Understanding Flavivirus Capsid Protein Functions: The Tip of the Iceberg
Abstract
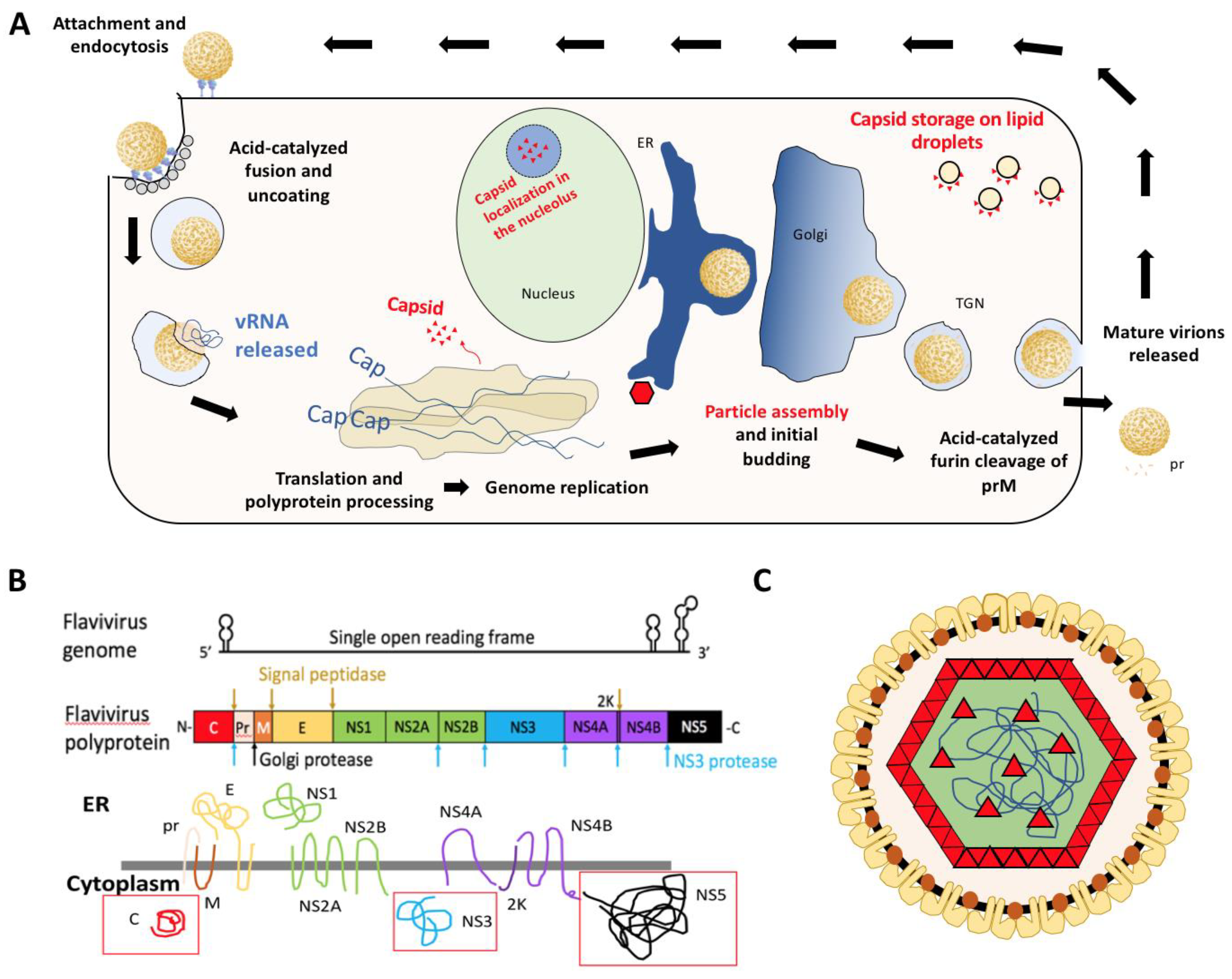
1. Introduction
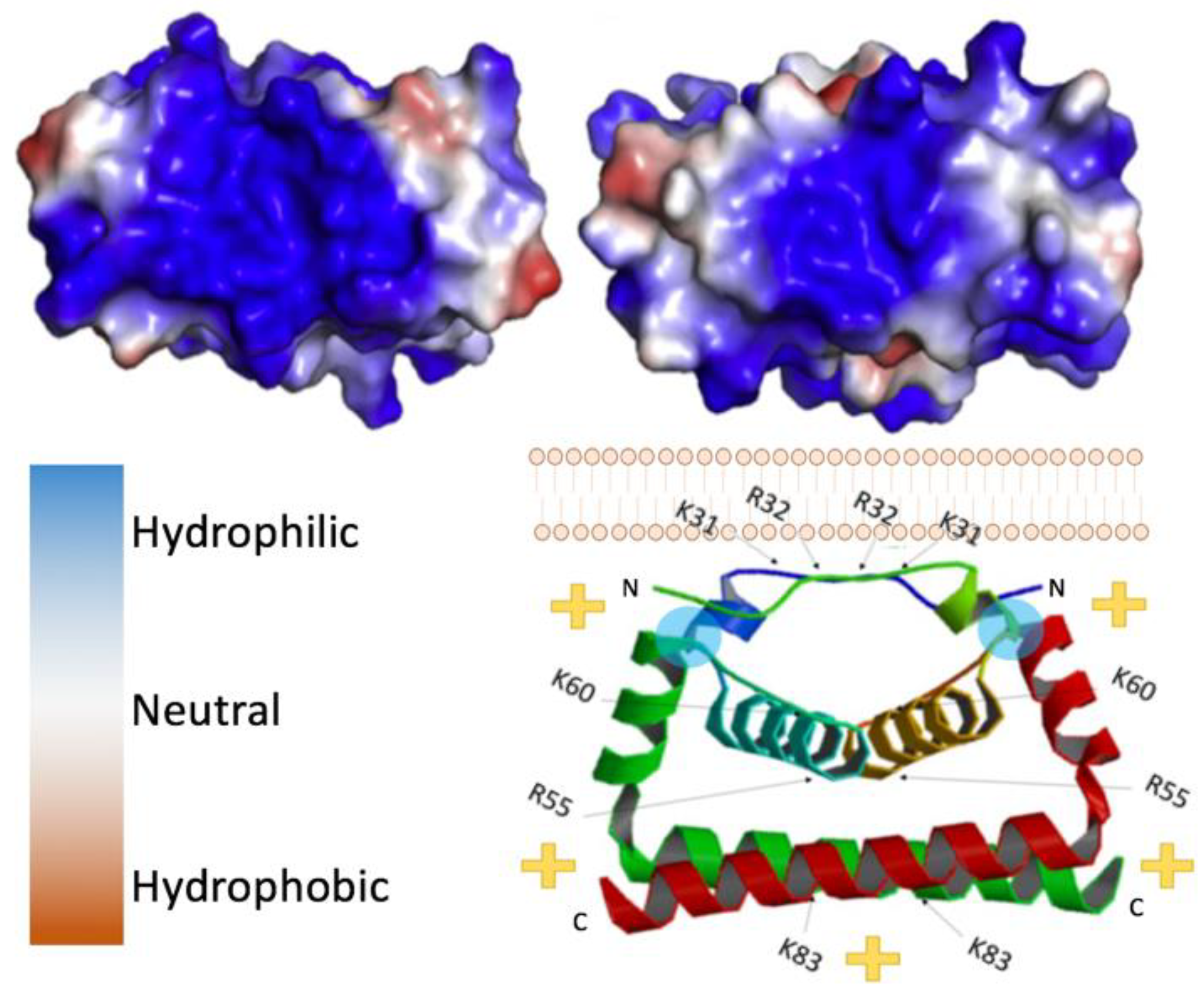
2. Structure of Flavivirus Capsid and Its Role in Packaging
3. Alternative Functions of Flavivirus Capsids
4. Indiscriminant Binding of Flavivirus Capsid to Nucleic Acids
5. Flavivirus Capsid as a Target for Antiviral Treatment
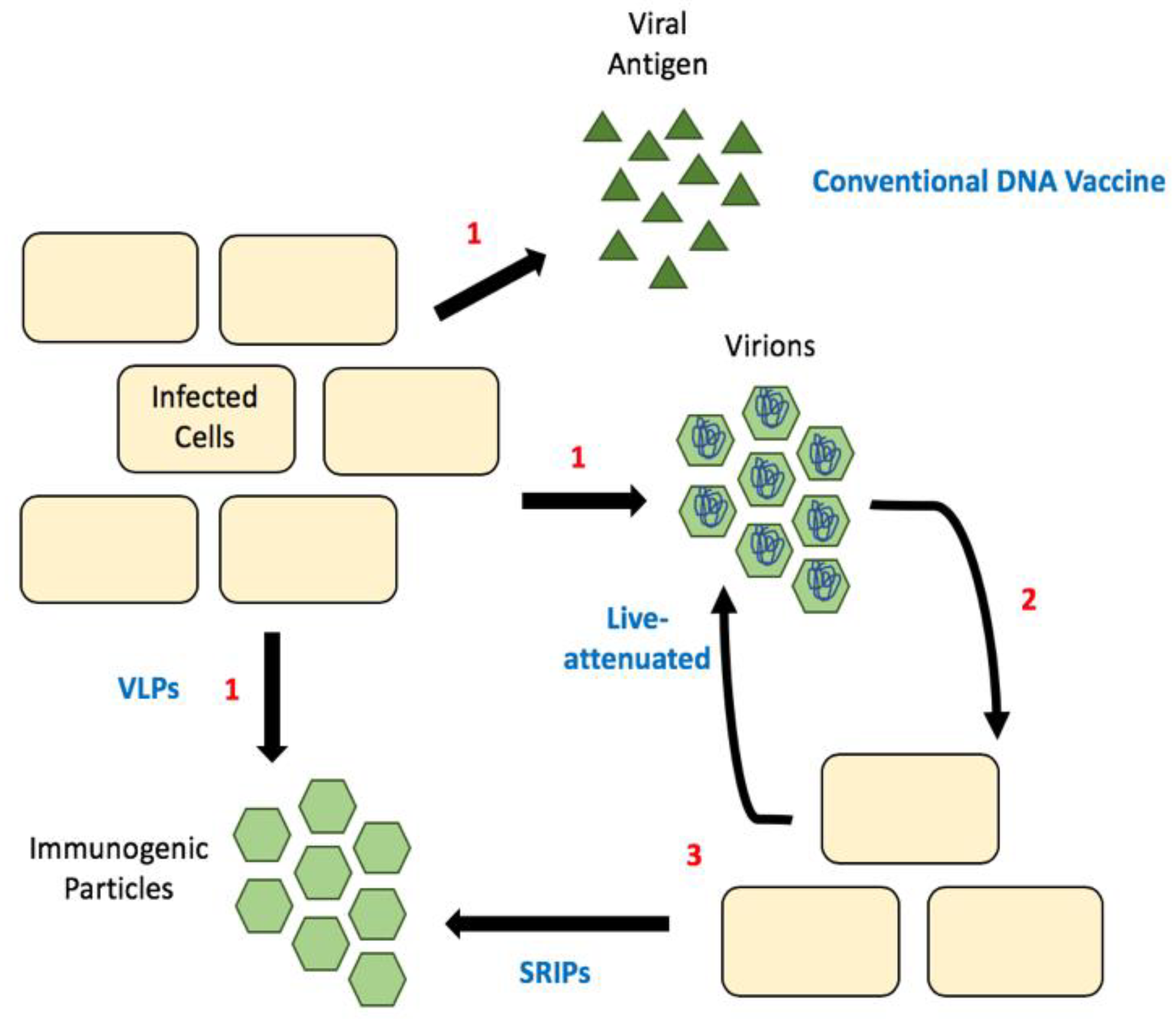
6. Considering Capsid in Flavivirus Vaccine Design
7. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Perera, R.; Khaliq, M.; Kuhn, R.J. Closing the door on flaviviruses: Entry as a target for antiviral drug design. Antivir. Res. 2008, 80, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Song, B.H.; Yun, S.I.; Woolley, M.; Lee, Y.M. Zika virus: History, epidemiology, transmission, and clinical presentation. J. Neuroimmunol. 2017, 308, 50–64. [Google Scholar] [CrossRef] [PubMed]
- Armstrong, P.M.; Andreadis, T.G.; Shepard, J.J.; Thomas, M.C. Northern Range Expansion of the Asian Tiger Mosquito (Aedes Albopictus): Analysis of Mosquito Data from Connecticut, USA. PLoS Negl. Trop. Dis. 2017, 11, e0005623. [Google Scholar] [CrossRef] [PubMed]
- Bartlow, A.W.; Manore, C.; Xu, C.; Kaufeld, K.A.; Del Valle, S.; Ziemann, A.; Fairchild, G.; Fair, J.M. Forecasting Zoonotic Infectious Disease Response to Climate Change: Mosquito Vectors and a Changing Environment. Vet. Sci. 2019, 6. [Google Scholar] [CrossRef]
- Kraemer, M.U.; Sinka, M.E.; Duda, K.A.; Mylne, A.Q.; Shearer, F.M.; Barker, C.M.; Moore, C.G.; Carvalho, R.G.; Coelho, G.E.; Van Bortel, W.; et al. The Global Distribution of the Arbovirus Vectors Aedes Aegypti and Ae. Albopictus. Elife 2015, 4, e08347. [Google Scholar] [CrossRef]
- Ming, G.L.; Tang, H.; Song, H. Advances in Zika Virus Research: Stem Cell Models, Challenges, and Opportunities. Cell Stem Cell 2016, 19, 690–702. [Google Scholar] [CrossRef]
- Villar, L.; Dayan, G.H.; Arredondo-García, J.L.; Rivera, D.M.; Cunha, R.; Deseda, C.; Reynales, H.; Costa, M.S.; Morales-Ramírez, J.O.; Carrasquilla, G.; et al. Efficacy of a Tetravalent Dengue Vaccine in Children in Latin America. N. Engl. J. Med. 2015, 372, 113–123. [Google Scholar] [CrossRef]
- Hadinegoro, S.R.; Arredondo-García, J.L.; Capeding, M.R.; Deseda, C.; Chotpitayasunondh, T.; Dietze, R.; Ismail, H.I.M.; Reynales, H.; Limkittikul, K.; Rivera-Medina, D.M.; et al. Efficacy and Long-Term Safety of a Dengue Vaccine in Regions of Endemic Disease. N. Engl. J. Med. 2015, 373, 1195–1206. [Google Scholar] [CrossRef]
- Dyer, O. Philippines halts dengue immunisation campaign owing to safety risk. BMJ 2017, 359, j5759. [Google Scholar] [CrossRef]
- Garcia-Blanco, M.A.; Vasudevan, S.G.; Bradrick, S.S.; Nicchitta, C. Flavivirus RNA transactions from viral entry to genome replication. Antivir. Res. 2016, 134, 244–249. [Google Scholar] [CrossRef]
- Neufeldt, C.J.; Cortese, M.; Acosta, E.G.; Bartenschlager, R. Rewiring cellular networks by members of the Flaviviridae family. Nat. Rev. Microbiol. 2018, 16, 125–142. [Google Scholar] [CrossRef] [PubMed]
- Blazevic, J.; Rouha, H.; Bradt, V.; Heinz, F.X.; Stiasny, K. Membrane Anchors of the Structural Flavivirus Proteins and Their Role in Virus Assembly. J. Virol. 2016, 90, 6365–6378. [Google Scholar] [CrossRef] [PubMed]
- Chang, G.J.; Davis, B.S.; Hunt, A.R.; Holmes, D.A.; Kuno, G. Flavivirus DNA vaccines: Current status and potential. Ann. N. Y. Acad. Sci. 2001, 951, 272–285. [Google Scholar] [CrossRef] [PubMed]
- Martins, I.C.; Gomes-Neto, F.; Faustino, A.F.; Carvalho, F.A.; Carneiro, F.A.; Bozza, P.T.; Mohana-Borges, R.; Castanho, M.A.R.B.; Almeida, F.C.L.; Santos, N.C.; et al. The disordered N-terminal region of dengue virus capsid protein contains a lipid-droplet-binding motif. Biochem. J. 2012, 444, 405–415. [Google Scholar] [CrossRef]
- Byk, L.A.; Gamarnik, A.V. Properties and Functions of the Dengue Virus Capsid Protein. Annu. Rev. Virol. 2016, 3, 263–281. [Google Scholar] [CrossRef]
- Iglesias, N.G.; Mondotte, J.A.; Byk, L.A.; De Maio, F.A.; Samsa, M.M.; Alvarez, C.; Gamarnik, A.V. Dengue Virus Uses a Non-Canonical Function of the Host GBF1-Arf-COPI System for Capsid Protein Accumulation on Lipid Droplets. Traffic 2015, 16, 962–977. [Google Scholar] [CrossRef]
- Samsa, M.M.; Mondotte, J.A.; Iglesias, N.G.; Assunção-Miranda, I.; Barbosa-Lima, G.; Da Poian, A.T.; Bozza, P.T.; Gamarnik, A.V. Dengue Virus Capsid Protein Usurps Lipid Droplets for Viral Particle Formation. PLoS Pathog. 2009, 5, e1000632. [Google Scholar] [CrossRef]
- Ishida, K.; Goto, S.; Ishimura, M.; Amanuma, M.; Hara, Y.; Suzuki, R.; Katoh, K.; Morita, E. Functional Correlation between Subcellular Localizations of Japanese Encephalitis Virus Capsid Protein and Virus Production. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Oh, W.; Yang, M.R.; Lee, E.W.; Park, K.M.; Pyo, S.; Yang, J.S.; Lee, H.W.; Song, J. Jab1 Mediates Cytoplasmic Localization and Degradation of West Nile Virus Capsid Protein. J. Boil. Chem. 2006, 281, 30166–30174. [Google Scholar] [CrossRef]
- Slomnicki, L.P.; Chung, N.H.; Parker, A.; Hermann, T.; Boyd, N.L.; Hetman, M. Ribosomal stress and Tp53-mediated neuronal apoptosis in response to capsid protein of the Zika virus. Sci. Rep. 2017, 7, 16652. [Google Scholar] [CrossRef]
- Tsuda, Y.; Mori, Y.; Abe, T.; Yamashita, T.; Okamoto, T.; Ichimura, T.; Moriishi, K.; Matsuura, Y. Nucleolar protein B23 interacts with Japanese encephalitis virus core protein and participates in viral replication. Microbiol. Immunol. 2006, 50, 225–234. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Anderson, R.; Hobman, T.C. The Capsid-Binding Nucleolar Helicase DDX56 Is Important for Infectivity of West Nile Virus. J. Virol. 2011, 85, 5571–5580. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.R.; Lee, S.R.; Oh, W.; Lee, E.W.; Yeh, J.Y.; Nah, J.J.; Joo, Y.S.; Shin, J.; Lee, H.W.; Pyo, S.; et al. West Nile Virus Capsid Protein Induces P53-Mediated Apoptosis Via the Sequestration of Hdm2 to the Nucleolus. Cell. Microbiol. 2008, 10, 165–176. [Google Scholar] [CrossRef] [PubMed]
- Gillespie, L.K.; Hoenen, A.; Morgan, G.; MacKenzie, J.M. The Endoplasmic Reticulum Provides the Membrane Platform for Biogenesis of the Flavivirus Replication Complex. J. Virol. 2010, 84, 10438–10447. [Google Scholar] [CrossRef] [PubMed]
- Junjhon, J.; Pennington, J.G.; Edwards, T.J.; Perera, R.; Lanman, J.; Kuhn, R.J. Ultrastructural Characterization and Three-Dimensional Architecture of Replication Sites in Dengue Virus-Infected Mosquito Cells. J. Virol. 2014, 88, 4687–4697. [Google Scholar] [CrossRef] [PubMed]
- Welsch, S.; Miller, S.; Romero-Brey, I.; Merz, A.; Bleck, C.K.; Walther, P.; Fuller, S.D.; Antony, C.; Krijnse-Locker, J.; Bartenschlager, R. Composition and Three-Dimensional Architecture of the Dengue Virus Replication and Assembly Sites. Cell Host Microbe 2009, 5, 365–375. [Google Scholar] [CrossRef]
- Cortese, M.; Goellner, S.; Acosta, E.G.; Neufeldt, C.J.; Oleksiuk, O.; Lampe, M.; Haselmann, U.; Funaya, C.; Schieber, N.; Ronchi, P.; et al. Ultrastructural Characterization of Zika Virus Replication Factories. Cell Rep. 2017, 18, 2113–2123. [Google Scholar] [CrossRef]
- Kümmerer, B.M.; Rice, C.M. Mutations in the Yellow Fever Virus Nonstructural Protein NS2A Selectively Block Production of Infectious Particles. J. Virol. 2002, 76, 4773–4784. [Google Scholar] [CrossRef]
- Liu, W.J.; Chen, H.B.; Khromykh, A.A. Molecular and Functional Analyses of Kunjin Virus Infectious cDNA Clones Demonstrate the Essential Roles for NS2A in Virus Assembly and for a Nonconservative Residue in NS3 in RNA Replication. J. Virol. 2003, 77, 7804–7813. [Google Scholar] [CrossRef]
- Zhang, X.W.; Xie, X.P.; Xia, H.J.; Zou, J.; Huang, L.F.; Popov, V.L.; Chen, X.W.; Shi, P.Y. Zika Virus NS2A-Mediated Virion Assembly. MBio 2019, 10. [Google Scholar] [CrossRef]
- Xie, X.; Zou, J.; Zhang, X.; Zhou, Y.; Routh, A.L.; Kang, C.; Popov, V.L.; Chen, X.; Wang, Q.Y.; Dong, H.; et al. Dengue NS2A Protein Orchestrates Virus Assembly. Cell Host Microbe 2019, 26, 606–622. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.C.; Huang, Z.S.; Chiang, P.L.; Chen, C.T.; Wu, H.N. Analysis of the nucleoside triphosphatase, RNA triphosphatase, and unwinding activities of the helicase domain of dengue virus NS3 protein. FEBS Lett. 2009, 583, 691–696. [Google Scholar] [CrossRef] [PubMed]
- Filomatori, C.V.; Lodeiro, M.F.; Alvarez, D.E.; Samsa, M.M.; Pietrasanta, L.; Gamarnik, A.V. A 5′ RNA element promotes dengue virus RNA synthesis on a circular genome. Genome Dev. 2006, 20, 2238–2249. [Google Scholar] [CrossRef] [PubMed]
- Pong, W.L.; Huang, Z.S.; Teoh, P.G.; Wang, C.C.; Wu, H.N. RNA binding property and RNA chaperone activity of dengue virus core protein and other viral RNA-interacting proteins. FEBS Lett. 2011, 585, 2575–2581. [Google Scholar] [CrossRef]
- Shang, Z.; Song, H.; Shi, Y.; Qi, J.; Gao, G.F. Crystal Structure of the Capsid Protein from Zika Virus. J. Mol. Biol. 2018, 430, 948–962. [Google Scholar] [CrossRef]
- Bhuvanakantham, R.; Chong, M.K.; Ng, M.L. Specific Interaction of Capsid Protein and Importin-Alpha/Beta Influences West Nile Virus Production. Biochem. Biophys. Res. Commun. 2009, 389, 63–69. [Google Scholar] [CrossRef]
- Yang, J.S.; Ramanathan, M.P.; Muthumani, K.; Choo, A.Y.; Jin, S.H.; Yu, Q.C.; Hwang, D.S.; Choo, D.K.; Lee, M.D.; Dang, K.; et al. Induction of Inflammation by West Nile virus Capsid through the Caspase-9 Apoptotic Pathway. Emerg. Infect. Dis. 2002, 8, 1379–1384. [Google Scholar] [CrossRef]
- Sessions, O.M.; Tan, Y.; Goh, K.C.; Liu, Y.; Tan, P.; Rozen, S.; Ooi, E.E. Host Cell Transcriptome Profile during Wild-Type and Attenuated Dengue Virus Infection. PLoS Negl. Trop. Dis. 2013, 7, e2107. [Google Scholar] [CrossRef]
- Singh, P.K.; Khatri, I.; Jha, A.; Pretto, C.D.; Spindler, K.R.; Arumugaswami, V.; Giri, S.; Kumar, A.; Bhasin, M.K. Determination of system level alterations in host transcriptome due to Zika virus (ZIKV) Infection in retinal pigment epithelium. Sci. Rep. 2018, 8, 11209. [Google Scholar] [CrossRef]
- Etebari, K.; Hegde, S.; Saldana, M.A.; Widen, S.G.; Wood, T.G.; Asgari, S.; Hughes, G.L. Global transcriptome analysis of Aedes aegypti mosquitoes in response to Zika virus infection. MSphere 2017, 2. [Google Scholar] [CrossRef]
- Yoshii, K. Epidemiology and pathological mechanisms of tick-borne encephalitis. J. Vet. Med. Sci. 2019, 81, 343–347. [Google Scholar] [CrossRef] [PubMed]
- Dokland, T.; Walsh, M.; Mackenzie, J.M.; Khromykh, A.A.; Ee, K.H.; Wang, S. West Nile Virus Core Protein; Tetramer Structure and Ribbon Formation. Structure 2004, 12, 1157–1163. [Google Scholar] [CrossRef] [PubMed]
- Poonsiri, T.; Wright, G.S.A.; Solomon, T.; Antonyuk, S.V. Crystal Structure of the Japanese Encephalitis Virus Capsid Protein. Viruses 2019, 11, 623. [Google Scholar] [CrossRef] [PubMed]
- Jones, C.T.; Ma, L.; Burgner, J.W.; Groesch, T.D.; Post, C.B.; Kuhn, R.J. Flavivirus Capsid Is a Dimeric Alpha-Helical Protein. J. Virol. 2003, 77, 7143–7149. [Google Scholar] [CrossRef]
- Samsa, M.M.; Mondotte, J.A.; Caramelo, J.J.; Gamarnik, A.V. Uncoupling Cis-Acting Rna Elements from Coding Sequences Revealed a Requirement of the N-Terminal Region of Dengue Virus Capsid Protein in Virus Particle Formation. J. Virol. 2012, 86, 1046–1058. [Google Scholar] [CrossRef]
- Ma, L.; Jones, C.T.; Groesch, T.D.; Kuhn, R.J.; Post, C.B. Solution structure of dengue virus capsid protein reveals another fold. Proc. Natl. Acad. Sci. USA 2004, 101, 3414–3419. [Google Scholar] [CrossRef]
- Patkar, C.G.; Jones, C.T.; Chang, Y.H.; Warrier, R.; Kuhn, R.J. Functional Requirements of the Yellow Fever Virus Capsid Protein. J. Virol. 2007, 81, 6471–6481. [Google Scholar] [CrossRef]
- Samuel, G.H.; Wiley, M.R.; Badawi, A.; Adelman, Z.N.; Myles, K.M. Yellow fever virus capsid protein is a potent suppressor of RNA silencing that binds double-stranded RNA. Proc. Natl. Acad. Sci. USA 2016, 113, 13863–13868. [Google Scholar] [CrossRef]
- Li, T.; Zhao, Q.; Yang, X.; Chen, C.; Yang, K.; Wu, C.; Zhang, T.; Duan, Y.; Xue, X.; Mi, K.; et al. Structural Insight into the Zika Virus Capsid Encapsulating the Viral Genome. Cell Res. 2018, 28, 497–499. [Google Scholar] [CrossRef]
- Allan, J. Participation of core histone “tails” in the stabilization of the chromatin solenoid. J. Cell Boil. 1982, 93, 285–297. [Google Scholar] [CrossRef]
- Requiao, R.D.; Carneiro, R.L.; Moreira, M.H.; Ribeiro-Alves, M.; Rossetto, S.; Palhano, F.L.; Domitrovic, T. Icosahedral viruses defined by their positively charged domains: A signature for viral identity and capsid assembly strategy. BioRxiv 2019. [Google Scholar] [CrossRef]
- Thiele, C.; Spandl, J. Cell biology of lipid droplets. Curr. Opin. Cell Biol. 2008, 20, 378–385. [Google Scholar] [CrossRef] [PubMed]
- Bulich, R.; Aaskov, J.G. Nuclear localization of dengue 2 virus core protein detected with monoclonal antibodies. J. Gen. Virol. 1992, 73, 2999–3003. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, E.R.; De Alencastro, R.B.; Horta, B.A.; Mohana-Borges, R. The flavivirus capsid protein: Structure, function and perspectives towards drug design. Virus Res. 2017, 227, 115–123. [Google Scholar] [CrossRef] [PubMed]
- Urbanowski, M.D.; Ilkow, C.S.; Hobman, T.C. Modulation of signaling pathways by RNA virus capsid proteins. Cell. Signal. 2008, 20, 1227–1236. [Google Scholar] [CrossRef] [PubMed]
- Balinsky, C.A.; Schmeisser, H.; Ganesan, S.; Singh, K.; Pierson, T.C.; Zoon, K.C. Nucleolin Interacts with the Dengue Virus Capsid Protein and Plays a Role in Formation of Infectious Virus Particles. J. Virol. 2013, 87, 13094–13106. [Google Scholar] [CrossRef]
- Varjak, M.; Donald, C.L.; Mottram, T.J.; Sreenu, V.B.; Merits, A.; Maringer, K.; Schnettler, E.; Kohl, A. Characterization of the Zika virus induced small RNA response in Aedes aegypti cells. PLoS Negl. Trop. Dis. 2017, 11, e0006010. [Google Scholar] [CrossRef]
- Fontaine, K.A.; Leon, K.E.; Khalid, M.M.; Tomar, S.; Jimenez-Morales, D.; Dunlap, M.; Kaye, J.A.; Shah, P.S.; Finkbeiner, S.; Krogan, N.J.; et al. The Cellular NMD Pathway Restricts Zika Virus Infection and Is Targeted by the Viral Capsid Protein. MBio 2018, 9. [Google Scholar] [CrossRef]
- Colpitts, T.M.; Barthel, S.; Wang, P.; Fikrig, E. Dengue Virus Capsid Protein Binds Core Histones and Inhibits Nucleosome Formation in Human Liver Cells. PLoS ONE 2011, 6, e24365. [Google Scholar] [CrossRef]
- Netsawang, J.; Noisakran, S.; Puttikhunt, C.; Kasinrerk, W.; Wongwiwat, W.; Malasit, P.; Yenchitsomanus, P.T.; Limjindaporn, T. Nuclear localization of dengue virus capsid protein is required for DAXX interaction and apoptosis. Virus Res. 2010, 147, 275–283. [Google Scholar] [CrossRef]
- Coyaud, E.; Ranadheera, C.; Cheng, D.T.; Goncalves, J.; Dyakov, B.J.A.; Laurent, E.M.N.; St-Germain, J.R.; Pelletier, L.; Gingras, A.C.; Brumell, J.H.; et al. Global Interactomics Uncovers Extensive Organellar Targeting by Zika Virus. Mol. Cell. Proteom. 2018, 17, 2242–2255. [Google Scholar] [CrossRef] [PubMed]
- Machara, A.; Lux, V.; Kožíšek, M.; Šašková, K.G.; Štěpánek, O.; Kotora, M.; Parkan, K.; Pávová, M.; Glass, B.; Sehr, P.; et al. Specific Inhibitors of HIV Capsid Assembly Binding to the C-Terminal Domain of the Capsid Protein: Evaluation of 2-Arylquinazolines as Potential Antiviral Compounds. J. Med. Chem. 2016, 59, 545–558. [Google Scholar] [CrossRef] [PubMed]
- Phelps, D.K.; Post, C.B. A novel basis of capsid stabilization by antiviral compounds. J. Mol. Biol. 1995, 254, 544–551. [Google Scholar] [CrossRef] [PubMed]
- Bocanegra, R.; Rodriguez-Huete, A.; Fuertes, M.Á.; Del Álamo, M.; Mateu, M.G. Molecular recognition in the human immunodeficiency virus capsid and antiviral design. Virus Res. 2012, 169, 388–410. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, M.; Kaur, R.; Saha, A.; Mudgal, R.; Yadav, R.; Dash, P.K.; Parida, M.; Kumar, P.; Tomar, S. Evaluation of antiviral activity of piperazine against Chikungunya virus targeting hydrophobic pocket of alphavirus capsid protein. Antivir. Res. 2017, 146, 102–111. [Google Scholar] [CrossRef]
- Tang, C.; Loeliger, E.; Kinde, I.; Kyere, S.; Mayo, K.; Barklis, E.; Sun, Y.; Huang, M.; Summers, M.F. Antiviral inhibition of the HIV-1 capsid protein. J. Mol. Biol. 2003, 327, 1013–1020. [Google Scholar] [CrossRef]
- Rossmann, M.G. Antiviral agents targeted to interact with viral capsid proteins and a possible application to human immunodeficiency virus. Proc. Natl. Acad. Sci. USA 1988, 85, 4625–4627. [Google Scholar] [CrossRef]
- Touret, F.; Baronti, C.; Goethals, O.; Van Loock, M.; De Lamballerie, X.; Querat, G. Phylogenetically based establishment of a dengue virus panel, representing all available genotypes, as a tool in dengue drug discovery. Antivir. Res. 2019, 168, 109–113. [Google Scholar] [CrossRef]
- Byrd, C.M.; Dai, D.; Grosenbach, D.W.; Berhanu, A.; Jones, K.F.; Cardwell, K.B.; Schneider, C.; Wineinger, K.A.; Page, J.M.; Harver, C.; et al. A Novel Inhibitor of Dengue Virus Replication That Targets the Capsid Protein. Antimicrob. Agents Chemother. 2013, 57, 15–25. [Google Scholar] [CrossRef]
- Scaturro, P.; Trist, I.M.L.; Paul, D.; Kumar, A.; Acosta, E.G.; Byrd, C.M.; Jordan, R.; Brancale, A.; Bartenschlager, R. Characterization of the Mode of Action of a Potent Dengue Virus Capsid Inhibitor. J. Virol. 2014, 88, 11540–11555. [Google Scholar] [CrossRef]
- Boon, P.L.S.; Martins, A.S.; Enguita, F.J.; Lim, X.N.; Santos, N.C.; Matsudaira, P.T.; Martins, I.C.; Yue, W.; Bond, P.J.; Huber, R.G. Genome-Wide Associations of Flavivirus Capsid Proteins. BioRxiv 2019. [Google Scholar] [CrossRef]
- Soto-Acosta, R.; Bautista-Carbajal, P.; Syed, G.H.; Siddiqui, A.; Del Angel, R.M. Nordihydroguaiaretic acid (NDGA) inhibits replication and viral morphogenesis of dengue virus. Antivir. Res. 2014, 109, 132–140. [Google Scholar] [CrossRef] [PubMed]
- Faizan, I.; Abdullah, M.; Ali, S.; Naqvi, I.H.; Ahmed, A.; Parveen, S. Zika Virus-Induced Microcephaly and Its Possible Molecular Mechanism. Intervirology 2016, 59, 152–158. [Google Scholar] [CrossRef] [PubMed]
- Mlakar, J.; Korva, M.; Tul, N.; Popovic, M.; Poljsak-Prijatelj, M.; Mraz, J.; Kolenc, M.; Rus, K.R.; Vipotnik, T.V.; Vodušek, V.F.; et al. Zika Virus Associated with Microcephaly. N. Engl. J. Med. 2016, 374, 951–958. [Google Scholar] [CrossRef] [PubMed]
- Seino, K.K.; Long, M.T.; Gibbs, E.P.J.; Bowen, R.A.; Beachboard, S.E.; Humphrey, P.P.; Dixon, M.A.; Bourgeois, M.A. Comparative Efficacies of Three Commercially Available Vaccines against West Nile Virus (WNV) in a Short-Duration Challenge Trial Involving an Equine WNV Encephalitis Model. Clin. Vaccine Immunol. 2007, 14, 1465–1471. [Google Scholar] [CrossRef] [PubMed]
- Ostlund, E.N.; Crom, R.L.; Pedersen, D.D.; Johnson, D.J.; Williams, W.O.; Schmitt, B.J. Equine West Nile Encephalitis, United States. Emerg. Infect. Dis. 2001, 7, 665–669. [Google Scholar] [CrossRef]
- Galler, R.; Freire, M.; Jabor, A.; Mann, G. The yellow fever 17D vaccine virus: Molecular basis of viral attenuation and its use as an expression vector. Braz. J. Med Biol. Res. 1997, 30, 157–168. [Google Scholar] [CrossRef]
- Jelinek, T. Ixiaro®: A new vaccine against Japanese encephalitis. Expert Rev. Vaccines 2009, 8, 1501–1511. [Google Scholar] [CrossRef]
- Vaccine Information Statement. Available online: https://www.cdc.gov/vaccines/hcp/vis/vis-statements/yf.html (accessed on 25 August 2019).
- Martin, M.; Tsai, T.F.; Cropp, B.; Chang, G.J.J.; Holmes, D.A.; Tseng, J.; Shieh, W.J.; Zaki, S.R.; Al-Sanouri, I.; Cutrona, A.F.; et al. Fever and multisystem organ failure associated with 17D-204 yellow fever vaccination: A report of four cases. Lancet 2001, 358, 98–104. [Google Scholar] [CrossRef]
- World Health Organization. A Global Strategy to Eliminate Yellow Fever Epidemics (Eye) 2017–2026; World Health Organization: Geneva, Switzerland, 2018. [Google Scholar]
- Collins, N. Contributions of Structural and Non-Structural Genes to the Attenuation of the 17d Vaccine. Ph.D. Thesis, University of Texas Medical Branch, Galveston, TX, USA, 2017. [Google Scholar]
- Beck, A.; Tesh, R.B.; Wood, T.G.; Widen, S.G.; Ryman, K.D.; Barrett, A.D. Comparison of the Live Attenuated Yellow Fever Vaccine 17d-204 Strain to Its Virulent Parental Strain Asibi by Deep Sequencing. J. Infect. Dis. 2014, 209, 334–344. [Google Scholar] [CrossRef]
- Villordo, S.M.; Gamarnik, A.V. Genome Cyclization as Strategy for Flavivirus Rna Replication. Virus Res. 2009, 139, 230–239. [Google Scholar] [CrossRef] [PubMed]
- Mandl, C.W. Flavivirus Immunization with Capsid-Deletion Mutants: Basics, Benefits, and Barriers. Viral Immunol. 2004, 17, 461–472. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.; Kum, D.B.; Xia, H.; Luo, H.; Shan, C.; Zou, J.; Muruato, A.E.; Medeiros, D.B.; Nunes, B.T.; Dallmeier, K.; et al. A Single-Dose Live-Attenuated Zika Virus Vaccine with Controlled Infection Rounds that Protects against Vertical Transmission. Cell Host Microbe 2018, 24, 487–499. [Google Scholar] [CrossRef] [PubMed]
- Schlick, P.; Kofler, R.M.; Schittl, B.; Taucher, C.; Nagy, E.; Meinke, A.; Mandl, C.W. Characterization of West Nile virus live vaccine candidates attenuated by capsid deletion mutations. Vaccine 2010, 28, 5903–5909. [Google Scholar] [CrossRef]
- Shan, C.; Xie, X.; Zou, J.; Züst, R.; Zhang, B.; Ambrose, R.; MacKenzie, J.; Fink, K.; Shi, P.Y. Using a virion assembly-defective dengue virus as a vaccine approach. J. Virol. 2018, 92. [Google Scholar] [CrossRef]
- Barrett, A.D.T. Flavivirus DNA vaccine with a kick. Nat. Biotechnol. 2008, 26, 525–526. [Google Scholar] [CrossRef]
- Barrett, A.D.T. Current status of Zika vaccine development: Zika vaccines advance into clinical evaluation. NPJ Vaccines 2018, 3, 24. [Google Scholar] [CrossRef]
- Beckett, C.G.; Tjaden, J.; Burgess, T.; Danko, J.R.; Tamminga, C.; Simmons, M.; Wu, S.J.; Sun, P.; Kochel, T.; Raviprakash, K.; et al. Evaluation of a prototype dengue-1 DNA vaccine in a Phase 1 clinical trial. Vaccine 2011, 29, 960–968. [Google Scholar] [CrossRef]
- Gaudinski, M.R.; Houser, K.V.; Morabito, K.M.; Hu, Z.; Yamshchikov, G.; Rothwell, R.S.; Berkowitz, N.; Mendoza, F.; Saunders, J.G.; Novik, L.; et al. Safety, Tolerability, and Immunogenicity of Two Zika Virus DNA Vaccine Candidates in Healthy Adults: Randomised, Open-Label, Phase 1 Clinical Trials. Lancet 2018, 391, 552–562. [Google Scholar] [CrossRef]
- Chang, D.C.; Liu, W.J.; Anraku, I.; Clark, D.C.; Pollitt, C.C.; Suhrbier, A.; Hall, R.A.; Khromykh, A.A. Single-round infectious particles enhance immunogenicity of a DNA vaccine against West Nile virus. Nat. Biotechnol. 2008, 26, 571–577. [Google Scholar] [CrossRef]
- Roby, J.A.; Bielefeldt-Ohmann, H.; Prow, N.A.; Chang, D.C.; Hall, R.A.; Khromykh, A.A. Increased expression of capsid protein in trans enhances production of single-round infectious particles by West Nile virus DNA vaccine candidate. J. Gen. Virol. 2014, 95, 2176–2191. [Google Scholar] [CrossRef] [PubMed]
- Dejnirattisai, W.; Supasa, P.; Wongwiwat, W.; Rouvinski, A.; Barba-Spaeth, G.; Duangchinda, T.; Sakuntabhai, A.; Cao-Lormeau, V.M.; Malasit, P.; Rey, F.A.; et al. Dengue virus sero-cross-reactivity drives antibody-dependent enhancement of infection with zika virus. Nat. Immunol. 2016, 17, 1102–1108. [Google Scholar] [CrossRef] [PubMed]
- Raboni, S.M.; Bonfim, C.; Almeida, B.M.; Zanluca, C.; Koishi, A.C.; Rodrigues, P.R.; Kay, C.K.; Ribeiro, L.L.; Scola, R.H.; Dos Santos, C.N.D. Flavivirus cross-reactivity in serological tests and Guillain-Barr? syndrome in a hematopoietic stem cell transplant patient: A case report. Transpl. Infect. Dis. 2017, 19, e12700. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Shen, H.; Jia, R.; Wang, M.; Chen, S.; Zhu, D.; Liu, M.; Zhao, X.; Yang, Q.; Wu, Y.; et al. Oral Vaccination with a DNA Vaccine Encoding Capsid Protein of Duck Tembusu Virus Induces Protection Immunity. Viruses 2018, 10, 180. [Google Scholar] [CrossRef]
- Lobigs, M.; Müllbacher, A.; Regner, M. Mhc Class I up-Regulation by Flaviviruses: Immune Interaction with Unknown Advantage to Host or Pathogen. Immunol. Cell Biol. 2003, 81, 217–223. [Google Scholar] [CrossRef]
- Lobigs, M.; Müllbacher, A.; Lee, E. Evidence That a Mechanism for Efficient Flavivirus Budding Upregulates Mhc Class I. Immunol. Cell Biol. 2004, 82, 184–188. [Google Scholar] [CrossRef]
- Momburg, F.; Müllbacher, A.; Lobigs, M. Modulation of Transporter Associated with Antigen Processing (TAP)-Mediated Peptide Import into the Endoplasmic Reticulum by Flavivirus Infection. J. Virol. 2001, 75, 5663–5671. [Google Scholar] [CrossRef]
- Müllbacher, A.; Lobigs, M. Up-Regulation of Mhc Class I by Flavivirus-Induced Peptide Translocation into the Endoplasmic Reticulum. Immunity 1995, 3, 207–214. [Google Scholar] [CrossRef]
- Ryu, W.S. Molecular Virology of Human Pathogenic Viruses; Elsevier, Inc.: Amsterdam, The Netherlands, 2017. [Google Scholar]
- Apte-Sengupta, S.; Sirohi, D.; Kuhn, R.J. Coupling of replication and assembly in flaviviruses. Curr. Opin. Virol. 2014, 9, 134–142. [Google Scholar] [CrossRef]
- Hasan, S.S.; Sevvana, M.; Kuhn, R.J.; Rossmann, M.G. Structural biology of Zika virus and other flaviviruses. Nat. Struct. Mol. Biol. 2018, 25, 13–20. [Google Scholar] [CrossRef]
- Paul, D.; Bartenschlager, R. Flaviviridae Replication Organelles: Oh, What a Tangled Web We Weave. Annu. Rev. Virol. 2015, 2, 289–310. [Google Scholar] [CrossRef] [PubMed]
| Location. | Pathway | Protein | Result | Pro- or Anti- viral | Antiviral treatment could | References |
|---|---|---|---|---|---|---|
| Endoplasmic Reticulum | Vesicular Transport | YKT6 SCFD1 USE1 VAPB STX8 TMEM85 SAR1B | Movement of capsid from site of cleavage to lipid droplets or nucleus | Pro | Prevent capsid storage on lipid droplets, entry into nucleus/nucleolus, prevent particle assembly | [61] |
| Signal Peptidase Complex | SPC52 SEC11A SPC53 | Cleavage of capsid from flavivirus polyprotein | ||||
| ER Stress Response | PARP16 DORGK1 UBXN8 RTN3 SLC33A1 | Cell survival or cell death | Anti | |||
| Nucleus | Importins | IPO7 IPO9 | Targeting of viral capsid to nucleus | Both | Prevent capsid interaction & entry into host nucleus | [19,20,30,36,61] |
| LINC Complex | SUN1 SYNE2 | Interaction with nuclear membrane | [61] | |||
| NPC | NUP35 NUPL2 TMEM48 POM121 | Entry of viral capsid into host nucleus | [61] | |||
| NMD Pathway | PABPC UPF1 | Prevent degradation of viral transcripts, increase degradation of host transcripts | Pro | Allow degradation of viral transcripts by host cells | [58] | |
| Host Transcription | DAXX | Prevent binding of host transcription factors | Prevent changes in host gene transcription | [60] | ||
| Core Histones | Change position of histones | [59] | ||||
| Nucleolus | p53 regulation | MDM2 | p53 induced apoptosis | Anti | [23] | |
| RNA methylation | NSUN2 | Modified methylation of tRNAs, mRNAs, and ncRNAs | Not yet understood | Prevent changes in transcript methylation | [61] | |
| Ribosome biogenesis | NOL8 NOP16 NOC4L B23 NPM1 nucleolin | Ribosomal stress | Anti | [61] | ||
| [21] | ||||||
| [56] | ||||||
| Protein turnover | Jab1 | Remove viral capsid protein | [19] | |||
| Lipid Metabolism | Lipid droplets | AGPAT6 LPCAT2 AUP1 MBOAT2 | Storage of accumulated capsid until particle assembly | Pro | Cause issues with particle formation; formation of VLPS | [61] |
| Sphingolipid Metabolism | KDSR TEX2 | Initial budding into ER | ||||
| Lipid binding | GRAMD1A GRAMD3 | Interaction with lipid membrane | ||||
| Ceramide Metabolism | SMPD4 CERS2 GBA2 | Initial budding into ER |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sotcheff, S.; Routh, A. Understanding Flavivirus Capsid Protein Functions: The Tip of the Iceberg. Pathogens 2020, 9, 42. https://doi.org/10.3390/pathogens9010042
Sotcheff S, Routh A. Understanding Flavivirus Capsid Protein Functions: The Tip of the Iceberg. Pathogens. 2020; 9(1):42. https://doi.org/10.3390/pathogens9010042
Chicago/Turabian StyleSotcheff, Stephanea, and Andrew Routh. 2020. "Understanding Flavivirus Capsid Protein Functions: The Tip of the Iceberg" Pathogens 9, no. 1: 42. https://doi.org/10.3390/pathogens9010042
APA StyleSotcheff, S., & Routh, A. (2020). Understanding Flavivirus Capsid Protein Functions: The Tip of the Iceberg. Pathogens, 9(1), 42. https://doi.org/10.3390/pathogens9010042





