CrossRef]
- Mater, D.D.; Langella, P.; Corthier, G.; Flores, M.J. A probiotic Lactobacillus strain can acquire vancomycin resistance during digestive transit in mice. J. Mol. Microbiol. Biotechnol. 2008, 14, 123–127. [Google Scholar] [CrossRef]
- Jalali, M.; Abedi, D.; Varshosaz, J.; Najjarzadeh, M.; Mirlohi, M.; Tavakoli, N. Stability evaluation of freeze-dried Lactobacillus paracasei subsp. tolerance and Lactobacillus delbrueckii subsp. bulgaricus in oral capsules. Res Pharm Sci. 2012, 7, 31–36. [Google Scholar]
- Generally Recognized as Safe (GRAS). Available online: http://www.fda.gov/food/IngredientspackagingLabeling/GRAS/ (accessed on 7 February 2019).
- Investigational new drug applications (INDs)-determining whether human research studies can be conducted without an IND. Available online: www.fda.gov/downloads/Drugs/GuidanceComplianceRegulatoryInformation/UCM229175.pdf (accessed on 30 September 2013).
- Duc, L.H.; Hong, H.A.; Barbosa, T.M.; Henriques, A.O.; Cutting, S.M. Characterization of Bacillus probiotics available for human use. Appl. Environ. Microbiol. 2004, 70, 2161–2171. [Google Scholar] [CrossRef]
- Green, D.H.; Wakeley, P.R.; Page, A.; Barnes, A.; Baccigalupi, L.; Ricca, E.; Cutting, S.M. Characterization of two Bacillus probiotics. Appl. Environ. Microbiol. 1999, 65, 4288–4291. [Google Scholar]
- Temmerman, R.; Scheirlinck, I.; Huys, G.; Swings, J. Culture-independent analysis of probiotic products by denaturing gradient gel electrophoresis. Appl. Environ. Microbiol. 2003, 69, 220–226. [Google Scholar] [CrossRef]
- Masco, L.; Huys, G.; De Brandt, E.; Temmerman, R.; Swings, J. Culture-dependent and culture-independent qualitative analysis of probiotic products claimed to contain bifidobacteria. Int. J. Food Microbiol. 2005, 102, 221–230. [Google Scholar] [CrossRef]
- Ouwehand, A.C. A review of dose-responses of probiotics in human studies. Benef. Microbes 2017, 8, 143–151. [Google Scholar] [CrossRef]
- Hoffmann, D.E. Health claim regulation of probiotics in the USA and the EU: Is there a middle way? Benef. Microbes 2013, 4, 109–115. [Google Scholar] [CrossRef]
- Lee, T.T.; Morisset, M.; Astier, C.; Moneret-Vautrin, D.A.; Cordebar, V.; Beaudouin, E.; Codreanu, F.; Bihain, B.E.; Kanny, G. Contamination of probiotic preparations with milk allergens can cause anaphylaxis in children with cow’s milk allergy. J. Allergy Clin. Immunol. 2007, 119, 746–747. [Google Scholar]
- Martín-Muñoz, M.F.; Fortuni, M.; Caminoa, M.; Belver, T.; Quirce, S.; Caballero, T. Anaphylactic reaction to probiotics. Cow’s milk and hen’s egg allergens in probiotic compounds. Pediatr. Allergy Immunol. 2012, 23, 778–784. [Google Scholar] [CrossRef]
- Allen, E.N.; Chandler, C.I.; Mandimika, N.; Leisegang, C.; Barnes, K. Eliciting adverse effects data from participants in clinical trials. Cochrane Database Syst. Rev. 2018, 1, MR000039. [Google Scholar] [CrossRef]
- Jin, Y.; Sanger, N.; Shams, I.; Luo, C.; Shahid, H.; Li, G.; Bhatt, M.; Zielinski, L.; Bantoto, B.; Wang, M.; et al. Does the medical literature remain inadequately described despite having reporting guidelines for 21 years?—A systematic review of reviews: An update. J. Multidiscip. Healthc. 2018, 11, 495–510. [Google Scholar] [CrossRef]
- Bafeta, A.; Koh, M.; Riveros, C.; Ravaud, P. Harms Reporting in Randomized Controlled Trials of Interventions Aimed at Modifying Microbiota: A Systematic Review. Ann. Intern. Med. 2018, 169, 240–247. [Google Scholar] [CrossRef]
- van den Nieuwboer, M.; Claassen, E.; Morelli, L.; Guarner, F.; Brummer, R.J. Probiotic and synbiotic safety in infants under two years of age. Benef. Microbes 2014, 5, 45–60. [Google Scholar] [CrossRef]
- Mugambi, M.N.; Musekiwa, A.; Lombard, M.; Young, T.; Blaauw, R. Association between funding source, methodological quality and research outcomes in randomized controlled trials of synbiotics, probiotics and prebiotics added to infant formula: A systematic review. BMC Med. Res. Methodol. 2013, 13, 137. [Google Scholar] [CrossRef]
- Aceti, A.; Beghetti, I.; Maggio, L.; Martini, S.; Faldella, G.; Corvaglia, L. Filling the Gaps: Current Research Directions for a Rational Use of Probiotics in Preterm Infants. Nutrients 2018, 10, 1472. [Google Scholar] [CrossRef]
- Huff, B.A. Caveat emptor. “Probiotics” might not be what they seem. Can. Fam. Physician 2004, 50, 583–587. [Google Scholar]
- Sanders, M. How do we know when something called” probiotic” is really a probiotic? A guideline for consumers and healthcare professionals. Funct. Food Rev. 2009, 1, 3–12. [Google Scholar]
- Sanders, M.E.; Klaenhammer, T.R.; Ouwehand, A.C.; Pot, B.; Johansen, E.; Heimbach, J.T.; Marco, M.L.; Tennilä, J.; Ross, R.P.; Franz, C.; et al. Effects of genetic, processing, or product formulation changes on efficacy and safety of probiotics. Ann. N. Y. Acad. Sci. 2014, 1309, 1–18. [Google Scholar] [CrossRef]
- Katan, M.B. Why the European Food Safety Authority was right to reject health claims for probiotics. Benef. Microbes 2012, 3, 85–89. [Google Scholar] [CrossRef]
- Smug, L.N.; Salminen, S.; Sanders, M.E.; Ebner, S. Yoghurt and probiotic bacteria in dietary guidelines of the member states of the European Union. Benef. Microbes 2014, 5, 61–66. [Google Scholar] [CrossRef]
- Dronkers, T.M.G.; Krist, L.; Van Overveld, F.J.; Rijkers, G.T. The ascent of the blessed: Regulatory issues on health effects and health claims for probiotics in Europe and the rest of the world. Benef. Microbes 2018, 9, 717–723. [Google Scholar] [CrossRef]
- Available online: www.law.umaryland.edu/media/SOL/...Law/FinalWhitePaper.pdf (accessed on 15 November 2012).
- Ebner, S.; Smug, L.N.; Kneifel, W.; Salminen, S.J.; Sanders, M.E. Probiotics in dietary guidelines and clinical recommendations outside the European Union. World J. Gastroenterol. 2014, 20, 16095–16100. [Google Scholar] [CrossRef]
- Lee, E.S.; Song, E.J.; Nam, Y.D.; Lee, S.Y. Probiotics in human health and disease: From nutribiotics to pharmabiotics. J. Microbiol. 2018, 56, 773–782. [Google Scholar] [CrossRef]
- Azad, M.A.K.; Sarker, M.; Li, T.; Yin, J. Probiotic Species in the Modulation of Gut Microbiota: An Overview. Biomed. Res. Int. 2018, 2018, 9478630. [Google Scholar] [CrossRef]
| Medical Applications 1 | Medical Applications 2 | Functional Applications |
|---|---|---|
| Lactose intolerance [20,22,23] | Chronic renal failure [22,24] | Functional digestive complains [22] |
| Hyperlipidemia [18,22] | HIV infection [25] | Mood and behavior changes [26] |
| Nephrolithiasis (oxalate stones) [27] | Cirrhosis, liver encephalopathy, NAFLD [15,23] | Memory improvement [28] |
| Inflammatory bowel disease [22,23,24] | Organ transplant [23] | Anxiety, fatigue, weakness, body or localized pains, nausea [26,29] |
| Irritable bowel syndrome [13,22,23,30] | Metabolic diseases [12,22] | Constipation/loose stools changes [22] |
| Eczema, allergic rhinitis, asthma [12,22,24,30] | Constipation [22] | Day care health [24] |
| Infectious diarrhea [22,23,24,30] | Periodontitis [22] | Working places health [24] |
| Respiratory tract infections [12,22,24] | Depression [26] | Wellbeing [17,24] |
| Traveler’s diarrhea [22,23,24] | Stay in Intensive care unit [31] | Anti-stress [29] |
| Necrotizing enterocolitis [13,24] | Prematurity [32] | Increase longevity [33] |
| Pouchitis [34] | Infant colic [13,15] | Improve sexuality [35] |
| Helicobacter pylori [22,30] | Autoimmune diseases [13,22,23,24,30] | Impaired “intestinal integrity” [22,24] |
| Neurological disorders [21] | Cystic fibrosis, pancreatitis [23,30] | |
| Overweight and obesity [18,21] | Ethanol-induced liver disease [23] | |
| Various cancers [22,23,30] | Small bowel bacterial overgrowth [22] | |
| Along or after antibiotics therapy [22] | Enhancement of oral vaccine administration [30] | |
| Clostridium difficile induced colitis [22,23,30] | Ischemic heart disease [18,22] | |
| Respiratory/urinary tract, rotavirus infections [22,23,24] | Hypertension [36] | |
| Vaginosis [24,30] | Neuropsychiatric/degenerative diseases [37,38] | |
| Dental caries [22,23,24,30] | Enhance growth [22,24,39] | |
| Diabetes type 2 [23] | Enhance weight gain [22,24,39] |
| Infectious/Gastrointestinal | Allergic | Genetic | Patho-Toxogenicity |
|---|---|---|---|
| Bacteremia [41,74,150] | Rhinitis [149] | Transfer of virulent factors: | Enhanced adhesion and protein aggregation [74,149] |
| Sepsis [41,74] | Wheezing bronchitis [151] | Antibiotic resistance [74,149,152,153,154] | Mucolysis/hemolysis [74,149] |
| Fungemia [41,155] | Rash [149] | Hemolysin [149,152] | Bile salt hydrolysis [74] |
| Endocarditis, meningitis, endometritis, peritonitis, pneumonia [150,156,157] | Gelatinase [149] | DNA degradation and proteolysis [149] | |
| Liver abscess [150] | Metabolic | DNAse [149] | Innate defense resistance [52,149] |
| Diarrhea, Abdominal cramps [74] | d-lactic acidosis [41,74,149] | Enolase activating plasminogen [149] | Food poisoning [149] |
| Nausea, vomiting, flatulence, taste disturbance [41,74] | Metalloendopeptidase [158] | Immune evasion or over stimulation [74,149] | |
| Low appetite [159] | Cytolysin modification, transport, activation [160] | Facilitated microbial conjugation/translocation [74,149] | |
| Sex pheromones [161] | Macrophage/monocyte chemotactism [162] | ||
| Nanoparticles: Lactomicroselenium [163] | |||
| Gastrointestinal ischemia [41,74] | |||
| Mechanical choking [74] | |||
| Peptide deamidation [164] | |||
| Epigenetic and mobilome manipulation [52] |
| Publication | Mal-Designed | Lack of Standardization | High Data Variance | Biased | High Withdrawal | Incomplete Reporting | Reference |
|---|---|---|---|---|---|---|---|
| Review | + | + | + | + | + | [154] | |
| Systemic review | + | + | + | + | + | [148] | |
| Review | + | + | + | + | + | [74] | |
| Systemic review | + | + | + | + | [166] | ||
| Systemic review | + | + | + | + | + | [148] | |
| Systemic review | + | + | + | + | + | [167] | |
| Meta-analysis | + | + | [168] | ||||
| Meta-analysis | + | + | + | [169] | |||
| Meta-analysis | + | + | + | + | [170] | ||
| Meta-analysis | + | + | [171] | ||||
| Meta-analysis | + | + | + | + | [172] | ||
| Meta-analysis | + | + | + | + | [173] | ||
| Systemic review | + | + | [174] | ||||
| Meta-analysis | + | + | + | + | [175] | ||
| Meta-analysis | + | + | + | [62] | |||
| Meta-analysis | + | + | + | [176] | |||
| Meta-analysis | + | + | [177] | ||||
| Meta-analysis | + | + | + | [178] | |||
| Meta-analysis | + | + | + | [179] | |||
| Meta-analysis | + | + | [180] | ||||
| Meta-analysis | + | + | + | + | [181] | ||
| Meta-analysis | + | + | + | [182] | |||
| Systemic review | + | + | + | + | [183] | ||
| Meta-analysis | + | + | + | + | [184] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lerner, A.; Shoenfeld, Y.; Matthias, T. Probiotics: If It Does Not Help It Does Not Do Any Harm. Really? Microorganisms 2019, 7, 104. https://doi.org/10.3390/microorganisms7040104
Lerner A, Shoenfeld Y, Matthias T. Probiotics: If It Does Not Help It Does Not Do Any Harm. Really? Microorganisms. 2019; 7(4):104. https://doi.org/10.3390/microorganisms7040104
Chicago/Turabian StyleLerner, Aaron, Yehuda Shoenfeld, and Torsten Matthias. 2019. "Probiotics: If It Does Not Help It Does Not Do Any Harm. Really?" Microorganisms 7, no. 4: 104. https://doi.org/10.3390/microorganisms7040104
APA StyleLerner, A., Shoenfeld, Y., & Matthias, T. (2019). Probiotics: If It Does Not Help It Does Not Do Any Harm. Really? Microorganisms, 7(4), 104. https://doi.org/10.3390/microorganisms7040104