Repeat-Induced Point Mutations Drive Divergence between Fusarium circinatum and Its Close Relatives
Abstract
1. Introduction
2. Results
2.1. Identification, Characterization and Potential Functionality of RIP-Associated Proteins
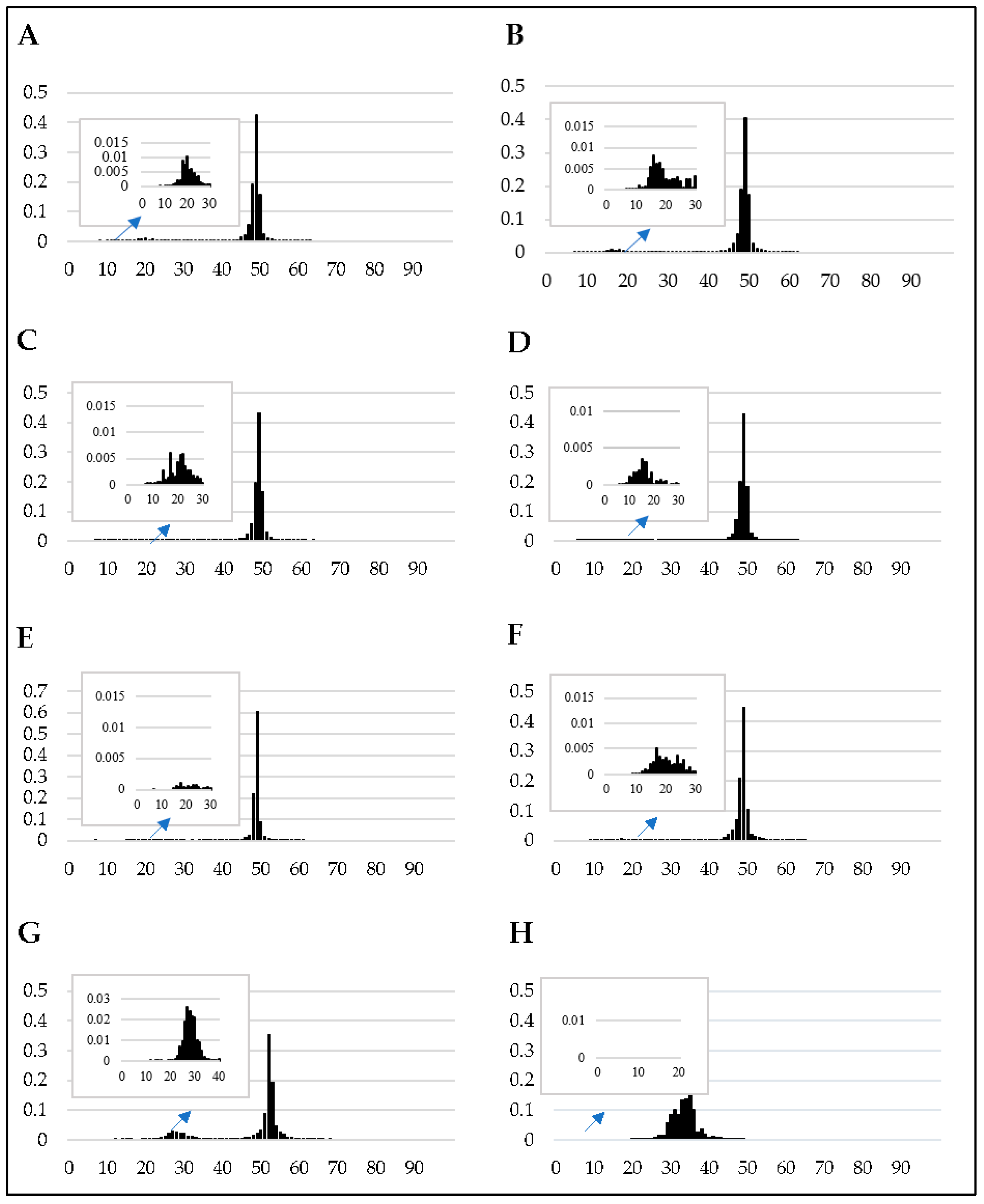
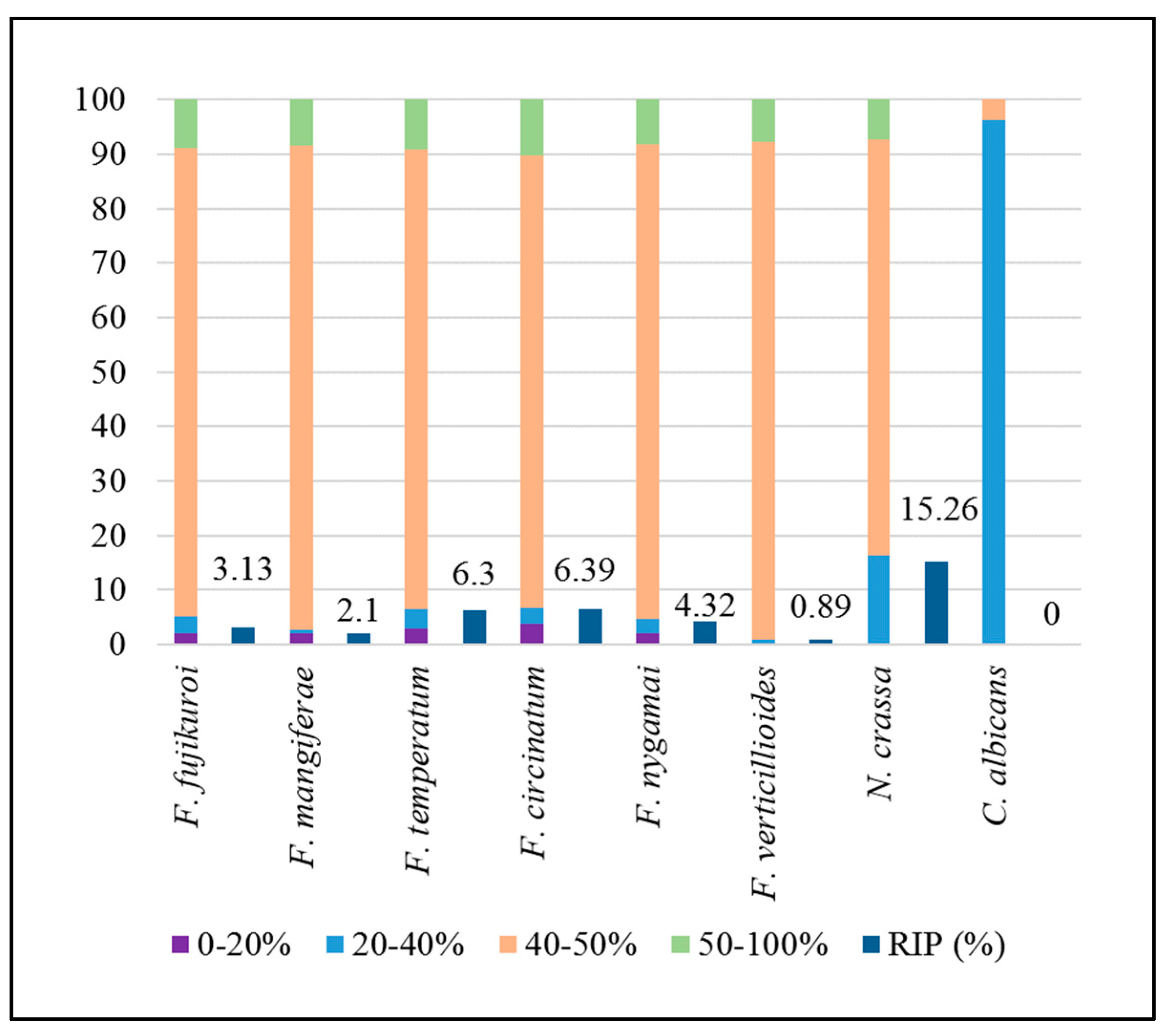
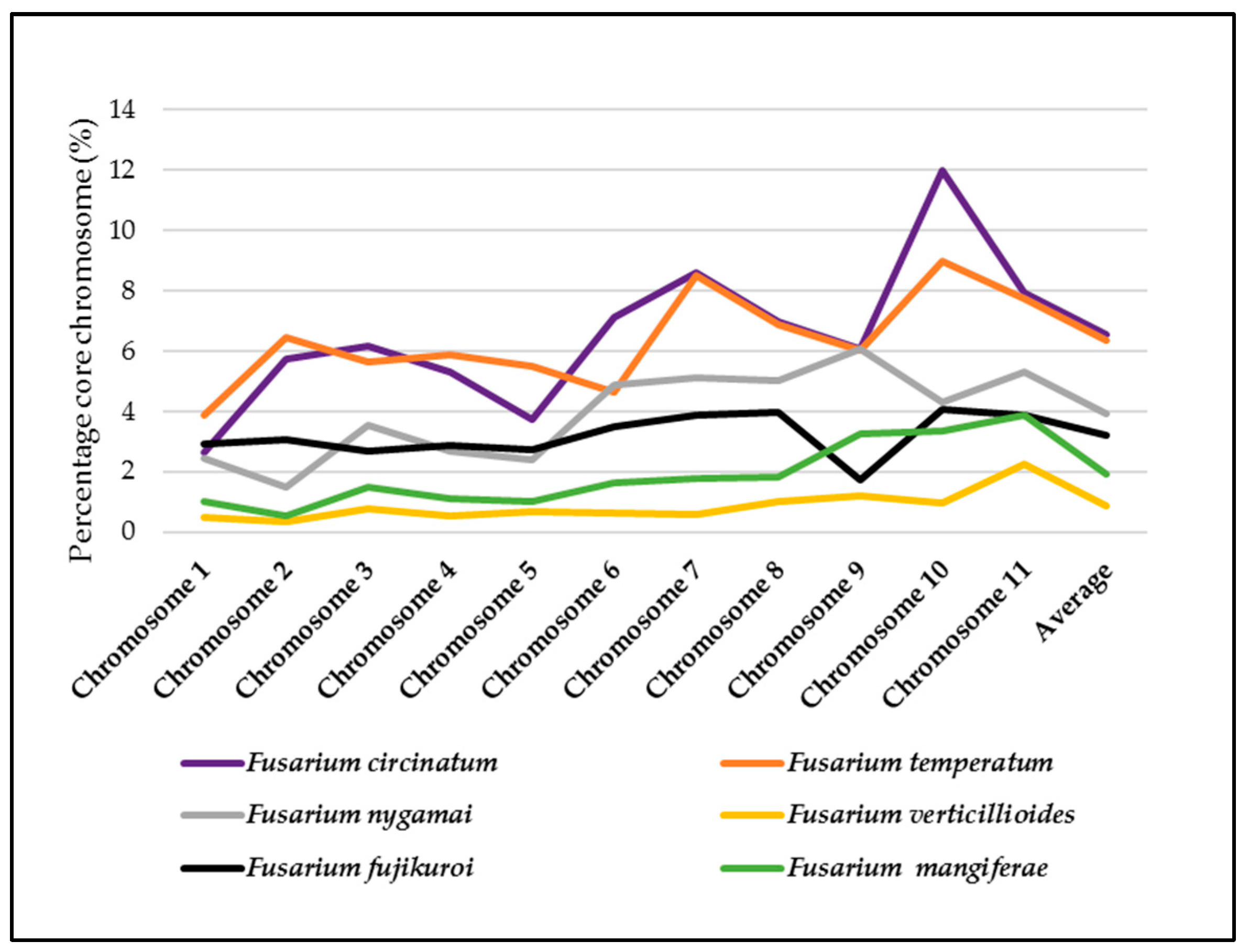
2.2. Genome-Wide in Silico RIP-Analysis
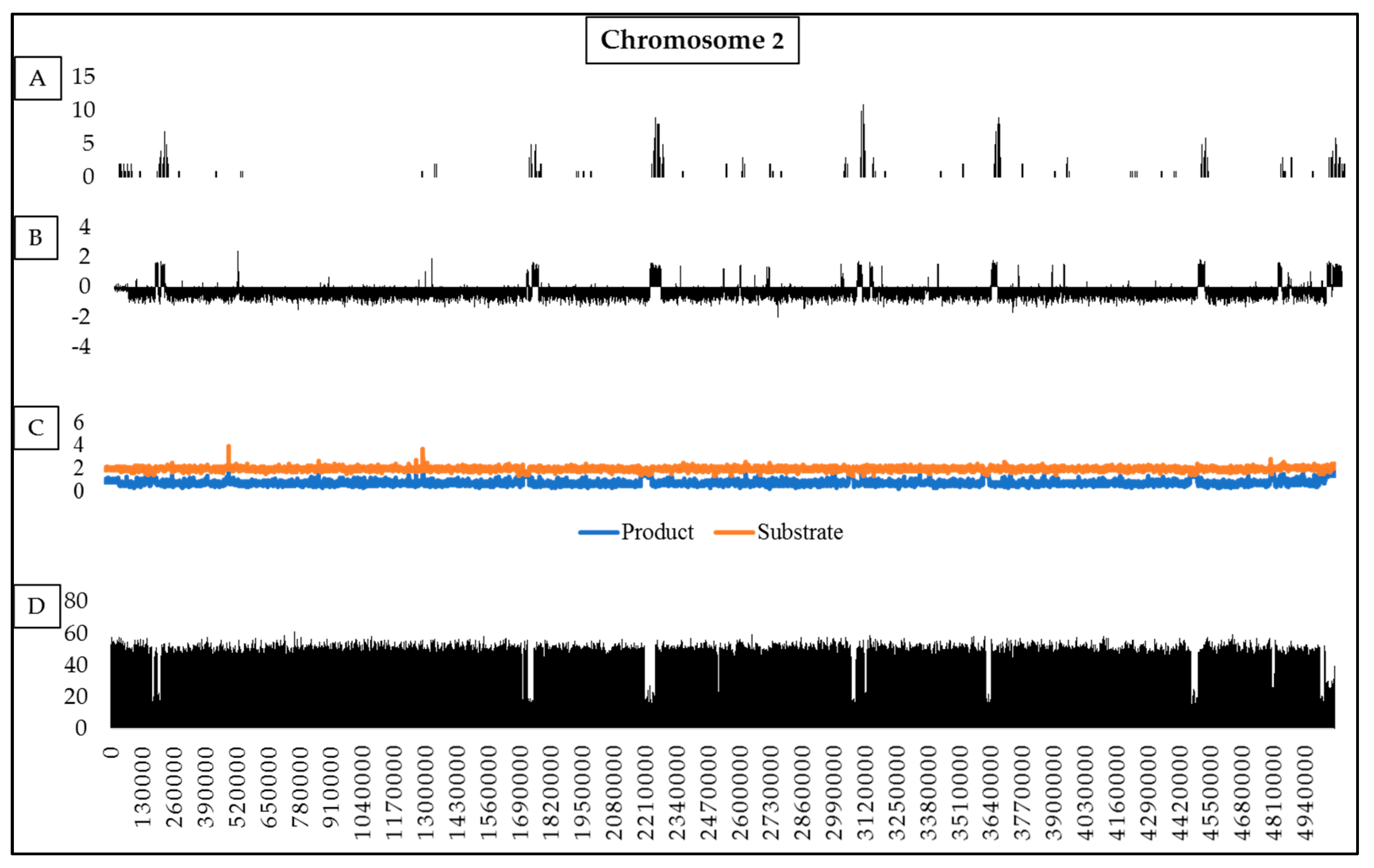
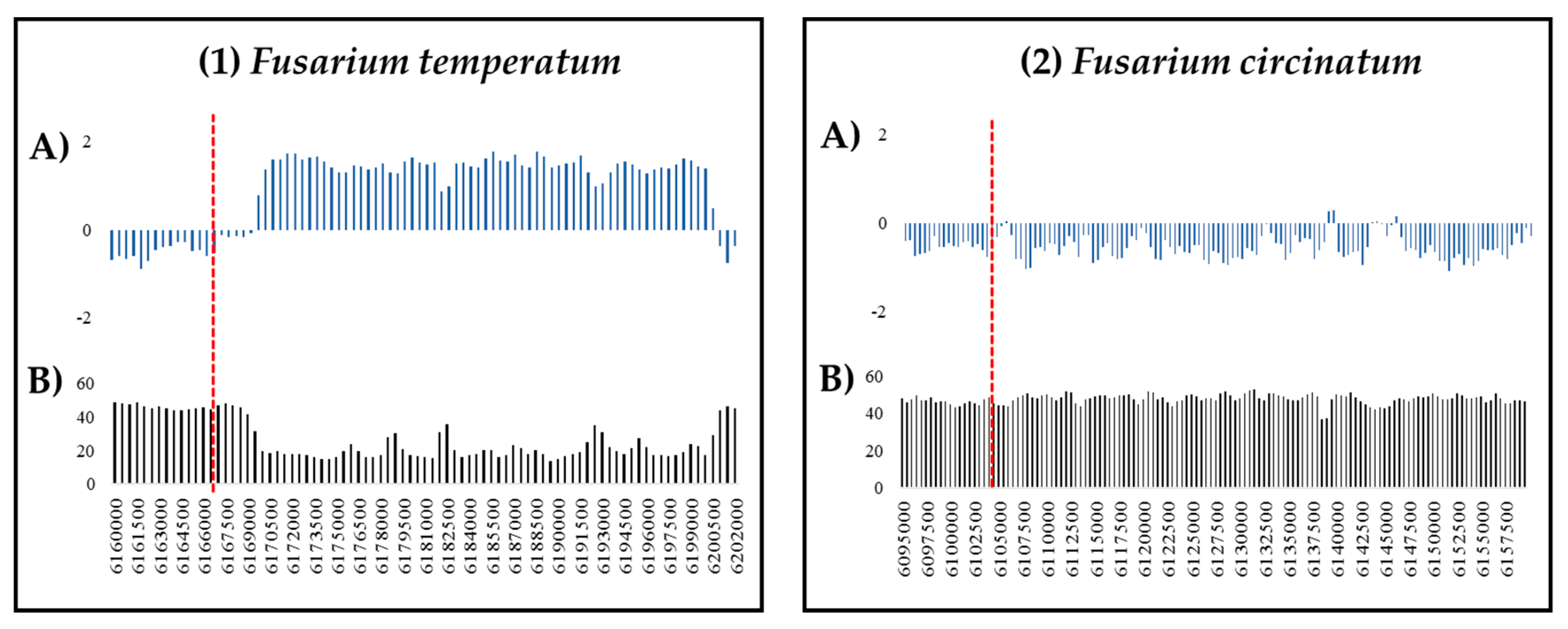
2.3. RIP Analysis of Regions Showing TRD
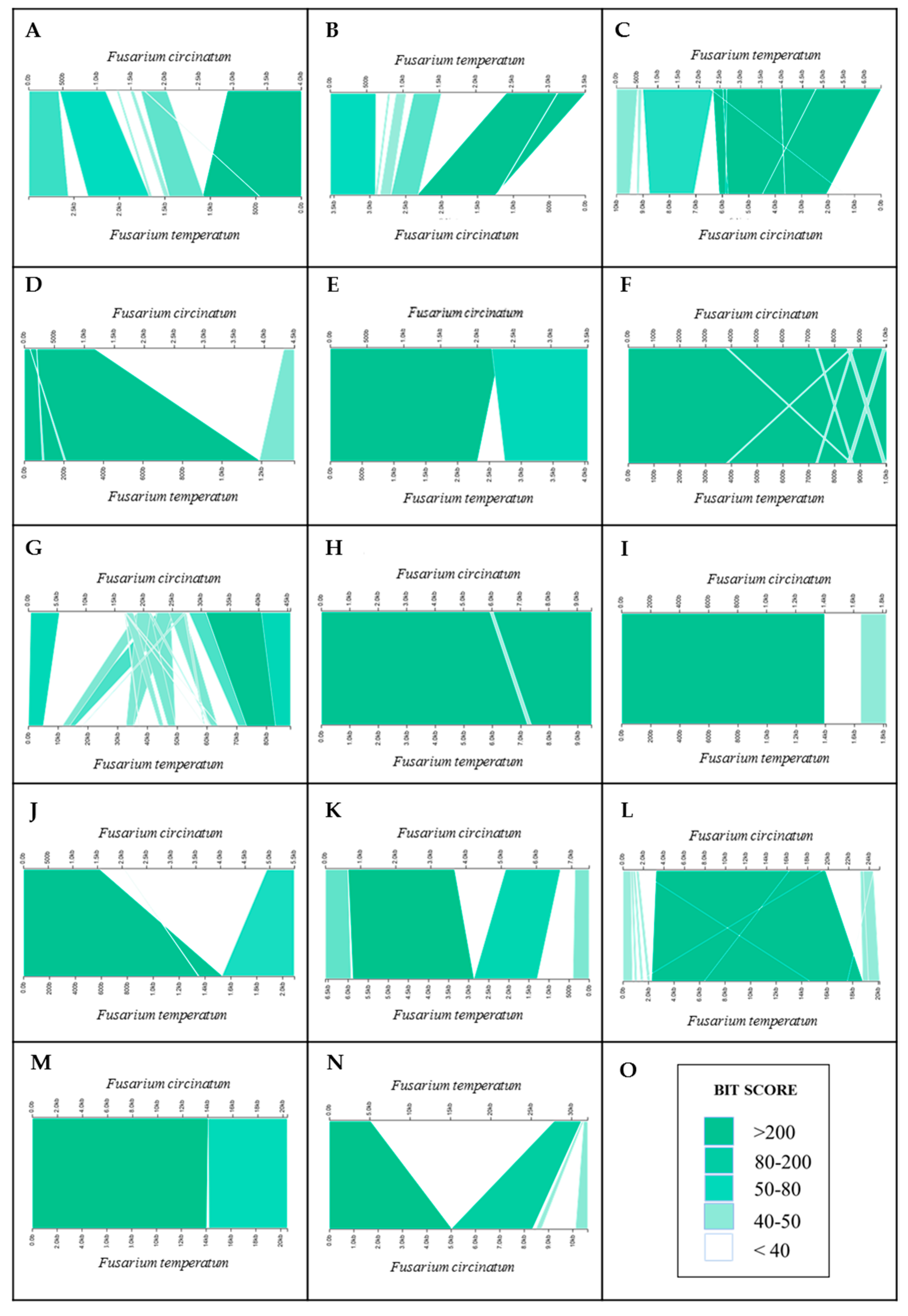
2.4. Pseudogenization and Loss of Homology
3. Discussion
3.1. The Genomes of F. circinatum and Its Relatives Harbor Evidence of RIP
3.2. RIP Can Change Genomic Environments and Gene Functionality
3.3. RIP May Contribute to Lineage-Divergence
4. Materials and Methods
4.1. Genome Sequences
4.2. Identification, Characterization and Potential Functionality of RIP-Associated proteins
4.3. Genome-Wide in Silico RIP-Analysis
4.4. RIP Analysis of Regions Showing TRD
4.5. Pseudogenization and Loss of Homology
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Leslie, J.F.; Summerell, B.A. The Fusarium Laboratory Manual, 1st ed.; John Wiley & Sons: Ames, IA, USA, 2008; pp. 1–369. [Google Scholar]
- Waalwijk, C.A.; Vanheule, K.; Audenaert, H.; Zhang, H.; Warris, S.; van de Geest, H.; van der Lee, T. Fusarium in the age of genomics. Trop. Plant Pathol. 2017, 42, 184–189. [Google Scholar] [CrossRef]
- Wingfield, M.; Hammerbacher, A.; Ganley, R.; Steenkamp, E.T.; Gordon, T.; Wingfield, B.; Coutinho, T. Pitch canker caused by Fusarium circinatum—A growing threat to pine plantations and forests worldwide. Australas. Plant Pathol. 2008, 37, 319–334. [Google Scholar] [CrossRef]
- Wingfield, B.D.; Steenkamp, E.T.; Santana, Q.C.; Coetzee, M.P.A.; Bam, S.; Barnes, I.; Beukes, C.W.; Chane, W.-Y.; De Vos, L.; Fourie, G.; et al. First fungal genome sequence from Africa: A preliminary analysis. S. Afr. J. Sci. 2012, 108, 104–122. [Google Scholar] [CrossRef]
- Wingfield, B.D.; Liu, M.; Nguyen, H.D.; Lane, F.A.; Morgan, S.W.; De Vos, L.; Wilken, P.M.; Duong, T.A.; Aylward, J.; Coetzee, M.P. Nine draft genome sequences of Claviceps purpurea s. lat.; including C. arundinis, C. humidiphila, and C. cf. spartinae, pseudomolecules for the pitch canker pathogen Fusarium circinatum, draft genome of Davidsoniella eucalypti, Grosmannia galeiformis, Quambalaria eucalypti, and Teratosphaeria destructans. IMA Fungus 2018, 9, 401–418. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.J.; Van Der Does, H.C.; Borkovich, K.A.; Coleman, J.J.; Daboussi, M.-J.; Di Pietro, A.; Dufresne, M.; Freitag, M.; Grabherr, M.; Henrissat, B.; et al. Comparative genomics reveals mobile pathogenicity chromosomes in Fusarium. Nature 2010, 464, 367–373. [Google Scholar] [CrossRef] [PubMed]
- Wiemann, P.; Sieber, C.M.; Von Bargen, K.W.; Studt, L.; Niehaus, E.M.; Espino, J.J.; Huß, K.; Michielse, C.B.; Albermann, S.; Wagner, D.; et al. Deciphering the cryptic genome: Genome-wide analyses of the rice pathogen Fusarium fujikuroi reveal complex regulation of secondary metabolism and novel metabolites. PLoS Pathog. 2013, 9, e1003475. [Google Scholar] [CrossRef]
- Niehaus, E.M.; Münsterkötter, M.; Proctor, R.H.; Brown, D.W.; Sharon, A.; Idan, Y.; Oren-Young, L.; Sieber, C.M.; Novák, O.; Pěnčík, A. Comparative “omics” in of the Fusarium fujikuroi species complex highlights differences in genetic potential and metabolite synthesis. Genome Biol. Evol. 2016, 8, 3574–3599. [Google Scholar] [CrossRef]
- Geiser, D.M.; Aoki, T.; Bacon, C.W.; Baker, S.E.; Bhattacharyya, M.K.; Brandt, M.E.; Brown, D.W.; Burgess, L.W.; Chulze, S.; Coleman, J.J.; et al. One fungus, one name: Defining the genus Fusarium in a scientifically robust way that preserves longstanding use. Phytopathology 2013, 103, 400–408. [Google Scholar] [CrossRef]
- O’Donnell, K.; Cigelnik, E.; Nirenberg, H.I. Molecular systematics and phylogeography of the Gibberella fujikuroi species complex. Mycologia 1998, 90, 465–493. [Google Scholar] [CrossRef]
- O’Donnell, K.; Rooney, A.P.; Proctor, R.H.; Brown, D.W.; McCormick, S.P.; Ward, T.J.; Frandsen, R.J.; Lysøe, E.; Rehner, S.A.; Aoki, T. Phylogenetic analyses of RPB1 and RPB2 support a middle Cretaceous origin for a clade comprising all agriculturally and medically important fusaria. Fungal Genet. Biol. 2013, 52, 20–31. [Google Scholar] [CrossRef]
- Kvas, M.; Marasas, J.W.O.; Wingfield, B.D.; Wingfield, M.J.; Steenkamp, E.T. Diversity and evolution of Fusarium species in the Gibberella fujikuroi complex. Fungal Divers. 2019, 34, 1–21. [Google Scholar]
- De Vos, L.; Steenkamp, E.T.; Martin, S.H.; Santana, Q.C.; Fourie, G.; van der Merwe, N.A.; Wingfield, M.J.; Wingfield, B.D. Genome-wide macrosynteny among Fusarium species in the Gibberella fujikuroi complex revealed by amplified fragment length polymorphisms. PLoS ONE 2014, 9, e114682. [Google Scholar] [CrossRef] [PubMed]
- Daboussi, M. Fungal transposable elements and genome evolution. Genetica 1997, 100, 253–260. [Google Scholar] [CrossRef] [PubMed]
- Daboussi, M.J.; Capy, P. Transposable elements in filamentous fungi. Annu. Rev. Microbiol. 2003, 57, 275–299. [Google Scholar] [CrossRef]
- Spanu, P.D.; Abbott, J.C.; Amselem, J.; Burgis, T.A.; Soanes, D.M.; Stüber, K.; van Themaat, E.V.L.; Brown, J.K.; Butcher, S.A.; Gurr, S.J. Genome expansion and gene loss in powdery mildew fungi reveal tradeoffs in extreme parasitism. Science 2010, 330, 1543–1546. [Google Scholar] [CrossRef]
- Gladyshev, E. Repeat-Induced Point mutation (RIP) and other genome defense mechanisms in fungi. Microbiol. Spectr. 2017, 5, 1–21. [Google Scholar] [CrossRef]
- Selker, E.U.; Stevens, J.N. DNA methylation at asymmetric sites is associated with numerous transition mutations. Proc. Natl. Acad. Sci. USA 1985, 82, 8114–8118. [Google Scholar] [CrossRef]
- Selker, E.U.; Cambareri, E.B.; Jensen, B.C.; Haack, K.R. Rearrangement of duplicated DNA in specialized cells of Neurospora. Cell 1987, 51, 741–752. [Google Scholar] [CrossRef]
- Clutterbuck, A.J. Genomic evidence of Repeat-Induced Point mutation (RIP) in filamentous ascomycetes. Fungal Genet. Biol. 2011, 48, 306–326. [Google Scholar] [CrossRef]
- Horns, F.; Petit, E.; Yockteng, R.; Hood, M.E. Patterns of Repeat-Induced Point mutation in transposable elements of basidiomycete fungi. GBE 2012, 4, 240–247. [Google Scholar] [CrossRef]
- Rouxel, T.; Grandaubert, J.; Hane, J.K.; Hoede, C.; van de Wouw, A.P.; Couloux, A.; Dominguez, V.; Anthouard, V.; Bally, P.; Bourras, S.; et al. Effector diversification within compartments of the Leptosphaeria maculans genome affected by Repeat-Induced Point mutations. Nat. Commun. 2011, 2, 202. [Google Scholar] [CrossRef] [PubMed]
- Amselem, J.; Lebrun, M.H.; Quesneville, H. Whole genome comparative analysis of transposable elements provides new insight into mechanisms of their inactivation in fungal genomes. BMC Genomics 2015, 16, 141–155. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Meerupati, T.; Andersson, K.-M.; Friman, E.; Kumar, D.; Tunlid, A.; Ahrén, D. Genomic mechanisms accounting for the adaptation to parasitism in nematode-trapping fungi. PLoS Genetics 2013, 9, e1003909. [Google Scholar] [CrossRef] [PubMed]
- Testa, A.C.; Oliver, R.P.; Hane, J.K. OcculterCut: A comprehensive survey of AT-rich regions in fungal genomes. GBE 2016, 8, 2044–2064. [Google Scholar] [CrossRef]
- Derbyshire, M.; Denton-Giles, M.; Hegedus, D.; Seifbarghy, S.; Rollins, J.; Van Kan, J.; Seidl, M.F.; Faino, L.; Mbengue, M.; Navaud, O. The complete genome sequence of the phytopathogenic fungus Sclerotinia sclerotiorum reveals insights into the genome architecture of broad host range pathogens. GBE 2017, 9, 593–618. [Google Scholar] [CrossRef]
- Van de Wouw, A.P.; Cozijnsen, A.J.; Hane, J.K.; Brunner, P.C.; McDonald, B.A.; Oliver, R.P.; Howlett, B.J. Evolution of linked avirulence effectors in Leptosphaeria maculans is affected by genomic environment and exposure to resistance genes in host plants. PLoS Pathog. 2010, 6, e1001180. [Google Scholar] [CrossRef]
- Daverdin, G.; Rouxel, T.; Gout, L.; Aubertot, J.-N.; Fudal, I.; Meyer, M.; Parlange, F.; Carpezat, J.; Balesdent, M.H. Genome structure and reproductive behaviour influence the evolutionary potential of a fungal phytopathogen. PLoS Pathog. 2012, 8, e1003020. [Google Scholar] [CrossRef]
- Dong, S.; Raffaele, S.; Kamoun, S. The two-speed genomes of filamentous pathogens: Waltz with plants. Curr. Opin. Genet. Dev. 2015, 35, 57–65. [Google Scholar] [CrossRef]
- Honda, S.; Lewis, Z.A.; Shimada, K.; Fischle, W.; Sack, R.; Selker, E.U. Heterochromatin protein 1 forms distinct complexes to direct histone deacetylation and DNA methylation. Nat. Struct. Mol. Biol. 2012, 19, 471–477. [Google Scholar] [CrossRef]
- Li, W.-C.; Huang, C.-H.; Chen, C.L.; Chuang, Y.-C.; Tung, S.-Y.; Wang, T.-F. Trichoderma reesei complete genome sequence, Repeat-Induced Point mutation, and partitioning of CAZyme gene clusters. Biotechnol. Biofuels 2017, 10, 170. [Google Scholar] [CrossRef]
- King, R.; Urban, M.; Hammond-Kosack, M.C.; Hassani-Pak, K.; Hammond-Kosack, K.E. The completed genome sequence of the pathogenic ascomycete fungus Fusarium graminearum. BMC Genomics 2015, 16, 544. [Google Scholar] [CrossRef] [PubMed]
- Coleman, J.J.; Rounsley, S.D.; Rodriguez-Carres, M.; Kuo, A.; Wasmann, C.C.; Grimwood, J.; Schmutz, J.; Taga, M.; White, G.J.; Zhou, S. The genome of Nectria haematococca: Contribution of supernumerary chromosomes to gene expansion. PLoS Genet. 2009, 5, e1000618. [Google Scholar] [CrossRef] [PubMed]
- Leslie, J.F.; Dickman, M. Fate of DNA encoding hygromycin resistance after meiosis in transformed strains of Gibberella fujikuroi (Fusarium moniliforme). Appl. Environ. Microb. 1991, 57, 1423–1429. [Google Scholar]
- De Vos, L.; van der Merwe, N.A.; Wingfield, M.J.; Myburg, A.A.; Wingfield, B.D. Transmission ratio distortion in an interspecific cross between Fusarium circinatum and Fusarium subglutinans. Genes Genom. 2013, 35, 177–183. [Google Scholar] [CrossRef]
- Kouzminova, E.; Selker, E.U. dim-2 encodes a DNA methyltransferase responsible for all known cytosine methylation in Neurospora. EMBO J. 2001, 20, 4309–4323. [Google Scholar] [CrossRef]
- Aramayo, R.; Selker, E.U. Neurospora crassa, a model system for epigenetics research. CHS Perspect. Biol. 2013, 5, 1–19. [Google Scholar] [CrossRef]
- Lewis, Z.A.; Adhvaryu, K.K.; Honda, S.; Shiver, A.L.; Selker, E.U. Identification of DIM-7, a protein required to target the DIM-5 H3 methyltransferase to chromatin. Proc. Natl. Acad. Sci. USA 2010, 107, 8310–8315. [Google Scholar] [CrossRef]
- Du, J.; Johnson, L.M.; Jacobsen, S.E.; Patel, D.J. DNA methylation pathways and their crosstalk with histone methylation. Nat. Rev. Mol. Cell Biol. 2015, 16, 519–532. [Google Scholar] [CrossRef]
- Margolin, B.S.; Garrett-Engele, P.W.; Stevens, J.N.; Fritz, D.Y.; Garrett-Engele, C.; Metzenberg, R.L.; Selker, E.U. A methylated Neurospora 5S rRNA pseudogene contains a transposable element inactivated by Repeat-Induced Point mutation. Genetics 1998, 149, 1787–1797. [Google Scholar]
- Selker, E.U.; Tountas, N.A.; Cross, S.H.; Margolin, B.S.; Murphy, J.G.; Bird, P.; Freitag, M. The methylated component of the Neurospora crassa genome. Nature 2003, 422, 893–897. [Google Scholar] [CrossRef]
- Lewis, Z.A.; Honda, S.; Khlafallah, T.K.; Jeffress, J.K.; Freitag, M.; Mohn, F.; Schübeler, D.; Selker, E.U. Relics of Repeat-Induced Point mutation direct heterochromatin formation in Neurospora crassa. Genome Res. 2009, 19, 427–437. [Google Scholar] [CrossRef] [PubMed]
- Van Wyk, S.; Harrison, C.H.; Wingfield, B.W.; De Vos, L.; van der Merwe, N.A.; Steenkamp, E.T. The RIPper, a web-based tool for genome-wide quantification of Repeat-Induced Point (RIP) mutations. PeerJ 2019, 7, e7447. [Google Scholar] [CrossRef] [PubMed]
- Tamaru, H.; Selker, E.U. A histone H3 methyltransferase controls DNA methylation in Neurospora crassa. Nature 2001, 414, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Freitag, M.; Williams, R.L.; Kothe, G.O.; Selker, E.U. A cytosine methyltransferase homologue is essential for repeat-induced point mutation in Neurospora crassa. Proc. Natl. Acad. Sci. USA 2002, 99, 8802–8807. [Google Scholar] [CrossRef]
- Freitag, M.; Hickey, P.C.; Khlafallah, T.K.; Read, N.D.; Selker, E.U. HP1 is essential for DNA methylation in Neurospora. Mol. Cell 2004, 13, 427–434. [Google Scholar] [CrossRef]
- Ellison, C.E.; Stajich, J.E.; Jacobson, D.J.; Natvig, D.O.; Lapidus, A.; Foster, B.; Aerts, A.; Riley, R.; Lindquist, E.A.; Grigoriev, I.V. Massive changes in genome architecture accompany the transition to self-fertility in the filamentous fungus Neurospora tetrasperma. Genetics 2011, 189, 55–69. [Google Scholar] [CrossRef]
- Galagan, J.E.; Calvo, S.E.; Borkovich, K.A.; Selker, E.U.; Read, N.D.; Jaffe, D.; Fitz Hugh, W.; Ma, L.J.; Smirnov, S.; Purcell, S. The genome sequence of the filamentous fungus Neurospora crassa. Nature 2003, 422, 859–868. [Google Scholar] [CrossRef]
- Gao, Q.; Jin, K.; Ying, S.-H.; Zhang, Y.; Xiao, G.; Shang, Y.; Duan, Z.; Hu, X.; Xie, X.-Q.; Zhou, G. Genome sequencing and comparative transcriptomics of the model entomopathogenic fungi Metarhizium anisopliae and M. acridum. PLoS Genet. 2011, 7, e1001264. [Google Scholar] [CrossRef]
- Braumann, I.; Van Den Berg, M.; Kempken, F. Repeat-Induced Point mutation in two asexual fungi, Aspergillus niger and Penicillium chrysogenum. Curr. Genet. 2008, 53, 287–297. [Google Scholar] [CrossRef]
- Hane, J.K.; Williams, A.H.; Taranto, A.P.; Solomon, P.S.; Oliver, R.P. Repeat-Induced Point Mutation: A Fungal-Specific, Endogenous Mutagenesis Process. Genetic Transformation Systems in Fungi, 2nd ed.; Springer: Basel, Switzerland, 2015; pp. 55–68. [Google Scholar] [CrossRef]
- Croll, D.; Zala, M.; McDonald, B.A. Breakage-fusion-bridge cycles and large insertions contribute to the rapid evolution of accessory chromosomes in a fungal pathogen. PLoS Genet. 2013, 9, e1003567. [Google Scholar] [CrossRef]
- Balesdent, M.H.; Fudal, I.; Ollivier, B.; Bally, P.; Grandaubert, J.; Eber, F.; Chèvre, A.M.; Leflon, M.; Rouxel, T. The dispensable chromosome of Leptosphaeria maculans shelters an effector gene conferring avirulence towards Brassica rapa. New Phytol. 2013, 198, 887–898. [Google Scholar] [CrossRef] [PubMed]
- Vanheule, A.; Audenaert, K.; Warris, S.; van de Geest, H.; Schijlen, E.; Höfte, M.; De Saeger, S.; Haesaert, G.; Waalwijk, C.; van der Lee, T. Living apart together: Crosstalk between the core and supernumerary genomes in a fungal plant pathogen. BMC Genom. 2016, 17, 670. [Google Scholar] [CrossRef] [PubMed]
- Galagan, J.E.; Selker, E.U. RIP: The evolutionary cost of genome defense. Trends Genet. 2004, 20, 417–423. [Google Scholar] [CrossRef] [PubMed]
- Fudal, I.; Ross, S.; Brun, H.; Besnard, A.-L.; Ermel, M.; Kuhn, M.-L.; Balesdent, M.-H.; Rouxel, T. Repeat-Induced Point mutation (RIP) as an alternative mechanism of evolution toward virulence in Leptosphaeria maculans. MPMI 2009, 22, 932–941. [Google Scholar] [CrossRef] [PubMed]
- Bender, J. Cytosine methylation of repeated sequences in eukaryotes: The role of DNA pairing. Trends Biochem. Sci. 1998, 23, 252–256. [Google Scholar] [CrossRef]
- Stukenbrock, E.H.; Croll, D. The evolving fungal genome. Fungal Biol. Rev. 2014, 28, 1–12. [Google Scholar] [CrossRef]
- Gallegos, A.; Jacobson, D.J.; Raju, N.B.; Skupski, M.P.; Natvig, D.O. Suppressed recombination and a pairing anomaly on the mating-type chromosome of Neurospora tetrasperma. Genetics 2000, 154, 623–633. [Google Scholar]
- Jacobson, D.J. Blocked recombination along the mating-type chromosomes of Neurospora tetrasperma involves both structural heterozygosity and autosomal genes. Genetics 2005, 171, 839–843. [Google Scholar] [CrossRef]
- Menkis, A.; Bastiaans, E.; Jacobson, D.; Johannesson, H. Phylogenetic and biological species diversity within the Neurospora tetrasperma complex. J. Evol. Biol. 2009, 22, 1923–1936. [Google Scholar] [CrossRef]
- Steenkamp, E.T.; Wingfield, M.J.; McTaggart, A.R.; Wingfield, B.D. Fungal species and their boundaries matter – Definitions, mechanisms and practical implications. Fungal Biol. Rev. 2018, 32, 104–116. [Google Scholar] [CrossRef]
- Coghlan, A.; Eichler, E.E.; Oliver, S.G.; Paterson, A.H.; Stein, L. Chromosome evolution in eukaryotes: A multi-kingdom perspective. Trends Genet. 2005, 21, 673–682. [Google Scholar] [CrossRef] [PubMed]
- Emelianov, I.; Marec, F.; Mallet, J. Genomic evidence for divergence with gene flow in host races of the larch budmoth. Proc. R. Soc. Lond. B 2004, 271, 97–105. [Google Scholar] [CrossRef] [PubMed]
- Wingfield, B.D.; Berger, D.K.; Steenkamp, E.T.; Lim, H.J.; Duong, T.A.; Bluhm, B.H.; De Beer, Z.W.; De Vos, L.; Fourie, G.; Naidoo, K.; et al. Draft genome of Cercospora zeina, Fusarium pininemorale, Hawksworthiomyces lignivorus, Huntiella decipiens and Ophiostoma ips. IMA Fungus 2017, 8, 385–396. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, R.M.; Seppey, M.; Simão, F.A.; Manni, M.; Ioannidis, P.; Klioutchnikov, G.; Kriventseva, E.V.; Zdobnov, E.M. BUSCO applications from quality assessments to gene prediction and phylogenomics. Mol. Biol. Evol. 2017, 35, 543–548. [Google Scholar] [CrossRef] [PubMed]
- Hoff, K.J.; Stanke, M. WebAUGUSTUS—A web service for training AUGUSTUS and predicting genes in eukaryotes. Nucleic Acids Res. 2013, 41, W123–W128. [Google Scholar] [CrossRef] [PubMed]
- Flutre, T.; Emmanuelle, P.; Quesneville, H. Transposable Element Annotation in Completely Sequenced Eukaryote Genomes, 3rd ed.; Springer: Berlin/Heidelberg, Germany, 2012; pp. 17–39. [Google Scholar]
- Jones, P.; Binns, D.; Chang, H.-Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 2017, 108, 1–7. [Google Scholar] [CrossRef]
- Wickramasinghe, S.; Rincon, G.; Islas-Trejo, A.; Medrano, J.F. Transcriptional profiling of bovine milk using RNA sequencing. BMC Genom. 2012, 13, 45–59. [Google Scholar] [CrossRef]
- Harris, R.S. Improved Pairwise Alignment of Genomic DNA. Ph.D. Thesis, Pennsylvania State University, Pennsylvania, PA, USA, 2007. [Google Scholar]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Conesa, A.; Götz, S.; García-Gómez, J.M.; Terol, J.; Talón, M.; Robles, M. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 2005, 21, 3674–3676. [Google Scholar] [CrossRef]
- Supek, F.; Bošnjak, M.; Škunca, N.; Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 2011, 6, e21800. [Google Scholar] [CrossRef] [PubMed]
- De Vos, L.; Myburg, A.A.; Wingfield, M.J.; Desjardins, A.E.; Gordon, T.R.; Wingfield, B.D. Complete genetic linkage maps from an interspecific cross between Fusarium circinatum and Fusarium subglutinans. Fungal Genet. Biol. 2007, 44, 701–714. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Randomizer. Available online: www.randomizer.org (accessed on 13 June 2019).
- Wintersinger, J.A.; Wasmuth, J.D. Kablammo: An interactive, web-based BLAST results visualizer. Bioinformatics 2015, 31, 1305–1306. [Google Scholar] [CrossRef] [PubMed]
| Species | Strain | Accession Number 1 | Percentage Completeness 2 |
|---|---|---|---|
| Fusarium circinatum | FSP34 | AYJV00000000.2 | 98.1 |
| Fusarium temperatum | Netza 9 | LJGR00000000.1 | 98.8 |
| Fusarium nygamai | MRC 8546 | LBNR00000000.1 | 98.7 |
| Fusarium fujikuroi | IMI 58289 | GCA_900079805.1 | 98.6 |
| Fusarium mangiferae | MRC 7560 | FCQH00000000.1 | 99.0 |
| Fusarium verticillioides | 7600 | AAIM00000000.2 | 98.3 |
| Neurospora crassa | OR74A | AABX00000000.3 | 98.1 |
| Candida albicans | SC5314 | GCA_000182965.3 | 97.9 |
| Protein Name 1 | Accession Number 2 | Gene Identifier (% Similarity to the Neurospora crassa homolog) 3 | ||||||
|---|---|---|---|---|---|---|---|---|
| Fc | Ft | Ff | Fm | Fv | Fn | Within FFSC % similarity | ||
| HP-1 | XP_957632.1 | g 30.102 (59.89%) | g 30.82 (59.32%) | FF_062931 (61.80%) | FMAN_09299 (61.80%) | FVEG_013876 (59.89%) | g 3888.1 (58.54%) | 90.58%–98.39% |
| RID | XP_011392925.1 | g 24.18 (36.08%) | g 25.67 (35.29%) | FF_06164 (36.27%) | FMAN_09165 (42.36%) | FVEG_02018 (35.4%) | g 3766 (35.22%) | 86.10%–97.68% |
| DIM-2 | XP_959891.1 | g 22.31 (48.8%) | g 23.9 (48.83%) | FF_08441 (47.1%) | FMAN_10524 (48.68%) | FVEG_11429 (49.44%) | g 4731.1 (50.07%) | 90.39%–97.78% |
| DIM-3 | XP_960652.1 | g 17.69 (89.07%) | g 16.59 (88.58%) | FF_02702.1 (91.49%) | FMAN_04997 (91.48%) | FVEG_08024 (91.49%) | g 4826t1 (88.58%) | 96.27%–99.56% |
| DIM-5 | XP_957479.2 | g 28.0 (69.36%) | g 29.67 (61.27%) | FF_07620.1 (62.43%) | FMAN_07768 (61.56%) | FVEG_08911 (61.88%) | g 2471.1 (62.43%) | 95.93%–97.97% |
| DIM-7 | XP_961308.2 | g 14.27 (28.87%) | g 14.74 (36.62%) | FF_02137.1 (34.36%) | FMAN_04175 (30.43%) | FVEG_07938 (35.26%) | g 4746.1 (27.05%) | 87.23%–96.12% |
| DIM-8 | XP_962347.1 | g 13.103 (52.63%) | g 14.50 (53.12%) | FF_04892.1 (53.02%) | FMAN_02500 (42.55%) | FVEG_06222 (53.64%) | g 1063.1 (53.39%) | 95.95%–97.72% |
| DIM-9 | XP_956278.3 | g 15.26 (74.74%) | g 15.10 (75.79%) | FF_0162701 (75.79%) | FMAN_00488 (77.89%) | FVEG_09499 (76.84%) | g 31951 (76.84%) | 87.77%–93.87% |
| RIP Statistics | Fusarium Species 1 | |||||
|---|---|---|---|---|---|---|
| Fc | Ft | Ff | Fm | Fv | Fn | |
| Number of RIP-affected windows | 5766 | 5703 | 2741 | 1946 | 741 | 4468 |
| Average RIP composite index | 1.33 | 1.38 | 1.54 | 1.14 | 1.17 | 1.20 |
| Average size of RIP-affected genomic regions (bp) | 15,537 | 17,065 | 6542 | 9764 | 6219 | 12,609 |
| Number of genes in RIP-affected regions | 36 | 34 | 11 | 2 | 3 | 5 |
| Number of LRARs 2 | 162 | 153 | 95 | 65 | 32 | 149 |
| Gene density in LRARs 3 | 1.5 (34.41) | 1.38 (31.16) | 0.62 (34.42) | 0.36 (31.59) | 1.06 (38.93) | 0.36 (30.57) |
| Largest LRAR (bp) | 62,500 | 66,000 | 79,000 | 27,000 | 10,000 | 55,500 |
| RIP-affected genomic proportion (bp) | 6.39% (2,881,899) | 6.30% (2,851,440) | 3.13% (1,371,951) | 2.10% (972,147) | 0.89% (372,420) | 4.32% (2,229,769) |
| Genomic proportion containing LRARs (bp) | 5.6% (2,517,015) | 5.8% (2,611,000) | 1.4% (621,500) | 1.4% (634,659) | 0.5% (199,000) | 3.6% (1,878,813) |
| Chromosome Statistics | Fusarium Species 1 | ||||
|---|---|---|---|---|---|
| Fc | Ff | Ft | Fm | Fn | |
| Chromosome size (bp) | 525,065 | 692,922 | 986,231 | 887,429 | 791,442 |
| GC content (%) | 46.36 | 41.05 | 42.30 | 43.02 | 41.7 |
| Number of genes per chromosome | 171 | 136 | 189 | 163 | 133 |
| RIP-affected proportion (bp) 2 | 47,112 | 85,922 | 242,613 | 74,367 | 108,149 |
| Percentage of chromosome that is RIP-affected (%) 2 | 4.85 | 8.23 | 18.25 | 8.38 | 20.66 |
| Number of LRARs per chromosome 3 | 1 | 2 | 6 | 5 | 13 |
| Number of genes in RIP-affected regions | 0 | 0 | 1 | 0 | 0 |
| RIP Statistics Associated with AFLP Markers | Species | ||
|---|---|---|---|
| F. circinatum | F. temperatum | ||
| TRD markers | Total number of markers | 30 | 31 |
| Number of RIP-targeted markers | 19 | 18 | |
| Number of markers not RIP-targeted | 11 | 13 | |
| Proportion of the RIP-targeted total genome (%) 2 | 2.17 | 2.30 | |
| Mendelian markers | Total number of markers | 29 | 26 |
| Number of RIP-targeted markers | 9 | 6 | |
| Number of markers not RIP-targeted | 20 | 20 | |
| Proportion of the RIP-targeted total genome (%) 2 | 0.53 | 1.74 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
van Wyk, S.; Wingfield, B.D.; De Vos, L.; van der Merwe, N.A.; Santana, Q.C.; Steenkamp, E.T. Repeat-Induced Point Mutations Drive Divergence between Fusarium circinatum and Its Close Relatives. Pathogens 2019, 8, 298. https://doi.org/10.3390/pathogens8040298
van Wyk S, Wingfield BD, De Vos L, van der Merwe NA, Santana QC, Steenkamp ET. Repeat-Induced Point Mutations Drive Divergence between Fusarium circinatum and Its Close Relatives. Pathogens. 2019; 8(4):298. https://doi.org/10.3390/pathogens8040298
Chicago/Turabian Stylevan Wyk, Stephanie, Brenda D. Wingfield, Lieschen De Vos, Nicolaas A. van der Merwe, Quentin C. Santana, and Emma T. Steenkamp. 2019. "Repeat-Induced Point Mutations Drive Divergence between Fusarium circinatum and Its Close Relatives" Pathogens 8, no. 4: 298. https://doi.org/10.3390/pathogens8040298
APA Stylevan Wyk, S., Wingfield, B. D., De Vos, L., van der Merwe, N. A., Santana, Q. C., & Steenkamp, E. T. (2019). Repeat-Induced Point Mutations Drive Divergence between Fusarium circinatum and Its Close Relatives. Pathogens, 8(4), 298. https://doi.org/10.3390/pathogens8040298






