Rapid Bacterial Community Changes during Vermicomposting of Grape Marc Derived from Red Winemaking
Abstract
1. Introduction
2. Materials and Methods
2.1. Grape Marc and Earthworm Species
2.2. Vermicomposting Set-up and Sampling Design
2.3. Microbial Activity
2.4. DNA Sequencing and Bioinformatic Analyses
2.5. Statistical Analysis
3. Results
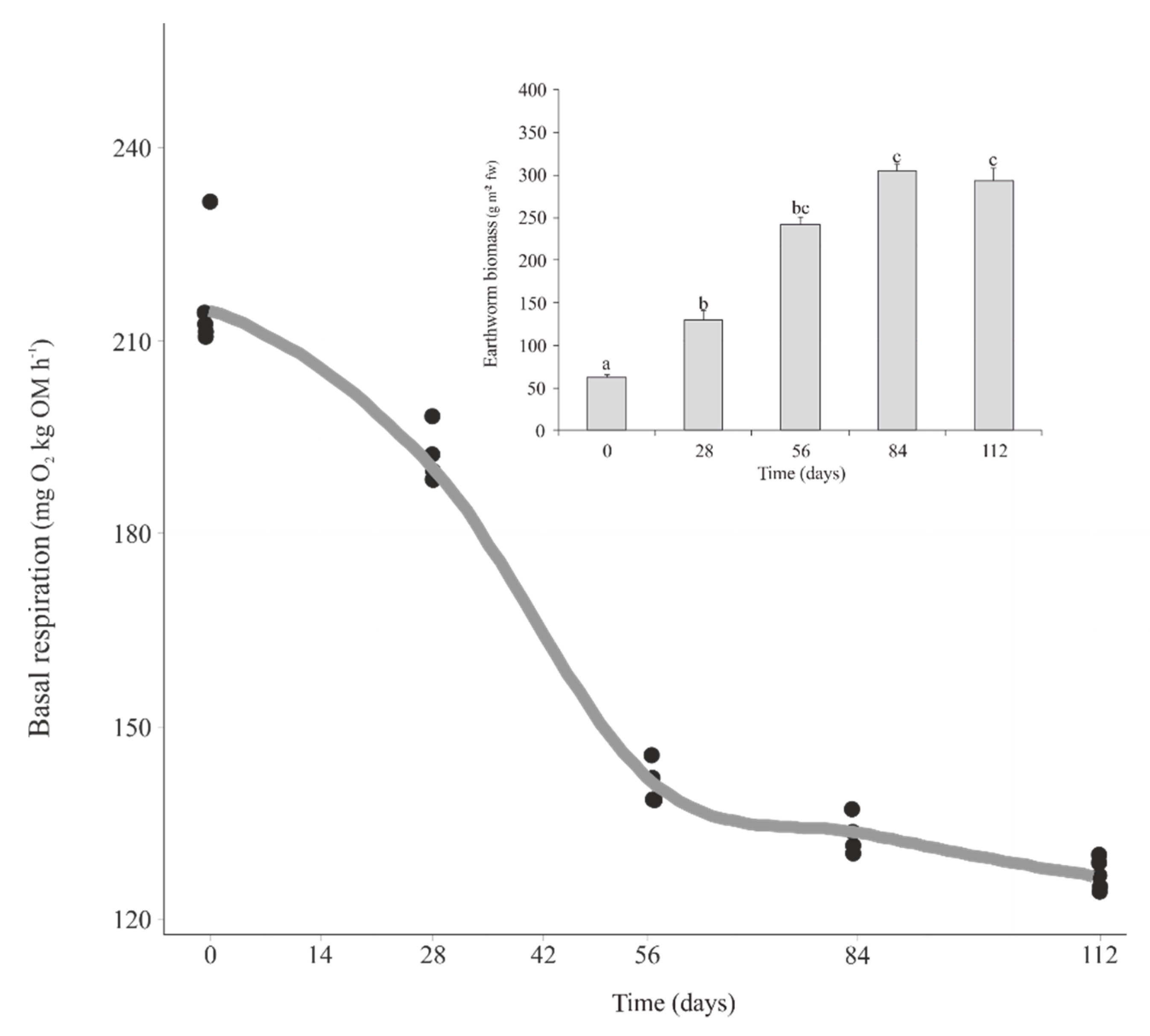
3.1. Earthworm Biomass and Microbial Activity during Vermicomposting of Mencía Grape Marc
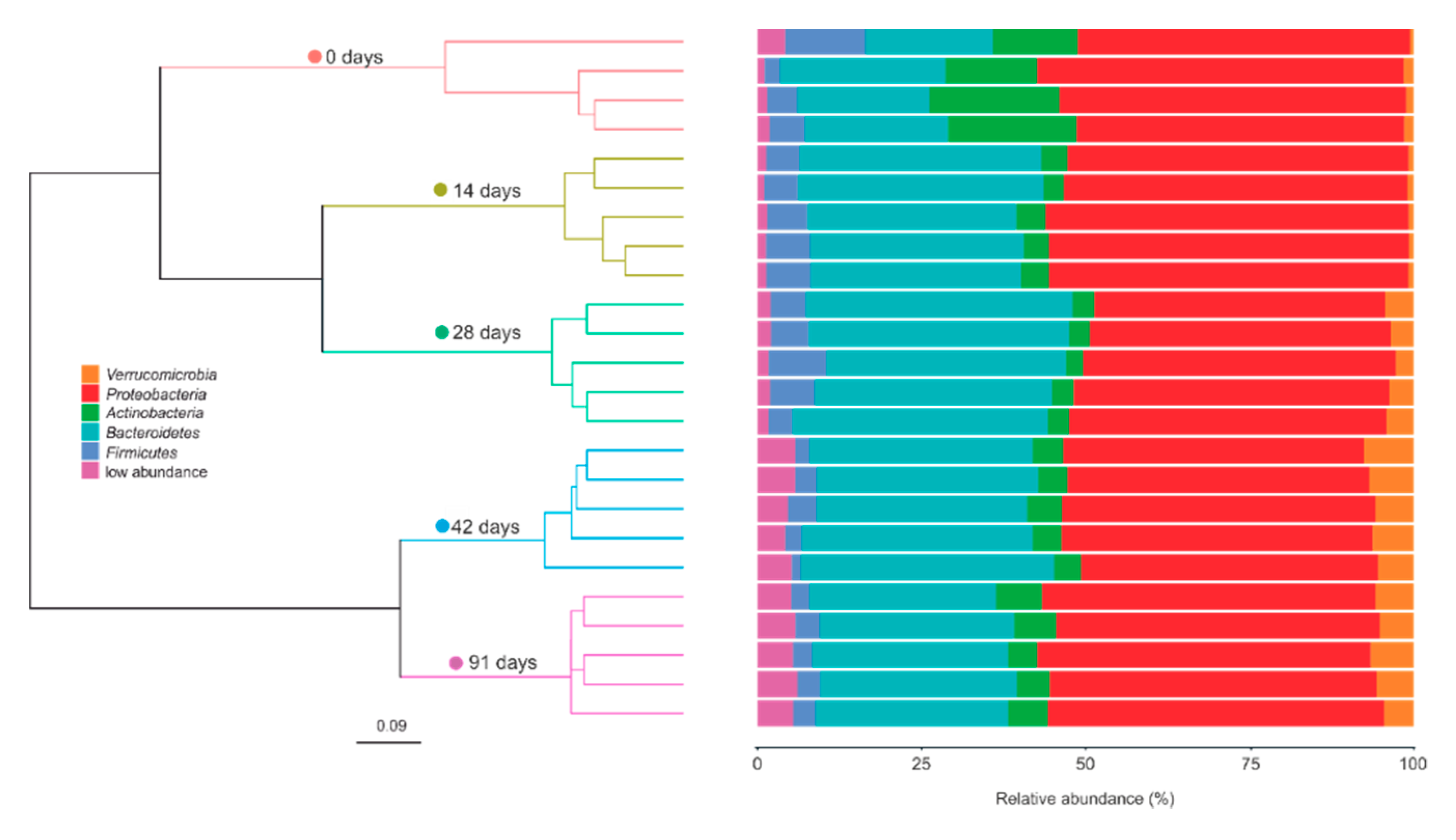
3.2. Changes in Bacterial Community Composition during Vermicomposting of Mencía Grape Marc
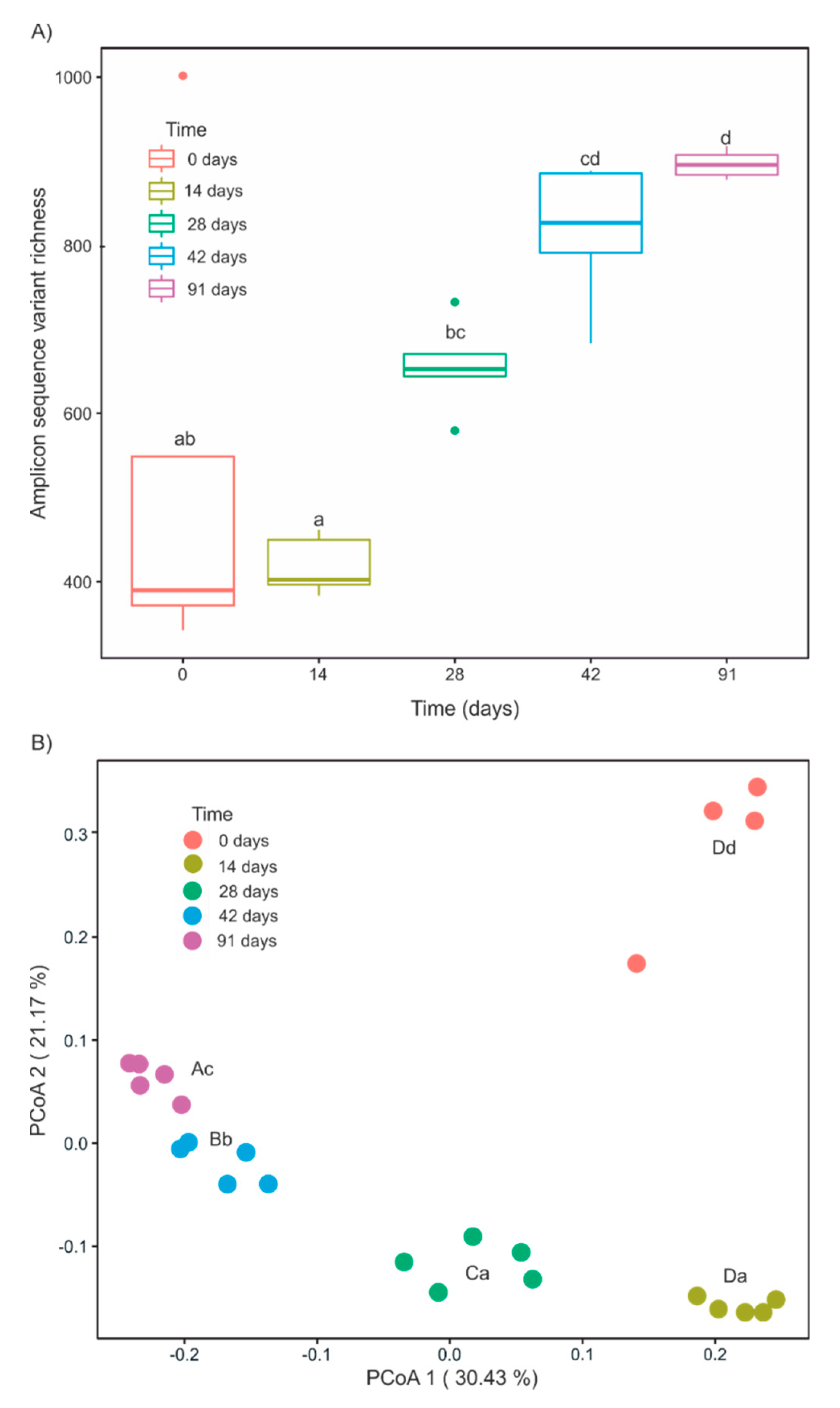
3.3. Changes in α- and β-Diversity during Vermicomposting of Mencía Grape Marc
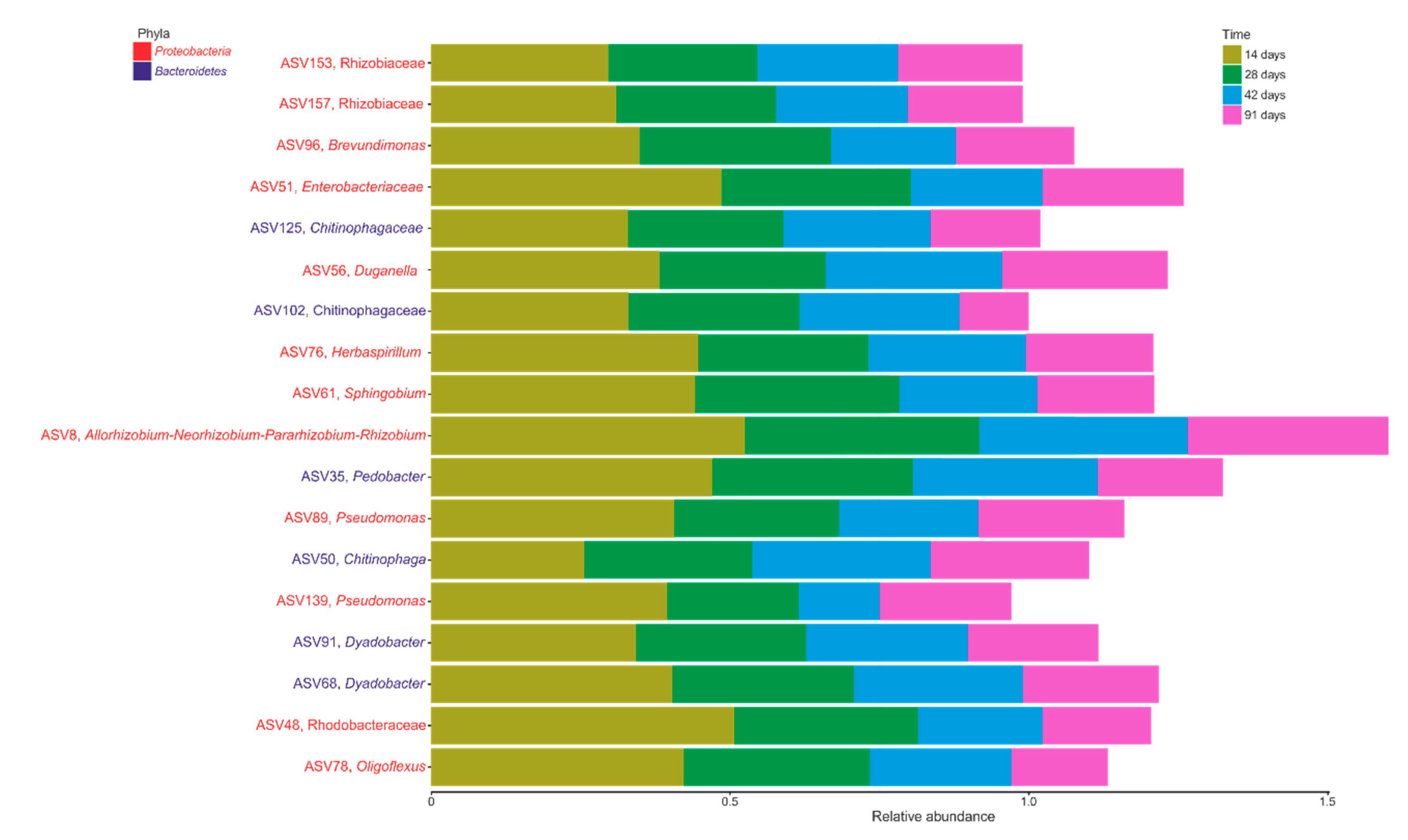
3.4. Core Microbiome during Vermicomposting of Mencía Grape Marc
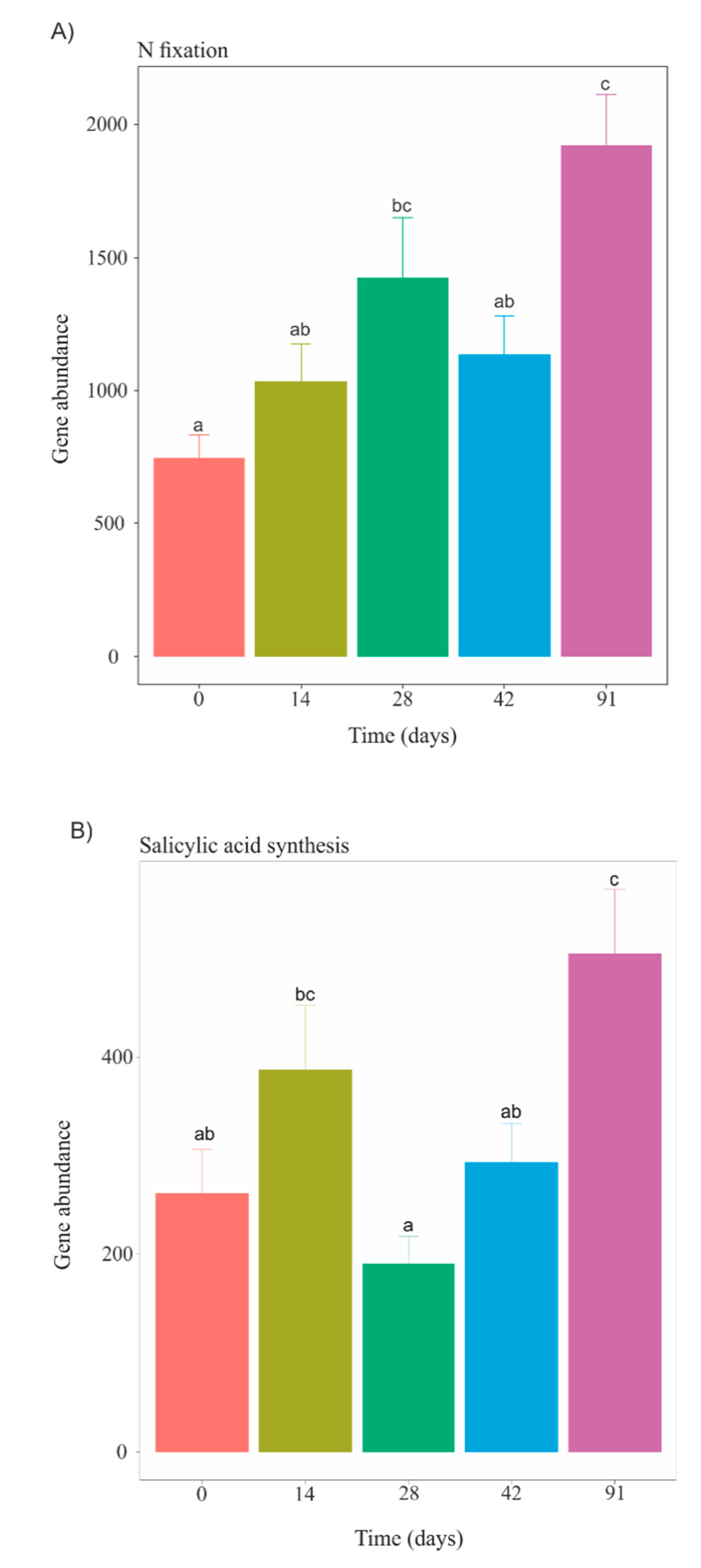
3.5. Functional Profiles during Vermicomposting of Mencía Grape Marc
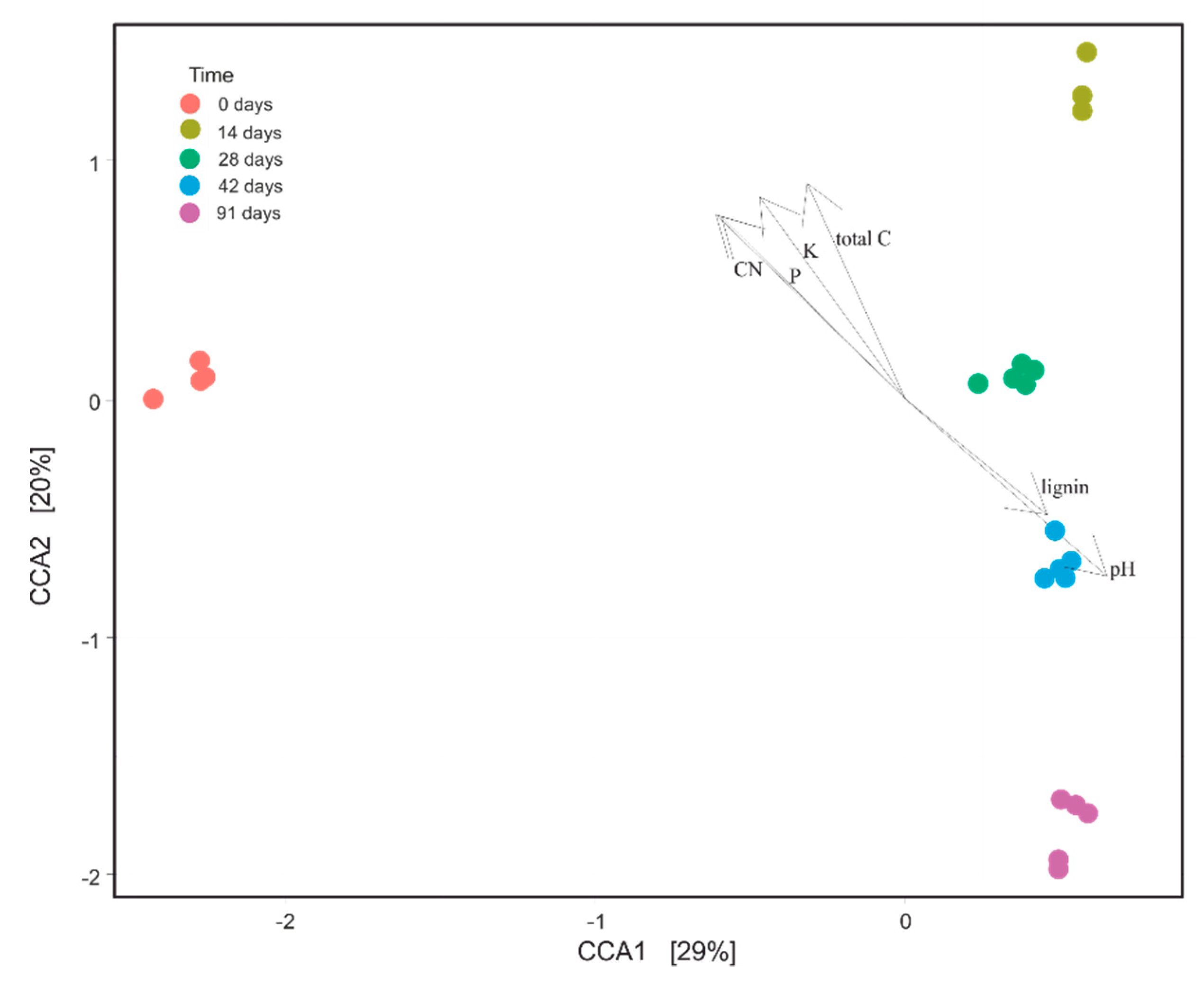
3.6. Influence of Physico-Chemical Variables on Bacterial Community Composition during Vermicomposting of Mencía Grape Marc
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Domínguez, J.; Aira, M.; Gómez-Brandón, M. Vermicomposting: Earthworms Enhance the Work of Microbes. In Microbes at Work; Insam, H., Franke-Whittle, I., Goberna, M., Eds.; Springer: Berlin/Heidelberg, Germany, 2010; pp. 93–114. [Google Scholar]
- Drake, H.L.; Horn, M.A. As the worm turns: The earthworm gut as a transient habitat for soil microbial biomes. Annu. Rev. Microbiol. 2007, 61, 169–189. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Brandón, M.; Aira, M.; Lores, M.; Dominguez, J. Epigeic earthworms exert a bottleneck effect on microbial communities through gut associated processes. PLoS ONE 2011, 6, e24786. [Google Scholar] [CrossRef] [PubMed]
- Aira, M.; Bybee, S.; Pérez-Losada, M.; Domínguez, J. Feeding on microbiomes: Effects of detritivory on the taxonomic and phylogenetic bacterial composition of animal manures. FEMS Microbiol. Ecol. 2015, 91, fiv117. [Google Scholar] [CrossRef] [PubMed]
- Aira, M.; Dominguez, J. Earthworm effects without earthworms: Inoculation of raw organic matter with worm-worked substrates alters microbial community functioning. PLoS ONE 2011, 6, e16354. [Google Scholar] [CrossRef] [PubMed]
- Aira, M.; Lazcano, C.; Gómez-Brandón, M.; Domínguez, J. Ageing effects of casts of Aporrectodea caliginosa on soil microbial community structure and activity. Appl. Soil Ecol. 2010, 46, 143–146. [Google Scholar] [CrossRef]
- Kearns, P.J.; Shade, A. Trait-based patterns of microbial dynamics in dormancy potential and heterotrophic strategy: Case studies of resource-based and post-press succession. ISME J. 2018, 12, 2575–2581. [Google Scholar] [CrossRef]
- Fierer, N.; Nemergut, D.; Knight, R.; Craine, J.M. Changes through time: Integrating microorganisms into the study of succession. Res. Microbiol. 2010, 161, 635–642. [Google Scholar] [CrossRef]
- Aira, M.; Pérez-Losada, M.; Domínguez, J. Microbiome dynamics during cast ageing in the earthworm Aporrectodea caliginosa. Appl. Soil Ecol. 2019, 139, 56–63. [Google Scholar] [CrossRef]
- Fernandez-Gomez, M.J.; Nogales, R.; Insam, H.; Romero, E.; Goberna, M. Use of DGGE and COMPOCHIP for investigating bacterial communities of various vermicomposts produced from different wastes under dissimilar conditions. Sci. Total Environ. 2012, 414, 664–671. [Google Scholar] [CrossRef]
- Fracchia, L.; Dohrmann, A.B.; Martinotti, M.G.; Tebbe, C.C. Bacterial diversity in a finished compost and vermicompost: Differences revealed by cultivation-independent analyses of PCR-amplified 16S rRNA genes. Appl. Microbiol. Biotechnol. 2006, 71, 942–952. [Google Scholar] [CrossRef]
- Gopal, M.; Bhute, S.S.; Gupta, A.; Prabhu, S.R.; Thomas, G.V.; Whitman, W.B.; Jangid, K. Changes in structure and function of bacterial communities during coconut leaf vermicomposting. Antonie Van Leeuwenhoek 2017, 110, 1339–1355. [Google Scholar] [CrossRef] [PubMed]
- Huang, K.; Li, F.; Wei, Y.; Chen, X.; Fu, X. Changes of bacterial and fungal community compositions during vermicomposting of vegetable wastes by Eisenia foetida. Bioresour. Technol. 2013, 150, 235–241. [Google Scholar] [CrossRef] [PubMed]
- Huang, K.; Xia, H.; Cui, G.; Li, F. Effects of earthworms on nitrification and ammonia oxidizers in vermicomposting systems for recycling of fruit and vegetable wastes. Sci. Total Environ. 2017, 578, 337–345. [Google Scholar] [CrossRef] [PubMed]
- Lv, B.; Xing, M.; Yang, J.; Zhang, L. Pyrosequencing reveals bacterial community differences in composting and vermicomposting on the stabilization of mixed sewage sludge and cattle dung. Appl. Microbiol. Biotechnol. 2015, 99, 10703–10712. [Google Scholar] [CrossRef]
- Vaz-Moreira, I.; Silva, M.E.; Manaia, C.M.; Nunes, O.C. Diversity of bacterial isolates from commercial and homemade composts. Microb. Ecol. 2008, 55, 714–722. [Google Scholar] [CrossRef]
- Vivas, A.; Moreno, B.; Garcia-Rodriguez, S.; Benitez, E. Assessing the impact of composting and vermicomposting on bacterial community size and structure, and microbial functional diversity of an olive-mill waste. Bioresour. Technol. 2009, 100, 1319–1326. [Google Scholar] [CrossRef]
- Gómez-Brandón, M.; Domínguez, J. Recycling of Solid Organic Wastes through Vermicomposting: Microbial Community Changes Throughout the Process and Use of Vermicompost as a Soil Amendment. Crit. Rev. Environ. Sci. Technol. 2014, 44, 1289–1312. [Google Scholar] [CrossRef]
- Hussain, M.; Cholette, S.; Castaldi, R.M. An Analysis of Globalization Forces in the Wine Industry. J. Glob. Market. 2008, 21, 33–47. [Google Scholar] [CrossRef]
- Domínguez, J.; Martínez-Cordeiro, H.; Alvarez-Casas, M.; Lores, M. Vermicomposting grape marc yields high quality organic biofertiliser and bioactive polyphenols. Waste Manag. Res. 2014, 32, 1235–1240. [Google Scholar] [CrossRef]
- Domínguez, J.; Martínez-Cordeiro, H.; Lores, M. Earthworms and Grape Marc: Simultaneous Production of a High-Quality Biofertilizer and Bioactive-Rich Seeds. In Grape and Wine Biotechnology; Morata, A., Loira, I., Eds.; Intech Open Science: Rijeka, Croatia, 2016; pp. 167–183. [Google Scholar]
- Domínguez, J.; Sánchez-Hernández, J.C.; Lores, M. Vermicomposting of Winemaking By-Products. In Handbook of Grape Processing By-Products; Galanakis, C.M., Ed.; Academic Press, Elsevier: London, UK, 2017; pp. 55–78. [Google Scholar]
- Gómez-Brandón, M.; Lores, M.; Insam, H.; Domínguez, J. Strategies for recycling and valorization of grape marc. Crit. Rev. Biotechnol. 2019, 39, 437–450. [Google Scholar] [CrossRef]
- Gómez-Brandón, M.; Lores, M.; Martinez-Cordeiro, H.; Dominguez, J. Effectiveness of vermicomposting for bioconversion of grape marc derived from red winemaking into a value-added product. Environ. Sci. Pollut. Res. 2019. [Google Scholar] [CrossRef]
- Částková, T.; Hanč, A. Change of parameters of layers in a large-scale grape marc vermicomposting system with continuous feeding. Waste Manag. Res. 2019, 37, 826–832. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Brandón, M.; Lazcano, C.; Lores, M.; Domínguez, J. Short-term stabilization of grape marc through earthworms. J. Hazard. Mater. 2011, 187, 291–295. [Google Scholar] [CrossRef] [PubMed]
- Kolbe, A.R.; Aira, M.; Gómez-Brandón, M.; Pérez-Losada, M.; Domínguez, J. Bacterial succession and functional diversity during vermicomposting of white grape marc (Vitis vinifera v. Albariño). Sci. Rep. 2019, 9, 7472. [Google Scholar] [CrossRef] [PubMed]
- Domínguez, J.; Edwards, C.A. Biology and Ecology of Earthworm Species Used for Vermicomposting. In Vermiculture Technology: Earthworms, Organic Waste and Environmental Management; Edwards, C.A., Arancon, N.Q., Sherman, R.L., Eds.; CRC Press: Boca Raton, FL, USA, 2011; pp. 25–37. [Google Scholar]
- Domínguez, J.; Aira, M.; Kolbe, A.R.; Gómez-Brandón, M.; Pérez-Losada, M. Changes in the composition and function of bacterial communities during vermicomposting may explain beneficial properties of vermicompost. Sci. Rep. 2019, 9, 9657. [Google Scholar] [CrossRef] [PubMed]
- International Organization for Standardization. ISO 16072: Soil Quality Laboratory Methods for Determination of Microbial Soil Respiration; International Organization for Standardization: Geneva, Switzerland, 2002. [Google Scholar]
- Kozich, J.J.; Westcott, S.L.; Baxter, N.T.; Highlander, S.K.; Schloss, P.D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 2013, 79, 5112–5120. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glockner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef]
- Langille, M.G.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Vega Thurber, R.L.; Knight, R.; et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef]
- Kanehisa, M.; Sato, Y.; Furumichi, M.; Morishima, K.; Tanabe, M. New approach for understanding genome variations in KEGG. Nucleic Acids Res. 2019, 47, 590–595. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- McMurdie, P.J.; Holmes, S. Phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef] [PubMed]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2—Approximately maximum-likelihood trees for large alignments. PLoS ONE 2010, 5, e9490. [Google Scholar] [CrossRef]
- Faith, D.P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 1992, 61, 1–10. [Google Scholar] [CrossRef]
- Lozupone, C.; Knight, R. UniFrac: A new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 2005, 71, 8228–8235. [Google Scholar] [CrossRef]
- Pinheiro, J.; Bates, D.; DebRoy, S.; Sarkar, D.; R Core Team. Nlme: Linear and Nonlinear Mixed Effects Models, In vol. R package version 3.1-120; The R Foundation: Vienna, Austria, 2015. [Google Scholar]
- Hothorn, T.; Bretz, F.; Westfall, P. Simultaneous inference in general parametric models. BIOM J. 2008, 50, 346–363. [Google Scholar] [CrossRef]
- Oksanen, J.; Blanchet, F.G.; Kindt, R.; Legendre, P.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; Stevens, M.H.H.; Wagner, H. Vegan: Community Ecology Package. R Packag Version 24–1. Available online: https://CRANR-project.org/package=vegan (2016) (accessed on 11 June 2019).
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2014. [Google Scholar]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016. [Google Scholar]
- Ali, U.; Sajid, N.; Khalid, A.; Riaz, L.; Rabbani, M.M.; Syed, J.H.; Malik, R.N. A review on vermicomposting of organic wastes. Environ. Prog. Sustain. Energy 2015, 34, 1050–1062. [Google Scholar] [CrossRef]
- Luo, G.; Li, L.; Friman, V.-P.; Guo, J.; Guo, S.; Shen, Q.; Ling, N. Organic amendments increase crop yields by improving microbe-mediated soil functioning of agroecosystems: A meta-analysis. Soil Biol. Biochem. 2018, 124, 105–115. [Google Scholar] [CrossRef]
- Ho, A.; Di Lonardo, D.P.; Bodelier, P.L. Revisiting life strategy concepts in environmental microbial ecology. FEMS Microbiol. Ecol. 2017, 93. [Google Scholar] [CrossRef]
- Sauvadet, M.; Fanin, N.; Chauvat, M.; Bertrand, I. Can the comparison of above- and below-ground litter decomposition improve our understanding of bacterial and fungal successions? Soil Biol. Biochem. 2019, 132, 24–27. [Google Scholar] [CrossRef]
- Shade, A.; Handelsman, J. Beyond the Venn diagram: The hunt for a core microbiome. Environ. Microbiol. 2012, 14, 4–12. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, T.L.; Balsemao-Pires, E.; Saraiva, R.M.; Ferreira, P.C.; Hemerly, A.S. Nitrogen signalling in plant interactions with associative and endophytic diazotrophic bacteria. J. Exp. Bot. 2014, 65, 5631–5642. [Google Scholar] [CrossRef] [PubMed]
- Danon, M.; Franke-Whittle, I.H.; Insam, H.; Chen, Y.; Hadar, Y. Molecular analysis of bacterial community succession during prolonged compost curing. FEMS Microbiol. Ecol. 2008, 65, 133–144. [Google Scholar] [CrossRef]
- Haichar, F.Z.; Marol, C.; Berge, O.; Rangel-Castro, J.I.; Prosser, J.I.; Balesdent, J.; Heulin, T.; Achouak, W. Plant host habitat and root exudates shape soil bacterial community structure. ISME J. 2008, 2, 1221–1230. [Google Scholar] [CrossRef]
- Marques, A.P.G.C.; Pires, C.; Moreira, H.; Rangel, A.; Castro, P.M.L. Assessment of the plant growth promotion abilities of six bacterial isolates using Zea mays as indicator plant. Soil Biol. Biochem. 2010, 42, 1229–1235. [Google Scholar] [CrossRef]
- Bedini, A.; Mercy, L.; Schneider, C.; Franken, P.; Lucic-Mercy, E. Unraveling the Initial Plant Hormone Signaling, Metabolic Mechanisms and Plant Defense Triggering the Endomycorrhizal Symbiosis Behavior. Front. Plant Sci. 2018, 9, 1800. [Google Scholar] [CrossRef] [PubMed]
- Lazcano, C.; Domínguez, J. The use of vermicompost in sustainable agriculture: Impact on plant growth and soil fertility. In Soil Nutrients; Miransari, M., Ed.; Nova Science Publishers: New York, NY, USA, 2011; pp. 230–254. [Google Scholar]
- Song, X.; Liu, M.; Wu, D.; Griffiths, B.S.; Jiao, J.; Li, H.; Hu, F. Interaction matters: Synergy between vermicompost and PGPR agents improves soil quality, crop quality and crop yield in the field. Appl. Soil Ecol. 2015, 89, 25–34. [Google Scholar] [CrossRef]
- Blouin, M.; Barrere, J.; Meyer, J.; Lartigue, S.; Barot, S.; Mathieu, J. Vermicompost significantly affects plant growth. A meta-analysis. Agron. Sustain. Dev. 2019, 39, 34. [Google Scholar] [CrossRef]
| Alpha Diversity | F4,19 | p(>F) | |
|---|---|---|---|
| Observed | 10.45 | 0.0003 | |
| Chao1 | 9.11 | <0.0001 | |
| Shannon | 39.50 | <0.0001 | |
| Faith PD | 13.34 | 0.0001 | |
| Beta diversity | F4,19 | p-value | |
| Unifrac–unweighted | PCoA1 | 203.58 | <0.0001 |
| PCoA2 | 114.67 | <0.0001 | |
| Unifrac-weighted | PCoA1 | 175.00 | <0.0001 |
| PCoA2 | 71.91 | <0.0001 | |
| Bray-Curtis | PCoA1 | 158.46 | <0.0001 |
| PCoA2 | 339.88 | <0.0001 | |
| Jaccard | PCoA1 | 254.34 | <0.0001 |
| PCoA2 | 146.48 | <0.0001 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gómez Brandón, M.; Aira, M.; Kolbe, A.R.; de Andrade, N.; Pérez-Losada, M.; Domínguez, J. Rapid Bacterial Community Changes during Vermicomposting of Grape Marc Derived from Red Winemaking. Microorganisms 2019, 7, 473. https://doi.org/10.3390/microorganisms7100473
Gómez Brandón M, Aira M, Kolbe AR, de Andrade N, Pérez-Losada M, Domínguez J. Rapid Bacterial Community Changes during Vermicomposting of Grape Marc Derived from Red Winemaking. Microorganisms. 2019; 7(10):473. https://doi.org/10.3390/microorganisms7100473
Chicago/Turabian StyleGómez Brandón, María, Manuel Aira, Allison R. Kolbe, Nariane de Andrade, Marcos Pérez-Losada, and Jorge Domínguez. 2019. "Rapid Bacterial Community Changes during Vermicomposting of Grape Marc Derived from Red Winemaking" Microorganisms 7, no. 10: 473. https://doi.org/10.3390/microorganisms7100473
APA StyleGómez Brandón, M., Aira, M., Kolbe, A. R., de Andrade, N., Pérez-Losada, M., & Domínguez, J. (2019). Rapid Bacterial Community Changes during Vermicomposting of Grape Marc Derived from Red Winemaking. Microorganisms, 7(10), 473. https://doi.org/10.3390/microorganisms7100473





