A Review on Biotechnological Approaches Applied for Marine Hydrocarbon Spills Remediation
Abstract
1. Introduction
2. Microorganisms Involved in the Removal of Oil Spills from Marine Surfaces
3. Bioremediation and Affecting Factors
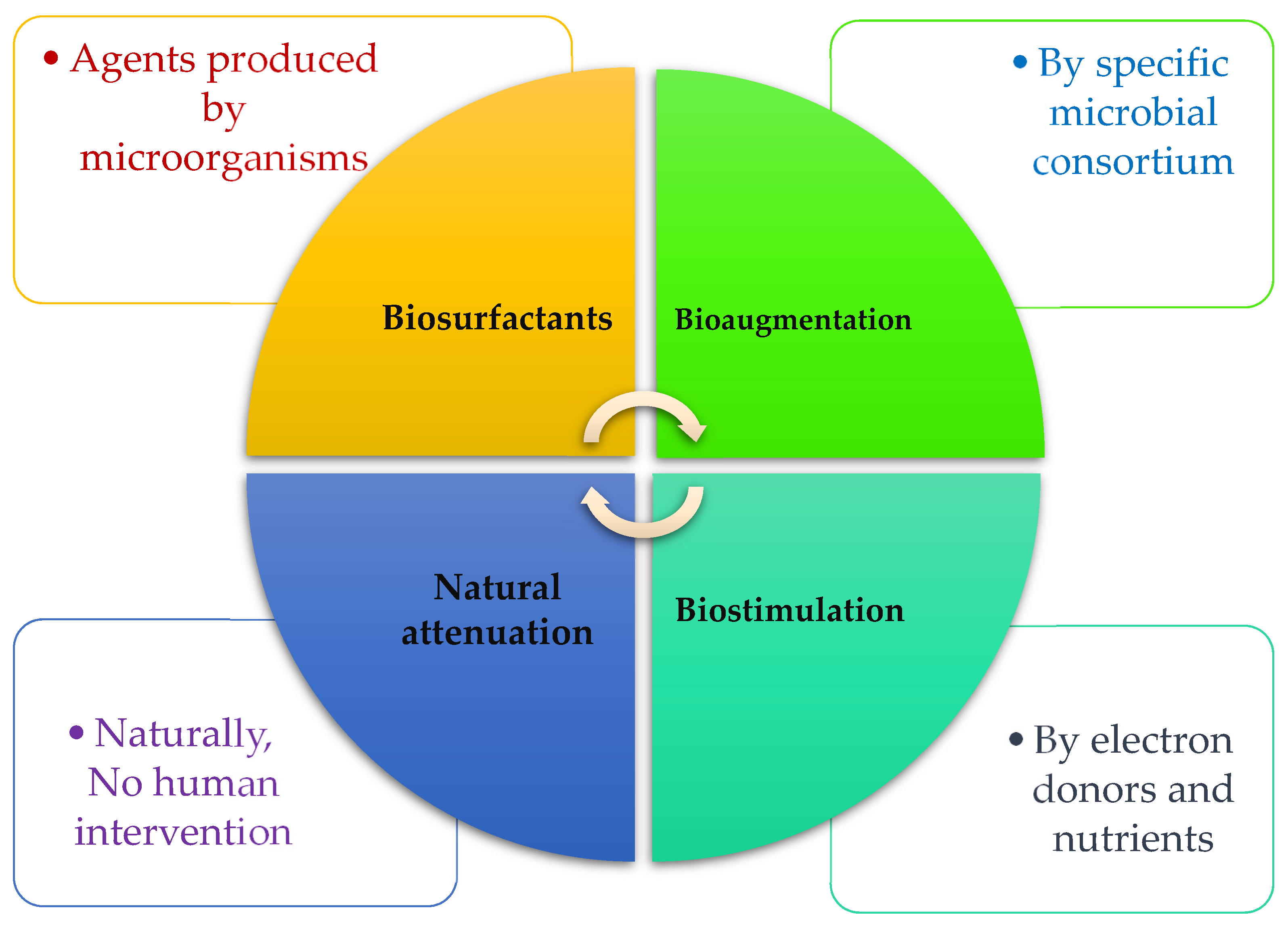
4. Biologically Based Solutions for Ashore and Marine Pollution
4.1. Bioaugmentation
4.2. Biostimulation
4.3. Biosurfactants
4.4. Cell Immobilization Techniques for Increasing Bioremediation Efficiency
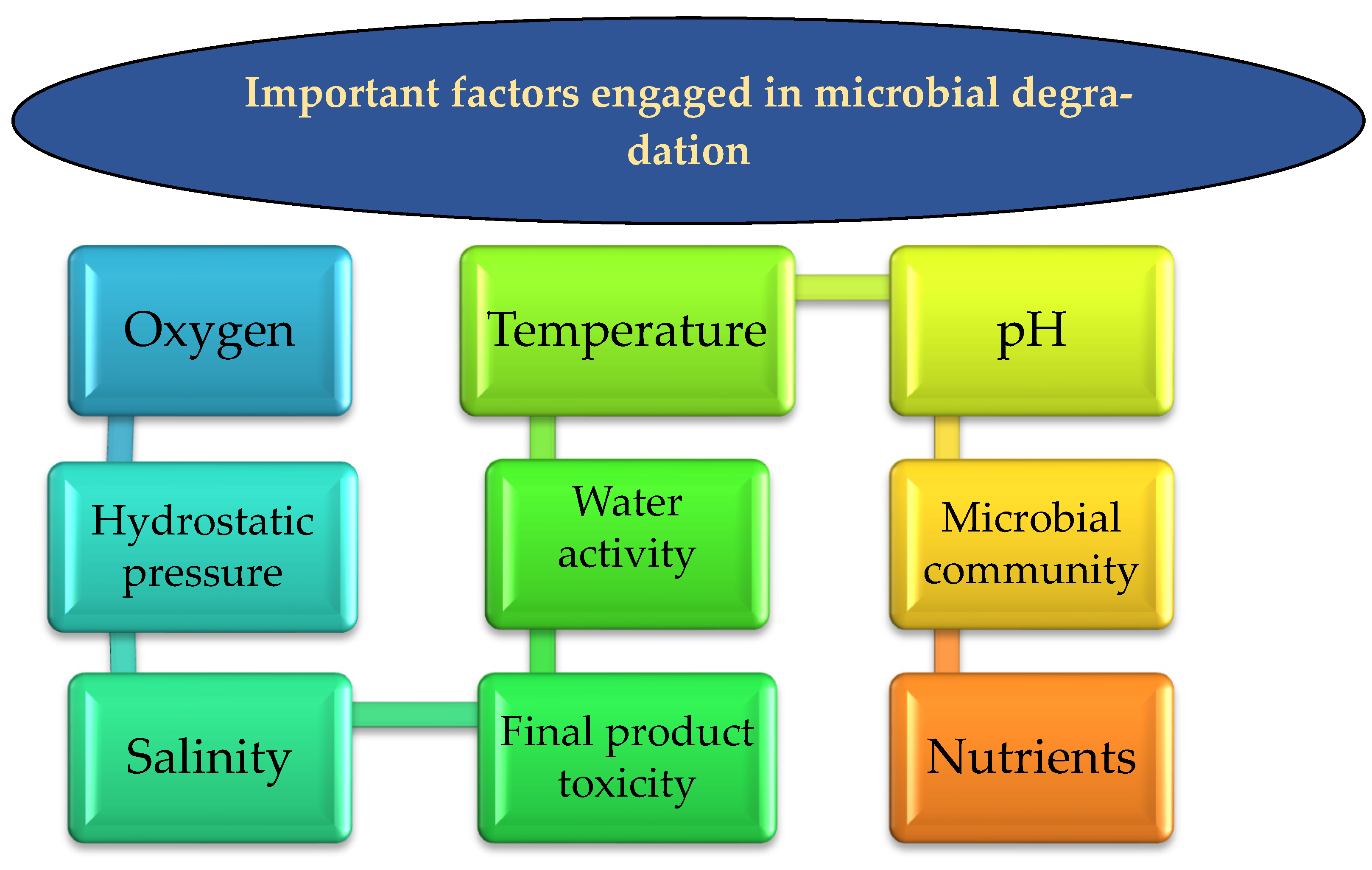
5. Factors Engaged in Microbial Degradation
5.1. Oxygen Bioavailability
5.2. Hydrostatic Pressure
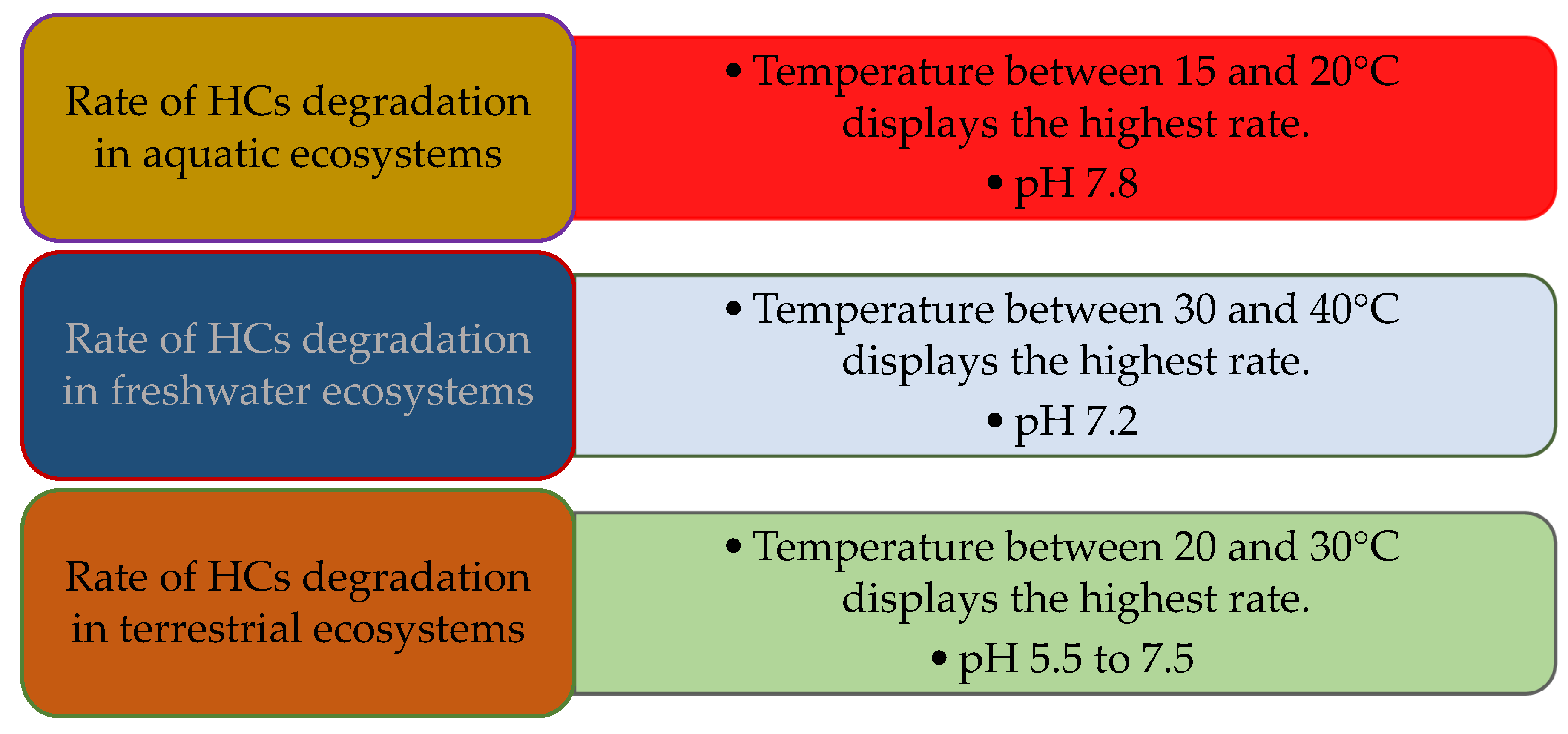
5.3. Temperature
5.4. Microbial Community
5.5. Product Toxicity of HCs Spills
6. Anaerobic and Aerobic Degradation of HCs and Involving Enzymes
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Mityagina, M.I.; Lavrova, O.Y.; Kostianoy, A.G. Main pattern of the Caspian Sea surface oil pollution revealed by satellite data. Ecol. Montenegrina 2019, 25, 91–105. [Google Scholar] [CrossRef]
- Li, P.; Cai, Q.; Lin, W.; Chen, B.; Zhang, B. Offshore oil spill response practices and emerging challenges. Mar. Pollut. Bull. 2016, 110, 6–27. [Google Scholar] [CrossRef] [PubMed]
- Ramseur, J.L. Deepwater Horizon Oil Spill: The Fate of the Oil; Congressional Research Service, Library of Congress: Washington, DC, USA, 2010.
- Chang, S.E.; Stone, J.; Demes, K.; Piscitelli, M. Consequences of oil spills: A review and framework for informing planning. Ecol. Soc. 2014, 19, 26. [Google Scholar] [CrossRef]
- Tango, M.; Islam, M. Potential of extremophiles for biotechnological and petroleum applications. Energy Sources 2002, 24, 543–559. [Google Scholar] [CrossRef]
- Mapelli, F.; Scoma, A.; Michoud, G.; Aulenta, F.; Boon, N.; Borin, S.; Kalogerakis, N.; Daffonchio, D. Biotechnologies for marine oil spill cleanup: Indissoluble ties with microorganisms. Trends Biotechnol. 2017, 35, 860–870. [Google Scholar] [CrossRef] [PubMed]
- Pandotra, P.; Raina, M.; Salgotra, R.; Ali, S.; Mir, Z.A.; Bhat, J.A.; Tyagi, A.; Upadhahy, D. Plant-bacterial partnership: A major pollutants remediation approach. In Modern Age Environmental Problems and Their Remediation; Springer: Berlin/Heidelberg, Germany, 2018; pp. 169–200. [Google Scholar]
- Bacosa, H.P.; Erdner, D.L.; Liu, Z. Differentiating the roles of photooxidation and biodegradation in the weathering of Light Louisiana Sweet crude oil in surface water from the Deepwater Horizon site. Mar. Pollut. Bull. 2015, 95, 265–272. [Google Scholar] [CrossRef]
- Bao, M.; Chen, Q.; Gong, Y.; Li, Y.; Wang, H.; Jiang, G. Removal efficiency of heavy oil by free and immobilised microorganisms on laboratory-scale. Can. J. Chem. Eng. 2013, 91, 1–8. [Google Scholar] [CrossRef]
- Calderoli, P.A.; Espínola, F.J.; Dionisi, H.M.; Gil, M.N.; Jansson, J.K.; Lozada, M. Predominance and high diversity of genes associated to denitrification in metagenomes of subantarctic coastal sediments exposed to urban pollution. PLoS ONE 2018, 13, e0207606. [Google Scholar] [CrossRef]
- Yakimov, M.M.; Timmis, K.N.; Golyshin, P.N. Obligate oil-degrading marine bacteria. Curr. Opin. Biotechnol. 2007, 18, 257–266. [Google Scholar] [CrossRef]
- Dombrowski, N.; Donaho, J.A.; Gutierrez, T.; Seitz, K.W.; Teske, A.P.; Baker, B.J. Reconstructing metabolic pathways of hydrocarbon-degrading bacteria from the Deepwater Horizon oil spill. Nat. Microbiol. 2016, 1, 16057. [Google Scholar] [CrossRef]
- Dellagnezze, B.; Vasconcellos, S.; Angelim, A.; Melo, V.; Santisi, S.; Cappello, S.; Oliveira, V. Bioaugmentation strategy employing a microbial consortium immobilized in chitosan beads for oil degradation in mesocosm scale. Mar. Pollut. Bull. 2016, 107, 107–117. [Google Scholar] [CrossRef] [PubMed]
- Dubinsky, E.A.; Conrad, M.E.; Chakraborty, R.; Bill, M.; Borglin, S.E.; Hollibaugh, J.T.; Mason, O.U.; Piceno, Y.M.; Reid, F.C.; Stringfellow, W.T.; et al. Succession of hydrocarbon-degrading bacteria in the aftermath of the Deepwater Horizon oil spill in the Gulf of Mexico. Environ. Sci. Technol. 2013, 47, 10860–10867. [Google Scholar] [CrossRef]
- Duran, R.; Cravo-Laureau, C. Role of environmental factors and microorganisms in determining the fate of polycyclic aromatic hydrocarbons in the marine environment. FEMS Microbiol. Rev. 2016, 40, 814–830. [Google Scholar] [CrossRef] [PubMed]
- Gittel, A.; Donhauser, J.; Røy, H.; Girguis, P.R.; Jørgensen, B.B.; Kjeldsen, K.U. Ubiquitous presence and novel diversity of anaerobic alkane degraders in cold marine sediments. Front. Microbiol. 2015, 6, 1414. [Google Scholar] [CrossRef]
- Gertler, C.; Bargiela, R.; Mapelli, F.; Han, X.; Chen, J.; Hai, T.; Amer, R.A.; Mahjoubi, M.; Malkawi, H.; Magagnini, M. Conversion of uric acid into ammonium in oil-degrading marine microbial communities: A possible role of Halomonads. Microb. Ecol. 2015, 70, 724–740. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, T.; Singleton, D.R.; Berry, D.; Yang, T.; Aitken, M.D.; Teske, A. Hydrocarbon-degrading bacteria enriched by the Deepwater Horizon oil spill identified by cultivation and DNA-SIP. ISME J. 2013, 7, 2091–2104. [Google Scholar] [CrossRef] [PubMed]
- Hecht, H.; Srebnik, S. Structural characterization of sodium alginate and calcium alginate. Biomacromolecules 2016, 17, 2160–2167. [Google Scholar] [CrossRef]
- Hazen, T.C.; Dubinsky, E.A.; DeSantis, T.Z.; Andersen, G.L.; Piceno, Y.M.; Singh, N.; Jansson, J.K.; Probst, A.; Borglin, S.E.; Fortney, J.L. Deep-sea oil plume enriches indigenous oil-degrading bacteria. Science 2010, 330, 204–208. [Google Scholar] [CrossRef]
- Hu, G.; Li, J.; Zeng, G. Recent development in the treatment of oily sludge from petroleum industry: A review. J. Hazard. Mater. 2013, 261, 470–490. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.-Y.; Dong, L.; Zhao, J.-K.; Hu, X.; Shen, C.; Qiao, Y.; Zhang, X.; Wang, Y.; Ismagilov, R.F.; Liu, S.-J. High-throughput single-cell cultivation on microfluidic streak plates. Appl. Environ. Microbiol. 2016, 82, 2210–2218. [Google Scholar] [CrossRef]
- Jin, H.M.; Jeong, H.I.; Kim, K.H.; Hahn, Y.; Madsen, E.L.; Jeon, C.O. Genome-wide transcriptional responses of Alteromonas naphthalenivorans SN2 to contaminated seawater and marine tidal flat sediment. Sci. Rep. 2016, 6, 21796. [Google Scholar] [CrossRef] [PubMed]
- Kimes, N.E.; Callaghan, A.V.; Aktas, D.F.; Smith, W.L.; Sunner, J.; Golding, B.T.; Drozdowska, M.; Hazen, T.C.; Suflita, J.M.; Morris, P.J. Metagenomic analysis and metabolite profiling of deep–sea sediments from the Gulf of Mexico following the Deepwater Horizon oil spill. Front. Microbiol. 2013, 4, 50. [Google Scholar] [CrossRef] [PubMed]
- Kostka, J.E.; Prakash, O.; Overholt, W.A.; Green, S.J.; Freyer, G.; Canion, A.; Delgardio, J.; Norton, N.; Hazen, T.C.; Huettel, M. Hydrocarbon-degrading bacteria and the bacterial community response in Gulf of Mexico beach sands impacted by the Deepwater Horizon oil spill. Appl. Environ. Microbiol. 2011, 77, 7962–7974. [Google Scholar] [CrossRef]
- Kube, M.; Chernikova, T.N.; Al-Ramahi, Y.; Beloqui, A.; Lopez-Cortez, N.; Guazzaroni, M.-E.; Heipieper, H.J.; Klages, S.; Kotsyurbenko, O.R.; Langer, I. Genome sequence and functional genomic analysis of the oil-degrading bacterium Oleispira antarctica. Nat. Commun. 2013, 4, 2156. [Google Scholar] [CrossRef] [PubMed]
- Lea-Smith, D.J.; Biller, S.J.; Davey, M.P.; Cotton, C.A.; Sepulveda, B.M.P.; Turchyn, A.V.; Scanlan, D.J.; Smith, A.G.; Chisholm, S.W.; Howe, C.J. Contribution of cyanobacterial alkane production to the ocean hydrocarbon cycle. Proc. Natl. Acad. Sci. USA 2015, 112, 13591–13596. [Google Scholar] [CrossRef]
- Liu, H.; Xu, J.; Liang, R.; Liu, J. Characterization of the medium-and long-chain n-alkanes degrading Pseudomonas aeruginosa strain SJTD-1 and its alkane hydroxylase genes. PLoS ONE 2014, 9, e105506. [Google Scholar] [CrossRef]
- Macgregor, R.B., Jr. The interactions of nucleic acids at elevated hydrostatic pressure. Biochim. Biophys. Acta (BBA)-Protein Struct. Mol. Enzymol. 2002, 1595, 266–276. [Google Scholar] [CrossRef]
- Martin, D.; Bartlett, D.H.; Roberts, M.F. Solute accumulation in the deep-sea bacterium Photobacterium profundum. Extremophiles 2002, 6, 507–514. [Google Scholar] [CrossRef]
- Mason, O.U.; Scott, N.M.; Gonzalez, A.; Robbins-Pianka, A.; Bælum, J.; Kimbrel, J.; Bouskill, N.J.; Prestat, E.; Borglin, S.; Joyner, D.C. Metagenomics reveals sediment microbial community response to Deepwater Horizon oil spill. ISME J. 2014, 8, 1464–1475. [Google Scholar] [CrossRef]
- Meng, L.; Liu, H.; Bao, M.; Sun, P. Microbial community structure shifts are associated with temperature, dispersants and nutrients in crude oil-contaminated seawaters. Mar. Pollut. Bull. 2016, 111, 203–212. [Google Scholar] [CrossRef]
- Messina, E.; Denaro, R.; Crisafi, F.; Smedile, F.; Cappello, S.; Genovese, M.; Genovese, L.; Giuliano, L.; Russo, D.; Ferrer, M. Genome sequence of obligate marine polycyclic aromatic hydrocarbons-degrading bacterium Cycloclasticus sp. 78-ME, isolated from petroleum deposits of the sunken tanker Amoco Milford Haven, Mediterranean Sea. Mar. Genom. 2016, 25, 11–13. [Google Scholar] [CrossRef]
- Nikolopoulou, M.; Pasadakis, N.; Kalogerakis, N. Evaluation of autochthonous bioaugmentation and biostimulation during microcosm-simulated oil spills. Mar. Pollut. Bull. 2013, 72, 165–173. [Google Scholar] [CrossRef] [PubMed]
- Mulet, M.; David, Z.; Nogales, B.; Bosch, R.; Lalucat, J.; García-Valdés, E. Pseudomonas diversity in crude-oil-contaminated intertidal sand samples obtained after the Prestige oil spill. Appl. Environ. Microbiol. 2011, 77, 1076–1085. [Google Scholar] [CrossRef] [PubMed]
- Saini, S.; Dhania, G. Cadmium as an environmental pollutant: Ecotoxicological effects, health hazards, and bioremediation approaches for its detoxification from contaminated sites. In Bioremediation of Industrial Waste for Environmental Safety; Springer: Berlin/Heidelberg, Germany, 2020; pp. 357–387. [Google Scholar]
- Xu, X.; Liu, W.; Tian, S.; Wang, W.; Qi, Q.; Jiang, P.; Gao, X.; Li, F.; Li, H.; Yu, H. Petroleum hydrocarbon-degrading bacteria for the remediation of oil pollution under aerobic conditions: A perspective analysis. Front. Microbiol. 2018, 9, 2885. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Li, J.; Liu, M.; Sun, H.; Bao, M. Study on the biodegradation of crude oil by free and immobilized bacterial consortium in marine environment. PLoS ONE 2017, 12, e0174445. [Google Scholar] [CrossRef]
- Khan, B.; Bilal Khan Niazi, M.; Samin, G.; Jahan, Z. Thermoplastic starch: A possible biodegradable food packaging material—a review. J. Food Process Eng. 2017, 40, e12447. [Google Scholar] [CrossRef]
- Saikia, K.; Rathankumar, A.K.; Vaithyanathan, V.K.; Cabana, H.; Vaidyanathan, V.K. Preparation of highly diffusible porous cross-linked lipase B from Candida antarctica conjugates: Advances in mass transfer and application in transesterification of 5-Hydroxymethylfurfural. Int. J. Biol. Macromol. 2021, 170, 583–592. [Google Scholar] [CrossRef]
- Atashgahi, S.; Sánchez-Andrea, I.; Heipieper, H.J.; van der Meer, J.R.; Stams, A.J.; Smidt, H. Prospects for harnessing biocide resistance for bioremediation and detoxification. Science 2018, 360, 743–746. [Google Scholar] [CrossRef]
- Passow, U.; Stout, S.A. Character and sedimentation of “lingering” Macondo oil to the deep-sea after the Deepwater Horizon oil spill. Mar. Chem. 2020, 218, 103733. [Google Scholar] [CrossRef]
- Yang, T.; Nigro, L.M.; Gutierrez, T.; Joye, S.B.; Highsmith, R.; Teske, A. Pulsed blooms and persistent oil-degrading bacterial populations in the water column during and after the Deepwater Horizon blowout. Deep. Sea Res. Part II Top. Stud. Oceanogr. 2016, 129, 282–291. [Google Scholar] [CrossRef]
- Ullrich, S.R.; Poehlein, A.; Tischler, J.S.; González, C.; Ossandon, F.J.; Daniel, R.; Holmes, D.S.; Schlömann, M.; Mühling, M. Genome analysis of the biotechnologically relevant acidophilic iron oxidising strain JA12 indicates phylogenetic and metabolic diversity within the novel genus “Ferrovum”. PLoS ONE 2016, 11, e0146832. [Google Scholar] [CrossRef] [PubMed]
- Warwick, R.; Clarke, K. Relearning the ABC: Taxonomic changes and abundance/biomass relationships in disturbed benthic communities. Mar. Biol. 1994, 118, 739–744. [Google Scholar] [CrossRef]
- Sanyal, O.; Shinde, V.L.; Meena, R.M.; Damare, S.; Shenoy, B.D. The ITS-based phylogeny of fungi associated with tarballs. Mar. Pollut. Bull. 2016, 113, 277–281. [Google Scholar] [CrossRef]
- Nazina, T.N.; Shestakova, N.M.; Semenova, E.M.; Korshunova, A.V.; Kostrukova, N.K.; Tourova, T.P.; Min, L.; Feng, Q.; Poltaraus, A.B. Diversity of metabolically active bacteria in water-flooded high-temperature heavy oil reservoir. Front. Microbiol. 2017, 8, 707. [Google Scholar] [CrossRef] [PubMed]
- Daccò, C.; Girometta, C.; Asemoloye, M.; Carpani, G.; Picco, A.; Tosi, S. Key fungal degradation patterns, enzymes and their applications for the removal of aliphatic hydrocarbons in polluted soils: A review. Int. Biodeterior. Biodegrad. 2020, 147, 104866. [Google Scholar] [CrossRef]
- Varjani, S.J.; Upasani, V.N. A new look on factors affecting microbial degradation of petroleum hydrocarbon pollutants. Int. Biodeterior. Biodegrad. 2017, 120, 71–83. [Google Scholar] [CrossRef]
- Fatima, K.; Imran, A.; Naveed, M.; Afzal, M. Plant-bacteria synergism: An innovative approach for the remediation of crude oil-contaminated soils. Soil Environ. 2017, 36, 93–113. [Google Scholar] [CrossRef]
- Fathepure, B.Z. Recent studies in microbial degradation of petroleum hydrocarbons in hypersaline environments. Front. Microbiol. 2014, 5, 173. [Google Scholar] [CrossRef]
- Ebadi, A.; Sima, N.A.K.; Olamaee, M.; Hashemi, M.; Nasrabadi, R.G. Effective bioremediation of a petroleum-polluted saline soil by a surfactant-producing Pseudomonas aeruginosa consortium. J. Adv. Res. 2017, 8, 627–633. [Google Scholar] [CrossRef]
- Hidayat, A.; Tachibana, S. Biodegradation of aliphatic hydrocarbon in three types of crude oil by Fusarium sp. F 092 under stress with artificial sea water. J. Environ. Sci. Technol. 2012, 5, 64–73. [Google Scholar] [CrossRef]
- AI-Jawhari, I.F.H. Ability of some soil fungi in biodegradation of petroleum hydrocarbon. J. Appl. Environ. Microbiol. 2014, 2, 46–52. [Google Scholar]
- Ravindran, A.; Sajayan, A.; Priyadharshini, G.B.; Selvin, J.; Kiran, G.S. Revealing the efficacy of thermostable biosurfactant in heavy metal bioremediation and surface treatment in vegetables. Front. Microbiol. 2020, 11, 222. [Google Scholar] [CrossRef] [PubMed]
- Srogi, K. Monitoring of environmental exposure to polycyclic aromatic hydrocarbons: A review. Environ. Chem. Lett. 2007, 5, 169–195. [Google Scholar] [CrossRef]
- Abdel-Shafy, H.I.; Mansour, M.S. A review on polycyclic aromatic hydrocarbons: Source, environmental impact, effect on human health and remediation. Egypt. J. Pet. 2016, 25, 107–123. [Google Scholar] [CrossRef]
- Gupte, A.; Tripathi, A.; Patel, H.; Rudakiya, D.; Gupte, S. Bioremediation of polycyclic aromatic hydrocarbon (PAHs): A perspective. Open Biotechnol. J. 2016, 10, 363–378. [Google Scholar] [CrossRef]
- Mojiri, A.; Zhou, J.L.; Ohashi, A.; Ozaki, N.; Kindaichi, T. Comprehensive review of polycyclic aromatic hydrocarbons in water sources, their effects and treatments. Sci. Total Environ. 2019, 696, 133971. [Google Scholar] [CrossRef]
- St. Helen, G.; Goniewicz, M.L.; Dempsey, D.; Wilson, M.; Jacob, P., III; Benowitz, N.L. Exposure and kinetics of polycyclic aromatic hydrocarbons (PAHs) in cigarette smokers. Chem. Res. Toxicol. 2012, 25, 952–964. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.; Liu, E.; Zhang, E.; Luo, W.; Chen, L.; Wang, C.; Lin, Q. Historical records and sources of polycyclic aromatic hydrocarbons (PAHs) and organochlorine pesticides (OCPs) in sediment from a representative plateau lake, China. Chemosphere 2017, 173, 78–88. [Google Scholar] [CrossRef]
- Johnsen, A.R.; Karlson, U. Diffuse PAH contamination of surface soils: Environmental occurrence, bioavailability, and microbial degradation. Appl. Microbiol. Biotechnol. 2007, 76, 533–543. [Google Scholar] [CrossRef]
- Kozak, K.; Ruman, M.; Kosek, K.; Karasiński, G.; Stachnik, Ł.; Polkowska, Ż. Impact of volcanic eruptions on the occurrence of PAHs compounds in the aquatic ecosystem of the southern part of West Spitsbergen (Hornsund Fjord, Svalbard). Water 2017, 9, 42. [Google Scholar] [CrossRef]
- Adams, G.O.; Fufeyin, P.T.; Okoro, S.E.; Ehinomen, I. Bioremediation, biostimulation and bioaugmention: A review. Int. J. Environ. Bioremediation Biodegrad. 2015, 3, 28–39. [Google Scholar] [CrossRef]
- Vogt, C.; Richnow, H.H. Bioremediation via in situ microbial degradation of organic pollutants. Geobiotechnology II 2013, 142, 123–146. [Google Scholar]
- Kumar, V.; Shahi, S.; Singh, S. Bioremediation: An eco-sustainable approach for restoration of contaminated sites. In Microbial Bioprospecting for Sustainable Development; Springer: Berlin/Heidelberg, Germany, 2018; pp. 123–146. [Google Scholar]
- Lovley, D.R. Live wires: Direct extracellular electron exchange for bioenergy and the bioremediation of energy-related contamination. Energy Environ. Sci. 2011, 4, 4896–4906. [Google Scholar] [CrossRef]
- Militon, C.; Jézéquel, R.; Gilbert, F.; Corsellis, Y.; Sylvi, L.; Cravo-Laureau, C.; Duran, R.; Cuny, P. Dynamics of bacterial assemblages and removal of polycyclic aromatic hydrocarbons in oil-contaminated coastal marine sediments subjected to contrasted oxygen regimes. Environ. Sci. Pollut. Res. 2015, 22, 15260–15272. [Google Scholar] [CrossRef]
- Mukherjee, A.K.; Das, K. Correlation between diverse cyclic lipopeptides production and regulation of growth and substrate utilization by Bacillus subtilis strains in a particular habitat. FEMS Microbiol. Ecol. 2005, 54, 479–489. [Google Scholar] [CrossRef]
- Sayed, K.; Baloo, L.; Sharma, N.K. Bioremediation of total petroleum hydrocarbons (TPH) by bioaugmentation and biostimulation in water with floating oil spill containment booms as bioreactor basin. Int. J. Environ. Res. Public Health 2021, 18, 2226. [Google Scholar] [CrossRef]
- Tyagi, M.; da Fonseca, M.M.R.; de Carvalho, C.C. Bioaugmentation and biostimulation strategies to improve the effectiveness of bioremediation processes. Biodegradation 2011, 22, 231–241. [Google Scholar] [CrossRef] [PubMed]
- El Fantroussi, S.; Agathos, S.N. Is bioaugmentation a feasible strategy for pollutant removal and site remediation? Curr. Opin. Microbiol. 2005, 8, 268–275. [Google Scholar] [CrossRef] [PubMed]
- Hazra, C.; Kundu, D.; Chaudhari, A. Biosurfactant-assisted bioaugmentation in bioremediation. In Microorganisms in Environmental Management; Satyanarayana, T., Johri, B., Eds.; Springer: Dordrecht, The Netherlands, 2012; pp. 631–664. [Google Scholar]
- Alisi, C.; Musella, R.; Tasso, F.; Ubaldi, C.; Manzo, S.; Cremisini, C.; Sprocati, A.R. Bioremediation of diesel oil in a co-contaminated soil by bioaugmentation with a microbial formula tailored with native strains selected for heavy metals resistance. Sci. Total Environ. 2009, 407, 3024–3032. [Google Scholar] [CrossRef]
- Rahman, K.; Banat, I.; Thahira, J.; Thayumanavan, T.; Lakshmanaperumalsamy, P. Bioremediation of gasoline contaminated soil by a bacterial consortium amended with poultry litter, coir pith and rhamnolipid biosurfactant. Bioresour. Technol. 2002, 81, 25–32. [Google Scholar] [CrossRef]
- Li, X.; Lin, X.; Li, P.; Liu, W.; Wang, L.; Ma, F.; Chukwuka, K. Biodegradation of the low concentration of polycyclic aromatic hydrocarbons in soil by microbial consortium during incubation. J. Hazard. Mater. 2009, 172, 601–605. [Google Scholar] [CrossRef]
- Goldstein, R.M.; Mallory, L.M.; Alexander, M. Reasons for possible failure of inoculation to enhance biodegradation. Appl. Environ. Microbiol. 1985, 50, 977–983. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.; Ke, L.; Wong, Y.; Tam, N. Degradation of polycyclic aromatic hydrocarbons by a bacterial consortium enriched from mangrove sediments. Environ. Int. 2005, 31, 149–154. [Google Scholar] [CrossRef] [PubMed]
- Daghio, M.; Tatangelo, V.; Franzetti, A.; Gandolfi, I.; Papacchini, M.; Careghini, A.; Sezenna, E.; Saponaro, S.; Bestetti, G. Hydrocarbon degrading microbial communities in bench scale aerobic biobarriers for gasoline contaminated groundwater treatment. Chemosphere 2015, 130, 34–39. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.; Liu, C.; Liu, L.; Zhang, Y.; Liu, Q.; Wu, W.-M. Selection of functional consortium for crude oil-contaminated soil remediation. Int. Biodeterior. Biodegrad. 2011, 65, 1244–1248. [Google Scholar] [CrossRef]
- Wittich, R.-M.; Wolff, P. Growth of the genetically engineered strain Cupriavidus necator RW112 with chlorobenzoates and technical chlorobiphenyls. Microbiology 2007, 153, 186–195. [Google Scholar] [CrossRef][Green Version]
- Varjani, S.J.; Rana, D.P.; Jain, A.K.; Bateja, S.; Upasani, V.N. Synergistic ex-situ biodegradation of crude oil by halotolerant bacterial consortium of indigenous strains isolated from on shore sites of Gujarat, India. Int. Biodeterior. Biodegrad. 2015, 103, 116–124. [Google Scholar] [CrossRef]
- Mancera-López, M.; Esparza-García, F.; Chávez-Gómez, B.; Rodríguez-Vázquez, R.; Saucedo-Castaneda, G.; Barrera-Cortés, J. Bioremediation of an aged hydrocarbon-contaminated soil by a combined system of biostimulation–bioaugmentation with filamentous fungi. Int. Biodeterior. Biodegrad. 2008, 61, 151–160. [Google Scholar] [CrossRef]
- Supaphol, S.; Panichsakpatana, S.; Trakulnaleamsai, S.; Tungkananuruk, N.; Roughjanajirapa, P.; O’Donnell, A.G. The selection of mixed microbial inocula in environmental biotechnology: Example using petroleum contaminated tropical soils. J. Microbiol. Methods 2006, 65, 432–441. [Google Scholar] [CrossRef]
- Silva, Í.S.; dos Santos, E.d.C.; de Menezes, C.R.; de Faria, A.F.; Franciscon, E.; Grossman, M.; Durrant, L.R. Bioremediation of a polyaromatic hydrocarbon contaminated soil by native soil microbiota and bioaugmentation with isolated microbial consortia. Bioresour. Technol. 2009, 100, 4669–4675. [Google Scholar] [CrossRef]
- Jacques, R.J.; Okeke, B.C.; Bento, F.M.; Teixeira, A.S.; Peralba, M.C.; Camargo, F.A. Microbial consortium bioaugmentation of a polycyclic aromatic hydrocarbons contaminated soil. Bioresour. Technol. 2008, 99, 2637–2643. [Google Scholar] [CrossRef] [PubMed]
- Das, K.; Mukherjee, A.K. Crude petroleum-oil biodegradation efficiency of Bacillus subtilis and Pseudomonas aeruginosa strains isolated from a petroleum-oil contaminated soil from North-East India. Bioresour. Technol. 2007, 98, 1339–1345. [Google Scholar] [CrossRef] [PubMed]
- Brzeszcz, J.; Kapusta, P.; Steliga, T.; Turkiewicz, A. Hydrocarbon removal by two differently developed microbial inoculants and comparing their actions with biostimulation treatment. Molecules 2020, 25, 661. [Google Scholar] [CrossRef] [PubMed]
- Hassanshahian, M.; Yakimov, M.M.; Denaro, R.; Genovese, M.; Cappello, S. Using Real-Time PCR to assess changes in the crude oil degrading microbial community in contaminated seawater mesocosms. Int. Biodeterior. Biodegrad. 2014, 93, 241–248. [Google Scholar] [CrossRef]
- Al-Hawash, A.B.; Dragh, M.A.; Li, S.; Alhujaily, A.; Abbood, H.A.; Zhang, X.; Ma, F. Principles of microbial degradation of petroleum hydrocarbons in the environment. Egypt. J. Aquat. Res. 2018, 44, 71–76. [Google Scholar] [CrossRef]
- Orji, F.A.; Ibiene, A.A.; Dike, E.N. Laboratory scale bioremediation of petroleum hydrocarbon–polluted mangrove swamps in the Niger Delta using cow dung. Malays. J. Microbiol. 2012, 8, 219–228. [Google Scholar]
- Ponsin, V.; Coulomb, B.; Guelorget, Y.; Maier, J.; Höhener, P. In situ biostimulation of petroleum hydrocarbon degradation by nitrate and phosphate injection using a dipole well configuration. J. Contam. Hydrol. 2014, 171, 22–31. [Google Scholar] [CrossRef]
- Shivlata, L.; Tulasi, S. Thermophilic and alkaliphilic Actinobacteria: Biology and potential applications. Front. Microbiol. 2015, 6, 1014. [Google Scholar] [CrossRef]
- Rodrigues, L.R. Microbial surfactants: Fundamentals and applicability in the formulation of nano-sized drug delivery vectors. J. Colloid Interface Sci. 2015, 449, 304–316. [Google Scholar] [CrossRef]
- Cheng, Y.; He, H.; Yang, C.; Zeng, G.; Li, X.; Chen, H.; Yu, G. Challenges and solutions for biofiltration of hydrophobic volatile organic compounds. Biotechnol. Adv. 2016, 34, 1091–1102. [Google Scholar] [CrossRef]
- Kim, L.H.; Jung, Y.; Yu, H.-W.; Chae, K.-J.; Kim, I.S. Physicochemical interactions between rhamnolipids and Pseudomonas aeruginosa biofilm layers. Environ. Sci. Technol. 2015, 49, 3718–3726. [Google Scholar] [CrossRef] [PubMed]
- Anjum, F.; Gautam, G.; Edgard, G.; Negi, S. Biosurfactant production through Bacillus sp. MTCC 5877 and its multifarious applications in food industry. Bioresour. Technol. 2016, 213, 262–269. [Google Scholar] [CrossRef] [PubMed]
- Gregorich, E.; Gillespie, A.; Beare, M.; Curtin, D.; Sanei, H.; Yanni, S. Evaluating biodegradability of soil organic matter by its thermal stability and chemical composition. Soil Biol. Biochem. 2015, 91, 182–191. [Google Scholar] [CrossRef]
- Patel, J.; Borgohain, S.; Kumar, M.; Rangarajan, V.; Somasundaran, P.; Sen, R. Recent developments in microbial enhanced oil recovery. Renew. Sustain. Energy Rev. 2015, 52, 1539–1558. [Google Scholar] [CrossRef]
- Itrich, N.R.; McDonough, K.M.; van Ginkel, C.G.; Bisinger, E.C.; LePage, J.N.; Schaefer, E.C.; Menzies, J.Z.; Casteel, K.D.; Federle, T.W. Widespread microbial adaptation to l-glutamate-N, N-diacetate (L-GLDA) following its market introduction in a consumer cleaning product. Environ. Sci. Technol. 2015, 49, 13314–13321. [Google Scholar] [CrossRef] [PubMed]
- Joy, S.; Rahman, P.K.; Sharma, S. Biosurfactant production and concomitant hydrocarbon degradation potentials of bacteria isolated from extreme and hydrocarbon contaminated environments. Chem. Eng. J. 2017, 317, 232–241. [Google Scholar] [CrossRef]
- Amani, H.; Müller, M.M.; Syldatk, C.; Hausmann, R. Production of microbial rhamnolipid by Pseudomonas aeruginosa MM1011 for ex situ enhanced oil recovery. Appl. Biochem. Biotechnol. 2013, 170, 1080–1093. [Google Scholar] [CrossRef]
- Goldman, S.; Shabtai, Y.; Rubinovitz, C.; Rosenberg, E.; Gutnick, D. Emulsan in Acinetobacter calcoaceticus RAG-1: Distribution of cell-free and cell-associated cross-reacting material. Appl. Environ. Microbiol. 1982, 44, 165–170. [Google Scholar] [CrossRef]
- Urum, K.; Pekdemir, T. Evaluation of biosurfactants for crude oil contaminated soil washing. Chemosphere 2004, 57, 1139–1150. [Google Scholar] [CrossRef] [PubMed]
- Morita, T.; Konishi, M.; Fukuoka, T.; Imura, T.; Kitamoto, D. Microbial conversion of glycerol into glycolipid biosurfactants, mannosylerythritol lipids, by a basidiomycete yeast, Pseudozyma antarctica JCM 10317T. J. Biosci. Bioeng. 2007, 104, 78–81. [Google Scholar] [CrossRef]
- Camacho-Chab, J.C.; Guézennec, J.; Chan-Bacab, M.J.; Ríos-Leal, E.; Sinquin, C.; Muñiz-Salazar, R.; Rosa-García, S.D.C.D.L.; Reyes-Estebanez, M.; Ortega-Morales, B.O. Emulsifying activity and stability of a non-toxic bioemulsifier synthesized by Microbacterium sp. MC3B-10. Int. J. Mol. Sci. 2013, 14, 18959–18972. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Y.; Xiao, F.; Wei, X.; Wen, Z.; Chen, S. Improvement of lichenysin production in Bacillus licheniformis by replacement of native promoter of lichenysin biosynthesis operon and medium optimization. Appl. Microbiol. Biotechnol. 2014, 98, 8895–8903. [Google Scholar] [CrossRef] [PubMed]
- Solaiman, D.K.; Ashby, R.D.; Zerkowski, J.A.; Foglia, T.A. Simplified soy molasses-based medium for reduced-cost production of sophorolipids by Candida bombicola. Biotechnol. Lett. 2007, 29, 1341–1347. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.-S.; Kim, E.-K. Lipopeptide production from Bacillus sp. GB16 using a novel oxygenation method. Enzym. Microb. Technol. 2004, 35, 639–647. [Google Scholar] [CrossRef]
- Rahmati, F. Microencapsulation of Lactobacillus acidophilus and Lactobacillus plantarum in Eudragit S100 and alginate chitosan under gastrointestinal and normal conditions. Appl. Nanosci. 2020, 10, 391–399. [Google Scholar] [CrossRef]
- Nwankwegu, A.S.; Onwosi, C.O. Microbial cell immobilization: A renaissance to bioaugmentation inadequacies. A review. Environ. Technol. Rev. 2017, 6, 186–198. [Google Scholar] [CrossRef]
- Rahmati, F. Impact of microencapsulation on two probiotic strains in alginate chitosan and Eudragit S100 under gastrointestinal and normal conditions. Open Biotechnol. J. 2019, 13, 59–67. [Google Scholar] [CrossRef]
- Moslemy, P.; Neufeld, R.J.; Guiot, S.R. Biodegradation of gasoline by gellan gum-encapsulated bacterial cells. Biotechnol. Bioeng. 2002, 80, 175–184. [Google Scholar] [CrossRef]
- Liu, Y.; Zhang, A.; Wang, X. Biodegradation of phenol by using free and immobilized cells of Acinetobacter sp. XA05 and Sphingomonas sp. FG03. Biochem. Eng. J. 2009, 44, 187–192. [Google Scholar] [CrossRef]
- Xu, Y.; Lu, M. Bioremediation of crude oil-contaminated soil: Comparison of different biostimulation and bioaugmentation treatments. J. Hazard. Mater. 2010, 183, 395–401. [Google Scholar] [CrossRef]
- Wang, S.; Li, X.; Liu, W.; Li, P.; Kong, L.; Ren, W.; Wu, H.; Tu, Y. Degradation of pyrene by immobilized microorganisms in saline-alkaline soil. J. Environ. Sci. 2012, 24, 1662–1669. [Google Scholar] [CrossRef]
- Simons, K.L.; Ansar, A.; Kadali, K.; Bueti, A.; Adetutu, E.M.; Ball, A.S. Investigating the effectiveness of economically sustainable carrier material complexes for marine oil remediation. Bioresour. Technol. 2012, 126, 202–207. [Google Scholar] [CrossRef] [PubMed]
- El Mahdi, A.M.; Aziz, H.A.; Abu Amr, S.S.; El-Gendy, N.S.; Nassar, H.N. Isolation and characterization of Pseudomonas sp. NAF1 and its application in biodegradation of crude oil. Environ. Earth Sci. 2016, 75, 380. [Google Scholar] [CrossRef]
- Ławniczak, Ł.; Woźniak-Karczewska, M.; Loibner, A.P.; Heipieper, H.J.; Chrzanowski, Ł. Microbial degradation of hydrocarbons—basic principles for bioremediation: A review. Molecules 2020, 25, 856. [Google Scholar] [CrossRef] [PubMed]
- Amenaghawon, A.N.; Osunbor, O.; Obahiagbon, K.O. Impact of nutrients, aeration and agitation on the bioremediation of crude oil polluted water using mixed microbial culture. Int. J. Sci. Res. Environ. Sci. 2014, 2, 43. [Google Scholar] [CrossRef]
- Martins, S.C.S.; Martins, C.M.; Fiúza, L.M.C.G.; Santaella, S.T. Immobilization of microbial cells: A promising tool for treatment of toxic pollutants in industrial wastewater. Afr. J. Biotechnol. 2013, 12, 4412–4418. [Google Scholar]
- Messing, R.; Oppermann, R.; Kolot, F. Pore dimensions for accumulating biomass. II. Microbes that form spores and exhibit mycelial growth. Biotechnol. Bioeng. 1979, 21, 59–67. [Google Scholar] [CrossRef]
- Meng, L.; Li, W.; Bao, M.; Sun, P. Promoting the treatment of crude oil alkane pollution through the study of enzyme activity. Int. J. Biol. Macromol. 2018, 119, 708–716. [Google Scholar] [CrossRef]
- Xu, A.; Di Wang, Y.D.; Zheng, Y.; Wang, B.; Wei, Q.; Wang, S.; Yang, L.; Ma, L.Z. Integrated comparative genomic analysis and phenotypic profiling of Pseudomonas aeruginosa isolates from crude oil. Front. Microbiol. 2020, 11, 519. [Google Scholar] [CrossRef]
- Salway, J.G. Metabolism at a Glance; John Wiley & Sons: Hoboken, NJ, USA, 2016. [Google Scholar]
- Cruz Viggi, C.; Presta, E.; Bellagamba, M.; Kaciulis, S.; Balijepalli, S.K.; Zanaroli, G.; Petrangeli Papini, M.; Rossetti, S.; Aulenta, F. The “Oil-Spill Snorkel”: An innovative bioelectrochemical approach to accelerate hydrocarbons biodegradation in marine sediments. Front. Microbiol. 2015, 6, 881. [Google Scholar] [CrossRef]
- Scoma, A.; Boon, N. Osmotic stress confers enhanced cell integrity to hydrostatic pressure but impairs growth in Alcanivorax borkumensis SK2. Front. Microbiol. 2016, 7, 729. [Google Scholar] [CrossRef] [PubMed]
- Scoma, A.; Garrido-Amador, P.; Nielsen, S.D.; Røy, H.; Kjeldsen, K.U. The polyextremophilic bacterium Clostridium paradoxum attains piezophilic traits by modulating its energy metabolism and cell membrane composition. Appl. Environ. Microbiol. 2019, 85, e00802-19. [Google Scholar] [CrossRef] [PubMed]
- Espínola, F.; Dionisi, H.M.; Borglin, S.; Brislawn, C.J.; Jansson, J.K.; Mac Cormack, W.P.; Carroll, J.; Sjöling, S.; Lozada, M. Metagenomic analysis of subtidal sediments from polar and subpolar coastal environments highlights the relevance of anaerobic hydrocarbon degradation processes. Microb. Ecol. 2018, 75, 123–139. [Google Scholar] [CrossRef] [PubMed]
- Gontikaki, E.; Potts, L.; Anderson, J.; Witte, U. Hydrocarbon-degrading bacteria in deep-water subarctic sediments (Faroe-Shetland channel). J. Appl. Microbiol. 2018, 125, 1040–1053. [Google Scholar] [CrossRef] [PubMed]
- Doyle, S.M.; Whitaker, E.A.; De Pascuale, V.; Wade, T.L.; Knap, A.H.; Santschi, P.H.; Quigg, A.; Sylvan, J.B. Rapid formation of microbe-oil aggregates and changes in community composition in coastal surface water following exposure to oil and the dispersant Corexit. Front. Microbiol. 2018, 9, 689. [Google Scholar] [CrossRef] [PubMed]
- Kujawinski, E.B.; Reddy, C.M.; Rodgers, R.P.; Thrash, J.C.; Valentine, D.L.; White, H.K. The first decade of scientific insights from the Deepwater Horizon oil release. Nat. Rev. Earth Environ. 2020, 1, 237–250. [Google Scholar] [CrossRef]
- Naether, D.J.; Slawtschew, S.; Stasik, S.; Engel, M.; Olzog, M.; Wick, L.Y.; Timmis, K.N.; Heipieper, H.J. Adaptation of the hydrocarbonoclastic bacterium Alcanivorax borkumensis SK2 to alkanes and toxic organic compounds: A physiological and transcriptomic approach. Appl. Environ. Microbiol. 2013, 79, 4282–4293. [Google Scholar] [CrossRef]
- Sorokin, D.Y.; Diender, M.; Merkel, A.Y.; Koenen, M.; Bale, N.J.; Pabst, M.; Sinninghe Damsté, J.S.; Sousa, D.Z. Natranaerofaba carboxydovora gen. nov., sp. nov., an extremely haloalkaliphilic CO-utilizing acetogen from a hypersaline soda lake representing a novel deep phylogenetic lineage in the class ‘Natranaerobiia’. Environ. Microbiol. 2020, 23, 3460–3476. [Google Scholar] [CrossRef]
- Kunová, N.; Ondrovičová, G.; Bauer, J.A.; Bellová, J.; Ambro, Ľ.; Martináková, L.; Kotrasová, V.; Kutejová, E.; Pevala, V. The role of Lon-mediated proteolysis in the dynamics of mitochondrial nucleic acid-protein complexes. Sci. Rep. 2017, 7, 631. [Google Scholar] [CrossRef]
- Sharma, I.M.; Woodson, S.A. RbfA and IF3 couple ribosome biogenesis and translation initiation to increase stress tolerance. Nucleic Acids Res. 2020, 48, 359–372. [Google Scholar] [CrossRef]
- Rahmati, F.; Hosseini, S.S.; Safai, S.M.; Lajayer, B.A.; Hatami, M. New insights into the role of nanotechnology in microbial food safety. 3 Biotech 2020, 10, 425. [Google Scholar] [CrossRef] [PubMed]
- Laursen, B.S.; Sørensen, H.P.; Mortensen, K.K.; Sperling-Petersen, H.U. Initiation of protein synthesis in bacteria. Microbiol. Mol. Biol. Rev. 2005, 69, 101–123. [Google Scholar] [CrossRef] [PubMed]
- Kusube, M.; Kyaw, T.S.; Tanikawa, K.; Chastain, R.A.; Hardy, K.M.; Cameron, J.; Bartlett, D.H. Colwellia marinimaniae sp. nov., a hyperpiezophilic species isolated from an amphipod within the Challenger Deep, Mariana Trench. Int. J. Syst. Evol. Microbiol. 2017, 67, 824–831. [Google Scholar] [CrossRef] [PubMed]
- Murali, R. Studies on the Cytochrome Bd-Type Oxygen Reductase Superfamily and the Discovery of a Novel Nitric Oxide Reductase; University of Illinois at Urbana-Champaign: Champaign, IL, USA, 2016. [Google Scholar]
- Boeuf, D.; Edwards, B.R.; Eppley, J.M.; Hu, S.K.; Poff, K.E.; Romano, A.E.; Caron, D.A.; Karl, D.M.; DeLong, E.F. Biological composition and microbial dynamics of sinking particulate organic matter at abyssal depths in the oligotrophic open ocean. Proc. Natl. Acad. Sci. USA 2019, 116, 11824–11832. [Google Scholar] [CrossRef] [PubMed]
- Jebbar, M.; Franzetti, B.; Girard, E.; Oger, P. Microbial diversity and adaptation to high hydrostatic pressure in deep-sea hydrothermal vents prokaryotes. Extremophiles 2015, 19, 721–740. [Google Scholar] [CrossRef]
- Schneiker, S.; Dos Santos, V.A.; Bartels, D.; Bekel, T.; Brecht, M.; Buhrmester, J.; Chernikova, T.N.; Denaro, R.; Ferrer, M.; Gertler, C. Genome sequence of the ubiquitous hydrocarbon-degrading marine bacterium Alcanivorax borkumensis. Nat. Biotechnol. 2006, 24, 997–1004. [Google Scholar] [CrossRef]
- Das, N.; Chandran, P. Microbial degradation of petroleum hydrocarbon contaminants: An overview. Biotechnol. Res. Int. 2011, 2011, 941810. [Google Scholar] [CrossRef]
- Venosa, A.D.; Zhu, X. Biodegradation of crude oil contaminating marine shorelines and freshwater wetlands. Spill Sci. Technol. Bull. 2003, 8, 163–178. [Google Scholar] [CrossRef]
- Acosta-González, A.; Marqués, S. Bacterial diversity in oil-polluted marine coastal sediments. Curr. Opin. Biotechnol. 2016, 38, 24–32. [Google Scholar] [CrossRef]
- Varjani, S.J. Microbial degradation of petroleum hydrocarbons. Bioresour. Technol. 2017, 223, 277–286. [Google Scholar] [CrossRef]
- Su, Z.; Dai, T.; Tang, Y.; Tao, Y.; Huang, B.; Mu, Q.; Wen, D. Sediment bacterial community structures and their predicted functions implied the impacts from natural processes and anthropogenic activities in coastal area. Mar. Pollut. Bull. 2018, 131, 481–495. [Google Scholar] [CrossRef] [PubMed]
- Kotoky, R.; Rajkumari, J.; Pandey, P. The rhizosphere microbiome: Significance in rhizoremediation of polyaromatic hydrocarbon contaminated soil. J. Environ. Manag. 2018, 217, 858–870. [Google Scholar] [CrossRef] [PubMed]
- Shlomi, T.; Cabili, M.N.; Herrgård, M.J.; Palsson, B.Ø.; Ruppin, E. Network-based prediction of human tissue-specific metabolism. Nat. Biotechnol. 2008, 26, 1003–1010. [Google Scholar] [CrossRef] [PubMed]
- Chaffron, S.; Rehrauer, H.; Pernthaler, J.; Von Mering, C. A global network of coexisting microbes from environmental and whole-genome sequence data. Genome Res. 2010, 20, 947–959. [Google Scholar] [CrossRef]
- Langille, M.G.I.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Thurber, R.L.V.; Knight, R. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef]
- Freeman, K.R.; Pescador, M.Y.; Reed, S.C.; Costello, E.K.; Robeson, M.S.; Schmidt, S.K. Soil CO2 flux and photoautotrophic community composition in high-elevation,‘barren’ soil. Environ. Microbiol. 2009, 11, 674–686. [Google Scholar] [CrossRef]
- Zafra, G.; Absalón, A.E.; Cortés-Espinosa, D.V. Morphological changes and growth of filamentous fungi in the presence of high concentrations of PAHs. Braz. J. Microbiol. 2015, 46, 937–941. [Google Scholar] [CrossRef]
- Kalantary, R.R.; Mohseni-Bandpei, A.; Esrafili, A.; Nasseri, S.; Ashmagh, F.R.; Jorfi, S.; Ja’Fari, M. Effectiveness of biostimulation through nutrient content on the bioremediation of phenanthrene contaminated soil. J. Environ. Health Sci. Eng. 2014, 12, 143. [Google Scholar] [CrossRef]
- Patel, A.B.; Shaikh, S.; Jain, K.R.; Desai, C.; Madamwar, D. Polycyclic aromatic hydrocarbons: Sources, toxicity and remediation approaches. Front. Microbiol. 2020, 11, 2675. [Google Scholar] [CrossRef]
- Vermelho, A.B.; Supuran, C.T.; Guisan, J.M. Microbial enzyme: Applications in industry and in bioremediation. Enzym. Res. 2012, 2012, 980681. [Google Scholar] [CrossRef][Green Version]
- Madigan, M.; Martinko, J.; Dunlap, P.; Clark, D. Brock Biology of Microorganisms; Benjamin Cummings: San Francisco, CA, USA,, 2010. [Google Scholar]
- Rabus, R.; Boll, M.; Heider, J.; Meckenstock, R.U.; Buckel, W.; Einsle, O.; Ermler, U.; Golding, B.T.; Gunsalus, R.P.; Kroneck, P.M. Anaerobic microbial degradation of hydrocarbons: From enzymatic reactions to the environment. J. Mol. Microbiol. Biotechnol. 2016, 26, 5–28. [Google Scholar] [CrossRef] [PubMed]
- Van Beilen, J.B.; Funhoff, E.G. Alkane hydroxylases involved in microbial alkane degradation. Appl. Microbiol. Biotechnol. 2007, 74, 13–21. [Google Scholar] [CrossRef] [PubMed]
- Van Hamme, J.D.; Singh, A.; Ward, O.P. Recent advances in petroleum microbiology. Microbiol. Mol. Biol. Rev. 2003, 67, 503–549. [Google Scholar] [CrossRef] [PubMed]
| A/AN | Genome | Microorganism | Phylogeny | Target Substrate | Habitat | References |
|---|---|---|---|---|---|---|
| A | Y | Alcanivorax borkumensis | γ-proteobacteria, Alcanivoracaceae | n-alkanes | Seawater, sediment, beach sand, coastal salt marsh | [8,9] |
| A | Y | Alcanivorax dieselolei | γ-proteobacteria, Alcanivoracaceae | n-alkanes | Seawater, sediment | [10] |
| A | Y | Marinobacter hydrocarbonoclasticus | γ-proteobacteria, Alteromonadaceae | n-alkanes, PAHs | Seawater, sediment | [11] |
| A | Y | Cycloclasticus pugetii | γ-proteobacteria, Piscirickettsiaceae | PAHs | Sediment | [12,13] |
| A | Y | Oleispira Antarctica | γ-proteobacteria, Oceanospirillaceae | n-alkanes | Seawater | [14] |
| A | N | Oleibacter marinus | γ-proteobacteria, Oceanospirillaceae | n-alkanes | Seawater | [15] |
| A | N | Oleiphilus messinensis | γ-proteobacteria, Oleiphilaceae | n-alkanes | Seawater, sediment | [11] |
| A/AN | Y | Pseudomonas pachastrellae | γ-proteobacteria, Pseudomonadaceae | n-alkanes, PAHs | Sediment, beach sand | [16,17] |
| A/AN | Y | Pseudomonas stutzeri | γ-proteobacteria, Pseudomonadaceae | n-alkanes, PAHs, BTEX | Seawater, marine sediments, beach sand | [18] |
| A | N | Halomonas halodurans; Halomonas organivorans | γ-proteobacteria, Halomonadaceae | n-alkanes | Seawater, sediment | [19,20] |
| A | Y | Thalassolituus oleivorans | γ-proteobacteria, Oceanospirillaceae | n-alkanes | Surface seawaters, sediments, coastal and estuarine areas | [21] |
| A | Y | Alteromonas naphthalenivorans | γ-proteobacteria, Alteromonadaceae | PAHs | Seawater, tidal flat sediment | [22] |
| A | Y | Acinetobacter venetianus | γ-proteobacteria, Moraxellaceae | n-alkanes | Surface water, sediment. | [23] |
| A | Y | Dietzia maris | Actinobacteria, Dietziaceae | n-alkanes, PAHs | Seawater, deep sea hydrothermal field | [24] |
| A | N | Rhodobacter sp. SS12.29; Rhodococcus sp. ice-oil-488 s | γ -proteobacteria, Rhodobacteraceae | PAHs | Seawater | [25] |
| A | N | Sphingopixis sp. | γ -proteobacteria, Sphingomonadaceae | PAHs | Seawater | [25] |
| AN | Y | Desulfatibacillum alkenivorans | γ -proteobacteria, Desulfobacteraceae | n-alkanes | Sediment | [26] |
| AN | N | Desulfosarcina-Desulfococcus cluster strains | γ -proteobacteria, Desulfobacteraceae | Short chain n-alkanes | Sediments of marine HC seeps | [2,27] |
| AN | N | Desulfococcus oleovorans | γ -proteobacteria, Desulfobacteraceae | n-alkanes, aromatic HCs | Sediment | [28] |
| A | Y | Bacillus pumilus | Bacilli, Bacillaceae | n-alkanes, PAHs | Sediment | [18,29] |
| A | N | Bacillus stratosphericus | Bacilli, Bacillaceae | PAHs, BTEX | Seawater | [6] |
| AN | Y | Archaeoglobus fulgidus | Euryarchaeota, Archaeoglobaceae | n-alkanes | Shallow marine hydrothermal system | [30] |
| AN | Y | Thermococcus sibiricus | Euryarchaeota, Thermococcaceae | n-alkanes | Oil reservoir | [31] |
| AN | Y | Ferroglobus placidus | Euryarchaeota, Archaeoglobaceae | Aromatic HCs | Shallow marine hydrothermal system | [32,33] |
| AN | N | Dothideomycetes-related taxa | Fungi | PAHs | Beach sediment, tarballs, salt marshes | [34,35] |
| Source | Examples | ||
|---|---|---|---|
| Mobile Sources | Vehicle Exhausts [55] | Aircraft Exhaust [56] | Oil Tankers [57] |
| Industrial Sources | Coke Production/Burning [58] | Cement Manufacturing [59] | Tyre Manufacturing [58] |
| Domestic Sources | Coal Cooking [56] | Wood Burning [55] | Cigarette/Tobacco Smoking [60] |
| Agricultural Sources | Agricultural Wastes [55] | Pesticides [61] | Fertilizers [61] |
| Natural Sources | Forest Fire [62] | Volcanic Eruptions [63] | Wild Fire [63] |
| Pollutant Type | Microorganisms | Reference |
|---|---|---|
| PAHs (fluorene, pyrene, phenanthrene) | Rhodococcus sp., Acinetobacter sp., Pseudomonas sp. | [78] |
| Gasoline | Methylibium petroleiphilum LMG22953 | [79] |
| Crude oil | Roseomonas sp., Bacillus marisflavi, Microbacterium oxydans | [80] |
| Crude oil | Alcanivorax borkumensis, Thalassolituus oleivorans | [81] |
| Crude oil | P. aeruginosa, Rhodococcus sp. CE461, Rhodococcus sp. CT451 | [82] |
| Petroleum HCs | Rhizopus sp., Penicillium funiculosum, Aspergillus sydowii, Rhizobiales sp., Pseudomonas sp., Brucella sp., Bacillus sp., Rhodococcus sp., Microbacterium sp. | [83] |
| Petroleum HCs | Pseudomonas oleovorans, Ochrobactrum sp., Stenotrophomonas maltophila | [84] |
| Mixture of PAHs (anthracene, naphthalene, phenanthrene, pyrene, dibenzo[a]anthracene | Bacillus strains B1F, B5A and B3G, Chromobacterium sp. 4015, Enterobacter aglomerans sp. B1A | [85] |
| PAHs (anthracene, phenanthrene, pyrene) | Mycobacterium fortuitum, Bacillus cereus, Microbacterium sp., Gornodia, Polyisoprenivorans, Microbacteriaceae, Bacterium, Fusarium oxysporium | [86] |
| Crude petroleum oil hydrocarbon | B. subtilis DM-04, P. aeruginosa M and NM | [87] |
| Microorganisms | Biosurfactant | Economic Significance | References |
|---|---|---|---|
| P. aeruginosa | Rhamno lipids | Bioremediation | [102] |
| Acinetobacter calcoaceticus | Emulsan Glycolipopeptide | Enhanced oil recovery by microbes | [103] |
| Rhodococcuserythropolis | Trehalose lipids | Dissolution of HCs | [104] |
| Ustilagomaydis | Cellobiose lipids | Antifungal compounds | [105] |
| Microbacterium | Microbactan Glycolipopeptide | Emulsifier | [106] |
| B. licheniformis | Lichenysin | Enhanced oil recovery by microbes | [107] |
| C. bombicola | Sophoro lipids | Antimicrobial property | [108] |
| B. subtilis | Surfactin | Antimicrobial property | [109] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rahmati, F.; Asgari Lajayer, B.; Shadfar, N.; van Bodegom, P.M.; van Hullebusch, E.D. A Review on Biotechnological Approaches Applied for Marine Hydrocarbon Spills Remediation. Microorganisms 2022, 10, 1289. https://doi.org/10.3390/microorganisms10071289
Rahmati F, Asgari Lajayer B, Shadfar N, van Bodegom PM, van Hullebusch ED. A Review on Biotechnological Approaches Applied for Marine Hydrocarbon Spills Remediation. Microorganisms. 2022; 10(7):1289. https://doi.org/10.3390/microorganisms10071289
Chicago/Turabian StyleRahmati, Farzad, Behnam Asgari Lajayer, Najmeh Shadfar, Peter M. van Bodegom, and Eric D. van Hullebusch. 2022. "A Review on Biotechnological Approaches Applied for Marine Hydrocarbon Spills Remediation" Microorganisms 10, no. 7: 1289. https://doi.org/10.3390/microorganisms10071289
APA StyleRahmati, F., Asgari Lajayer, B., Shadfar, N., van Bodegom, P. M., & van Hullebusch, E. D. (2022). A Review on Biotechnological Approaches Applied for Marine Hydrocarbon Spills Remediation. Microorganisms, 10(7), 1289. https://doi.org/10.3390/microorganisms10071289









