Pathogenetic Impact of Bacterial–Fungal Interactions
Abstract
1. Introduction
2. Bacterial–Fungal Interactions in the Context of the Microbiome
2.1. Impact of Microbiome Dysbiosis on the Bacterial–Fungal Equilibrium
2.2. Microbial Metabolites—the Good and the Bad
3. Communication between Bacteria and Fungi Mediated by Proteins and Small Molecules
3.1. Bacterial–Fungal Interactions
3.1.1. Candida Species and Different Bacteria
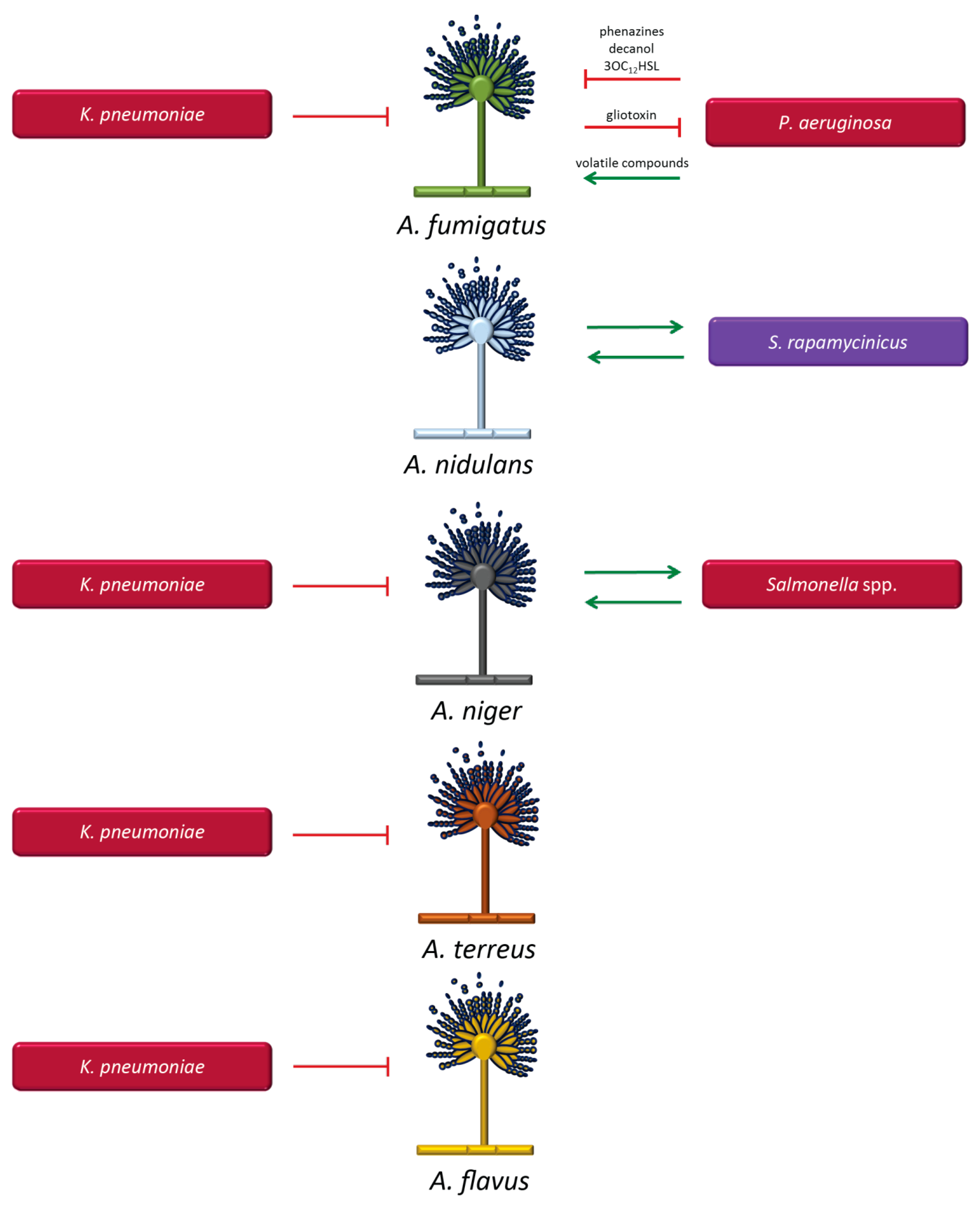
3.1.2. Aspergillus Species and Bacteria
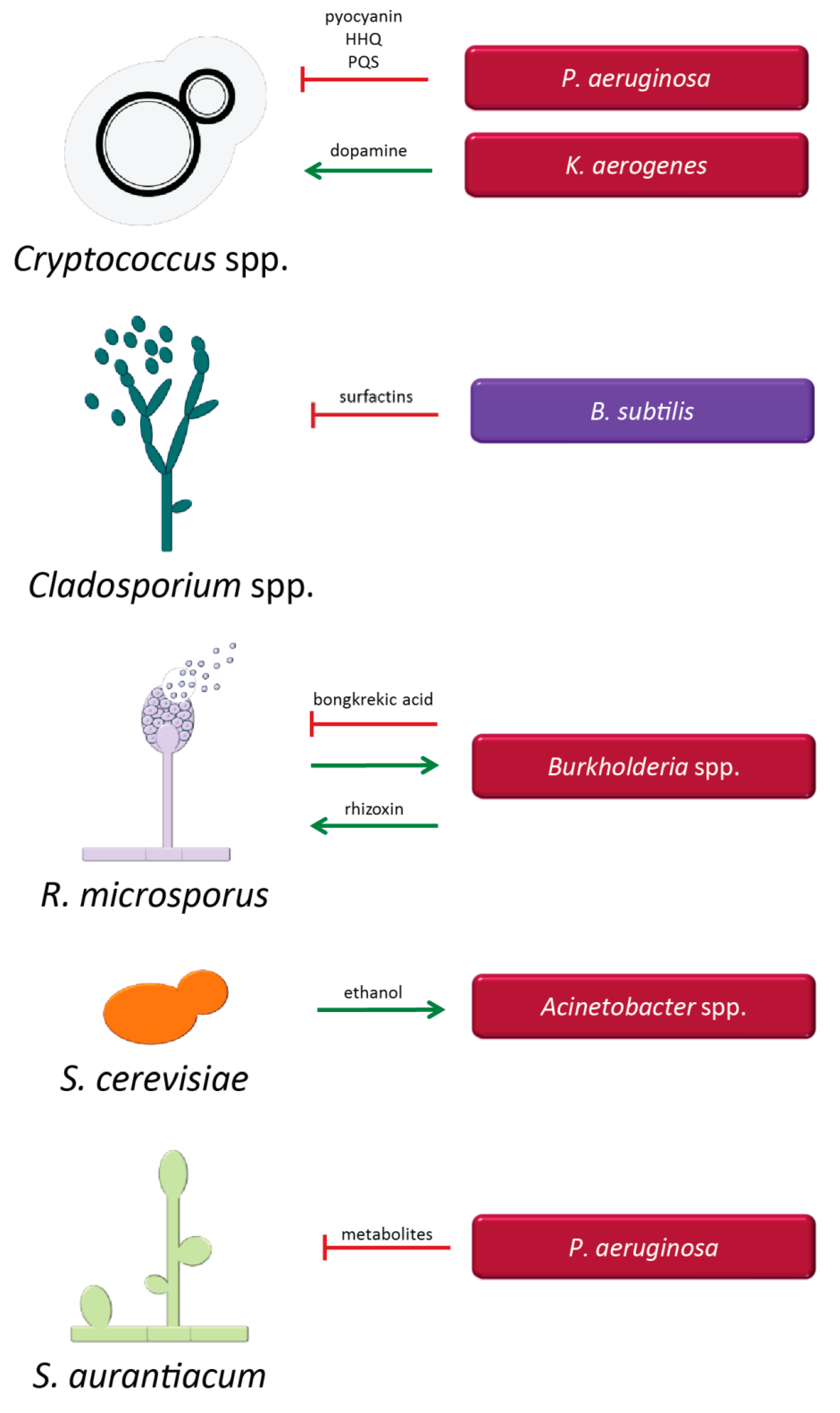
3.1.3. Cryptococcus Species and Bacteria
3.1.4. Interaction of Other Fungal Species with Bacteria
4. Host Immune Response to Bacterial and Fungal (Co-)Infections
5. In Vivo Models of Bacterial and Fungal (Co)-Infections
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Neely, A.N.; Law, E.J.; Holder, I.A. Increased susceptibility to lethal Candida infections in burned mice preinfected with Pseudomonas aeruginosa or pretreated with proteolytic enzymes. Infect. Immun. 1986, 52, 200–204. [Google Scholar] [PubMed]
- Bergeron, A.C.; Seman, B.G.; Hammond, J.H.; Archambault, L.S.; Hogan, D.A.; Wheeler, R.T. Candida and Pseudomonas interact to enhance virulence of mucosal infection in transparent zebrafish. Infect. Immun. 2017. [Google Scholar] [CrossRef]
- Kim, S.H.; Yoon, Y.K.; Kim, M.J.; Sohn, J.W. Risk factors for and clinical implications of mixed Candida/bacterial bloodstream infections. Clin. Microbiol. Infect. 2013, 19, 62–68. [Google Scholar] [CrossRef] [PubMed]
- Wargo, M.J.; Hogan, D.A. Fungal--bacterial interactions: a mixed bag of mingling microbes. Curr. Opin. Microbiol. 2006, 9, 359–364. [Google Scholar] [CrossRef] [PubMed]
- Peters, B.M.; Jabra-Rizk, M.A.; O’May, G.A.; Costerton, J.W.; Shirtliff, M.E. Polymicrobial interactions: impact on pathogenesis and human disease. Clin. Microbiol. Rev. 2012, 25, 193–213. [Google Scholar] [CrossRef] [PubMed]
- Diaz, P.I.; Strausbaugh, L.D.; Dongari-Bagtzoglou, A. Fungal-bacterial interactions and their relevance to oral health: linking the clinic and the bench. Front. Cell. Infect. Microbiol. 2014, 4, 101. [Google Scholar] [CrossRef] [PubMed]
- Hermann, C.; Hermann, J.; Munzel, U.; Ruchel, R. Bacterial flora accompanying Candida yeasts in clinical specimens. Mycoses 1999, 42, 619–627. [Google Scholar] [CrossRef]
- Morales, D.K.; Hogan, D.A. Candida albicans interactions with bacteria in the context of human health and disease. Plos Pathog. 2010, 6, 1000886. [Google Scholar] [CrossRef]
- Kett, D.H.; Azoulay, E.; Echeverria, P.M.; Vincent, J.L. Candida bloodstream infections in intensive care units: analysis of the extended prevalence of infection in intensive care unit study. Crit. Care. Med. 2011, 39, 665–670. [Google Scholar] [CrossRef]
- Thorn, J.L.; Gilchrist, K.B.; Sobonya, R.E.; Gaur, N.K.; Lipke, P.N.; Klotz, S.A. Postmortem Candidemia: Marker of disseminated disease. J. Clin. Pathol. 2010, 63, 337–340. [Google Scholar] [CrossRef]
- De Sordi, L.; Muhlschlegel, F.A. Quorum sensing and fungal-bacterial interactions in Candida albicans: A communicative network regulating microbial coexistence and virulence. Fems Yeast Res. 2009, 9, 990–999. [Google Scholar] [CrossRef] [PubMed]
- Dixon, E.F.; Hall, R.A. Noisy neighbourhoods: quorum sensing in fungal-polymicrobial infections. Cell Microbiol. 2015, 17, 1431–1441. [Google Scholar] [CrossRef] [PubMed]
- Braga, R.M.; Dourado, M.N.; Araujo, W.L. Microbial interactions: ecology in a molecular perspective. Braz. J. Microbiol. 2016, 47, 86–98. [Google Scholar] [CrossRef] [PubMed]
- Fischbach, M.A. Microbiome: Focus on Causation and Mechanism. Cell 2018, 174, 785–790. [Google Scholar] [CrossRef] [PubMed]
- Knight, R.; Vrbanac, A.; Taylor, B.C.; Aksenov, A.; Callewaert, C.; Debelius, J.; Gonzalez, A.; Kosciolek, T.; McCall, L.I.; McDonald, D.; et al. Best practices for analysing microbiomes. Nat. Rev. Microbiol. 2018, 16, 410–422. [Google Scholar] [CrossRef] [PubMed]
- Garrett, W.S. Cancer and the microbiota. Science 2015, 348, 80–86. [Google Scholar] [CrossRef] [PubMed]
- Kruger, W.; Vielreicher, S.; Kapitan, M.; Jacobsen, I.D.; Niemiec, M.J. Fungal-Bacterial Interactions in Health and Disease. Pathogens 2019, 8, 70. [Google Scholar] [CrossRef] [PubMed]
- Marsland, B.J.; Gollwitzer, E.S. Host-microorganism interactions in lung diseases. Nat. Rev. Immunol. 2014, 14, 827–835. [Google Scholar] [CrossRef]
- Nash, A.K.; Auchtung, T.A.; Wong, M.C.; Smith, D.P.; Gesell, J.R.; Ross, M.C.; Stewart, C.J.; Metcalf, G.A.; Muzny, D.M.; Gibbs, R.A.; et al. The gut mycobiome of the Human Microbiome Project healthy cohort. Microbiome 2017, 5, 153. [Google Scholar] [CrossRef]
- Witherden, E.A.; Shoaie, S.; Hall, R.A.; Moyes, D.L. The Human Mucosal Mycobiome and Fungal Community Interactions. J. Fungi (Basel) 2017, 3, 56. [Google Scholar] [CrossRef]
- Jo, J.H.; Kennedy, E.A.; Kong, H.H. Topographical and physiological differences of the skin mycobiome in health and disease. Virulence 2017, 8, 324–333. [Google Scholar] [CrossRef] [PubMed]
- Statovci, D.; Aguilera, M.; MacSharry, J.; Melgar, S. The Impact of Western Diet and Nutrients on the Microbiota and Immune Response at Mucosal Interfaces. Front. Immunol. 2017, 8, 838. [Google Scholar] [CrossRef] [PubMed]
- Clarke, G.; Stilling, R.M.; Kennedy, P.J.; Stanton, C.; Cryan, J.F.; Dinan, T.G. Minireview: Gut microbiota: the neglected endocrine organ. Mol. Endocrinol. 2014, 28, 1221–1238. [Google Scholar] [CrossRef]
- Belkaid, Y.; Hand, T.W. Role of the microbiota in immunity and inflammation. Cell 2014, 157, 121–141. [Google Scholar] [CrossRef]
- Caballero, S.; Pamer, E.G. Microbiota-mediated inflammation and antimicrobial defense in the intestine. Annu. Rev. Immunol. 2015, 33, 227–256. [Google Scholar] [CrossRef]
- Selber-Hnatiw, S.; Rukundo, B.; Ahmadi, M.; Akoubi, H.; Al-Bizri, H.; Aliu, A.F.; Ambeaghen, T.U.; Avetisyan, L.; Bahar, I.; Baird, A. Human Gut Microbiota: Toward an Ecology of Disease. Front. Microbiol. 2017, 8, 1265. [Google Scholar] [CrossRef]
- Tojo, R.; Suarez, A.; Clemente, M.G.; de los Reyes-Gavilan, C.G.; Margolles, A.; Gueimonde, M.; Ruas-Madiedo, P. Intestinal microbiota in health and disease: role of bifidobacteria in gut homeostasis. World J. Gastroenterol. 2014, 20, 15163–15176. [Google Scholar] [CrossRef]
- Candela, M.; Turroni, S.; Biagi, E.; Carbonero, F.; Rampelli, S.; Fiorentini, C.; Brigidi, P. Inflammation and colorectal cancer, when microbiota-host mutualism breaks. World J. Gastroenterol. 2014, 20, 908–922. [Google Scholar] [CrossRef]
- Galloway-Pena, J.; Brumlow, C.; Shelburne, S. Impact of the Microbiota on Bacterial Infections during Cancer Treatment. Trends Microbiol. 2017, 25, 992–1004. [Google Scholar] [CrossRef]
- Heisel, T.; Montassier, E.; Johnson, A.; Al-Ghalith, G.; Lin, Y.W.; Wei, L.N.; Knights, D.; Gale, C.A. High-Fat Diet Changes Fungal Microbiomes and Interkingdom Relationships in the Murine Gut. mSphere 2017, 2. [Google Scholar] [CrossRef]
- Lof, M.; Janus, M.M.; Krom, B.P. Metabolic Interactions between Bacteria and Fungi in Commensal Oral Biofilms. J. Fungi (Basel) 2017, 3, 40. [Google Scholar] [CrossRef]
- Raskov, H.; Burcharth, J.; Pommergaard, H.C. Linking Gut Microbiota to Colorectal Cancer. J. Cancer 2017, 8, 3378–3395. [Google Scholar] [CrossRef] [PubMed]
- Tsilimigras, M.C.; Fodor, A.; Jobin, C. Carcinogenesis and therapeutics: the microbiota perspective. Nat. Microbiol. 2017, 2, 17008. [Google Scholar] [CrossRef] [PubMed]
- Zitvogel, L.; Daillere, R.; Roberti, M.P.; Routy, B.; Kroemer, G. Anticancer effects of the microbiome and its products. Nat. Rev. Microbiol. 2017, 15, 465–478. [Google Scholar] [CrossRef]
- Schwabe, R.F.; Jobin, C. The microbiome and cancer. Nat. Rev. Cancer 2013, 13, 800–812. [Google Scholar] [CrossRef]
- Liguori, G.; Lamas, B.; Richard, M.L.; Brandi, G.; da Costa, G.; Hoffmann, T.W.; Di Simone, M.P.; Calabrese, C.; Poggioli, G.; Langella, P.; et al. Fungal Dysbiosis in Mucosa-associated Microbiota of Crohn’s Disease Patients. J. Crohns. Colitis. 2016, 10, 296–305. [Google Scholar] [CrossRef]
- Pope, J.L.; Tomkovich, S.; Yang, Y.; Jobin, C. Microbiota as a mediator of cancer progression and therapy. Transl. Res. 2017, 179, 139–154. [Google Scholar] [CrossRef]
- Gagniere, J.; Raisch, J.; Veziant, J.; Barnich, N.; Bonnet, R.; Buc, E.; Bringer, M.A.; Pezet, D.; Bonnet, M. Gut microbiota imbalance and colorectal cancer. World J. Gastroenterol. 2016, 22, 501–518. [Google Scholar] [CrossRef]
- DeGruttola, A.K.; Low, D.; Mizoguchi, A.; Mizoguchi, E. Current Understanding of Dysbiosis in Disease in Human and Animal Models. Inflamm. Bowel Dis. 2016, 22, 1137–1150. [Google Scholar] [CrossRef]
- Williams, S.C. Gnotobiotics. Proc. Natl. Acad. Sci. USA 2014, 111, 1661. [Google Scholar] [CrossRef]
- Gordon, H.A.; Pesti, L. The gnotobiotic animal as a tool in the study of host microbial relationships. Bacteriol. Rev. 1971, 35, 390–429. [Google Scholar] [PubMed]
- Turnbaugh, P.J.; Ridaura, V.K.; Faith, J.J.; Rey, F.E.; Knight, R.; Gordon, J.I. The effect of diet on the human gut microbiome: A metagenomic analysis in humanized gnotobiotic mice. Sci. Transl. Med. 2009, 1, 6ra14. [Google Scholar] [CrossRef] [PubMed]
- Gill, S.R.; Pop, M.; Deboy, R.T.; Eckburg, P.B.; Turnbaugh, P.J.; Samuel, B.S.; Gordon, J.I.; Relman, D.A.; Fraser-Liggett, C.M.; Nelson, K.E. Metagenomic analysis of the human distal gut microbiome. Science 2006, 312, 1355–1359. [Google Scholar] [CrossRef] [PubMed]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef]
- Backhed, F.; Ding, H.; Wang, T.; Hooper, L.V.; Koh, G.Y.; Nagy, A.; Semenkovich, C.F.; Gordon, J.I. The gut microbiota as an environmental factor that regulates fat storage. Proc. Natl. Acad. Sci. USA 2004, 101, 15718–15723. [Google Scholar] [CrossRef]
- Cash, H.L.; Whitham, C.V.; Behrendt, C.L.; Hooper, L.V. Symbiotic bacteria direct expression of an intestinal bactericidal lectin. Science 2006, 313, 1126–1130. [Google Scholar] [CrossRef]
- Umesaki, Y. Use of gnotobiotic mice to identify and characterize key microbes responsible for the development of the intestinal immune system. Proc. Jpn. Acad. Ser. B. 2014, 90, 313–332. [Google Scholar] [CrossRef]
- Burmolle, M.; Ren, D.; Bjarnsholt, T.; Sorensen, S.J. Interactions in multispecies biofilms: do they actually matter? Trends Microbiol. 2014, 22, 84–91. [Google Scholar] [CrossRef]
- de Vos, W.M. Microbial biofilms and the human intestinal microbiome. Npj Biofilms Microbiomes 2015, 1, 15005. [Google Scholar] [CrossRef]
- Raskov, H.; Kragh, K.N.; Bjarnsholt, T.; Alamili, M.; Gogenur, I. Bacterial biofilm formation inside colonic crypts may accelerate colorectal carcinogenesis. Clin. Transl. Med. 2018, 7, 018–0209. [Google Scholar] [CrossRef]
- Leclair, L.W.; Hogan, D.A. Mixed bacterial-fungal infections in the, C.F. respiratory tract. Med. Mycol. 2010, 48, 521522. [Google Scholar] [CrossRef][Green Version]
- Chotirmall, S.H.; O’Donoghue, E.; Bennett, K.; Gunaratnam, C.; O’Neill, S.J.; McElvaney, N.G. Sputum Candida albicans presages, F.E.V(1) decline and hospital-treated exacerbations in cystic fibrosis. Chest 2010, 138, 1186–1195. [Google Scholar] [CrossRef] [PubMed]
- Reece, E.; Segurado, R.; Jackson, A.; McClean, S.; Renwick, J.; Greally, P. Co-colonisation with Aspergillus fumigatus and Pseudomonas aeruginosa is associated with poorer health in cystic fibrosis patients: an Irish registry analysis. BMC Pulm. Med. 2017, 17, 70. [Google Scholar] [CrossRef] [PubMed]
- Fujimura, K.E.; Sitarik, A.R.; Havstad, S.; Lin, D.L.; Levan, S.; Fadrosh, D.; Panzer, A.R.; LaMere, B.; Rackaityte, E.; Lukacs, N.W.; et al. Neonatal gut microbiota associates with childhood multisensitized atopy and T cell differentiation. Nat. Med. 2016, 22, 1187–1191. [Google Scholar] [CrossRef] [PubMed]
- Arrieta, M.C.; Stiemsma, L.T.; Dimitriu, P.A.; Thorson, L.; Russell, S.; Yurist-Doutsch, S.; Kuzeljevic, B.; Gold, M.J.; Britton, H.M.; Lefebvre, D.L.; et al. Early infancy microbial and metabolic alterations affect risk of childhood asthma. Sci. Transl. Med. 2015, 7, 307ra152. [Google Scholar] [CrossRef]
- Sobel, J.D. Vulvovaginal candidosis. Lancet 2007, 369, 1961–1971. [Google Scholar] [CrossRef]
- van de Wijgert, J.H.; Borgdorff, H.; Verhelst, R.; Crucitti, T.; Francis, S.; Verstraelen, H.; Jespers, V. The vaginal microbiota: what have we learned after a decade of molecular characterization? PLoS ONE 2014, 9, e105998. [Google Scholar] [CrossRef]
- Hall, R.A.; Noverr, M.C. Fungal interactions with the human host: exploring the spectrum of symbiosis. Curr. Opin. Microbiol. 2017, 40, 58–64. [Google Scholar] [CrossRef]
- Burd, R.S.; Raymond, C.S.; Dunn, D.L. Endotoxin promotes synergistic lethality during concurrent Escherichia coli and Candida albicans infection. J. Surg. Res. 1992, 52, 537–542. [Google Scholar] [CrossRef]
- Ikeda, T.; Suegara, N.; Abe, S.; Yamaguchi, H. Efficacy of antibacterial drugs in mice with complex infection by Candida albicans and Escherichia coli. J. Antibiot. 1999, 52, 552–558. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Standaert-Vitse, A.; Sendid, B.; Joossens, M.; Francois, N.; Vandewalle-El Khoury, P.; Branche, J.; Van Kruiningen, H.; Jouault, T.; Rutgeerts, P.; Gower-Rousseau, C.; et al. Candida albicans colonization and, A.S.CA in familial Crohn’s disease. Am. J. Gastroenterol. 2009, 104, 1745–1753. [Google Scholar] [CrossRef] [PubMed]
- Chiaro, T.R.; Soto, R.; Zac Stephens, W.; Kubinak, J.L.; Petersen, C.; Gogokhia, L.; Bell, R.; Delgado, J.C.; Cox, J.; Voth, W.; et al. A member of the gut mycobiota modulates host purine metabolism exacerbating colitis in mice. Sci. Transl. Med. 2017, 9. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Zhang, F.; Yang, X.; Wu, N.; Jiang, W.; Li, X.; Liu, Y. Changes in the composition of intestinal fungi and their role in mice with dextran sulfate sodium-induced colitis. Sci. Rep. 2015, 5, 10416. [Google Scholar] [CrossRef] [PubMed]
- Noverr, M.C.; Noggle, R.M.; Toews, G.B.; Huffnagle, G.B. Role of antibiotics and fungal microbiota in driving pulmonary allergic responses. Infect. Immun. 2004, 72, 4996–5003. [Google Scholar] [CrossRef]
- Wheeler, M.L.; Limon, J.J.; Bar, A.S.; Leal, C.A.; Gargus, M.; Tang, J.; Brown, J.; Funari, V.A.; Wang, H.L.; Crother, T.R.; et al. Immunological Consequences of Intestinal Fungal Dysbiosis. Cell Host Microbe. 2016, 19, 865–873. [Google Scholar] [CrossRef]
- Sommer, F.; Backhed, F. The gut microbiota--masters of host development and physiology. Nat. Rev. Microbiol. 2013, 11, 227–238. [Google Scholar] [CrossRef]
- Singh, N.; Gurav, A.; Sivaprakasam, S.; Brady, E.; Padia, R.; Shi, H.; Thangaraju, M.; Prasad, P.D.; Manicassamy, S.; Munn, D.H.; et al. Activation of Gpr109a, receptor for niacin and the commensal metabolite butyrate, suppresses colonic inflammation and carcinogenesis. Immunity 2014, 40, 128–139. [Google Scholar] [CrossRef]
- Louis, P.; Flint, H.J. Diversity, metabolism and microbial ecology of butyrate-producing bacteria from the human large intestine. FEMS Microbiol. Lett. 2009, 294, 1–8. [Google Scholar] [CrossRef]
- Li, J.; Butcher, J.; Mack, D.; Stintzi, A. Functional impacts of the intestinal microbiome in the pathogenesis of inflammatory bowel disease. Inflamm. Bowel Dis. 2015, 21, 139–153. [Google Scholar] [CrossRef]
- Sam, Q.H.; Chang, M.W.; Chai, L.Y. The Fungal Mycobiome and Its Interaction with Gut Bacteria in the Host. Int. J. Mol. Sci. 2017, 18, 330. [Google Scholar] [CrossRef]
- Nguyen, G.C. Editorial: bugs and drugs: insights into the pathogenesis of inflammatory bowel disease. Am. J. Gastroenterol. 2011, 106, 2143–2145. [Google Scholar] [CrossRef] [PubMed]
- Schulz, M.D.; Atay, C.; Heringer, J.; Romrig, F.K.; Schwitalla, S.; Aydin, B.; Ziegler, P.K.; Varga, J.; Reindl, W.; Pommerenke, C.; et al. High-fat-diet-mediated dysbiosis promotes intestinal carcinogenesis independently of obesity. Nature 2014, 514, 508–512. [Google Scholar] [CrossRef] [PubMed]
- Belcheva, A.; Irrazabal, T.; Robertson, S.J.; Streutker, C.; Maughan, H.; Rubino, S.; Moriyama, E.H.; Copeland, J.K.; Surendra, A.; Kumar, S.; et al. Gut microbial metabolism drives transformation of, M.S.H2-deficient colon epithelial cells. Cell 2014, 158, 288–299. [Google Scholar] [CrossRef] [PubMed]
- Lupton, J.R. Microbial degradation products influence colon cancer risk: the butyrate controversy. J. Nutr. 2004, 134, 479–482. [Google Scholar] [CrossRef] [PubMed]
- Palmer, K.L.; Aye, L.M.; Whiteley, M. Nutritional cues control Pseudomonas aeruginosa multicellular behavior in cystic fibrosis sputum. J. Bacteriol. 2007, 189, 8079–8087. [Google Scholar] [CrossRef] [PubMed]
- Gainza-Cirauqui, M.L.; Nieminen, M.T.; Novak Frazer, L.; Aguirre-Urizar, J.M.; Moragues, M.D.; Rautemaa, R. Production of carcinogenic acetaldehyde by Candida albicans from patients with potentially malignant oral mucosal disorders. J. Oral Pathol. Med. 2013, 42, 243–249. [Google Scholar] [CrossRef]
- Meurman, J.H.; Uittamo, J. Oral micro-organisms in the etiology of cancer. Acta. Odontol. Scand. 2008, 66, 321–326. [Google Scholar] [CrossRef]
- Whiteley, M.; Diggle, S.P.; Greenberg, E.P. Progress in and promise of bacterial quorum sensing research. Nature 2017, 551, 313–320. [Google Scholar] [CrossRef]
- Rio, R.V.M. Don’t Bite the Hand that Feeds You. Cell Host Microbe. 2017, 21, 552–554. [Google Scholar] [CrossRef]
- Polke, M.; Jacobsen, I.D. Quorum sensing by farnesol revisited. Curr. Genet. 2017, 63, 791–797. [Google Scholar] [CrossRef]
- Papenfort, K.; Bassler, B.L. Quorum sensing signal-response systems in Gram-negative bacteria. Nat. Rev. Microbiol. 2016, 14, 576–588. [Google Scholar] [CrossRef] [PubMed]
- O’Toole, G.A. Classic Spotlight: Quorum Sensing and the Multicellular Life of Unicellular Organisms. J. Bacteriol. 2016, 198, 601. [Google Scholar] [CrossRef] [PubMed]
- Hofer, U. Biofilms: Turning tides for quorum sensing. Nat. Rev. Microbiol. 2016, 14, 64. [Google Scholar] [CrossRef] [PubMed]
- Pammi, M.; Liang, R.; Hicks, J.; Mistretta, T.A.; Versalovic, J. Biofilm extracellular, D.N.A enhances mixed species biofilms of Staphylococcus epidermidis and Candida albicans. BMC Microbiol. 2013, 13, 1471–2180. [Google Scholar] [CrossRef]
- Smith, K.; Rajendran, R.; Kerr, S.; Lappin, D.F.; Mackay, W.G.; Williams, C.; Ramage, G. Aspergillus fumigatus enhances elastase production in Pseudomonas aeruginosa co-cultures. Med. Mycol. 2015, 53, 645–655. [Google Scholar] [CrossRef]
- Mear, J.B.; Kipnis, E.; Faure, E.; Dessein, R.; Schurtz, G.; Faure, K.; Guery, B. Candida albicans and Pseudomonas aeruginosa interactions: more than an opportunistic criminal association? Med. Mal. Infect. 2013, 43, 146–151. [Google Scholar] [CrossRef]
- Tan, C.H.; Koh, K.S.; Xie, C.; Zhang, J.; Tan, X.H.; Lee, G.P.; Zhou, Y.; Ng, W.J.; Rice, S.A.; Kjelleberg, S. Community quorum sensing signalling and quenching: microbial granular biofilm assembly. Npj Biofilms Microbiomes 2015, 1, 15006. [Google Scholar] [CrossRef]
- Dong, Y.H.; Xu, J.L.; Li, X.Z.; Zhang, L.H. AiiA, an enzyme that inactivates the acylhomoserine lactone quorum-sensing signal and attenuates the virulence of Erwinia carotovora. Proc. Natl. Acad. Sci. USA 2000, 97, 3526–3531. [Google Scholar] [CrossRef]
- Defoirdt, T. Quorum-Sensing Systems as Targets for Antivirulence Therapy. Trends Microbiol. 2017. [Google Scholar] [CrossRef]
- Abraham, W.R. Going beyond the Control of Quorum-Sensing to Combat Biofilm Infections. Antibiotics (Basel) 2016, 5, 3. [Google Scholar] [CrossRef]
- Thompson, J.A.; Oliveira, R.A.; Djukovic, A.; Ubeda, C.; Xavier, K.B. Manipulation of the quorum sensing signal, A.I.-2 affects the antibiotic-treated gut microbiota. Cell Rep. 2015, 10, 1861–1871. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Contreras, R. Is Quorum Sensing Interference a Viable Alternative to Treat Pseudomonas aeruginosa Infections? Front. Microbiol. 2016, 7, 1454. [Google Scholar] [CrossRef] [PubMed]
- Guo, Q.; Wei, Y.; Xia, B.; Jin, Y.; Liu, C.; Pan, X.; Shi, J.; Zhu, F.; Li, J.; Qian, L.; et al. Identification of a small molecule that simultaneously suppresses virulence and antibiotic resistance of Pseudomonas aeruginosa. Sci. Rep. 2016, 6, 19141. [Google Scholar] [CrossRef] [PubMed]
- Muimhneacháin, E.Ó.; Reen, F.J.; O’Gara, F.; McGlacken, G.P. Analogues of Pseudomonas aeruginosa signalling molecules to tackle infections. Org. Biomol. Chem. 2018, 16, 169–179. [Google Scholar] [CrossRef]
- Smith, A.C.; Rice, A.; Sutton, B.; Gabrilska, R.; Wessel, A.K.; Whiteley, M.; Rumbaugh, K.P. Albumin Inhibits Pseudomonas aeruginosa Quorum Sensing and Alters Polymicrobial Interactions. Infect. Immun. 2017, 85, 116. [Google Scholar] [CrossRef]
- Imperi, F.; Leoni, L.; Visca, P. Antivirulence activity of azithromycin in Pseudomonas aeruginosa. Front. Microbiol. 2014, 5, 178. [Google Scholar] [CrossRef]
- Ramage, G.; Saville, S.P.; Wickes, B.L.; Lopez-Ribot, J.L. Inhibition of Candida albicans Biofilm Formation by Farnesol, a Quorum-Sensing Molecule. Appl. Environ. Microbiol. 2002, 68, 5459–5463. [Google Scholar] [CrossRef]
- Cordeiro, R.A.; Teixeira, C.E.; Brilhante, R.S.; Castelo-Branco, D.S.; Paiva, M.A.; Giffoni Leite, J.J.; Lima, D.T.; Monteiro, A.J.; Sidrim, J.J.; Rocha, M.F. Minimum inhibitory concentrations of amphotericin B, azoles and caspofungin against Candida species are reduced by farnesol. Med. Mycol. 2013, 51, 53–59. [Google Scholar] [CrossRef]
- Mear, J.B.; Gosset, P.; Kipnis, E.; Faure, E.; Dessein, R.; Jawhara, S.; Fradin, C.; Faure, K.; Poulain, D.; Sendid, B.; et al. Candida albicans airway exposure primes the lung innate immune response against Pseudomonas aeruginosa infection through innate lymphoid cell recruitment and interleukin-22-associated mucosal response. Infect. Immun. 2014, 82, 306–315. [Google Scholar] [CrossRef]
- Peleg, A.Y.; Hogan, D.A.; Mylonakis, E. Medically important bacterial-fungal interactions. Nat. Rev. Microbiol. 2010, 8, 340–349. [Google Scholar] [CrossRef]
- Gibson, J.; Sood, A.; Hogan, D.A. Pseudomonas aeruginosa-Candida albicans interactions: localization and fungal toxicity of a phenazine derivative. Appl. Env. Microbiol. 2009, 75, 504–513. [Google Scholar] [CrossRef] [PubMed]
- Hogan, D.A.; Vik, A.; Kolter, R. A Pseudomonas aeruginosa quorum-sensing molecule influences Candida albicans morphology. Mol. Microbiol. 2004, 54, 1212–1223. [Google Scholar] [CrossRef] [PubMed]
- Hogan, D.A. Talking to themselves: autoregulation and quorum sensing in fungi. Eukaryot. Cell. 2006, 5, 613–619. [Google Scholar] [CrossRef] [PubMed]
- Purschke, F.G.; Hiller, E.; Trick, I.; Rupp, S. Flexible survival strategies of Pseudomonas aeruginosa in biofilms result in increased fitness compared with Candida albicans. Mol. Cell Proteomics. 2012, 11, 1652–1669. [Google Scholar] [CrossRef] [PubMed]
- Kerr, J.R. Suppression of fungal growth exhibited by Pseudomonas aeruginosa. J. Clin. Microbiol. 1994, 32, 525–527. [Google Scholar]
- Kerr, J.R.; Taylor, G.W.; Rutman, A.; Hoiby, N.; Cole, P.J.; Wilson, R. Pseudomonas aeruginosa pyocyanin and 1-hydroxyphenazine inhibit fungal growth. J. Clin. Pathol. 1999, 52, 385–387. [Google Scholar] [CrossRef]
- Charlton, T.S.; de Nys, R.; Netting, A.; Kumar, N.; Hentzer, M.; Givskov, M.; Kjelleberg, S. A novel and sensitive method for the quantification of N-3-oxoacyl homoserine lactones using gas chromatography-mass spectrometry: application to a model bacterial biofilm. Env. Microbiol. 2000, 2, 530–541. [Google Scholar] [CrossRef]
- Hogan, D.A.; Kolter, R. Pseudomonas-Candida interactions: an ecological role for virulence factors. Science 2002, 296, 2229–2232. [Google Scholar] [CrossRef]
- Cugini, C.; Calfee, M.W.; Farrow, J.M., 3rd; Morales, D.K.; Pesci, E.C.; Hogan, D.A. Farnesol, a common sesquiterpene, inhibits, P.Q.S production in Pseudomonas aeruginosa. Mol. Microbiol. 2007, 65, 896–906. [Google Scholar] [CrossRef]
- Chen, A.I.; Dolben, E.F.; Okegbe, C.; Harty, C.E.; Golub, Y.; Thao, S.; Ha, D.G.; Willger, S.D.; O’Toole, G.A.; Harwood, C.S.; et al. Candida albicans ethanol stimulates Pseudomonas aeruginosa WspR-controlled biofilm formation as part of a cyclic relationship involving phenazines. PloS Pathog. 2014, 10, e1004480. [Google Scholar] [CrossRef]
- Fourie, R.; Pohl, C.H. Beyond Antagonism: The Interaction Between Candida Species and Pseudomonas aeruginosa. J. Fungi (Basel) 2019, 5, 34. [Google Scholar] [CrossRef] [PubMed]
- Cavalcanti, I.M.; Del Bel Cury, A.A.; Jenkinson, H.F.; Nobbs, A.H. Interactions between Streptococcus oralis, Actinomyces oris, and Candida albicans in the development of multispecies oral microbial biofilms on salivary pellicle. Mol. Oral Microbiol. 2017, 32, 60–73. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Sobue, T.; Thompson, A.; Xie, Z.; Poon, K.; Ricker, A.; Cervantes, J.; Diaz, P.I.; Dongari-Bagtzoglou, A. Streptococcal co-infection augments Candida pathogenicity by amplifying the mucosal inflammatory response. Cell Microbiol. 2014, 16, 214–231. [Google Scholar] [CrossRef]
- Holmes, A.R.; Gopal, P.K.; Jenkinson, H.F. Adherence of Candida albicans to a cell surface polysaccharide receptor on Streptococcus gordonii. Infect. Immun. 1995, 63, 1827–1834. [Google Scholar]
- Holmes, A.R.; McNab, R.; Jenkinson, H.F. Candida albicans binding to the oral bacterium Streptococcus gordonii involves multiple adhesin-receptor interactions. Infect. Immun. 1996, 64, 4680–4685. [Google Scholar]
- Nobbs, A.H.; Vickerman, M.M.; Jenkinson, H.F. Heterologous expression of Candida albicans cell wall-associated adhesins in Saccharomyces cerevisiae Reveals differential specificities in adherence and biofilm formation and in binding oral Streptococcus gordonii. Eukaryot. Cell 2010, 9, 1622–1634. [Google Scholar] [CrossRef]
- Jenkinson, H.F.; Lala, H.C.; Shepherd, M.G. Coaggregation of Streptococcus sanguis and other streptococci with Candida albicans. Infect. Immun. 1990, 58, 1429–1436. [Google Scholar]
- O’Sullivan, J.M.; Jenkinson, H.F.; Cannon, R.D. Adhesion of Candida albicans to oral streptococci is promoted by selective adsorption of salivary proteins to the streptococcal cell surface. Microbiology 2000, 146, 41–48. [Google Scholar] [CrossRef]
- Kim, D.; Sengupta, A.; Niepa, T.H.; Lee, B.H.; Weljie, A.; Freitas-Blanco, V.S.; Murata, R.M.; Stebe, K.J.; Lee, D.; Koo, H. Candida albicans stimulates Streptococcus mutans microcolony development via cross-kingdom biofilm-derived metabolites. Sci. Rep. 2017, 7, 41332. [Google Scholar] [CrossRef]
- Montelongo-Jauregui, D.; Saville, S.P.; Lopez-Ribot, J.L. Contributions of Candida albicans Dimorphism, Adhesive Interactions, and Extracellular Matrix to the Formation of Dual-Species Biofilms with Streptococcus gordonii. Mbio 2019, 10, e01179-19. [Google Scholar] [CrossRef]
- Holmes, A.R.; van der Wielen, P.; Cannon, R.D.; Ruske, D.; Dawes, P. Candida albicans binds to saliva proteins selectively adsorbed to silicone. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. Endod. 2006, 102, 488–494. [Google Scholar] [CrossRef] [PubMed]
- Bamford, C.V.; d’Mello, A.; Nobbs, A.H.; Dutton, L.C.; Vickerman, M.M.; Jenkinson, H.F. Streptococcus gordonii modulates Candida albicans biofilm formation through intergeneric communication. Infect. Immun. 2009, 77, 3696–3704. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.Y.; Fu, F.; Kong, W.N.; Xuan, Q.K.; Wen, D.H.; Chen, X.Q.; He, Y.M.; He, L.H.; Guo, J.; Zhou, A.P.; et al. Streptococcus agalactiae Inhibits Candida albicans Hyphal Development and Diminishes Host Vaginal Mucosal, T.H.17 Response. Front. Microbiol. 2018, 9, 198. [Google Scholar] [CrossRef] [PubMed]
- Vilchez, R.; Lemme, A.; Ballhausen, B.; Thiel, V.; Schulz, S.; Jansen, R.; Sztajer, H.; Wagner-Döbler, I. Streptococcus mutans inhibits Candida albicans hyphal formation by the fatty acid signaling molecule trans-2-decenoic acid (SDSF). Chembiochem 2010, 11, 1552–1562. [Google Scholar] [CrossRef]
- Jarosz, L.M.; Deng, D.M.; van der Mei, H.C.; Crielaard, W.; Krom, B.P. Streptococcus mutans competence-stimulating peptide inhibits Candida albicans hypha formation. Eukaryot. Cell 2009, 8, 1658–1664. [Google Scholar] [CrossRef]
- Ahn, S.J.; Wen, Z.T.; Burne, R.A. Multilevel control of competence development and stress tolerance in Streptococcus mutans, U.A.159. Infect. Immun. 2006, 74, 1631–1642. [Google Scholar] [CrossRef]
- Kong, E.F.; Tsui, C.; Kucharikova, S.; Andes, D.; Van Dijck, P.; Jabra-Rizk, M.A. Commensal Protection of Staphylococcus aureus against Antimicrobials by Candida albicans Biofilm Matrix. Mbio 2016, 7. [Google Scholar] [CrossRef]
- Pate, J.C.; Jones, D.B.; Wilhelmus, K.R. Prevalence and spectrum of bacterial co-infection during fungal keratitis. Br. J. Ophthalmol. 2006, 90, 289–292. [Google Scholar] [CrossRef]
- Gupta, N.; Haque, A.; Mukhopadhyay, G.; Narayan, R.P.; Prasad, R. Interactions between bacteria and Candida in the burn wound. Burns 2005, 31, 375–378. [Google Scholar] [CrossRef]
- Klotz, S.A.; Chasin, B.S.; Powell, B.; Gaur, N.K.; Lipke, P.N. Polymicrobial bloodstream infections involving Candida species: analysis of patients and review of the literature. Diagn. Microbiol. Infect. Dis. 2007, 59, 401–406. [Google Scholar] [CrossRef]
- Schlecht, L.M.; Peters, B.M.; Krom, B.P.; Freiberg, J.A.; Hansch, G.M.; Filler, S.G.; Jabra-Rizk, M.A.; Shirtliff, M.E. Systemic Staphylococcus aureus infection mediated by Candida albicans hyphal invasion of mucosal tissue. Microbiology 2015, 161, 168–181. [Google Scholar] [CrossRef] [PubMed]
- Krause, J.; Geginat, G.; Tammer, I. Prostaglandin E2 from Candida albicans Stimulates the Growth of Staphylococcus aureus in Mixed Biofilms. PLoS ONE 2015, 10, e0135404. [Google Scholar] [CrossRef] [PubMed]
- Shirtliff, M.E.; Peters, B.M.; Jabra-Rizk, M.A. Cross-kingdom interactions: Candida albicans and bacteria. FEMS Microbiol. Lett. 2009, 299, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Kong, E.F.; Tsui, C.; Kucharikova, S.; Van Dijck, P.; Jabra-Rizk, M.A. Modulation of Staphylococcus aureus Response to Antimicrobials by the Candida albicans Quorum Sensing Molecule Farnesol. Antimicrob. Agents Chemother. 2017, 61, e01573-17. [Google Scholar] [CrossRef]
- Inoue, Y.; Shiraishi, A.; Hada, T.; Hirose, K.; Hamashima, H.; Shimada, J. The antibacterial effects of terpene alcohols on Staphylococcus aureus and their mode of action. FEMS Microbiol. Lett. 2004, 237, 325–331. [Google Scholar] [CrossRef]
- Jabra-Rizk, M.A.; Meiller, T.F.; James, C.E.; Shirtliff, M.E. Effect of farnesol on Staphylococcus aureus biofilm formation and antimicrobial susceptibility. Antimicrob. Agents Chemother. 2006, 50, 1463–1469. [Google Scholar] [CrossRef]
- Camarillo-Marquez, O.; Cordova-Alcantara, I.M.; Hernandez-Rodriguez, C.H.; Garcia-Perez, B.E.; Martinez-Rivera, M.A.; Rodríguez-Tovar, A.V. Antagonistic Interaction of Staphylococcus aureus Toward Candida glabrata During in vitro Biofilm Formation Is Caused by an Apoptotic Mechanism. Front. Microbiol. 2018, 9, 2031. [Google Scholar] [CrossRef]
- Garsin, D.A.; Lorenz, M.C. Candida albicans and Enterococcus faecalis in the gut: synergy in commensalism? Gut Microbes 2013, 4, 409–415. [Google Scholar] [CrossRef]
- Kovac, J.; Kovac, D.; Slobodnikova, L.; Kotulova, D. Enterococcus faecalis and Candida albicans in the dental root canal and periapical infections. Bratisl. Lek. Listy. 2013, 114, 716–720. [Google Scholar]
- Cruz, M.R.; Graham, C.E.; Gagliano, B.C.; Lorenz, M.C.; Garsin, D.A. Enterococcus faecalis inhibits hyphal morphogenesis and virulence of Candida albicans. Infect. Immun. 2013, 81, 189–200. [Google Scholar] [CrossRef]
- Graham, C.E.; Cruz, M.R.; Garsin, D.A.; Lorenz, M.C. Enterococcus faecalis bacteriocin EntV inhibits hyphal morphogenesis, biofilm formation, and virulence of Candida albicans. Proc. Natl. Acad. Sci. USA 2017, 114, 4507–4512. [Google Scholar] [CrossRef] [PubMed]
- Shekh, R.M.; Roy, U. Biochemical characterization of an anti-Candida factor produced by Enterococcus faecalis. BMC Microbiol. 2012, 12, 1471–2180. [Google Scholar] [CrossRef] [PubMed]
- Dutton, L.C.; Jenkinson, H.F.; Lamont, R.J.; Nobbs, A.H. Role of Candida albicans secreted aspartyl protease Sap9 in interkingdom biofilm formation. Pathog. Dis. 2016, 74, 14. [Google Scholar] [CrossRef] [PubMed]
- Boris, S.; Barbes, C. Role played by lactobacilli in controlling the population of vaginal pathogens. Microbes Infect. 2000, 2, 543–546. [Google Scholar] [CrossRef]
- Noverr, M.C.; Huffnagle, G.B. Regulation of Candida albicans morphogenesis by fatty acid metabolites. Infect. Immun. 2004, 72, 6206–6210. [Google Scholar] [CrossRef] [PubMed]
- Vylkova, S.; Carman, A.J.; Danhof, H.A.; Collette, J.R.; Zhou, H.; Lorenz, M.C. The fungal pathogen Candida albicans autoinduces hyphal morphogenesis by raising extracellular pH. Mbio 2011, 2, e00055-11. [Google Scholar] [CrossRef]
- Wang, S.; Wang, Q.; Yang, E.; Yan, L.; Li, T.; Zhuang, H. Antimicrobial Compounds Produced by Vaginal Lactobacillus crispatus Are Able to Strongly Inhibit Candida albicans Growth, Hyphal Formation and Regulate Virulence-related Gene Expressions. Front. Microbiol. 2017, 8, 564. [Google Scholar] [CrossRef]
- Grimaudo, N.J.; Nesbitt, W.E.; Clark, W.B. Coaggregation of Candida albicans with oral Actinomyces species. Oral Microbiol. Immunol. 1996, 11, 59–61. [Google Scholar] [CrossRef]
- Arzmi, M.H.; Dashper, S.; Catmull, D.; Cirillo, N.; Reynolds, E.C.; McCullough, M. Coaggregation of Candida albicans, Actinomyces naeslundii and Streptococcus mutans is Candida albicans strain dependent. FEMS Yeast Res. 2015, 15, 7. [Google Scholar] [CrossRef]
- Arzmi, M.H.; Alnuaimi, A.D.; Dashper, S.; Cirillo, N.; Reynolds, E.C.; McCullough, M. Polymicrobial biofilm formation by Candida albicans, Actinomyces naeslundii, and Streptococcus mutans is Candida albicans strain and medium dependent. Med. Mycol. 2016, 54, 856–864. [Google Scholar] [CrossRef]
- Deng, L.; Li, W.; He, Y.; Wu, J.; Ren, B.; Zou, L. Cross-kingdom interaction of Candida albicans and Actinomyces viscosus elevated cariogenic virulence. Arch. Oral Biol. 2019, 100, 106–112. [Google Scholar] [CrossRef] [PubMed]
- Peleg, A.Y.; Seifert, H.; Paterson, D.L. Acinetobacter baumannii: emergence of a successful pathogen. Clin. Microbiol. Rev. 2008, 21, 538–582. [Google Scholar] [CrossRef] [PubMed]
- Peleg, A.Y.; Tampakakis, E.; Fuchs, B.B.; Eliopoulos, G.M.; Moellering, R.C., Jr.; Mylonakis, E. Prokaryote-eukaryote interactions identified by using Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 2008, 105, 14585–14590. [Google Scholar] [CrossRef] [PubMed]
- Gaddy, J.A.; Tomaras, A.P.; Actis, L.A. The Acinetobacter baumannii 19606 OmpA protein plays a role in biofilm formation on abiotic surfaces and in the interaction of this pathogen with eukaryotic cells. Infect. Immun. 2009, 77, 3150–3160. [Google Scholar] [CrossRef]
- Kostoulias, X.; Murray, G.L.; Cerqueira, G.M.; Kong, J.B.; Bantun, F.; Mylonakis, E.; Khoo, C.A.; Peleg, A.Y. Impact of a Cross-Kingdom Signaling Molecule of Candida albicans on Acinetobacter baumannii Physiology. Antimicrob. Agents Chemother. 2015, 60, 161–167. [Google Scholar] [CrossRef]
- Liu, C.Y.; Liao, C.H.; Chen, Y.C.; Chang, S.C. Changing epidemiology of nosocomial bloodstream infections in 11 teaching hospitals in Taiwan between 1993 and 2006. J. Microbiol. Immunol. Infect. 2010, 43, 416–429. [Google Scholar] [CrossRef]
- Bachtiar, E.W.; Bachtiar, B.M.; Jarosz, L.M.; Amir, L.R.; Sunarto, H.; Ganin, H.; Meijler, M.M.; Krom, B.P. AI-2 of Aggregatibacter actinomycetemcomitans inhibits Candida albicans biofilm formation. Front. Cell Infect. Microbiol. 2014, 4, 94. [Google Scholar] [CrossRef]
- Brusca, M.I.; Rosa, A.; Albaina, O.; Moragues, M.D.; Verdugo, F.; Pontón, J. The impact of oral contraceptives on women’s periodontal health and the subgingival occurrence of aggressive periodontopathogens and Candida species. J. Periodontol. 2010, 81, 1010–1018. [Google Scholar] [CrossRef]
- Rickard, A.H.; Campagna, S.R.; Kolenbrander, P.E. Autoinducer-2 is produced in saliva-fed flow conditions relevant to natural oral biofilms. J. Appl. Microbiol. 2008, 105, 2096–2103. [Google Scholar] [CrossRef][Green Version]
- Carlson, E. Enhancement by Candida albicans of Staphylococcus aureus, Serratia marcescens, and Streptococcus faecalis in the establishment of infection in mice. Infect. Immun. 1983, 39, 193–197. [Google Scholar]
- Bagg, J.; Silverwood, R.W. Coagglutination reactions between Candida albicans and oral bacteria. J. Med. Microbiol. 1986, 22, 165–169. [Google Scholar] [CrossRef] [PubMed]
- Grimaudo, N.J.; Nesbitt, W.E. Coaggregation of Candida albicans with oral Fusobacterium species. Oral Microbiol. Immunol. 1997, 12, 168–173. [Google Scholar] [CrossRef] [PubMed]
- Jabra-Rizk, M.A.; Falkler, W.A., Jr.; Merz, W.G.; Kelley, J.I.; Baqui, A.A.; Meiller, T.F. Coaggregation of Candida dubliniensis with Fusobacterium nucleatum. J. Clin. Microbiol. 1999, 37, 1464–1468. [Google Scholar] [PubMed]
- Bor, B.; Cen, L.; Agnello, M.; Shi, W.; He, X. Morphological and physiological changes induced by contact-dependent interaction between Candida albicans and Fusobacterium nucleatum. Sci. Rep. 2016, 6, 27956. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Cen, L.; Kaplan, C.; Zhou, X.; Lux, R.; Shi, W.; He, X. Cellular Components Mediating Coadherence of Candida albicans and Fusobacterium nucleatum. J. Dent. Res. 2015, 94, 1432–1438. [Google Scholar] [CrossRef]
- Baldwin, A.; Mahenthiralingam, E.; Drevinek, P.; Vandamme, P.; Govan, J.R.; Waine, D.J.; LiPuma, J.J.; Chiarini, L.; Dalmastri, C.; Henry, D.A.; et al. Environmental Burkholderia cepacia complex isolates in human infections. Emerg. Infect. Dis. 2007, 13, 458–461. [Google Scholar] [CrossRef]
- Sousa, S.A.; Ramos, C.G.; Leitao, J.H. Burkholderia cepacia Complex: Emerging Multihost Pathogens Equipped with a Wide Range of Virulence Factors and Determinants. Int. J. Microbiol. 2011, 10, 3. [Google Scholar] [CrossRef]
- Boon, C.; Deng, Y.; Wang, L.H.; He, Y.; Xu, J.L.; Fan, Y.; Pan, S.Q.; Zhang, L.H. A novel, D.S.F-like signal from Burkholderia cenocepacia interferes with Candida albicans morphological transition. ISME J. 2008, 2, 27–36. [Google Scholar] [CrossRef]
- Tian, J.; Weng, L.X.; Zhang, Y.Q.; Wang, L.H. BDSF inhibits Candida albicans adherence to urinary catheters. Microb. Pathog. 2013, 64, 33–38. [Google Scholar] [CrossRef]
- Fox, E.P.; Cowley, E.S.; Nobile, C.J.; Hartooni, N.; Newman, D.K.; Johnson, A.D. Anaerobic bacteria grow within Candida albicans biofilms and induce biofilm formation in suspension cultures. Curr. Biol. 2014, 24, 2411–2416. [Google Scholar] [CrossRef]
- van Leeuwen, P.T.; van der Peet, J.M.; Bikker, F.J.; Hoogenkamp, M.A.; Oliveira Paiva, A.M.; Kostidis, S.; Mayboroda, O.A.; Smits, W.K.; Krom, B.P. Interspecies Interactions between Clostridium difficile and Candida albicans. Msphere 2016, 1, e00187-16. [Google Scholar] [CrossRef] [PubMed]
- Somerville, G.A.; Proctor, R.A. Cultivation conditions and the diffusion of oxygen into culture media: the rationale for the flask-to-medium ratio in microbiology. BMC Microbiol. 2013, 13, 1471–2180. [Google Scholar] [CrossRef] [PubMed]
- Dione, N.; Khelaifia, S.; Lagier, J.C.; Raoult, D. The aerobic activity of metronidazole against anaerobic bacteria. Int. J. Antimicrob. Agents 2015, 45, 537–540. [Google Scholar] [CrossRef] [PubMed]
- Janus, M.M.; Crielaard, W.; Volgenant, C.M.; van der Veen, M.H.; Brandt, B.W.; Krom, B.P. Candida albicans alters the bacterial microbiome of early in vitro oral biofilms. J. Oral. Microbiol. 2017, 9, 1270613. [Google Scholar] [CrossRef] [PubMed]
- Tampakakis, E.; Peleg, A.Y.; Mylonakis, E. Interaction of Candida albicans with an intestinal pathogen, Salmonella enterica serovar Typhimurium. Eukaryot. Cell. 2009, 8, 732–737. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Mylonakis, E. Killing of Candida albicans filaments by Salmonella enterica serovar Typhimurium is mediated by sopB effectors, parts of a type, I.I.I secretion system. Eukaryot. Cell. 2011, 10, 782–790. [Google Scholar] [CrossRef] [PubMed]
- Briard, B.; Mislin, G.L.A.; Latge, J.P.; Beauvais, A. Interactions between Aspergillus fumigatus and Pulmonary Bacteria: Current State of the Field, New Data, and Future Perspective. J. Fungi (Basel) 2019, 5, 48. [Google Scholar] [CrossRef]
- Sass, G.; Nazik, H.; Penner, J.; Shah, H.; Ansari, S.R.; Clemons, K.V.; Groleau, M.C.; Dietl, A.M.; Visca; Haas, H.; et al. Aspergillus-Pseudomonas interaction, relevant to competition in airways. Med Mycol 2019, 57, S228–S232. [Google Scholar] [CrossRef]
- Sass, G.; Nazik, H.; Penner, J.; Shah, H.; Ansari, S.R.; Clemons, K.V.; Groleau, M.C.; Dietl, A.M.; Visca, P.; Haas, H.; et al. Studies of Pseudomonas aeruginosa Mutants Indicate Pyoverdine as the Central Factor in Inhibition of Aspergillus fumigatus Biofilm. J. Bacteriol. 2017, 200, e00345-17. [Google Scholar] [CrossRef]
- Reece, E.; Doyle, S.; Greally, P.; Renwick, J.; McClean, S. Aspergillus fumigatus Inhibits Pseudomonas aeruginosa in Co-culture: Implications of a Mutually Antagonistic Relationship on Virulence and Inflammation in the, C.F. Airway. Front. Microbiol. 2018, 9, 1205. [Google Scholar] [CrossRef]
- Mowat, E.; Rajendran, R.; Williams, C.; McCulloch, E.; Jones, B.; Lang, S.; Ramage, G. Pseudomonas aeruginosa and their small diffusible extracellular molecules inhibit Aspergillus fumigatus biofilm formation. FEMS Microbiol. Lett. 2010, 313, 96–102. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, J.A.; Penner, J.C.; Moss, R.B.; Haagensen, J.A.; Clemons, K.V.; Spormann, A.M.; Nazik, H.; Cohen, K.; Banaei, N.; Carolino, E.; et al. Inhibition of Aspergillus fumigatus and Its Biofilm by Pseudomonas aeruginosa Is Dependent on the Source, Phenotype and Growth Conditions of the Bacterium. PLoS ONE 2015, 10, e0134692. [Google Scholar] [CrossRef] [PubMed]
- Sass, G.; Ansari, S.R.; Dietl, A.M.; Deziel, E.; Haas, H.; Stevens, D.A. Intermicrobial interaction: Aspergillus fumigatus siderophores protect against competition by Pseudomonas aeruginosa. PLoS ONE 2019, 14, e0216085. [Google Scholar] [CrossRef] [PubMed]
- Briard, B.; Heddergott, C.; Latge, J.P. Volatile Compounds Emitted by Pseudomonas aeruginosa Stimulate Growth of the Fungal Pathogen Aspergillus fumigatus. Mbio 2016, 7, e00219. [Google Scholar] [CrossRef]
- Nutzmann, H.W.; Reyes-Dominguez, Y.; Scherlach, K.; Schroeckh, V.; Horn, F.; Gacek, A.; Schümann, J.; Hertweck, C.; Strauss, J.; Brakhage, A.A. Bacteria-induced natural product formation in the fungus Aspergillus nidulans requires Saga/Ada-mediated histone acetylation. Proc. Natl. Acad. Sci. USA 2011, 108, 14282–14287. [Google Scholar] [CrossRef] [PubMed]
- Schroeckh, V.; Scherlach, K.; Nutzmann, H.W.; Shelest, E.; Schmidt-Heck, W.; Schuemann, J.; Martin, K.; Hertweck, C.; Brakhage, A.A. Intimate bacterial-fungal interaction triggers biosynthesis of archetypal polyketides in Aspergillus nidulans. Proc. Natl. Acad. Sci. USA 2009, 106, 14558–14563. [Google Scholar] [CrossRef]
- Fischer, J.; Muller, S.Y.; Netzker, T.; Jager, N.; Gacek-Matthews, A.; Scherlach, K.; Stroe, M.C.; García-Altares, M.; Pezzini, F.; Schoeler, H.; et al. Chromatin mapping identifies BasR, a key regulator of bacteria-triggered production of fungal secondary metabolites. Elife 2018, 12, 40969. [Google Scholar] [CrossRef]
- Brandl, M.T.; Carter, M.Q.; Parker, C.T.; Chapman, M.R.; Huynh, S.; Zhou, Y. Salmonella biofilm formation on Aspergillus niger involves cellulose--chitin interactions. PLoS ONE 2011, 6, e25553. [Google Scholar] [CrossRef]
- Nogueira, M.F.; Pereira, L.; Jenull, S.; Kuchler, K.; Lion, T. Klebsiella pneumoniae prevents spore germination and hyphal development of Aspergillus species. Sci. Rep. 2019, 9, 018–36524. [Google Scholar] [CrossRef]
- Rella, A.; Yang, M.W.; Gruber, J.; Montagna, M.T.; Luberto, C.; Zhang, Y.M.; Del Poeta, M. Pseudomonas aeruginosa inhibits the growth of Cryptococcus species. Mycopathologia 2012, 173, 451–461. [Google Scholar] [CrossRef]
- Teoh-Chan, H.; Chau, P.Y.; Ng, M.H.; Wong, P.C. Inhibition of Cryptococcus neoformans by Pseudomonas aeruginosa. J. Med. Microbiol. 1975, 8, 77–81. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Frases, S.; Chaskes, S.; Dadachova, E.; Casadevall, A. Induction by Klebsiella aerogenes of a melanin-like pigment in Cryptococcus neoformans. Appl. Env. Microbiol. 2006, 72, 1542–1550. [Google Scholar] [CrossRef] [PubMed]
- Frases, S.; Salazar, A.; Dadachova, E.; Casadevall, A. Cryptococcus neoformans can utilize the bacterial melanin precursor homogentisic acid for fungal melanogenesis. Appl. Env. Microbiol. 2007, 73, 615–621. [Google Scholar] [CrossRef] [PubMed]
- Rick, E.M.; Woolnough, K.; Pashley, C.H.; Wardlaw, A.J. Allergic Fungal Airway Disease. J. Investig. Allergol. Clin. Immunol. 2016, 26, 344–354. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Pan, C.; Wang, K.; Chen, X.; Wu, X.; Chen, C.A.; Wu, B. Synthetic multispecies microbial communities reveals shifts in secondary metabolism and facilitates cryptic natural product discovery. Env. Microbiol. 2017, 19, 3606–3618. [Google Scholar] [CrossRef] [PubMed]
- Butt, A.T.; Thomas, M.S. Iron Acquisition Mechanisms and Their Role in the Virulence of Burkholderia Species. Front. Cell Infect. Microbiol. 2017, 7, 460. [Google Scholar] [CrossRef]
- Ross, C.; Opel, V.; Scherlach, K.; Hertweck, C. Biosynthesis of antifungal and antibacterial polyketides by Burkholderia gladioli in coculture with Rhizopus microsporus. Mycoses 2014, 3, 48–55. [Google Scholar] [CrossRef]
- Moebius, N.; Ross, C.; Scherlach, K.; Rohm, B.; Roth, M.; Hertweck, C. Biosynthesis of the respiratory toxin bongkrekic acid in the pathogenic bacterium Burkholderia gladioli. Chem. Biol. 2012, 19, 1164–1174. [Google Scholar] [CrossRef]
- Lackner, G.; Moebius, N.; Partida-Martinez, L.P.; Boland, S.; Hertweck, C. Evolution of an endofungal lifestyle: Deductions from the Burkholderia rhizoxinica genome. BMC Genomics 2011, 12, 1471–2164. [Google Scholar] [CrossRef]
- Smith, M.G.; Des Etages, S.G.; Snyder, M. Microbial synergy via an ethanol-triggered pathway. Mol. Cell Biol. 2004, 24, 3874–3884. [Google Scholar] [CrossRef]
- Gandhi, J.A.; Ekhar, V.V.; Asplund, M.B.; Abdulkareem, A.F.; Ahmadi, M.; Coelho, C.; Martinez, L.R. Alcohol enhances Acinetobacter baumannii-associated pneumonia and systemic dissemination by impairing neutrophil antimicrobial activity in a murine model of infection. PLoS ONE 2014, 9, e95707. [Google Scholar] [CrossRef] [PubMed]
- Kaur, J.; Pethani, B.P.; Kumar, S.; Kim, M.; Sunna, A.; Kautto, L.; Penesyan, A.; Paulsen, I.T.; Nevalainen, H. Pseudomonas aeruginosa inhibits the growth of Scedosporium aurantiacum, an opportunistic fungal pathogen isolated from the lungs of cystic fibrosis patients. Front. Microbiol. 2015, 6, 866. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.C.; Patel, S.; Meyer, W.; Chapman, B.; Yu, H.; Byth, K.; Middleton, P.G.; Nevalainen, H.; Sorrell, T.C. Pseudomonas aeruginosa Inhibits the Growth of Scedosporium and Lomentospora In Vitro. Mycopathologia 2018, 183, 251–261. [Google Scholar] [CrossRef] [PubMed]
- Pappas, P.G.; Lionakis, M.S.; Arendrup, M.C.; Ostrosky-Zeichner, L.; Kullberg, B.J. Invasive candidiasis. Nat. Rev. Dis. Primers 2018, 4, 18026. [Google Scholar] [CrossRef]
- Chin, V.K.; Lee, T.Y.; Rusliza, B.; Chong, P.P. Dissecting Candida albicans Infection from the Perspective of C. albicans Virulence and Omics Approaches on Host-Pathogen Interaction: A Review. Int. J. Mol. Sci. 2016, 17, 1643. [Google Scholar] [CrossRef]
- Patin, E.C.; Thompson, A.; Orr, S.J. Pattern recognition receptors in fungal immunity. Semin. Cell Dev. Biol. 2018, 8, 30541–30544. [Google Scholar] [CrossRef]
- Espinosa, V.; Rivera, A. Cytokines and the regulation of fungus-specific, C.D.4 T cell differentiation. Cytokine 2012, 58, 100–106. [Google Scholar] [CrossRef]
- Pichard, D.C.; Freeman, A.F.; Cowen, E.W. Primary immunodeficiency update: Part, I.I.. Syndromes associated with mucocutaneous candidiasis and noninfectious cutaneous manifestations. J. Am. Acad. Dermatol. 2015, 73, 367–381. [Google Scholar] [CrossRef]
- Chaplin, D.D. Overview of the immune response. J. Allergy Clin. Immunol. 2010, 125, 980. [Google Scholar] [CrossRef]
- Lewis, M.L.; Surewaard, B.G.J. Neutrophil evasion strategies by Streptococcus pneumoniae and Staphylococcus aureus. Cell Tissue Res. 2018, 371, 489–503. [Google Scholar] [CrossRef]
- Hernandez-Chavez, M.J.; Perez-Garcia, L.A.; Nino-Vega, G.A.; Mora-Montes, H.M. Fungal Strategies to Evade the Host Immune Recognition. J. Fungi (Basel) 2017, 3, 51. [Google Scholar] [CrossRef] [PubMed]
- Chow, S.H.; Deo, P.; Naderer, T. Macrophage cell death in microbial infections. Cell Microbiol. 2016, 18, 466–474. [Google Scholar] [CrossRef] [PubMed]
- Ricciardi, B.F.; Muthukrishnan, G.; Masters, E.; Ninomiya, M.; Lee, C.C.; Schwarz, E.M. Staphylococcus aureus Evasion of Host Immunity in the Setting of Prosthetic Joint Infection: Biofilm and Beyond. Curr. Rev. Musculoskelet Med. 2018, 9, 018–9501. [Google Scholar] [CrossRef] [PubMed]
- Byndloss, M.X.; Tsolis, R.M. Chronic Bacterial Pathogens: Mechanisms of Persistence. Microbiol. Spectr. 2016, 4, 0020–2015. [Google Scholar] [CrossRef]
- Bernal-Bayard, J.; Ramos-Morales, F. Molecular Mechanisms Used by Salmonella to Evade the Immune System. Curr. Issues Mol. Biol. 2018, 25, 133–168. [Google Scholar] [CrossRef] [PubMed]
- Roux, D.; Gaudry, S.; Dreyfuss, D.; El-Benna, J.; de Prost, N.; Denamur, E.; Saumon, G.; Ricard, J.D. Candida albicans impairs macrophage function and facilitates Pseudomonas aeruginosa pneumonia in rat. Crit. Care Med. 2009, 37, 1062–1067. [Google Scholar] [CrossRef]
- Xu, H.; Sobue, T.; Bertolini, M.; Thompson, A.; Vickerman, M.; Nobile, C.J.; Dongari-Bagtzoglou, A. S. oralis activates the Efg1 filamentation pathway in C. albicans to promote cross-kingdom interactions and mucosal biofilms. Virulence 2017, 8, 1602–1617. [Google Scholar] [CrossRef]
- Nash, E.E.; Peters, B.M.; Fidel, P.L.; Noverr, M.C. Morphology-Independent Virulence of Candida Species during Polymicrobial Intra-abdominal Infections with Staphylococcus aureus. Infect. Immun. 2015, 84, 90–98. [Google Scholar] [CrossRef]
- Kean, R.; Rajendran, R.; Haggarty, J.; Townsend, E.M.; Short, B.; Burgess, K.E.; Lang, S.; Millington, O.; Mackay, W.G.; Williams, C.; et al. Candida albicans Mycofilms Support Staphylococcus aureus Colonization and Enhances Miconazole Resistance in Dual-Species Interactions. Front. Microbiol. 2017, 8, 258. [Google Scholar] [CrossRef]
- Holt, J.E.; Houston, A.; Adams, C.; Edwards, S.; Kjellerup, B.V. Role of extracellular polymeric substances in polymicrobial biofilm infections of Staphylococcus epidermidis and Candida albicans modelled in the nematode Caenorhabditis elegans. Pathog Dis. 2017, 75. [Google Scholar] [CrossRef]
- Ader, F.; Jawhara, S.; Nseir, S.; Kipnis, E.; Faure, K.; Vuotto, F.; Chemani, C.; Sendid, B.; Poulain, D.; Guery, B. Short term Candida albicans colonization reduces Pseudomonas aeruginosa-related lung injury and bacterial burden in a murine model. Crit. Care 2011, 15, R150. [Google Scholar] [CrossRef]
- Lopez-Medina, E.; Fan, D.; Coughlin, L.A.; Ho, E.X.; Lamont, I.L.; Reimmann, C.; Hooper, L.V.; Koh, A.Y. Candida albicans Inhibits Pseudomonas aeruginosa Virulence through Suppression of Pyochelin and Pyoverdine Biosynthesis. Plos Pathog. 2015, 11, e1005129. [Google Scholar] [CrossRef] [PubMed]
- Diaz, P.I.; Xie, Z.; Sobue, T.; Thompson, A.; Biyikoglu, B.; Ricker, A.; Ikonomou, L.; Dongari-Bagtzoglou, A. Synergistic interaction between Candida albicans and commensal oral streptococci in a novel in vitro mucosal model. Infect. Immun. 2012, 80, 620–632. [Google Scholar] [CrossRef] [PubMed]
- Nash, E.E.; Peters, B.M.; Palmer, G.E.; Fidel, P.L.; Noverr, M.C. Morphogenesis is not required for Candida albicans-Staphylococcus aureus intra-abdominal infection-mediated dissemination and lethal sepsis. Infect. Immun. 2014, 82, 3426–3435. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ermolaeva, M.A.; Schumacher, B. Insights from the worm: the C. elegans model for innate immunity. Semin. Immunol. 2014, 26, 303–309. [Google Scholar] [CrossRef] [PubMed]
- Fehrmann, C.; Jurk, K.; Bertling, A.; Seidel, G.; Fegeler, W.; Kehrel, B.E.; Peters, G.; Becker, K.; Heilmann, C. Role for the fibrinogen-binding proteins coagulase and Efb in the Staphylococcus aureus-Candida interaction. Int. J. Med. Microbiol. 2013, 303, 230–238. [Google Scholar] [CrossRef] [PubMed]
- Villena, J.; Salva, S.; Aguero, G.; Alvarez, S. Immunomodulatory and protective effect of probiotic Lactobacillus casei against Candida albicans infection in malnourished mice. Microbiol. Immunol. 2011, 55, 434–445. [Google Scholar] [CrossRef]
- Zelante, T.; Iannitti, R.G.; Cunha, C.; De Luca, A.; Giovannini, G.; Pieraccini, G.; Zecchi, R.; D’Angelo, C.; Massi-Benedetti, C.; Fallarino, F.; et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity 2013, 39, 372–385. [Google Scholar] [CrossRef]
- Bork, P.; Beckmann, G. The, C.U.B domain. A widespread module in developmentally regulated proteins. J. Mol. Biol. 1993, 231, 539–545. [Google Scholar] [CrossRef]
- Arvanitis, M.; Mylonakis, E. Characteristics, Clinical Relevance, and the Role of Echinocandins in Fungal-Bacterial Interactions. Clin. Infect. Dis. 2015, 61, S630–634. [Google Scholar] [CrossRef]

| Bacterial–Fungal Interaction | Host | Immune Response | References | ||
|---|---|---|---|---|---|
| In Vivo Model | Mortality | Cytokines/Chemokines/Molecules | Effect | ||
| Candida albicans and Pseudomonas aeruginosa | Rat | Elevated pro-inflammatory cytokines: TNF-α, IFN-γ, IL-6 | Higher bacterial loads in the lungs; impaired macrophage function in the lungs | [216] | |
| Mouse | ↑ | High mortality mostly due to protease activity of P. aeruginosa | [1] | ||
| Mouse | ↓ | IL-22; IL-17 pathway; AMPs | Colonization by C. albicans leads to protection against P. aeruginosa-associated pneumonia; lower bacterial loads and decreased epithelial injury | [99,221] | |
| Mouse | ↓ | Factors secreted by C. albicans inhibit P. aeruginosa siderophores via cytotoxic molecules reducing the bacterial virulence | [222] | ||
| Zebrafish | ↑ | Elevated pro-inflammatory cytokine: IL-6; Elevated neutrophil chemoattractant: IL-8 | Increased C. albicans–mediated pathogenicity and virulence; increased inflammatory response; no excessive neutrophil infiltration | [2] | |
| C. albicans and Streptococcus spp. | Mouse | Toll-like receptor (TLR)-2 signalling; cytokines IL-17C, CXCL1, MIP-2/CXCL2, TNF, IL-1α, IL-1β; neutrophil protein CD177, CD14, MMP8 | Increased C. albicans–mediated pathogenicity and virulence; increased inflammatory response; increased neutrophil infiltration; hypervirulent biofilms; hyphal gene EFG1 required for robust mixed biofilms | [113,223] | |
| Candida spp. and Staphylococcus spp. | Mouse | ↑ | IL-6; PGE2; IL-1β; TNF-α | Yeast-to-hyphae transition of Candida does not influence dissemination and lethal sepsis | [218,224] |
| C. elegans | ↑ | C-type lectins; CUB domain containing factors; AMPs | Increased virulence of both species | [220,225] | |
| Galleria mellonella | ↑ | Increased pathogenicity; Staphylococcus aureus helps C. albicans circumvent the IS, contributing to its persistence | [219,226] | ||
| C. albicans and Enterococcus faecalis | C. elegans | ↓ | Exposure to E. faecalis primes the IS to better cope with later exposure to C. albicans; hyphae are inhibited; reduced tissue damage | [138,140] | |
| C. albicans and Escherichia coli | Mouse | ↑ | Endotoxin mediating synergistic lethality | Currently unknown | [59,60] |
| C. albicans and Lactobacillus spp. | Mouse | ↓ | TNF-α; IFN-γ; IL-6; IL-10; IL-22 | Bacterial treatment followed by C. albicans infection improved survival and resistance of the mouse | [227,228] |
| C. albicans and Acinetobacter baumannii | C. elegans | ↓ | C. albicans pathogenicity is decreased; hyphae are inhibited; C. albicans proliferation in the gut is reduced | [153] | |
| Aspergillus fumigatus and P. aeruginosa | G. mellonella | ↑ | Activation of mitogen-activated protein kinases (MAPKs) ERK and p38 | No additive of the co-infection on inflammation; lack of synergistic inflammatory response; saturation of signaling pathways | [180] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nogueira, F.; Sharghi, S.; Kuchler, K.; Lion, T. Pathogenetic Impact of Bacterial–Fungal Interactions. Microorganisms 2019, 7, 459. https://doi.org/10.3390/microorganisms7100459
Nogueira F, Sharghi S, Kuchler K, Lion T. Pathogenetic Impact of Bacterial–Fungal Interactions. Microorganisms. 2019; 7(10):459. https://doi.org/10.3390/microorganisms7100459
Chicago/Turabian StyleNogueira, Filomena, Shirin Sharghi, Karl Kuchler, and Thomas Lion. 2019. "Pathogenetic Impact of Bacterial–Fungal Interactions" Microorganisms 7, no. 10: 459. https://doi.org/10.3390/microorganisms7100459
APA StyleNogueira, F., Sharghi, S., Kuchler, K., & Lion, T. (2019). Pathogenetic Impact of Bacterial–Fungal Interactions. Microorganisms, 7(10), 459. https://doi.org/10.3390/microorganisms7100459





