Clinical Escherichia coli: From Biofilm Formation to New Antibiofilm Strategies
Abstract
:1. Introduction
2. Biofilm Formation of E. coli
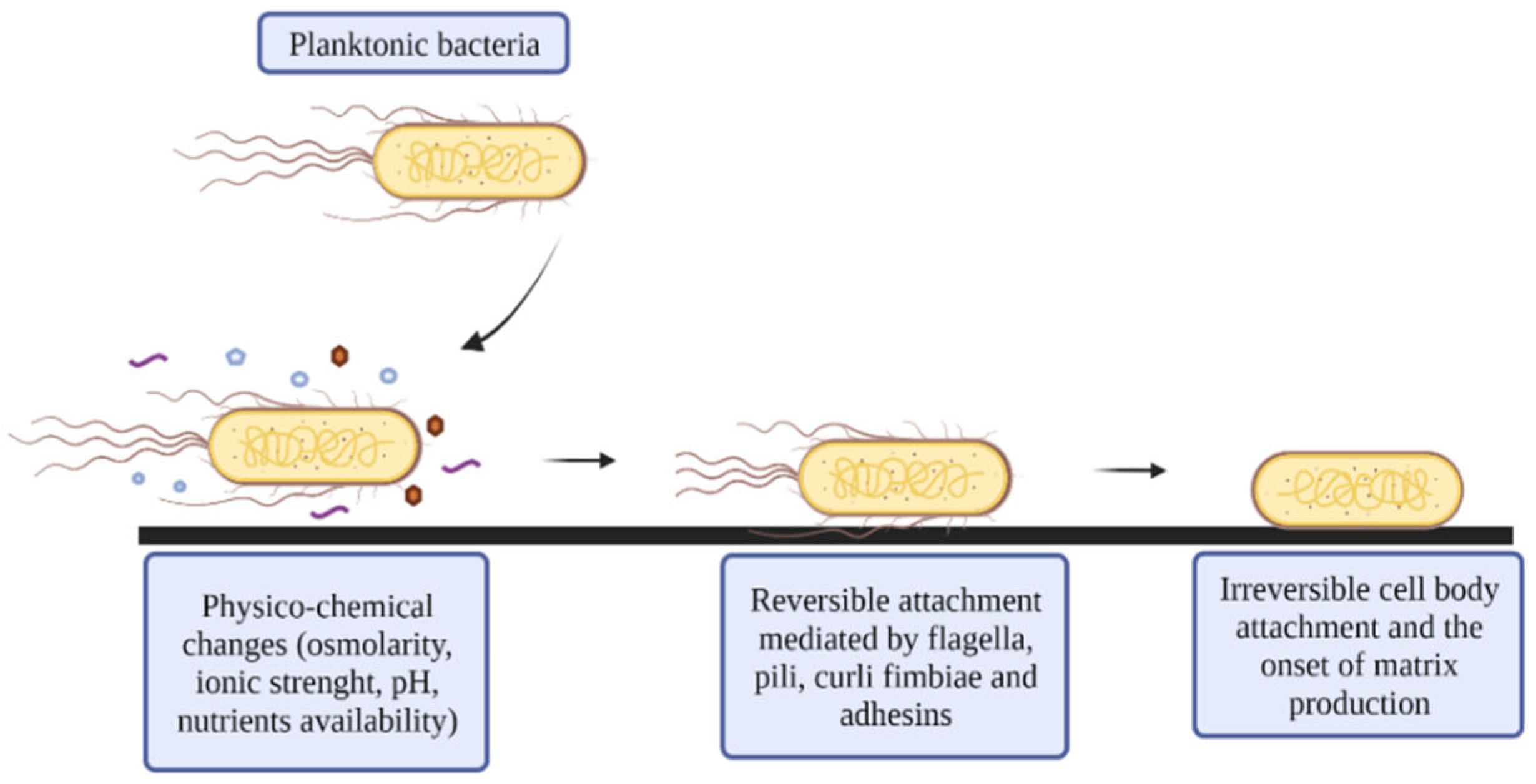
2.1. Reversible Attachment
2.2. Irreversible Attachment
- Conjugative pili or F-pili: are encoded on the F-plasmid and promote horizontal gene transference (HGT) between cells, cell-to-cell contact, aggregation, and nonspecific binding to abiotic surfaces, thus stabilizing the biofilm structure [24,25]. This plasmid stimulates the production of curli and colanic acid (CA), which are related to adhesion and biofilm maturation [26]. Other conjugative plasmids belonging to different incompatibility groups may also contribute to biofilm development [24].
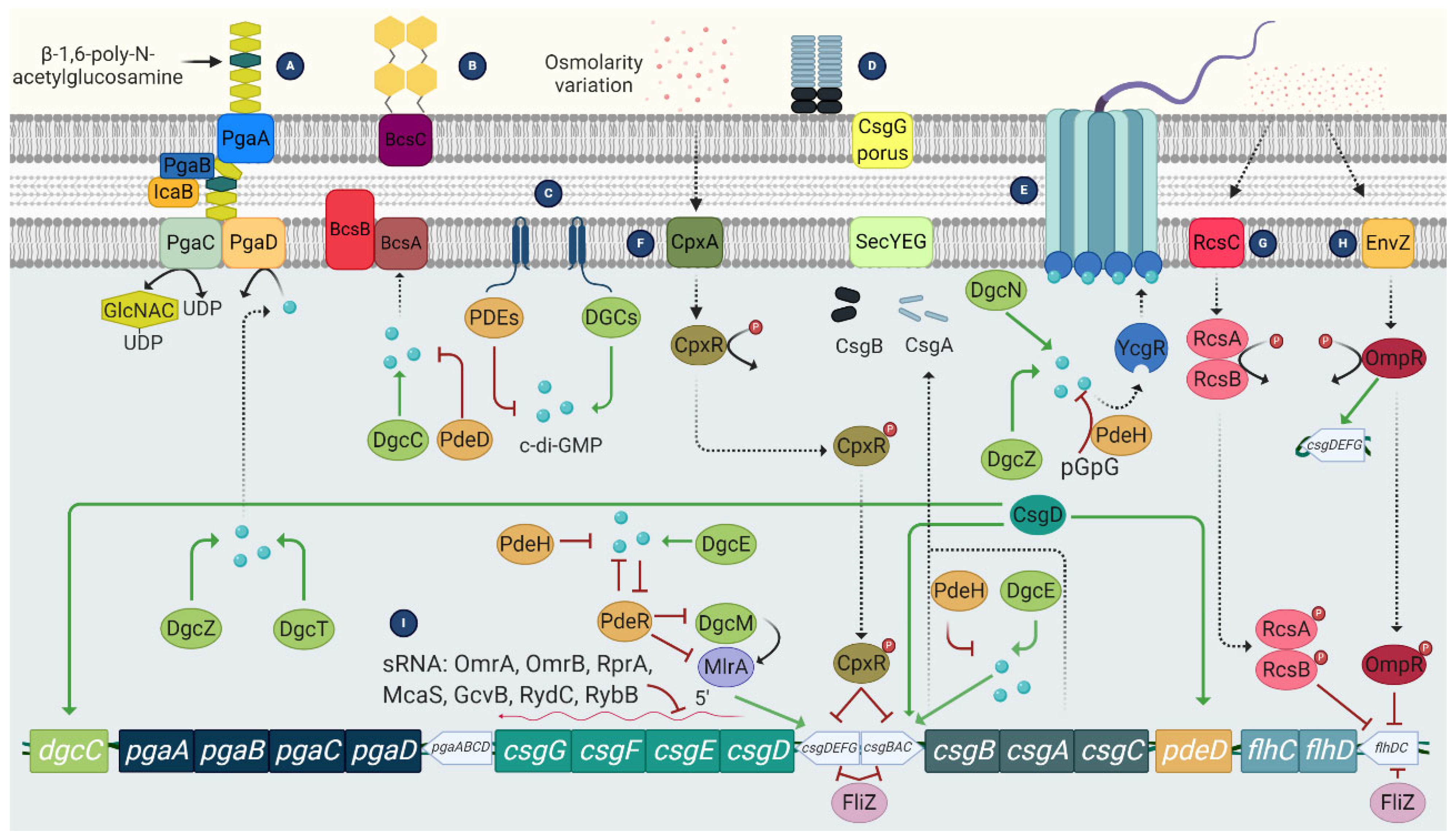
- Curli fimbriae: are amyloid structures that promote cell aggregation and attachment to abiotic surfaces [27]. These fibers account for up to 85% of the biomass of E. coli biofilms [28]. Curli fimbriae are encoded by two operons, the csgBAC operon, which encodes the structural components of the fiber, and the csgDEFG operon, which encodes the transcriptional regulatory protein CsgD and the machinery required to export the fimbriae, CsgEG [29]. The CsgD protein is also involved in cellulose synthesis [30]. Curli synthesis is triggered by various factors, from post-transcriptional changes via sRNA to environmental conditions (temperature less than 32 °C, osmolarity changes, nutrient limitation, or reduced oxygen levels) [31].
- Fimbriae 1 or type 1 pili: are considered major players in the initial steps of E. coli biofilm formation. They are encoded by the fimAICDFGH operon. FimA is the major subunit in type 1 fimbriae, and a variable number of these subunits (500–3000) forms the rod. FimC has chaperone activity in the periplasm and binds to the SecYEG translocon subunits of FimD, which are anchored in the outer membrane. The FimF, FimG, and FimH proteins are located at the tip of the fimbriae [32]. The FimH protein can bind to mannose due to the presence of a lectin domain in its structure, allowing E. coli to bind to mannose present in eukaryotic cells, EPS, and abiotic surfaces [27]. Finally, the FimI protein appears to be the terminator subunit in this type of pili [33].
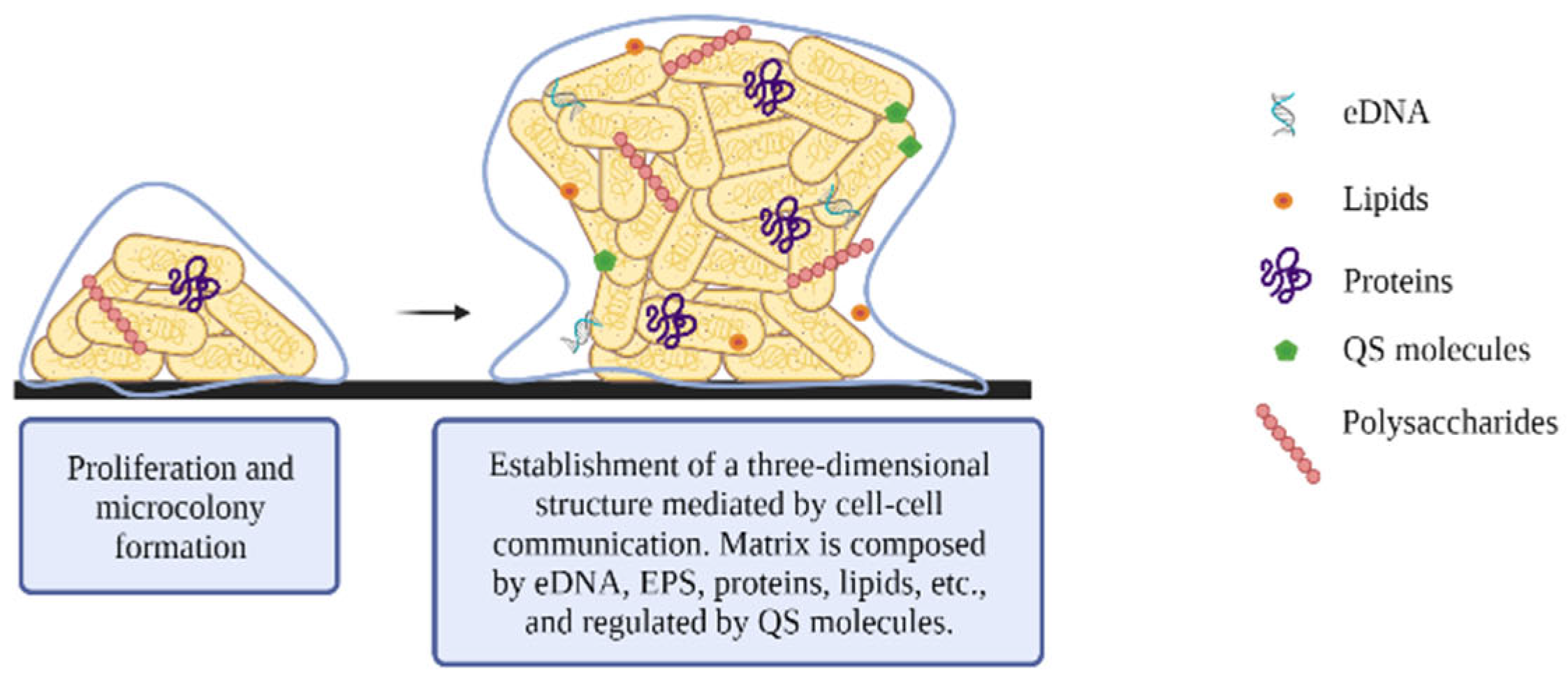
2.3. Maturation
- Poly-β-1,6- N-acetyl-D-glucosamine (PGA) is a positively charged linear homoglycan that promotes cell-to-cell adhesion and surface attachment. The pgaABCD operon encodes two glycosyltransferases: PgaC, which is involved in polymerization, and PgaD, which increases PGA production. Both proteins are essential for PGA biosynthesis. The other two proteins encoded by this operon, PgaA and PgaB, are involved in PGA export to the outer membrane [44,45]. Regulation of biosynthesis is mediated in part by c-di-GMP, in charge of the post-translational activation of PGA [44].
- Cellulose consists of a linear homopolysaccharide of β-1→4 bound to D-glucose and forming fibrils. Its synthesis is encoded by the yhjR-bcsQABZC and bcsEFG operons [46], which synthesize the bacterial cellulose synthase (Bcs) complex. This complex is formed by two proteins, the BcsA and BcsB proteins, which are anchored in the cytoplasmic membrane.
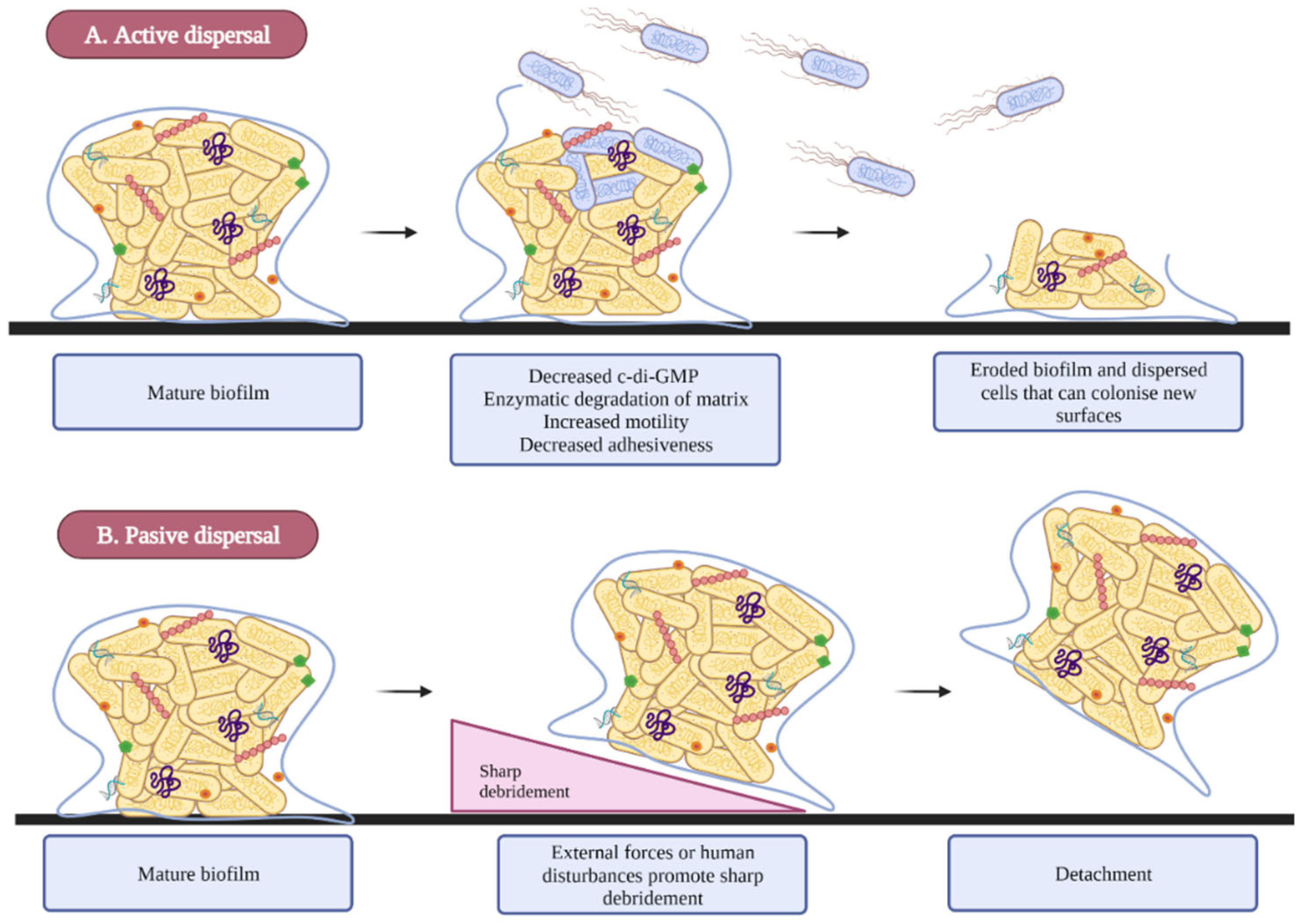
2.4. Dispersion
3. Regulation of E. coli Biofilms
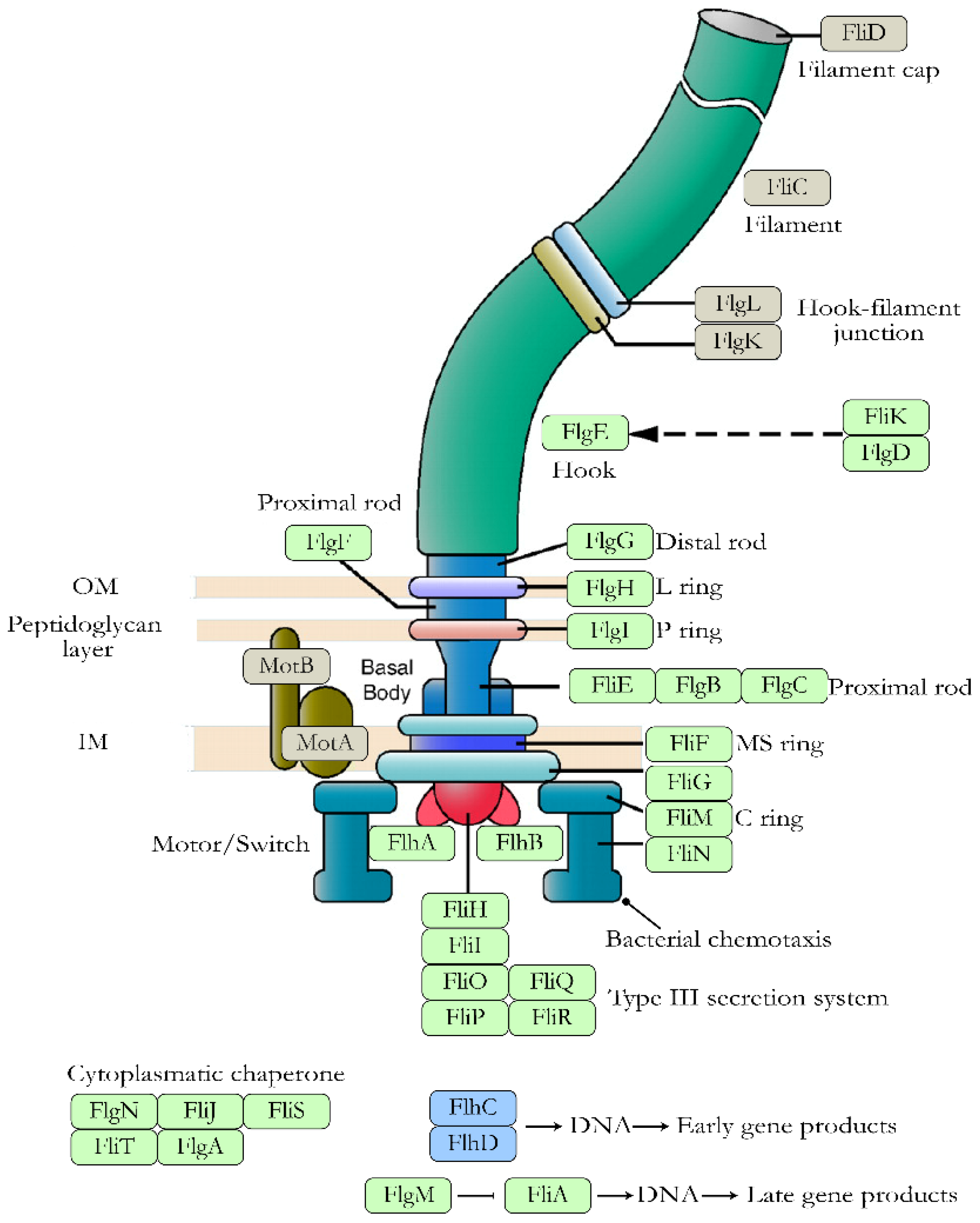
- 3′,5′-cyclic diguanylic acid (c-di-GMP): It is a secondary messenger synthesized by diguanylate cyclases and degraded to pGpG by specific phosphodiesterases. In E. coli, c-di-GMP plays an essential role both in flagellar motility and in the synthesis of curli, cellulose, and PGA. In the case of flagellar motility, this is controlled by c-di-GMP but also by the YcgR protein. Thus, a high level of c-di-GMP activates YCgR and blocks one of the flagellar proteins, FliG. As a result, the bacteria become immobile. Inactivation of YcgR by the protein PdeH leads to a decrease in c-di-GMP levels and thus to activation of the flagella so that the bacteria become mobile again [53].
- Two-component signaling systems (TCS) (Figure 5): They are widely distributed in bacteria. This system consists of a histidine kinase that acts as a sensor for external signals and a response regulator that modulates gene transcription in response to the external signal, allowing a rapid response. In the case of biofilm-related TCS, the most important in E. coli is: CpxA/CpxR, which modifies the chemical content of cell surfaces by activating OmpC, thus contributing to the inhibition of chemotaxis and flagellar activity [54]. In addition, CPxR can inhibit the expression of curli by binding to the operons that encode this type of fibers [55]; and the EnvZ/OmpR system, which is activated at low osmolarity and causes inhibition of flagella [56].
- The RcsCDB regulator: It consists of three regulators, the RcsC, RcsB, and RcsD proteins, which are involved in the synthesis of the capsule, but also regulate the synthesis of the CA and the expression of some genes related to the synthesis of the flagella and adhesion structures such as curli and Ag43 [57].
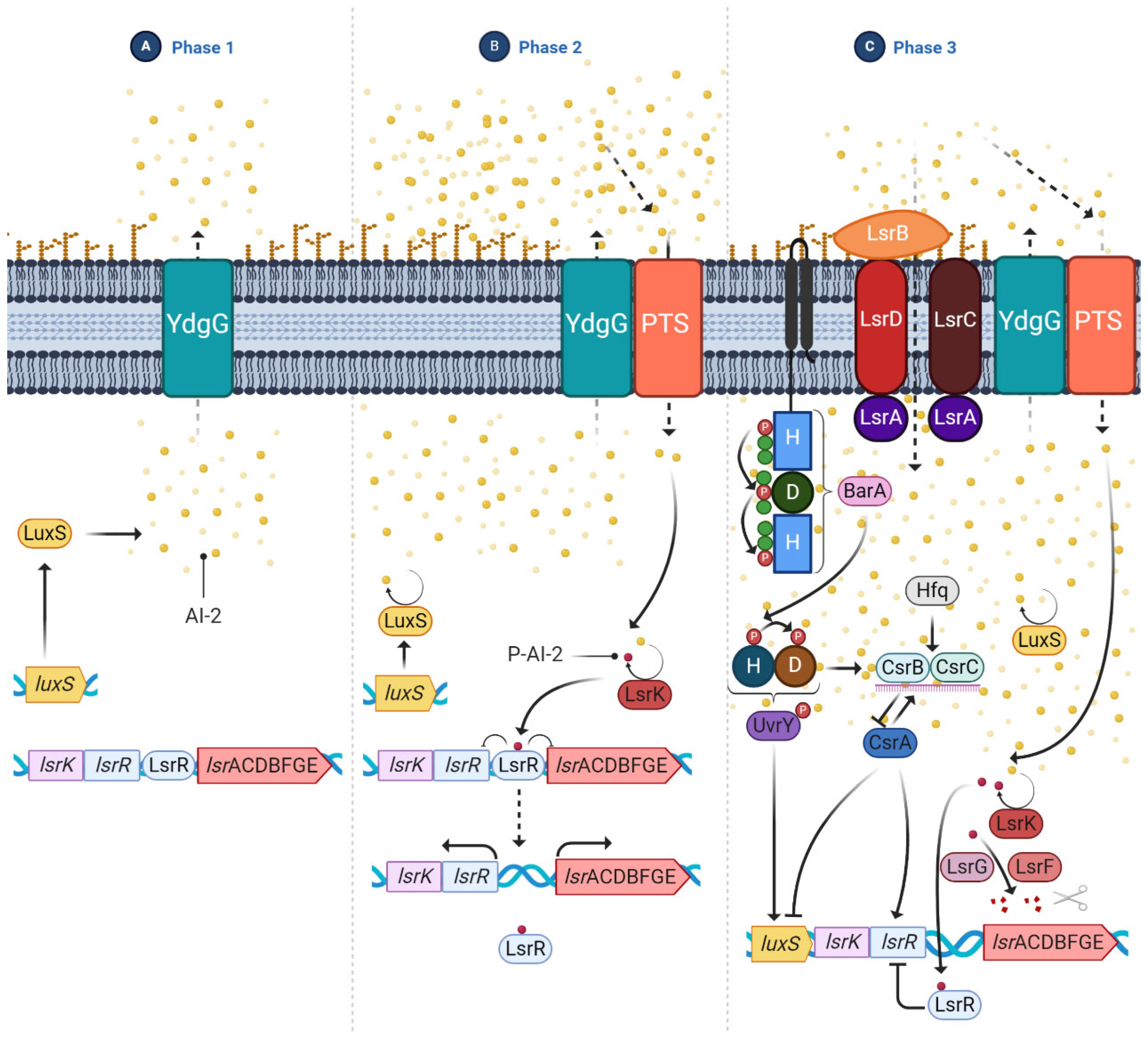
- Quorum sensing (QS): It is a cell-density-dependent chemical signaling system that allows individual cells to release small signal molecules to the surroundings to make their presence known. The small-signal molecules, also called autoinducers (AI), coordinate cell-density-dependent gene expression. QS is used to coordinate gene expression and regulate numerous processes involved in virulence, such as motility and biofilm formation, which are necessary for planktonic bacteria to adopt the biofilm phenotype [35,58,59]. AIs are present in Gram-negative and Gram-positive bacteria. In the case of E. coli, the most studied AI is AI-2, which is produced by the LuxS enzyme related to biofilm formation. The production of this AI is upregulated and it is rapidly secreted to the outside via the LSR transporters. Once optimal bacterial density is achieved, luxS is downregulated, inhibiting the production of AI-2 [59] (Figure 6).
4. Mechanisms of Antimicrobial Tolerance in E. coli Biofilms
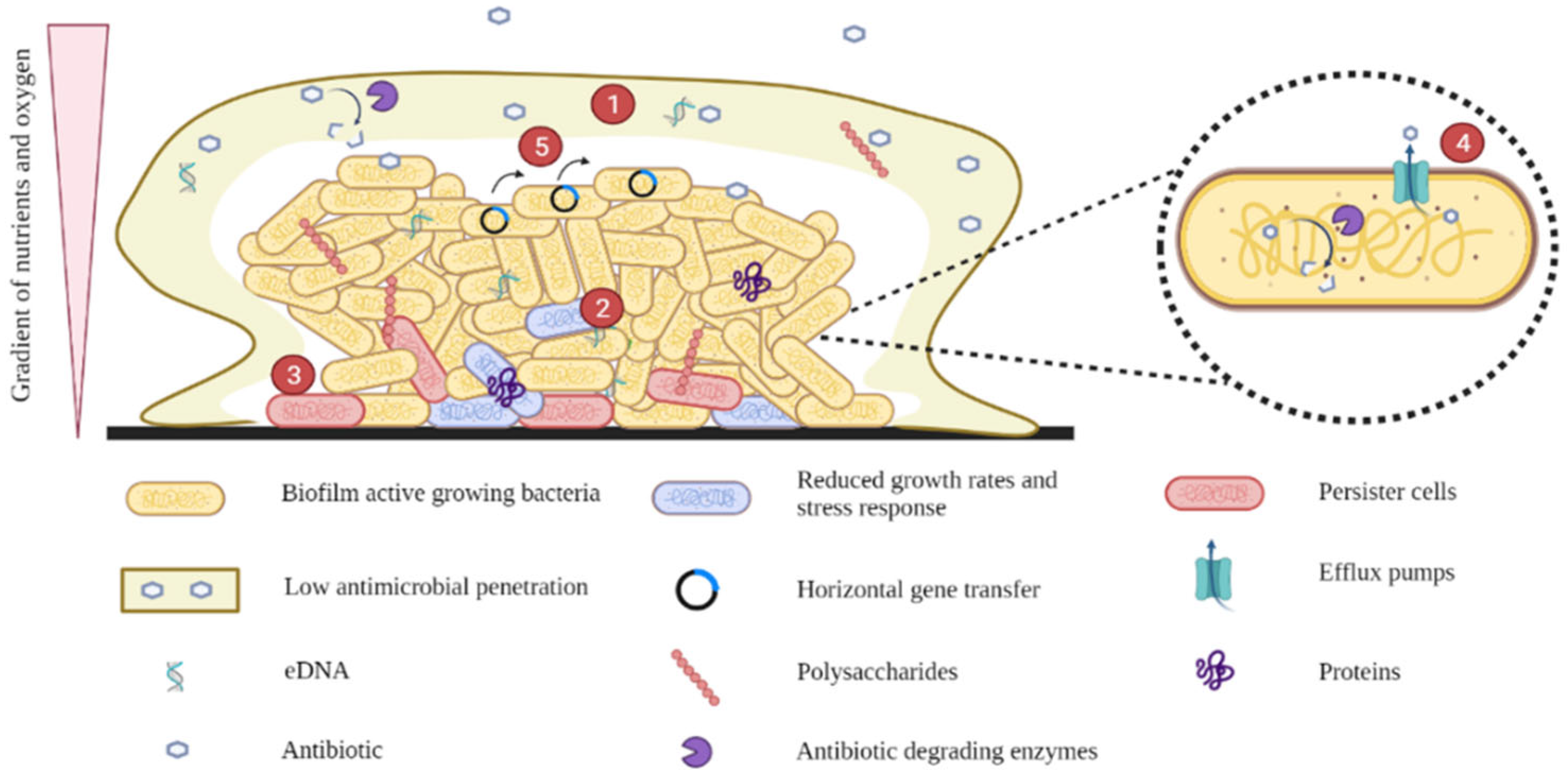
4.1. Low Antimicrobial Penetration
4.2. Reduced Growth Rates and Stress Responses
4.3. Persister Cells
4.4. Efflux Pumps
4.5. Horizontal Gene Transfer (HGT)
5. Role of Biofilms on Different Infections Caused by E. coli
5.1. Urinary Tract Infections (UTIs)
5.2. Bloodstream Infections (BSIs)
6. New Biofilm Treatments for E. coli Biofilms
6.1. Anti-Adhesion Agents
6.2. Inhibition of the QS Pathway
6.3. Phage Therapy
6.4. Antimicrobial Peptides
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Vert, M.; Doi, Y.; Hellwich, K.; Hess, M.; Hodge, P.; Kubisa, P.; Rinaudo, M.; Schué, F. Terminology for Biorelated Polymers and Applications. Chem. Int. Newsmag. IUPAC 2011, 33, 377–410. [Google Scholar] [CrossRef]
- Coenye, T. Response of Sessile Cells to Stress: From Changes in Gene Expression to Phenotypic Adaptation. FEMS Immunol. Med. Microbiol. 2010, 59, 239–252. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Costerton, J.W. Bacterial Biofilms: A Common Cause of Persistent Infections. Science 1999, 284, 1318–1322. [Google Scholar] [CrossRef] [Green Version]
- Konopka, A. What Is Microbial Community Ecology? ISME J. 2009, 3, 1223–1230. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lerch, T.Z.; Chenu, C.; Dignac, M.F.; Barriuso, E.; Mariotti, A. Biofilm vs. Planktonic Lifestyle: Consequences for Pesticide 2,4-D Metabolism by Cupriavidus Necator JMP134. Front. Microbiol. 2017, 8, 1–11. [Google Scholar] [CrossRef]
- Welin-Neilands, J.; Svensäter, G. Acid Tolerance of Biofilm Cells of Streptococcus mutans. Appl. Environ. Microbiol. 2007, 73, 5633–5638. [Google Scholar] [CrossRef] [Green Version]
- Davies, D. Understanding Biofilm Resistance to Antibacterial Agents. Nat. Rev. Drug Discov. 2003, 2, 114–122. [Google Scholar] [CrossRef]
- Donlan, R.M.; Costerton, J.W. Biofilms: Survival Mechanisms of Clinically Relevant Microorganisms. Clin. Microbiol. Rev. 2002, 15, 167–193. [Google Scholar] [CrossRef] [Green Version]
- Fernández-Barat, L.; Torres, A. Biofilms in Ventilator-Associated Pneumonia. Future Microbiol. 2016, 11, 1599–1610. [Google Scholar] [CrossRef]
- Fux, C.A.; Stoodley, P.; Hall-Stoodley, L.; Costerton, J.W. Bacterial Biofilms: A Diagnostic and Therapeutic Challenge. Expert Rev. Anti. Infect. Ther. 2003, 1, 667–683. [Google Scholar] [CrossRef]
- Gbejuade, H.O.; Lovering, A.M.; Webb, J.C. The Role of Microbial Biofilms in Prosthetic Joint Infections: A Review. Acta Orthop. 2015, 86, 147–158. [Google Scholar] [CrossRef] [PubMed]
- Khatoon, Z.; McTiernan, C.D.; Suuronen, E.J.; Mah, T.-F.; Alarcon, E.I. Bacterial Biofilm Formation on Implantable Devices and Approaches to Its Treatment and Prevention. Heliyon 2018, 4, e01067. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lohse, M.B.; Gulati, M.; Johnson, A.D.; Nobile, C.J. Development and Regulation of Single- and Multi-Species Candida albicans Biofilms. Nat. Rev. Microbiol. 2018, 16, 19–31. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Veerachamy, S.; Yarlagadda, T.; Manivasagam, G.; Yarlagadda, P.K. Bacterial Adherence and Biofilm Formation on Medical Implants: A Review. Proc. Inst. Mech. Eng. Part H J. Eng. Med. 2014, 228, 1083–1099. [Google Scholar] [CrossRef]
- Høiby, N.; Bjarnsholt, T.; Givskov, M.; Molin, S.; Ciofu, O. Antibiotic Resistance of Bacterial Biofilms. Int. J. Antimicrob. Agents 2010, 35, 322–332. [Google Scholar] [CrossRef] [Green Version]
- González, J.F.; Hahn, M.M.; Gunn, J.S. Chronic Biofilm-Based Infections: Skewing of the Immune Response. Pathog. Dis. 2018, 76, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Belas, R. Biofilms, Flagella, and Mechanosensing of Surfaces by Bacteria. Trends Microbiol. 2014, 22, 517–527. [Google Scholar] [CrossRef]
- Apel, D.; Surette, M.G. Bringing Order to a Complex Molecular Machine: The Assembly of the Bacterial Flagella. Biochim. Biophys. Acta Biomembr. 2008, 1778, 1851–1858. [Google Scholar] [CrossRef] [Green Version]
- Fitzgerald, D.M.; Bonocora, R.P.; Wade, J.T. Comprehensive Mapping of the Escherichia coli Flagellar Regulatory Network. PLoS Genet. 2014, 10, e1004649. [Google Scholar] [CrossRef] [Green Version]
- Macnab, R. Genetics and Biogenesis of Bacterial Flagella. Annu. Rev. Genet. 1992, 26, 131–158. [Google Scholar] [CrossRef]
- Kim, J.M.; Garcia-Alcala, M.; Balleza, E.; Cluzel, P. Stochastic Transcriptional Pulses Orchestrate Flagellar Biosynthesis in Escherichia coli. Sci. Adv. 2020, 6, eaax0947. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vidal, O.; Longin, R.; Prigent-Combaret, C.; Dorel, C.; Hooreman, M.; Lejeune, P. Isolation of an Escherichia coli K-12 Mutant Strain Able To Form Biofilms on Inert Surfaces: Involvement of a New OmpR Allele That Increases Curli Expression. J. Bacteriol. 1998, 180, 2442–2449. [Google Scholar] [CrossRef] [Green Version]
- Goller, C.C.; Romeo, T. Environmental Influences on Biofilm Development. In Bacterial Biofilms. Current Topics in Microbiology and Immunology; Springer: Berlin/Heidelberg, Germany, 2008; pp. 37–66. [Google Scholar]
- Ghigo, J.-M. Natural Conjugative Plasmids Induce Bacterial Biofilm Development. Nature 2001, 412, 442–445. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Khara, P.; Christie, P.J. Structural Bases for F Plasmid Conjugation and F Pilus Biogenesis in Escherichia coli. Proc. Natl. Acad. Sci. USA 2019, 116, 14222–14227. [Google Scholar] [CrossRef] [Green Version]
- May, T.; Okabe, S. Escherichia coli Harboring a Natural IncF Conjugative F Plasmid Develops Complex Mature Biofilms by Stimulating Synthesis of Colanic Acid and Curli. J. Bacteriol. 2008, 190, 7479–7490. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beloin, C.; Roux, A.; Ghigo, J.-M. Escherichia coli Biofilms. In Bacterial Biofilms. Current Topics in Microbiology and Immunology; Springer: Berlin/Heidelberg, Germany, 2008; Volume 322, pp. 249–289. ISBN 9783540754176. [Google Scholar]
- Salinas, N.; Povolotsky, T.L.; Landau, M.; Kolodkin-Gal, I. Emerging Roles of Functional Bacterial Amyloids in Gene Regulation, Toxicity, and Immunomodulation. Microbiol. Mol. Biol. Rev. 2021, 85, 1–17. [Google Scholar] [CrossRef]
- Barnhart, M.M.; Chapman, M.R. Curli Biogenesis and Function. Annu. Rev. Microbiol. 2006, 60, 131–147. [Google Scholar] [CrossRef] [Green Version]
- Gualdi, L.; Tagliabue, L.; Bertagnoli, S.; Ieranò, T.; De Castro, C.; Landini, P. Cellulose Modulates Biofilm Formation by Counteracting Curli-Mediated Colonization of Solid Surfaces in Escherichia coli. Microbiology 2008, 154, 2017–2024. [Google Scholar] [CrossRef] [Green Version]
- Lüthje, P.; Brauner, A. Virulence Factors of Uropathogenic E. coli and Their Interaction with the Host. Adv. Microb. Physiol. 2014, 65, 337–372. [Google Scholar] [CrossRef]
- Capitani, G.; Eidam, O.; Glockshuber, R.; Grütter, M.G. Structural and Functional Insights into the Assembly of Type 1 Pili from Escherichia Coli. Microbes Infect. 2006, 8, 2284–2290. [Google Scholar] [CrossRef]
- Valenski, M.L.; Harris, S.L.; Spears, P.A.; Horton, J.R.; Orndorff, P.E. The Product of the fimI Gene Is Necessary for Escherichia coli Type 1 Pilus Biosynthesis. J. Bacteriol. 2003, 185, 5007–5011. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Klemm, P.; Schembri, M. Type 1 Fimbriae, Curli, and Antigen 43: Adhesion, Colonization, and Biofilm Formation. EcoSal Plus 2013, 1, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Laganenka, L.; Colin, R.; Sourjik, V. Chemotaxis towards Autoinducer 2 Mediates Autoaggregation in Escherichia coli. Nat. Commun. 2016, 7, 12984. [Google Scholar] [CrossRef] [Green Version]
- Van der Woude, M.W.; Henderson, I.R. Regulation and Function of Ag43 (Flu). Annu. Rev. Microbiol. 2008, 62, 153–169. [Google Scholar] [CrossRef] [PubMed]
- Schooling, S.R.; Beveridge, T.J. Membrane Vesicles: An Overlooked Component of the Matrices of Biofilms. J. Bacteriol. 2006, 188, 5945–5957. [Google Scholar] [CrossRef] [Green Version]
- Rabin, N.; Zheng, Y.; Opoku-Temeng, C.; Du, Y.; Bonsu, E.; Sintim, H.O. Biofilm Formation Mechanisms and Targets for Developing Antibiofilm Agents. Future Med. Chem. 2015, 7, 493–512. [Google Scholar] [CrossRef]
- Flemming, H.-C.; Wingender, J. The Biofilm Matrix. Nat. Rev. Microbiol. 2010, 8, 623–633. [Google Scholar] [CrossRef] [PubMed]
- Sharma, G.; Sharma, S.; Sharma, P.; Chandola, D.; Dang, S.; Gupta, S.; Gabrani, R. Escherichia coli Biofilm: Development and Therapeutic Strategies. J. Appl. Microbiol. 2016, 121, 309–319. [Google Scholar] [CrossRef] [Green Version]
- Limoli, D.H.; Jones, C.J.; Wozniak, D.J. Bacterial Extracellular Polysaccharides in Biofilm Formation and Function. Microbiol. Spectr. 2015, 3, 1191–1200. [Google Scholar] [CrossRef] [Green Version]
- Sharma, D.; Misba, L.; Khan, A.U. Antibiotics versus Biofilm: An Emerging Battleground in Microbial Communities. Antimicrob. Resist. Infect. Control 2019, 8, 76. [Google Scholar] [CrossRef]
- Campoccia, D.; Montanaro, L.; Arciola, C.R. Extracellular DNA (eDNA). a Major Ubiquitous Element of the Bacterial Biofilm Architecture. Int. J. Mol. Sci. 2021, 22, 9100. [Google Scholar] [CrossRef] [PubMed]
- Itoh, Y.; Rice, J.D.; Goller, C.; Pannuri, A.; Taylor, J.; Meisner, J.; Beveridge, T.J.; Preston, J.F.; Romeo, T. Roles of pgaABCD Genes in Synthesis, Modification, and Export of the Escherichia coli Biofilm Adhesin Poly-β-1,6-N-Acetyl-D-Glucosamine. J. Bacteriol. 2008, 190, 3670–3680. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.; Preston, J.F.; Romeo, T. The PgaABCD Locus of Escherichia coli Promotes the Synthesis of a Polysaccharide Adhesin Required for Biofilm Formation. J. Bacteriol. 2004, 186, 2724–2734. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Serra, D.O.; Richter, A.M.; Hengge, R. Cellulose as an Architectural Element in Spatially Structured Escherichia coli Biofilms. J. Bacteriol. 2013, 195, 5540–5554. [Google Scholar] [CrossRef] [Green Version]
- Navasa, N.; Rodríguez-Aparicio, L.; Martínez-Blanco, H.; Arcos, M.; Ferrero, M.Á. Temperature Has Reciprocal Effects on Colanic Acid and Polysialic Acid Biosynthesis in E. coli K92. Appl. Microbiol. Biotechnol. 2009, 82, 721–729. [Google Scholar] [CrossRef]
- Hall-Stoodley, L.; Costerton, J.W.; Stoodley, P. Bacterial Biofilms: From the Natural Environment to Infectious Diseases. Nat. Rev. Microbiol. 2004, 2, 95–108. [Google Scholar] [CrossRef]
- Donlan, R.M. Biofilm Formation: A Clinically Relevant Microbiological Process. Clin. Infect. Dis. 2001, 33, 1387–1392. [Google Scholar] [CrossRef] [Green Version]
- Kostakioti, M.; Hadjifrangiskou, M.; Hultgren, S.J. Bacterial Biofilms: Development, Dispersal, and Therapeutic Strategies in the Dawn of the Postantibiotic Era. Cold Spring Harb. Perspect. Med. 2013, 3, 1–23. [Google Scholar] [CrossRef] [Green Version]
- Rumbaugh, K.P.; Sauer, K. Biofilm Dispersion. Nat. Rev. Microbiol. 2020, 18, 571–586. [Google Scholar] [CrossRef]
- Singh, S.; Datta, S.; Narayanan, K.B.; Rajnish, K.N. Bacterial Exo-Polysaccharides in Biofilms: Role in Antimicrobial Resistance and Treatments. J. Genet. Eng. Biotechnol. 2021, 19, 1–19. [Google Scholar] [CrossRef]
- Römling, U.; Galperin, M.Y.; Gomelsky, M. Cyclic Di-GMP: The First 25 Years of a Universal Bacterial Second Messenger. Microbiol. Mol. Biol. Rev. 2013, 77, 1–52. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Markova, J.A.; Anganova, E.V.; Turskaya, A.L.; Bybin, V.A.; Savilov, E.D. Regulation of Escherichia coli Biofilm Formation (Review). Appl. Biochem. Microbiol. 2018, 54, 1–11. [Google Scholar] [CrossRef]
- Prigent-Combaret, C.; Brombacher, E.; Vidal, O.; Ambert, A.; Lejeune, P.; Landini, P.; Dorel, C. Complex Regulatory Network Controls Initial Adhesion and Biofilm Formation in Escherichia coli via Regulation of the CsgD Gene. J. Bacteriol. 2001, 183, 7213–7223. [Google Scholar] [CrossRef] [Green Version]
- Prüß, B.M.; Besemann, C.; Denton, A.; Wolfe, A.J. A Complex Transcription Network Controls the Early Stages of Biofilm Development by Escherichia coli. J. Bacteriol. 2006, 188, 3731–3739. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Majdalani, N.; Gottesman, S. The Rcs Phosphorelay: A Complex Signal Transduction System. Annu. Rev. Microbiol. 2005, 59, 379–405. [Google Scholar] [CrossRef]
- Li, J.; Attila, C.; Wang, L.; Wood, T.K.; Valdes, J.J.; Bentley, W.E. Quorum Sensing in Escherichia coli Is Signaled by AI-2/LsrR: Effects on Small RNA and Biofilm Architecture. J. Bacteriol. 2007, 189, 6011–6020. [Google Scholar] [CrossRef] [Green Version]
- Pereira, C.S.; Thompson, J.A.; Xavier, K.B. AI-2-Mediated Signalling in Bacteria. FEMS Microbiol. Rev. 2013, 37, 156–181. [Google Scholar] [CrossRef] [Green Version]
- Rather, M.A.; Gupta, K.; Mandal, M. Microbial Biofilm: Formation, Architecture, Antibiotic Resistance, and Control Strategies. Braz. J. Microbiol. 2021, 52, 1701–1718. [Google Scholar] [CrossRef]
- Dincer, S.; Masume, F.; Uslu, D.; Delik, A. Antibiotic Resistance in Biofilm. In Bacterial Biofilms; IntechOpen: London, UK; BoD-Books on Demand: Norderstedt, Germany, 2020. [Google Scholar]
- Lewis, K. Riddle of Biofilm Resistance. Antimicrob. Agents Chemother. 2001, 45, 999–1007. [Google Scholar] [CrossRef] [Green Version]
- Mah, T.F.C.; O’Toole, G.A. Mechanisms of Biofilm Resistance to Antimicrobial Agents. Trends Microbiol. 2001, 9, 34–39. [Google Scholar] [CrossRef]
- Butt, A.; Khan, A. Antibiotics Resistance of Bacterial Biofilms. Middle East J. Bus. 2015, 10, 38–45. [Google Scholar] [CrossRef]
- Alav, I.; Sutton, J.M.; Rahman, K.M. Role of Bacterial Efflux Pumps in Biofilm Formation. J. Antimicrob. Chemother. 2018, 73, 2003–2020. [Google Scholar] [CrossRef] [Green Version]
- Soto, S.M. Role of Efflux Pumps in the Antibiotic Resistance of Bacteria Embedded in a Biofilm. Virulence 2013, 4, 223–229. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matsumura, K.; Furukawa, S.; Ogihara, H.; Morinaga, Y. Roles of Multidrug Efflux Pumps on the Biofilm Formation of Escherichia coli K-12. Biocontrol Sci. 2011, 16, 69–72. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Uruén, C.; Chopo-Escuin, G.; Tommassen, J.; Mainar-Jaime, R.C.; Arenas, J. Biofilms as Promoters of Bacterial Antibiotic Resistance and Tolerance. Antibiotics 2021, 10, 3. [Google Scholar] [CrossRef]
- Flores-Mireles, A.L.; Walker, J.N.; Caparon, M.; Hultgren, S.J. Urinary Tract Infections: Epidemiology, Mechanisms of Infection and Treatment Options. Nat. Rev. Microbiol. 2015, 13, 269–284. [Google Scholar] [CrossRef]
- Fazly Bazzaz, B.S.; Darvishi Fork, S.; Ahmadi, R.; Khameneh, B. Deep Insights into Urinary Tract Infections and Effective Natural Remedies. Afr. J. Urol. 2021, 27, 1–13. [Google Scholar] [CrossRef]
- Cave, J. Prevention of Recurrent Urinary Tract Infections in Women. Drug Ther. Bull. 2013, 51, 69–74. [Google Scholar]
- Soto, S.M.; Marco, F.; Guiral, E.; Vila, J. Biofilm Formation in Uropathogenic Escherichia coli Strains: Relationship with Urovirulence Factors and Antimicrobial Resistance. In Clinical Management of Complicated Urinary Tract Infection; Nikibakhsh, A., Ed.; IntechOpen: London, UK; BoD-Books on Demand: Norderstedt, Germany, 2011; ISBN 978-953-307-393-4. [Google Scholar]
- Soto, S.M.; Smithson, A.; Horcajada, J.P.; Martinez, J.A.; Mensa, J.P.; Vila, J. Implication of Biofilm Formation in the Persistence of Urinary Tract Infection Caused by Uropathogenic Escherichia coli. Clin. Microbiol. Infect. 2006, 12, 1034–1036. [Google Scholar] [CrossRef] [Green Version]
- Tambyah, P.A.; Maki, D.G. Catheter-Associated Urinary Tract Infection Is Rarely Symptomatic: A Prospective Study of 1497 Catheterized Patients. Arch. Intern. Med. 2000, 160, 678–682. [Google Scholar] [CrossRef] [Green Version]
- Nicolle, L.E. Catheter Associated Urinary Tract Infections. Antimicrob. Resist. Infect. Control 2014, 3, 23. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guggenbichler, J.P.; Assadian, O.; Boeswald, M.; Kramer, A. Incidence and Clinical Implication of Nosocomial Infections Associated with Implantable Biomaterials—Catheters, Ventilator-Associated Pneumonia, Urinary Tract Infections. GMS Krankenhhyg. Interdiszip. 2011, 6, 1–19. [Google Scholar] [CrossRef]
- Pai, K.C.; Wang, M.S.; Chen, Y.F.; Tseng, C.H.; Liu, P.Y.; Chen, L.C.; Sheu, R.K.; Wu, C.L. An Artificial Intelligence Approach to Bloodstream Infections Prediction. J. Clin. Med. 2021, 10, 2901. [Google Scholar] [CrossRef] [PubMed]
- Wolcott, R. Biofilm and Catheter-Related Bloodstream Infections. Br. J. Nurs. 2021, 30, S4–S9. [Google Scholar] [CrossRef] [PubMed]
- Erbay, A.; Ergönül, Ö.; Stoddard, G.J.; Samore, M.H. Recurrent Catheter-Related Bloodstream Infections: Risk Factors and Outcome. Int. J. Infect. Dis. 2006, 10, 396–400. [Google Scholar] [CrossRef] [Green Version]
- Pinto, H.; Simões, M.; Borges, A. Prevalence and Impact of Biofilms on Bloodstream and Urinary Tract Infections: A Systematic Review and Meta-Analysis. Antibiotics 2021, 10, 825. [Google Scholar] [CrossRef]
- Lu, L.; Hu, W.; Tian, Z.; Yuan, D.; Yi, G.; Zhou, Y.; Cheng, Q.; Zhu, J.; Li, M. Developing Natural Products as Potential Anti-Biofilm Agents. Chin. Med. 2019, 14, 1–17. [Google Scholar] [CrossRef] [Green Version]
- Lee, J.H.; Kim, Y.G.; Ryu, S.Y.; Cho, M.H.; Lee, J. Ginkgolic Acids and Ginkgo Biloba Extract Inhibit Escherichia coli O157: H7 and Staphylococcus aureus Biofilm Formation. Int. J. Food Microbiol. 2014, 174, 47–55. [Google Scholar] [CrossRef]
- Lee, J.H.; Regmi, S.C.; Kim, J.A.; Cho, M.H.; Yun, H.; Lee, C.S.; Lee, J. Apple Flavonoid Phloretin Inhibits Escherichia coli O157:H7 Biofilm Formation and Ameliorates Colon Inflammation in Rats. Infect. Immun. 2011, 79, 4819–4827. [Google Scholar] [CrossRef] [Green Version]
- Kim, Y.G.; Lee, J.H.; Gwon, G.; Kim, S.I.; Park, J.G.; Lee, J. Essential Oils and Eugenols Inhibit Biofilm Formation and the Virulence of Escherichia coli O157:H7. Sci. Rep. 2016, 6, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Sun, J.; Marais, J.P.J.; Khoo, C.; LaPlante, K.; Vejborg, R.M.; Givskov, M.; Tolker-Nielsen, T.; Seeram, N.P.; Rowley, D.C. Cranberry (Vaccinium Macrocarpon) Oligosaccharides Decrease Biofilm Formation by Uropathogenic Escherichia coli. J. Funct. Foods 2015, 17, 235–242. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saurav, K.; Costantino, V.; Venturi, V.; Steindler, L. Quorum Sensing Inhibitors from the Sea Discovered Using Bacterial N-Acyl-Homoserine Lactone-Based Biosensors. Mar. Drugs 2017, 15, 53. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, L.; Zhang, Y.; Ge, Y.; Zhu, X.; Pan, J. Regulatory Mechanisms and Promising Applications of Quorum Sensing-Inhibiting Agents in Control of Bacterial Biofilm Formation. Front. Microbiol. 2020, 11, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Vikram, A.; Jesudhasan, P.R.; Pillai, S.D.; Patil, B.S. Isolimonic Acid Interferes with Escherichia coli O157:H7 Biofilm and TTSS in QseBC and QseA Dependent Fashion. BMC Microbiol. 2012, 12, 1. [Google Scholar] [CrossRef] [Green Version]
- Ghosh, S.; Lahiri, D.; Nag, M.; Dey, A.; Pandit, S.; Sarkar, T.; Pati, S.; Kari, Z.A.; Ishak, A.R.; Edinur, H.A.; et al. Phytocompound Mediated Blockage of Quorum Sensing Cascade in ESKAPE Pathogens. Antibiotics 2022, 11, 61. [Google Scholar] [CrossRef] [PubMed]
- Vikram, A.; Jayaprakasha, G.K.; Jesudhasan, P.R.; Pillai, S.D.; Patil, B.S. Suppression of Bacterial Cell-Cell Signalling, Biofilm Formation and Type III Secretion System by Citrus Flavonoids. J. Appl. Microbiol. 2010, 109, 515–527. [Google Scholar] [CrossRef]
- Srinivasan, R.; Santhakumari, S.; Poonguzhali, P.; Geetha, M.; Dyavaiah, M.; Xiangmin, L. Bacterial Biofilm Inhibition: A Focused Review on Recent Therapeutic Strategies for Combating the Biofilm Mediated Infections. Front. Microbiol. 2021, 12, 1–19. [Google Scholar] [CrossRef]
- Yang, L.; Liu, Y.; Wu, H.; Song, Z.; Høiby, N.; Molin, S.; Givskov, M. Combating Biofilms. FEMS Immunol. Med. Microbiol. 2012, 65, 146–157. [Google Scholar] [CrossRef]
- Gu, Y.; Xu, Y.; Xu, J.; Yu, X.; Huang, X.; Liu, G.; Liu, X. Identification of Novel Bacteriophage VB_EcoP-EG1 with Lytic Activity against Planktonic and Biofilm Forms of Uropathogenic Escherichia coli. Appl. Microbiol. Biotechnol. 2019, 103, 315–326. [Google Scholar] [CrossRef]
- Chibeu, A.; Lingohr, E.J.; Masson, L.; Manges, A.; Harel, J.; Ackermann, H.W.; Kropinski, A.M.; Boerlin, P. Bacteriophages with the Ability to Degrade Uropathogenic Escherichia coli Biofilms. Viruses 2012, 4, 471–487. [Google Scholar] [CrossRef] [Green Version]
- Valério, N.; Oliveira, C.; Jesus, V.; Branco, T.; Pereira, C.; Moreirinha, C.; Almeida, A. Effects of Single and Combined Use of Bacteriophages and Antibiotics to Inactivate Escherichia coli. Virus Res. 2017, 240, 8–17. [Google Scholar] [CrossRef]
- Moradpour, Z.; Yousefi, N.; Sadeghi, D.; Ghasemian, A. Synergistic Bactericidal Activity of a Naturally Isolated Phage and Ampicillin against Urinary Tract Infecting Escherichia coli O157. Iran. J. Basic Med. Sci. 2020, 23, 257–263. [Google Scholar] [CrossRef] [PubMed]
- Comeau, A.M.; Tétart, F.; Trojet, S.N.; Prère, M.F.; Krisch, H.M. Phage-Antibiotic Synergy (PAS): β-Lactam and Quinolone Antibiotics Stimulate Virulent Phage Growth. PLoS ONE 2007, 2, 8–11. [Google Scholar] [CrossRef] [Green Version]
- Fleming, D.; Rumbaugh, K.P. Approaches to Dispersing Medical Biofilms. Microorganisms 2017, 5, 15. [Google Scholar] [CrossRef] [PubMed]
- Roy, R.; Tiwari, M.; Donelli, G.; Tiwari, V. Strategies for Combating Bacterial Biofilms: A Focus on Anti-Biofilm Agents and Their Mechanisms of Action. Virulence 2018, 9, 522–554. [Google Scholar] [CrossRef]
- Fleitas Martínez, O.; Cardoso, M.H.; Ribeiro, S.M.; Franco, O.L. Recent Advances in Anti-Virulence Therapeutic Strategies with a Focus on Dismantling Bacterial Membrane Microdomains, Toxin Neutralization, Quorum-Sensing Interference and Biofilm Inhibition. Front. Cell. Infect. Microbiol. 2019, 9, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Algburi, A.; Comito, N.; Kashtanov, D.; Dicks, L.M.T.; Chikindas, M.L. Control of Biofilm Formation: Antibiotics and Beyond. Appl. Environ. Microbiol. 2017, 83, 1–16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De la Fuente-Núñez, C.; Reffuveille, F.; Haney, E.F.; Straus, S.K.; Hancock, R.E.W. Broad-Spectrum Anti-Biofilm Peptide That Targets a Cellular Stress Response. PLoS Pathog. 2014, 10, e1004152. [Google Scholar] [CrossRef] [Green Version]
- Qvortrup, K.; Hultqvist, L.D.; Nilsson, M.; Jakobsen, T.H.; Jansen, C.U.; Uhd, J.; Andersen, J.B.; Nielsen, T.E.; Givskov, M.; Tolker-Nielsen, T. Small Molecule Anti-Biofilm Agents Developed on the Basis of Mechanistic Understanding of Biofilm Formation. Front. Chem. 2019, 7, 742. [Google Scholar] [CrossRef] [Green Version]
- Defraine, V.; Fauvart, M.; Michiels, J. Fighting Bacterial Persistence: Current and Emerging Anti-Persister Strategies and Therapeutics. Drug Resist. Updat. 2018, 38, 12–26. [Google Scholar] [CrossRef]
- Tetz, G.V.; Artemenko, N.K.; Tetz, V.V. Effect of DNase and Antibiotics on Biofilm Characteristics. Antimicrob. Agents Chemother. 2009, 53, 1204–1209. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rabin, N.; Zheng, Y.; Opoku-Temeng, C.; Du, Y.; Bonsu, E.; Sintim, H.O. Agents That Inhibit Bacterial Biofilm Formation. Future Med. Chem. 2015, 7, 647–671. [Google Scholar] [CrossRef] [PubMed]
- Verderosa, A.D.; Totsika, M.; Fairfull-Smith, K.E. Bacterial Biofilm Eradication Agents: A Current Review. Front. Chem. 2019, 7, 1–17. [Google Scholar] [CrossRef] [Green Version]
- Rosato, A.; Vitali, C.; De Laurentis, N.; Armenise, D.; Antonietta Milillo, M. Antibacterial Effect of Some Essential Oils Administered Alone or in Combination with Norfloxacin. Phytomedicine 2007, 14, 727–732. [Google Scholar] [CrossRef] [PubMed]
| Matrix Compound | Stage of Biofilm Formation in Which They Are Involved | Function in Biofilms | Reference |
|---|---|---|---|
| Polysaccharides | Adhesion | Binding and colonization of biotic and abiotic surfaces | [39] |
| Favor transitory cell immobilization and development of high cell densities | [39] | ||
| Promote cell-cell adhesion | [40] | ||
| Maturation | Encourage microbial interactions | [41] | |
| Provide shape and structural support to the biofilm | [40] | ||
| Favor tolerance to desiccation | [39] | ||
| Provide resistance to host defense and tolerance to antimicrobial agents | [39] | ||
| Facilitate interaction between the bacterial cells and the environment | [40] | ||
| Assist in sorption of organic and inorganic compound | [39] | ||
| Facilitate nutrient supply (carbon, nitrogen and phosphorus) | [42] | ||
| Proteins | Adhesion | Binding and colonization of biotic and abiotic surfaces | [39] |
| Favor transitory cell immobilization, development of high cell densities | [39] | ||
| Maturation | Provide shape and structural support to the biofilm | [40] | |
| Favor tolerance to desiccation | [39] | ||
| Provide resistance to host defense and tolerance to antimicrobial agents | [39] | ||
| Assist in sorption of organic and inorganic compound | [39] | ||
| Facilitate nutrient supply (carbon, nitrogen and phosphorus) | [42] | ||
| Encourage redox activity | [39] | ||
| Dispersion | Promote enzymatic degradation of matrix for cell spreading | [39] | |
| DNA | Adhesion | Binding and colonization of biotic and abiotic surfaces | [39] |
| Favor transitory cell immobilization, development of high cell densities | [39] | ||
| Maturation | Provides shape and structural support to the biofilm | [40] | |
| Exchange of virulence factors/antimicrobial resistance genes | [39] | ||
| Nutrient supply (carbon, nitrogen and phosphorus) | [42] | ||
| Contributes to bacterial aggregation promoting intercellular adhesion | [43] | ||
| Cation binding and sequestration | [43] |
| Action | Antibiofilm Molecules | References |
|---|---|---|
| Inhibition QS pathway | Tyrail derivatives | [100] |
| 1,5-dihydropyrrol-2-ones analogs | [100] | |
| Triphenyl scaffold-based hybrid compounds | [100] | |
| Non-native AHL | [100] | |
| EGCG, tannic acid, ellagic acid (polyphenols) | [99] | |
| Inhibition of (p)ppGpp regulated stringent response | 1018 peptide | [99,101,102] |
| ppGpp analogs | [103] | |
| Relacin | [104] | |
| Dispersion of EPS of biofilm | DNase I | [99,105] |
| Biofilm disassembly | Indole-triazole-amide analogs | [106] |
| Mitomycin C | [107] | |
| Pelargonium graveolens EO | [101,108] | |
| Norspermidine | [106] | |
| Cis-2-decenoic acid | [106] | |
| Zosteric acid derivatives | [100] | |
| D-amino acids, Polyamine nor-spermidine | [42,99,106] | |
| Indolicidin, PR-39 | [99] | |
| Ciprofloxacin-nitroxide hybrid, QACs | [107] | |
| 5-dydroxyindole, Isolimonic acid, Resveratrol | [106] | |
| ε-viniferin | [106] | |
| Neutralization/disaggregation of LPS | PMAP-23 peptide, Polymyxin B | [99] |
| Alteration of membrane permeabilization | Lytic peptides (PTP-7) | [99] |
| Inhibition of cell division or cell survival | Microcin B17 | [99] |
| Pyrrhocoricin | [106] | |
| Inhibition of c-di-GMP signaling system | Azathioprine, Sulfathiazole | [103] |
| C-di-GMP analogs | [103] | |
| Inhibition of appendages biosynthesis | AA-861 (benzoquinone derivative) | [99,106] |
| Acyl sulphonamides, Analogs of FN075 and BibC6 of ring-fused 2-pyridones, Bicyclic 2-pyridone pilus, Hydroxamic acids, Tetrazoles | [99,103] | |
| Ginkgolic acid C15:1, Phloretin | [106] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ballén, V.; Cepas, V.; Ratia, C.; Gabasa, Y.; Soto, S.M. Clinical Escherichia coli: From Biofilm Formation to New Antibiofilm Strategies. Microorganisms 2022, 10, 1103. https://doi.org/10.3390/microorganisms10061103
Ballén V, Cepas V, Ratia C, Gabasa Y, Soto SM. Clinical Escherichia coli: From Biofilm Formation to New Antibiofilm Strategies. Microorganisms. 2022; 10(6):1103. https://doi.org/10.3390/microorganisms10061103
Chicago/Turabian StyleBallén, Victoria, Virginio Cepas, Carlos Ratia, Yaiza Gabasa, and Sara M. Soto. 2022. "Clinical Escherichia coli: From Biofilm Formation to New Antibiofilm Strategies" Microorganisms 10, no. 6: 1103. https://doi.org/10.3390/microorganisms10061103
APA StyleBallén, V., Cepas, V., Ratia, C., Gabasa, Y., & Soto, S. M. (2022). Clinical Escherichia coli: From Biofilm Formation to New Antibiofilm Strategies. Microorganisms, 10(6), 1103. https://doi.org/10.3390/microorganisms10061103







