Staphylococcus saprophyticus Proteomic Analyses Elucidate Differences in the Protein Repertories among Clinical Strains Related to Virulence and Persistence
Abstract
1. Introduction
2. Results
2.1. Evaluation of S. saprophyticus Strains Cell Growth
2.2. Proteomic Profile of S. saprophyticus Strains
2.3. Differentially Abundant Proteins Among S. saprophyticus Strains
3. Discussion
4. Material and Methods
4.1. S. saprophyticus Strain Maintenance and Culture Conditions
4.2. Cell Growth Curve of S. saprophyticus Strains
4.3. Obtaining the Proteins Extracts from S. saprophyticus
4.4. Digestion of Protein Extracts for Nano-ESI-UPLC-MSE Acquisition
4.5. Data Processing and Protein Identification
4.6. Statistical Analysis of Proteomic Data
4.7. Urease Activity Assays
4.8. Thioredoxin Assay
4.9. S. saprophyticus Interaction Assay with Macrophage Cells
5. Conclusions
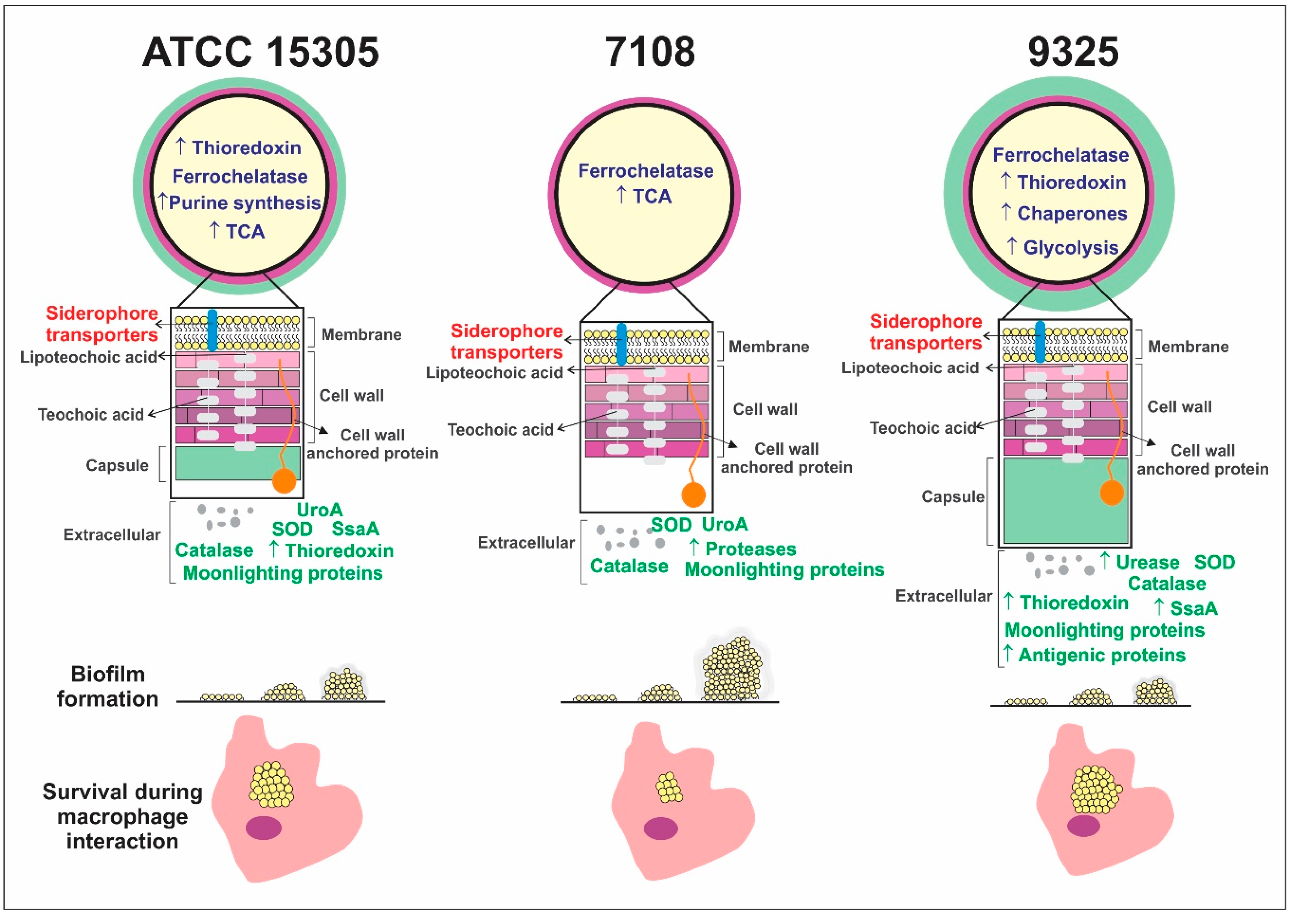
- Different bacterial strains from S. saprophyticus presenting phenotypic and genotypic variations also present proteome global content.
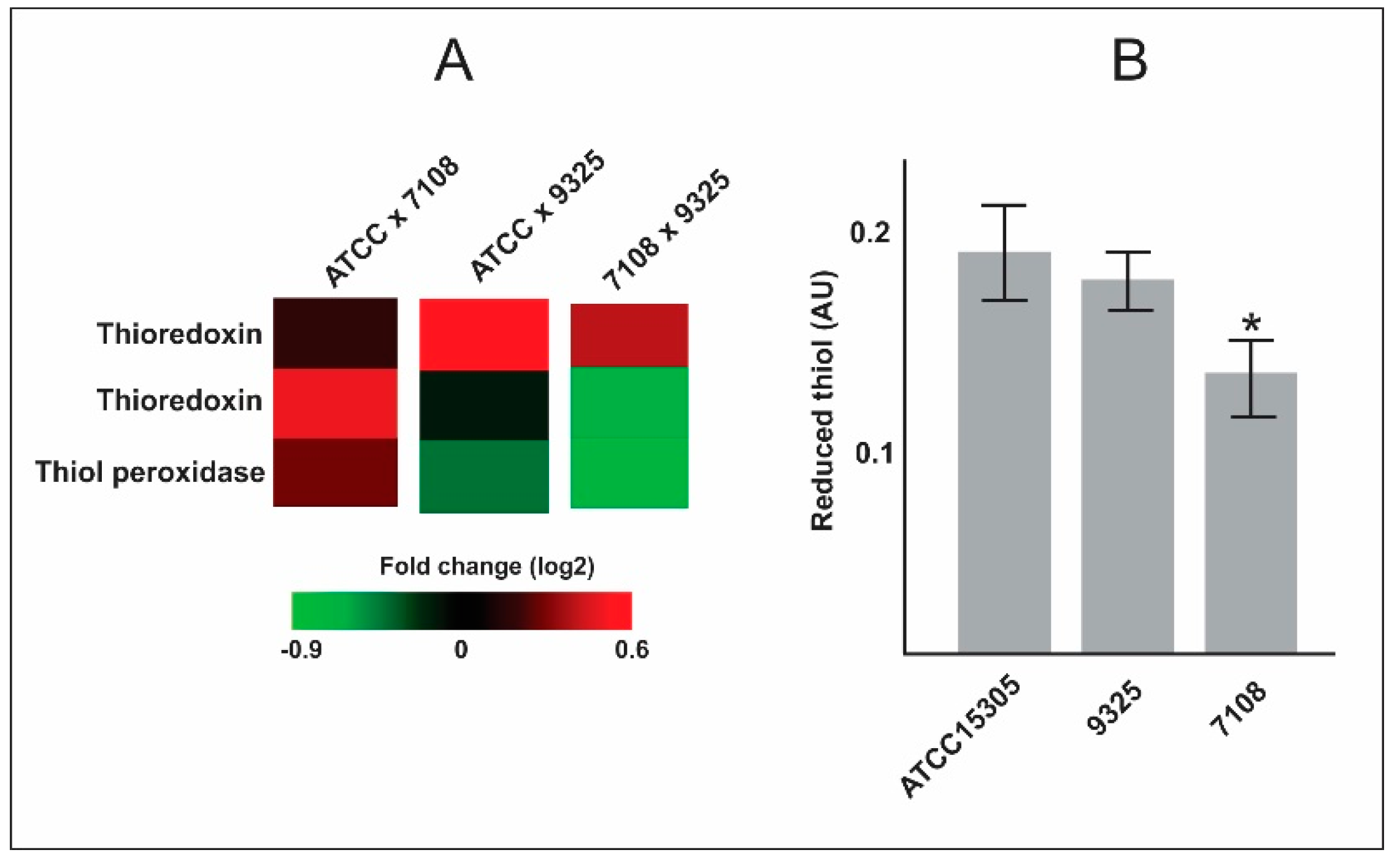
- The capsular strains ATCC 15305 and 9325 possess higher amounts of thioredoxins and reductases in comparison with the 7108, which could reflect in the ability to combat oxidative stress and to survive during interaction with host cells.
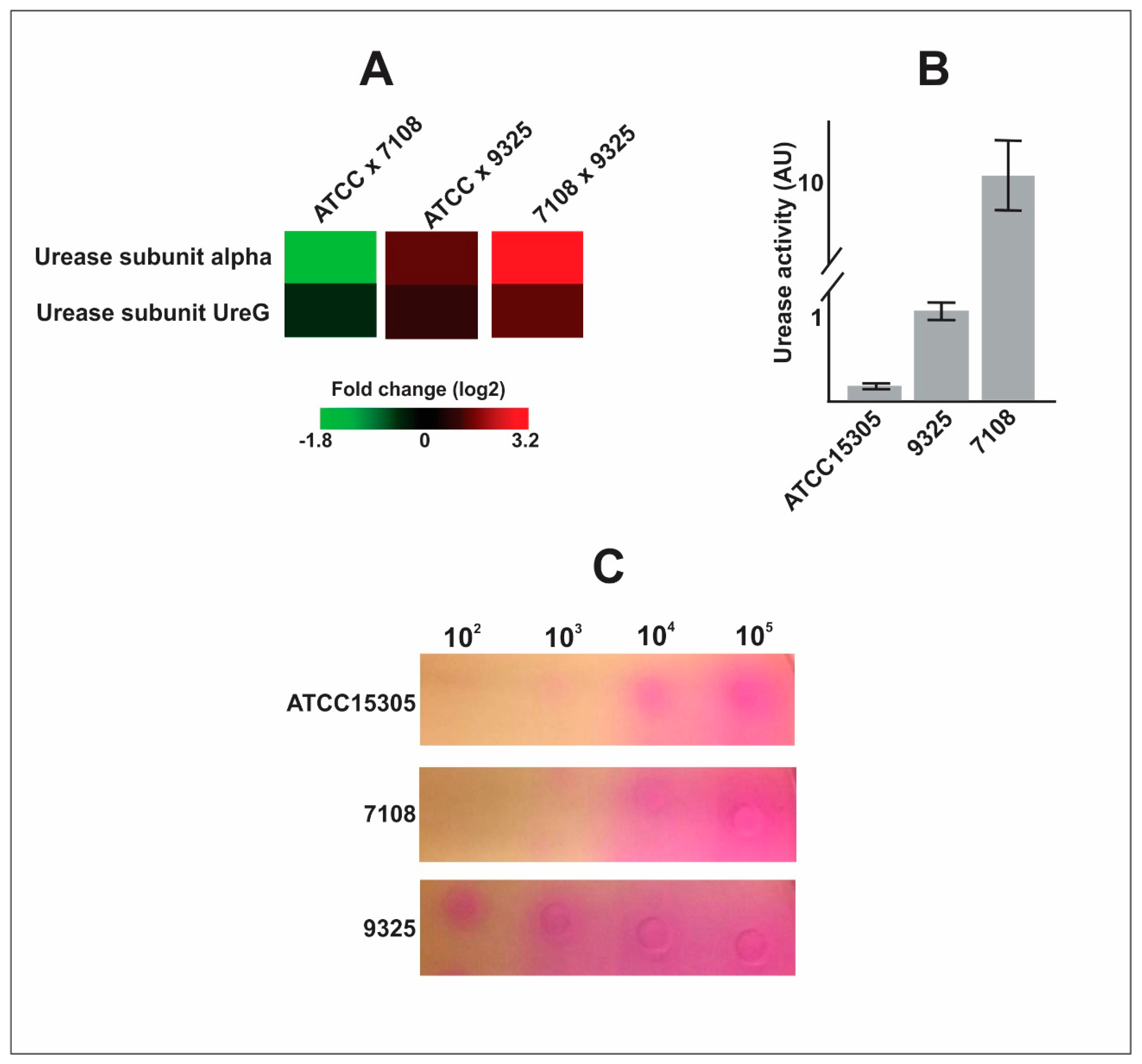
- The capsular strains 9325 and ATCC 15305 are more efficient in exporting urease, which can enhance ability to survive in the presence of urine.
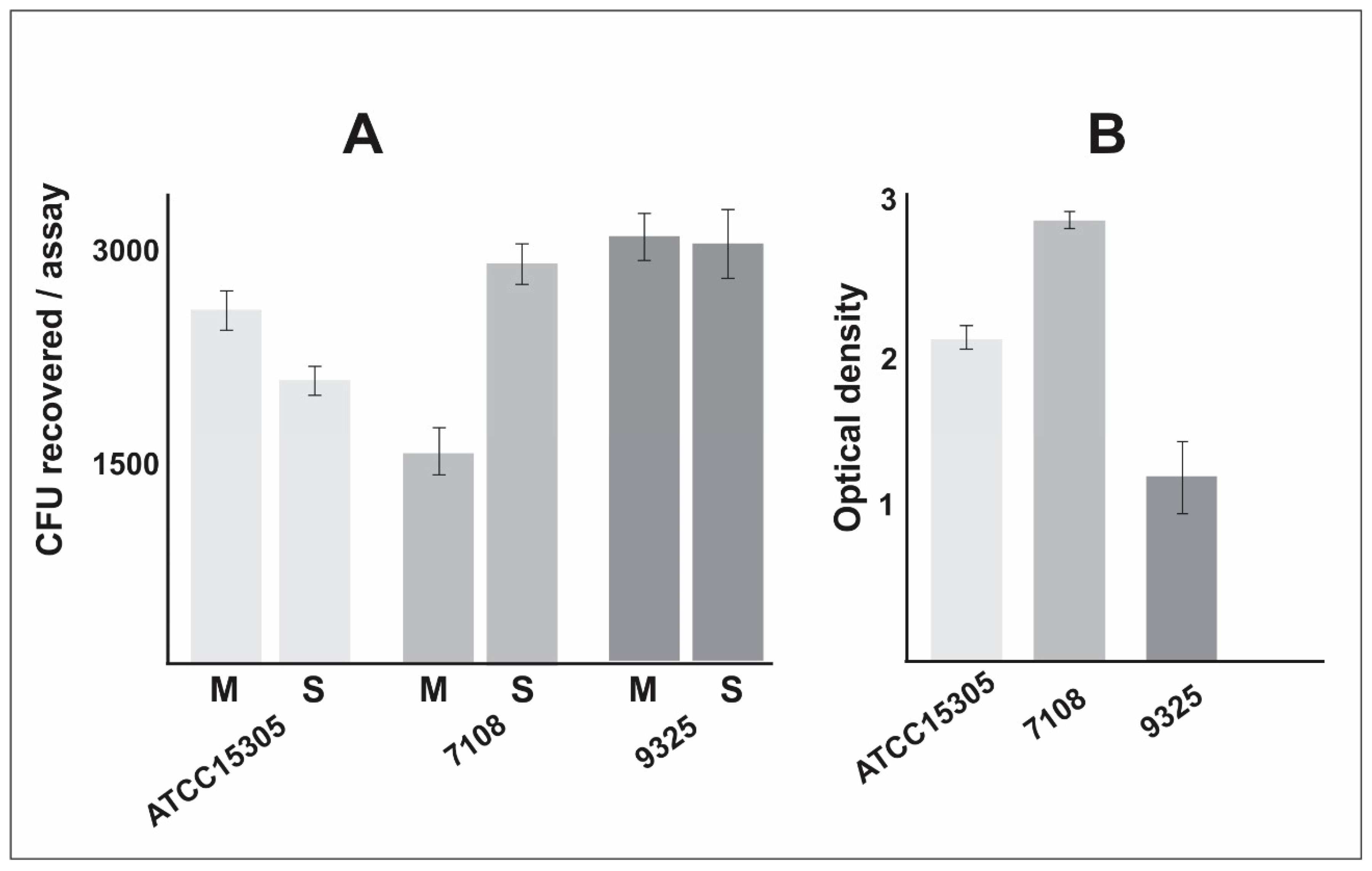
- The non-capsular strain 7108 presented higher ability to form biofilm, which is a particularly important characteristic that enhances the ability to persist in the host and in the environment.
- The proteomic approach can be used to detect and describe proteomic flexibility related to virulence, pathogenicity, and persistence of pathogens.
6. Disclosures
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Raz, R.; Colodner, R.; Kunin, C.M. Who are you—Staphylococcus saprophyticus? Clin. Infect. Dis. 2005, 40, 896–898. [Google Scholar] [CrossRef] [PubMed]
- Kloos, W.E.; Bannerman, T.L. Update on clinical significance of coagulase-negative staphylococci. Clin. Microbiol. Rev. 1994, 7, 117–140. [Google Scholar] [CrossRef] [PubMed]
- Martykanova, D.S.; Davletova, N.C.; Zemlenuhin, I.A.; Volchkova, V.I.; Mugallimov, S.M.; Ahatov, A.M.; Laikov, A.V.; Markelova, M.I.; Boulygina, E.A.; Lopukhov, L.V.; et al. Skin microbiota in contact sports athletes and selection of antiseptics for professional hygiene. Biomed. Res. Int. 2019, 2019, 9843781. [Google Scholar] [CrossRef]
- Gatermann, S.; John, J.; Marre, R. Staphylococcus saprophyticus urease: Characterization and contribution to uropathogenicity in unobstructed urinary tract infection of rats. Infect. Immun. 1989, 57, 110–116. [Google Scholar] [CrossRef]
- Loes, A.N.; Ruyle, L.; Arvizu, M.; Gresko, K.E.; Wilson, A.L.; Deutch, C.E. Inhibition of urease activity in the urinary tract pathogen Staphylococcus saprophyticus. Lett. Appl. Microbiol. 2014, 58, 31–41. [Google Scholar] [CrossRef]
- Gatermann, S.; Meyer, H.G.; Marre, R.; Wanner, G. Identification and characterization of surface proteins from Staphylococcus saprophyticus. Zent. Bakteriol. 1993, 278, 258–274. [Google Scholar] [CrossRef]
- Korte-Berwanger, M.; Sakinc, T.; Kline, K.; Nielsen, H.V.; Hultgren, S.; Gatermann, S.G. Significance of the d-serine-deaminase and d-serine metabolism of Staphylococcus saprophyticus for virulence. Infect. Immun. 2013, 81, 4525–4533. [Google Scholar] [CrossRef]
- Argemi, X.; Hansmann, Y.; Prola, K.; Prevost, G. Coagulase-negative staphylococci pathogenomics. Int. J. Mol. Sci. 2019, 20, 1215. [Google Scholar] [CrossRef]
- Lee, J.H.; Heo, S.; Jeong, M.; Jeong, D.W. Transfer of a mobile Staphylococcus saprophyticus plasmid isolated from fermented seafood that confers tetracycline resistance. PLoS ONE 2019, 14, e0213289. [Google Scholar] [CrossRef]
- De Sousa, V.S.; Da-Silva, A.P.S.; Sorenson, L.; Paschoal, R.P.; Rabello, R.F.; Campana, E.H.; Pinheiro, M.S.; Dos Santos, L.O.F.; Martins, N.; Botelho, A.C.N.; et al. Staphylococcus saprophyticus recovered from humans, food, and recreational waters in Rio de Janeiro, Brazil. Int. J. Microbiol. 2017, 2017, 4287547. [Google Scholar] [CrossRef]
- Li, H.; Xu, L.; Wang, J.; Wen, Y.; Vuong, C.; Otto, M.; Gao, Q. Conversion of Staphylococcus epidermidis strains from commensal to invasive by expression of the ica locus encoding production of biofilm exopolysaccharide. Infect. Immun. 2005, 73, 3188–3191. [Google Scholar] [CrossRef] [PubMed]
- Aguila-Arcos, S.; Alvarez-Rodriguez, I.; Garaiyurrebaso, O.; Garbisu, C.; Grohmann, E.; Alkorta, I. Biofilm-forming clinical Staphylococcus isolates harbor horizontal transfer and antibiotic resistance genes. Front. Microbiol. 2017, 8, 2018. [Google Scholar] [CrossRef] [PubMed]
- Martins, K.B.; Ferreira, A.M.; Pereira, V.C.; Pinheiro, L.; De Oliveira, A.; Da Cunha, M. In vitro effects of antimicrobial agents on planktonic and biofilm forms of Staphylococcus saprophyticus isolated from patients with urinary tract infections. Front. Microbiol. 2019, 10, 40. [Google Scholar] [CrossRef] [PubMed]
- Kleine, B.; Gatermann, S.; Sakinc, T. Genotypic and phenotypic variation among Staphylococcus saprophyticus from human and animal isolates. BMC Res. Notes 2010, 3, 163. [Google Scholar] [CrossRef]
- Oliveira, A.S.; Rosa, I.I.R.; Novaes, E.; Oliveira, L.S.; Baeza, L.C.; Borges, C.L.; Marlinghaus, L.; Soares, C.M.A.; Giambiagi-deMarval, M.; Parente-Rocha, J.A. The exoproteome profiles of three Staphylococcus saprophyticus strains reveal diversity in protein secretion contents. Microbiol. Res. 2018, 216, 85–96. [Google Scholar] [CrossRef]
- Souza, B.S.V.; Silva, K.C.S.; Parente, A.F.A.; Borges, C.L.; Paccez, J.D.; Pereira, M.; Soares, C.M.A.; Giambiagi-deMarval, M.; Silva-Bailao, M.G.; Parente-Rocha, J.A. The influence of pH on Staphylococcus saprophyticus iron metabolism and the production of siderophores. Microbes Infect. 2019, 21, 456–463. [Google Scholar] [CrossRef]
- Martinez-Martin, N. Technologies for proteome-wide discovery of extracellular host-pathogen interactions. J. Immunol. Res. 2017, 2017, 2197615. [Google Scholar] [CrossRef]
- Beltran, P.M.J.; Federspiel, J.D.; Sheng, X.; Cristea, I.M. Proteomics and integrative omic approaches for understanding host-pathogen interactions and infectious diseases. Mol. Syst. Biol. 2017, 13, 922. [Google Scholar] [CrossRef]
- Hecker, M.; Mader, U.; Volker, U. From the genome sequence via the proteome to cell physiology—Pathoproteomics and pathophysiology of Staphylococcus aureus. Int. J. Med. Microbiol. 2018, 308, 545–557. [Google Scholar] [CrossRef]
- Bonar, E.; Wojcik, I.; Wladyka, B. Proteomics in studies of Staphylococcus aureus virulence. Acta Biochim. Pol. 2015, 62, 367–381. [Google Scholar] [CrossRef]
- Arner, E.S.; Holmgren, A. Physiological functions of thioredoxin and thioredoxin reductase. Eur. J. Biochem. 2000, 267, 6102–6109. [Google Scholar] [CrossRef] [PubMed]
- Uziel, O.; Borovok, I.; Schreiber, R.; Cohen, G.; Aharonowitz, Y. Transcriptional regulation of the Staphylococcus aureus thioredoxin and thioredoxin reductase genes in response to oxygen and disulfide stress. J. Bacteriol. 2004, 186, 326–334. [Google Scholar] [CrossRef] [PubMed]
- Shaffer, C.L.; Zhang, E.W.; Dudley, A.G.; Dixon, B.; Guckes, K.R.; Breland, E.J.; Floyd, K.A.; Casella, D.P.; Algood, H.M.S.; Clayton, D.B.; et al. Purine biosynthesis metabolically constrains intracellular survival of uropathogenic Escherichia coli. Infect. Immun. 2017, 85. [Google Scholar] [CrossRef] [PubMed]
- Andersen-Civil, A.I.S.; Ahmed, S.; Guerra, P.R.; Andersen, T.E.; Hounmanou, Y.M.G.; Olsen, J.E.; Herrero-Fresno, A. The impact of inactivation of the purine biosynthesis genes, purN and purT, on growth and virulence in uropathogenic E. coli. Mol. Biol. Rep. 2018, 45, 2707–2716. [Google Scholar] [CrossRef] [PubMed]
- Yoshioka, S.; Newell, P.D. Disruption of de novo purine biosynthesis in Pseudomonas fluorescens Pf0-1 leads to reduced biofilm formation and a reduction in cell size of surface-attached but not planktonic cells. PeerJ 2016, 4, e1543. [Google Scholar] [CrossRef]
- Munoz-Elias, E.J.; Marcano, J.; Camilli, A. Isolation of Streptococcus pneumoniae biofilm mutants and their characterization during nasopharyngeal colonization. Infect. Immun. 2008, 76, 5049–5061. [Google Scholar] [CrossRef]
- Fey, P.D.; Olson, M.E. Current concepts in biofilm formation of Staphylococcus epidermidis. Future Microbiol. 2010, 5, 917–933. [Google Scholar] [CrossRef]
- Park, S.; Kelley, K.A.; Vinogradov, E.; Solinga, R.; Weidenmaier, C.; Misawa, Y.; Lee, J.C. Characterization of the structure and biological functions of a capsular polysaccharide produced by Staphylococcus saprophyticus. J. Bacteriol. 2010, 192, 4618–4626. [Google Scholar] [CrossRef]
- Murad, A.M.; Rech, E.L. NanoUPLC-MSE proteomic data assessment of soybean seeds using the Uniprot database. BMC Biotechnol. 2012, 12, 82. [Google Scholar] [CrossRef]
- Perez-Riverol, Y.; Csordas, A.; Bai, J.; Bernal-Llinares, M.; Hewapathirana, S.; Kundu, D.J.; Inuganti, A.; Griss, J.; Mayer, G.; Eisenacher, M.; et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res 2019, 47, D442–D450. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- Phipson, B.; Lee, S.; Majewski, I.J.; Alexander, W.S.; Smyth, G.K. Robust hyperparameter estimation protects against hypervariable genes and improves power to detect differential expression. Ann. Appl. Stat. 2016, 10, 946–963. [Google Scholar] [CrossRef]
- Okyay, T.O.; Rodrigues, D.F. High throughput colorimetric assay for rapid urease activity quantification. J. Microbiol. Methods 2013, 95, 324–326. [Google Scholar] [CrossRef] [PubMed]
- Christensen, W.B. Urea decomposition as a means of differentiating Proteus and paracolon cultures from each other and from Salmonella and Shigella types. J. Bacteriol. 1946, 52, 461–466. [Google Scholar] [CrossRef] [PubMed]
| Accession Number 1 | Protein Description | Log FC 2 | p-Value 3 | ||||
|---|---|---|---|---|---|---|---|
| ATCC * vs. 7108 | ATCC * vs. 9325 | 7108 vs. 9325 | ATCC * vs. 7108 | ATCC * vs. 9325 | 7108 vs. 9325 | ||
| Oxidative Stress | |||||||
| Q49WR2 | Thioredoxin | 0.494 | −0.133 | −0.627 | 0.011 | 0.480 | 0.001 |
| Q49YE4 | Probable thiol peroxidase | 0.280 | −0.419 | −0.699 | 0.376 | 0.085 | 0.005 |
| Q49UT8 | Alkyl hydroperoxide reductase subunit C | 0.490 | −0.408 | −0.899 | 0.026 | 0.040 | 0.000 |
| Q49XC1 | Catalase | 3.691 | 3.811 | 0.120 | 0.000 | 0.000 | 0.735 |
| Q49XN4 | Peptide methionine sulfoxide reductase MsrB | 0.228 | 0.460 | 0.232 | 0.449 | 0.044 | 0.339 |
| Q49XZ6 | Superoxide dismutase [Mn/Fe] | 0.338 | −0.406 | −0.744 | 0.128 | 0.038 | 0.000 |
| Q49YE0 | Putative universal stress protein SSP1056 | 2.055 | 2.055 | 0.000 | 0.000 | 0.000 | 1.000 |
| Q49UU5 | Nitronate monooxygenase | −0.339 | 1.063 | 1.402 | 0.402 | 0.001 | 0.000 |
| Nitrogen Metabolism | |||||||
| Q4A0J5 | Urease subunit alpha | −1.782 | 1.338 | 3.120 | 0.034 | 0.068 | 0.000 |
| Q4A0J8 | Urease accessory protein UreG | −0.416 | 0.936 | 1.352 | 0.464 | 0.038 | 0.003 |
| De Novo Purine Biosynthetic Pathway | |||||||
| Q49WI9 | N5-carboxyaminoimidazole ribonucleotide synthase PurK | 2.377 | 2.377 | 0.000 | 0.000 | 0.000 | 1.000 |
| Q49WJ0 | Phosphoribosylaminoimidazole-succinocarboxamide synthase PurC | 2.184 | 2.184 | 0.000 | 0.000 | 0.000 | 1.000 |
| Q49WJ1 | Phosphoribosylformylglycinamidine synthase subunit PurS | 1.035 | 1.035 | 0.000 | 0.000 | 0.000 | 1.000 |
| Q49WJ2 | Phosphoribosylformylglycinamidine synthase subunit PurQ | 1.006 | 1.006 | 0.000 | 0.011 | 0.007 | 1.000 |
| Q49WJ3 | Phosphoribosylformylglycinamidine synthase subunit PurL | 3.345 | 3.345 | 0.000 | 0.000 | 0.000 | 1.000 |
| Q49WJ5 | Phosphoribosylformylglycinamidine cyclo-ligase PurM | 2.365 | 2.365 | 0.000 | 0.000 | 0.000 | 1.000 |
| Q49WJ7 | Bifunctional purine biosynthesis protein PurH | 4.249 | 4.249 | 0.000 | 0.000 | 0.000 | 1.000 |
| Q49WJ8 | Phosphoribosylamine-glycine ligase PurD | 3.038 | 3.038 | 0.000 | 0.000 | 0.000 | 1.000 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Silva, K.C.S.; Silva, L.O.S.; Silva, G.A.A.; Borges, C.L.; Novaes, E.; Paccez, J.D.; Fontes, W.; Giambiagi-deMarval, M.; Soares, C.M.d.A.; Parente-Rocha, J.A. Staphylococcus saprophyticus Proteomic Analyses Elucidate Differences in the Protein Repertories among Clinical Strains Related to Virulence and Persistence. Pathogens 2020, 9, 69. https://doi.org/10.3390/pathogens9010069
Silva KCS, Silva LOS, Silva GAA, Borges CL, Novaes E, Paccez JD, Fontes W, Giambiagi-deMarval M, Soares CMdA, Parente-Rocha JA. Staphylococcus saprophyticus Proteomic Analyses Elucidate Differences in the Protein Repertories among Clinical Strains Related to Virulence and Persistence. Pathogens. 2020; 9(1):69. https://doi.org/10.3390/pathogens9010069
Chicago/Turabian StyleSilva, Karla Christina Sousa, Lana O’Hara Souza Silva, Guilherme Algusto Alves Silva, Clayton Luiz Borges, Evandro Novaes, Juliano Domiraci Paccez, Wagner Fontes, Marcia Giambiagi-deMarval, Célia Maria de Almeida Soares, and Juliana Alves Parente-Rocha. 2020. "Staphylococcus saprophyticus Proteomic Analyses Elucidate Differences in the Protein Repertories among Clinical Strains Related to Virulence and Persistence" Pathogens 9, no. 1: 69. https://doi.org/10.3390/pathogens9010069
APA StyleSilva, K. C. S., Silva, L. O. S., Silva, G. A. A., Borges, C. L., Novaes, E., Paccez, J. D., Fontes, W., Giambiagi-deMarval, M., Soares, C. M. d. A., & Parente-Rocha, J. A. (2020). Staphylococcus saprophyticus Proteomic Analyses Elucidate Differences in the Protein Repertories among Clinical Strains Related to Virulence and Persistence. Pathogens, 9(1), 69. https://doi.org/10.3390/pathogens9010069






