Formation and Maintenance of Tissue Resident Memory CD8+ T Cells after Viral Infection
Abstract
1. Introduction
2. Markers Used to Define TRM
3. Functions of Memory T Cell Markers
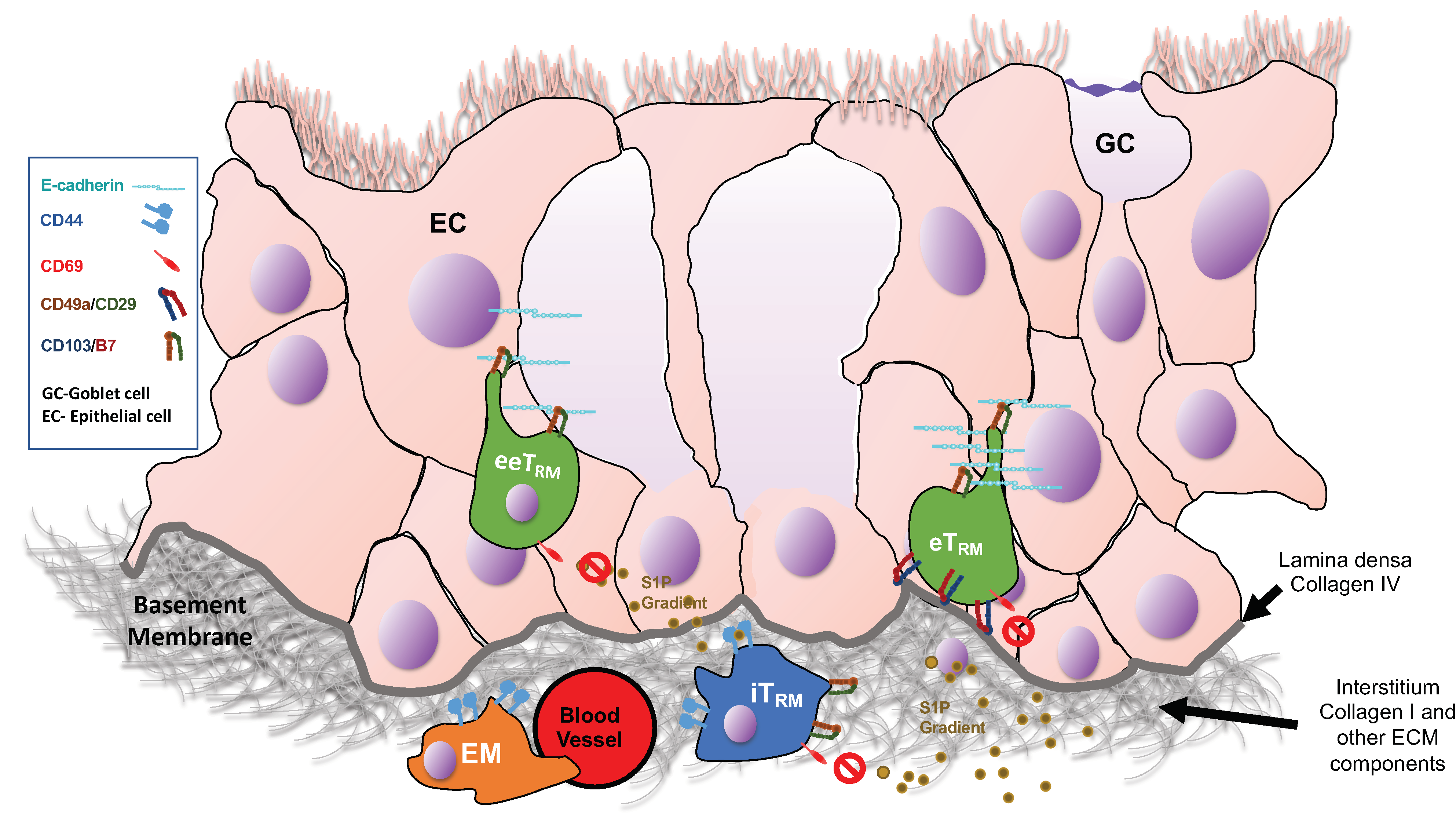
4. Formation of CD8 TRM and Interactions with Other Cells in the Tissue
5. Metabolic Changes Associated with TRM
6. Mechanisms that Suppress TRM Activation May Have other Functions Important to TRM Establishment and Maintenance
7. Concluding Remarks
Author Contributions
Funding
Conflicts of Interest
References
- Liang, S.; Mozdzanowska, K.; Palladino, G.; Gerhard, W. Heterosubtypic immunity to influenza type A virus in mice. Effector mechanisms and their longevity. J. Immunol. 1994, 152, 1653–1661. [Google Scholar] [PubMed]
- Hogan, R.J.; Usherwood, E.J.; Zhong, W.; Roberts, A.A.; Dutton, R.W.; Harmsen, A.G.; Woodland, D.L. Activated antigen-specific CD8+ T cells persist in the lungs following recovery from respiratory virus infections. J. Immunol. 2001, 166, 1813–1822. [Google Scholar] [CrossRef] [PubMed]
- Masopust, D.; Vezys, V.; Marzo, A.L.; Lefrancois, L. Preferential localization of effector memory cells in nonlymphoid tissue. Science 2001, 291, 2413–2417. [Google Scholar] [CrossRef] [PubMed]
- Ray, S.J.; Franki, S.N.; Pierce, R.H.; Dimitrova, S.; Koteliansky, V.; Sprague, A.G.; Doherty, P.C.; de Fougerolles, A.R.; Topham, D.J. The collagen binding alpha1beta1 integrin VLA-1 regulates CD8 T cell-mediated immune protection against heterologous influenza infection. Immunity 2004, 20, 167–179. [Google Scholar] [CrossRef]
- Teijaro, J.R.; Turner, D.; Pham, Q.; Wherry, E.J.; Lefrancois, L.; Farber, D.L. Cutting edge: Tissue-retentive lung memory CD4 T cells mediate optimal protection to respiratory virus infection. J. Immunol. 2011, 187, 5510–5514. [Google Scholar] [CrossRef]
- Sallusto, F.; Lenig, D.; Forster, R.; Lipp, M.; Lanzavecchia, A. Two subsets of memory T lymphocytes with distinct homing potentials and effector functions [see comments]. Nature 1999, 401, 708–712. [Google Scholar] [CrossRef]
- Woodberry, T.; Suscovich, T.J.; Henry, L.M.; August, M.; Waring, M.T.; Kaur, A.; Hess, C.; Kutok, J.L.; Aster, J.C.; Wang, F.; et al. Alpha E beta 7 (CD103) expression identifies a highly active, tonsil-resident effector-memory CTL population. J. Immunol. 2005, 175, 4355–4362. [Google Scholar] [CrossRef]
- Wakim, L.M.; Woodward-Davis, A.; Bevan, M.J. Memory T cells persisting within the brain after local infection show functional adaptations to their tissue of residence. Proc. Natl. Acad. Sci. USA 2010, 107, 17872–17879. [Google Scholar] [CrossRef]
- Kumar, B.V.; Ma, W.; Miron, M.; Granot, T.; Guyer, R.S.; Carpenter, D.J.; Senda, T.; Sun, X.; Ho, S.H.; Lerner, H.; et al. Human Tissue-Resident Memory T Cells Are Defined by Core Transcriptional and Functional Signatures in Lymphoid and Mucosal Sites. Cell Rep. 2017, 20, 2921–2934. [Google Scholar] [CrossRef]
- Mackay, L.K.; Minnich, M.; Kragten, N.A.; Liao, Y.; Nota, B.; Seillet, C.; Zaid, A.; Man, K.; Preston, S.; Freestone, D.; et al. Hobit and Blimp1 instruct a universal transcriptional program of tissue residency in lymphocytes. Science 2016, 352, 459–463. [Google Scholar] [CrossRef]
- Walsh, D.A.; Borges da Silva, H.; Beura, L.K.; Peng, C.; Hamilton, S.E.; Masopust, D.; Jameson, S.C. The Functional Requirement for CD69 in Establishment of Resident Memory CD8(+) T Cells Varies with Tissue Location. J. Immunol. 2019, 203, 946–955. [Google Scholar] [CrossRef] [PubMed]
- Reilly, E.C.; Lambert-Emo, K.; Reilly, N.; Chaves, F.A.; Oakes, P.; Topham, D.J. TRM Integrins CD103 and CD49a Differentially Support Adherence and Motility After Resolution of Influenza Virus Infection. Proc. Natl. Acad. Sci. USA 2019, in press. [Google Scholar]
- Topham, D.J.; Reilly, E.C. Tissue-Resident Memory CD8(+) T Cells: From Phenotype to Function. Front. Immunol. 2018, 9, 515. [Google Scholar] [CrossRef] [PubMed]
- Sallusto, F.; Geginat, J.; Lanzavecchia, A. Central memory and effector memory T cell subsets: function, generation, and maintenance. Annu. Rev. Immunol. 2004, 22, 745–763. [Google Scholar] [CrossRef] [PubMed]
- Spertini, O.; Luscinskas, F.W.; Kansas, G.S.; Munro, J.M.; Griffin, J.D.; Gimbrone, M.A., Jr.; Tedder, T.F. Leukocyte adhesion molecule-1 (LAM-1, L-selectin) interacts with an inducible endothelial cell ligand to support leukocyte adhesion. J. Immunol. 1991, 147, 2565–2573. [Google Scholar]
- Sallusto, F.; Kremmer, E.; Palermo, B.; Hoy, A.; Ponath, P.; Qin, S.; Forster, R.; Lipp, M.; Lanzavecchia, A. Switch in chemokine receptor expression upon TCR stimulation reveals novel homing potential for recently activated T cells. Eur. J. Immunol. 1999, 29, 2037–2045. [Google Scholar] [CrossRef]
- Debes, G.F.; Hopken, U.E.; Hamann, A. In vivo differentiated cytokine-producing CD4(+) T cells express functional CCR7. J. Immunol. 2002, 168, 5441–5447. [Google Scholar] [CrossRef]
- Lazaar, A.L.; Albelda, S.M.; Pilewski, J.M.; Brennan, B.; Pure, E.; Panettieri, R.A., Jr. T lymphocytes adhere to airway smooth muscle cells via integrins and CD44 and induce smooth muscle cell DNA synthesis. J. Exp. Med. 1994, 180, 807–816. [Google Scholar] [CrossRef]
- Mrass, P.; Kinjyo, I.; Ng, L.G.; Reiner, S.L.; Pure, E.; Weninger, W. CD44 mediates successful interstitial navigation by killer T cells and enables efficient antitumor immunity. Immunity 2008, 29, 971–985. [Google Scholar] [CrossRef]
- Bank, I.; Book, M.; Ware, R. Functional role of VLA-1 (CD49A) in adhesion, cation-dependent spreading, and activation of cultured human T lymphocytes. Cell Immunol. 1994, 156, 424–437. [Google Scholar] [CrossRef]
- Richter, M.V.; Topham, D.J. The alpha1beta1 integrin and TNF receptor II protect airway CD8+ effector T cells from apoptosis during influenza infection. J. Immunol. 2007, 179, 5054–5063. [Google Scholar] [CrossRef] [PubMed]
- Richter, M.; Ray, S.J.; Chapman, T.J.; Austin, S.J.; Rebhahn, J.; Mosmann, T.R.; Gardner, H.; Kotelianski, V.; de Fougerolles, A.R.; Topham, D.J. Collagen distribution and expression of collagen-binding alpha1beta1 (VLA-1) and alpha2beta1 (VLA-2) integrins on CD4 and CD8 T cells during influenza infection. J. Immunol. 2007, 178, 4506–4516. [Google Scholar] [CrossRef]
- Lee, Y.T.; Suarez-Ramirez, J.E.; Redman, J.M.; Aguila, C.C.; Hadley, G.A.; Cauley, L.S. CD69 and CD103 cooperatively regulate CD8 T cell responses in the lungs after viral infection. J. Immunol. 2009, 182. [Google Scholar]
- Lee, Y.T.; Suarez-Ramirez, J.E.; Wu, T.; Redman, J.M.; Bouchard, K.; Hadley, G.A.; Cauley, L.S. Environmental and antigen receptor-derived signals support sustained surveillance of the lungs by pathogen-specific cytotoxic T lymphocytes. J. Virol. 2011, 85, 4085–4094. [Google Scholar] [CrossRef] [PubMed]
- Cibrian, D.; Sanchez-Madrid, F. CD69: from activation marker to metabolic gatekeeper. Eur. J. Immunol. 2017, 47, 946–953. [Google Scholar] [CrossRef] [PubMed]
- Pauls, K.; Schon, M.; Kubitza, R.C.; Homey, B.; Wiesenborn, A.; Lehmann, P.; Ruzicka, T.; Parker, C.M.; Schon, M.P. Role of integrin alphaE(CD103)beta7 for tissue-specific epidermal localization of CD8+ T lymphocytes. J. Invest. Dermatol. 2001, 117, 569–575. [Google Scholar] [CrossRef] [PubMed]
- Cepek, K.L.; Rimm, D.L.; Brenner, M.B. Expression of a candidate cadherin in T lymphocytes. Proc. Natl. Acad. Sci. USA 1996, 93, 6567–6571. [Google Scholar] [CrossRef]
- Sung, S.S.J.; Fu, S.M.; Rose, C.E.; Gaskin, F.; Ju, S.T.; Beaty, S.R. A Major Lung CD103 ( E)- 7 Integrin-Positive Epithelial Dendritic Cell Population Expressing Langerin and Tight Junction Proteins. J. Immunol. 2006, 176, 2161–2172. [Google Scholar] [CrossRef]
- Takamura, S.; Yagi, H.; Hakata, Y.; Motozono, C.; McMaster, S.R.; Masumoto, T.; Fujisawa, M.; Chikaishi, T.; Komeda, J.; Itoh, J.; et al. Specific niches for lung-resident memory CD8+ T cells at the site of tissue regeneration enable CD69-independent maintenance. J. Exp. Med. 2016, 213, 3057–3073. [Google Scholar] [CrossRef]
- Casey, K.A.; Fraser, K.A.; Schenkel, J.M.; Moran, A.; Abt, M.C.; Beura, L.K.; Lucas, P.J.; Artis, D.; Wherry, E.J.; Hogquist, K.; et al. Antigen-independent differentiation and maintenance of effector-like resident memory T cells in tissues. J. Immunol. 2012, 188, 4866–4875. [Google Scholar] [CrossRef]
- Mackay, L.K.; Wynne-Jones, E.; Freestone, D.; Pellicci, D.G.; Mielke, L.A.; Newman, D.M.; Braun, A.; Masson, F.; Kallies, A.; Belz, G.T.; et al. T-box Transcription Factors Combine with the Cytokines TGF-beta and IL-15 to Control Tissue-Resident Memory T Cell Fate. Immunity 2015, 43, 1101–1111. [Google Scholar] [CrossRef] [PubMed]
- McMaster, S.R.; Wein, A.N.; Dunbar, P.R.; Hayward, S.L.; Cartwright, E.K.; Denning, T.L.; Kohlmeier, J.E. Pulmonary antigen encounter regulates the establishment of tissue-resident CD8 memory T cells in the lung airways and parenchyma. Mucosal Immunol. 2018, 11, 1071–1078. [Google Scholar] [CrossRef] [PubMed]
- Bromley, S.K.; Thomas, S.Y.; Luster, A.D. Chemokine receptor CCR7 guides T cell exit from peripheral tissues and entry into afferent lymphatics. Nat. Immunol. 2005, 6, 895–901. [Google Scholar] [CrossRef] [PubMed]
- Debes, G.F.; Arnold, C.N.; Young, A.J.; Krautwald, S.; Lipp, M.; Hay, J.B.; Butcher, E.C. Chemokine receptor CCR7 required for T lymphocyte exit from peripheral tissues. Nat. Immunol. 2005, 6, 889–894. [Google Scholar] [CrossRef]
- Jennrich, S.; Lee, M.H.; Lynn, R.C.; Dewberry, K.; Debes, G.F. Tissue exit: a novel control point in the accumulation of antigen-specific CD8 T cells in the influenza a virus-infected lung. J. Virol. 2012, 86, 3436–3445. [Google Scholar] [CrossRef]
- Thompson, E.A.; Darrah, P.A.; Foulds, K.E.; Hoffer, E.; Caffrey-Carr, A.; Norenstedt, S.; Perbeck, L.; Seder, R.A.; Kedl, R.M.; Lore, K. Monocytes Acquire the Ability to Prime Tissue-Resident T Cells via IL-10-Mediated TGF-beta Release. Cell. Rep. 2019, 28, 1127–1135 e1124. [Google Scholar] [CrossRef]
- Zhang, N.; Bevan, M.J. Transforming growth factor-beta signaling controls the formation and maintenance of gut-resident memory T cells by regulating migration and retention. Immunity 2013, 39, 687–696. [Google Scholar] [CrossRef]
- Nath, A.P.; Braun, A.; Ritchie, S.C.; Carbone, F.R.; Mackay, L.K.; Gebhardt, T.; Inouye, M. Comparative analysis reveals a role for TGF-beta in shaping the residency-related transcriptional signature in tissue-resident memory CD8+ T cells. PLoS ONE 2019, 14, e0210495. [Google Scholar] [CrossRef]
- McKinstry, K.K.; Strutt, T.M.; Buck, A.; Curtis, J.D.; Dibble, J.P.; Huston, G.; Tighe, M.; Hamada, H.; Sell, S.; Dutton, R.W.; et al. IL-10 deficiency unleashes an influenza-specific Th17 response and enhances survival against high-dose challenge. J. Immunol. 2009, 182, 7353–7363. [Google Scholar] [CrossRef]
- Zhou, A.C.; Batista, N.V.; Watts, T.H. 4-1BB Regulates Effector CD8 T Cell Accumulation in the Lung Tissue through a TRAF1-, mTOR-, and Antigen-Dependent Mechanism to Enhance Tissue-Resident Memory T Cell Formation during Respiratory Influenza Infection. J. Immunol. 2019, 202, 2482–2492. [Google Scholar] [CrossRef]
- Konjar, S.; Veldhoen, M. Dynamic Metabolic State of Tissue Resident CD8 T Cells. Front. Immunol. 2019, 10, 1683. [Google Scholar] [CrossRef] [PubMed]
- Fox, C.J.; Hammerman, P.S.; Thompson, C.B. Fuel feeds function: energy metabolism and the T-cell response. Nat. Rev. Immunol. 2005, 5, 844–852. [Google Scholar] [CrossRef] [PubMed]
- Doedens, A.L.; Phan, A.T.; Stradner, M.H.; Fujimoto, J.K.; Nguyen, J.V.; Yang, E.; Johnson, R.S.; Goldrath, A.W. Hypoxia-inducible factors enhance the effector responses of CD8(+) T cells to persistent antigen. Nat. Immunol. 2013, 14, 1173–1182. [Google Scholar] [CrossRef] [PubMed]
- Macintyre, A.N.; Finlay, D.; Preston, G.; Sinclair, L.V.; Waugh, C.M.; Tamas, P.; Feijoo, C.; Okkenhaug, K.; Cantrell, D.A. Protein kinase B controls transcriptional programs that direct cytotoxic T cell fate but is dispensable for T cell metabolism. Immunity 2011, 34, 224–236. [Google Scholar] [CrossRef]
- Wang, R.; Dillon, C.P.; Shi, L.Z.; Milasta, S.; Carter, R.; Finkelstein, D.; McCormick, L.L.; Fitzgerald, P.; Chi, H.; Munger, J.; et al. The transcription factor Myc controls metabolic reprogramming upon T lymphocyte activation. Immunity 2011, 35, 871–882. [Google Scholar] [CrossRef]
- Maekawa, Y.; Ishifune, C.; Tsukumo, S.; Hozumi, K.; Yagita, H.; Yasutomo, K. Notch controls the survival of memory CD4+ T cells by regulating glucose uptake. Nat. Med. 2015, 21, 55–61. [Google Scholar] [CrossRef]
- Van der Windt, G.J.; Everts, B.; Chang, C.H.; Curtis, J.D.; Freitas, T.C.; Amiel, E.; Pearce, E.J.; Pearce, E.L. Mitochondrial respiratory capacity is a critical regulator of CD8+ T cell memory development. Immunity 2012, 36, 68–78. [Google Scholar] [CrossRef]
- Fahrer, A.M.; Konigshofer, Y.; Kerr, E.M.; Ghandour, G.; Mack, D.H.; Davis, M.M.; Chien, Y.H. Attributes of gammadelta intraepithelial lymphocytes as suggested by their transcriptional profile. Proc. Natl. Acad. Sci. USA 2001, 98, 10261–10266. [Google Scholar] [CrossRef]
- Pan, Y.; Tian, T.; Park, C.O.; Lofftus, S.Y.; Mei, S.; Liu, X.; Luo, C.; O’Malley, J.T.; Gehad, A.; Teague, J.E.; et al. Survival of tissue-resident memory T cells requires exogenous lipid uptake and metabolism. Nature 2017, 543, 252–256. [Google Scholar] [CrossRef]
- Konjar, S.; Frising, U.C.; Ferreira, C.; Hinterleitner, R.; Mayassi, T.; Zhang, Q.; Blankenhaus, B.; Haberman, N.; Loo, Y.; Guedes, J.; et al. Mitochondria maintain controlled activation state of epithelial-resident T lymphocytes. Sci. Immunol. 2018, 3. [Google Scholar] [CrossRef]
- Benador, I.Y.; Veliova, M.; Liesa, M.; Shirihai, O.S. Mitochondria Bound to Lipid Droplets: Where Mitochondrial Dynamics Regulate Lipid Storage and Utilization. Cell Metab 2019, 29, 827–835. [Google Scholar] [CrossRef] [PubMed]
- Benador, I.Y.; Veliova, M.; Mahdaviani, K.; Petcherski, A.; Wikstrom, J.D.; Assali, E.A.; Acin-Perez, R.; Shum, M.; Oliveira, M.F.; Cinti, S.; et al. Mitochondria Bound to Lipid Droplets Have Unique Bioenergetics, Composition, and Dynamics that Support Lipid Droplet Expansion. Cell Metab. 2018, 27, 869–885 e866. [Google Scholar] [CrossRef] [PubMed]
- Prasad, S.; Hu, S.; Sheng, W.S.; Chauhan, P.; Singh, A.; Lokensgard, J.R. The PD-1: PD-L1 pathway promotes development of brain-resident memory T cells following acute viral encephalitis. J. Neuroinflammation 2017, 14, 82. [Google Scholar] [CrossRef] [PubMed]
- Shwetank; Frost, E.L.; Mockus, T.E.; Ren, H.M.; Toprak, M.; Lauver, M.D.; Netherby-Winslow, C.S.; Jin, G.; Cosby, J.M.; Evavold, B.D.; et al. PD-1 Dynamically Regulates Inflammation and Development of Brain-Resident Memory CD8 T Cells During Persistent Viral Encephalitis. Front. Immunol. 2019, 10, 783. [Google Scholar] [CrossRef]
- Peggs, K.S.; Quezada, S.A. PD-1 blockade: promoting endogenous anti-tumor immunity. Expert Rev. Anticancer Ther. 2012, 12, 1279–1282. [Google Scholar] [CrossRef]
- Borges da Silva, H.; Beura, L.K.; Wang, H.; Hanse, E.A.; Gore, R.; Scott, M.C.; Walsh, D.A.; Block, K.E.; Fonseca, R.; Yan, Y.; et al. The purinergic receptor P2RX7 directs metabolic fitness of long-lived memory CD8(+) T cells. Nature 2018, 559, 264–268. [Google Scholar] [CrossRef] [PubMed]
- Konjar, S.; Ferreira, C.; Blankenhaus, B.; Veldhoen, M. Intestinal Barrier Interactions with Specialized CD8 T Cells. Front. Immunol. 2017, 8, 1281. [Google Scholar] [CrossRef] [PubMed]
- Waggoner, S.N.; Kumar, V. Evolving role of 2B4/CD244 in T and NK cell responses during virus infection. Front. Immunol. 2012, 3, 377. [Google Scholar] [CrossRef] [PubMed]
- Edwards, J.; Wilmott, J.S.; Madore, J.; Gide, T.N.; Quek, C.; Tasker, A.; Ferguson, A.; Chen, J.; Hewavisenti, R.; Hersey, P.; et al. CD103(+) Tumor-Resident CD8(+) T Cells Are Associated with Improved Survival in Immunotherapy-Naive Melanoma Patients and Expand Significantly During Anti-PD-1 Treatment. Clin. Cancer Res. Offic. J. Am. Assoc. Cancer Res. 2018, 24, 3036–3045. [Google Scholar] [CrossRef]
- Horvath, K.M.; Brighton, L.E.; Herbst, M.; Noah, T.L.; Jaspers, I. Live attenuated influenza virus (LAIV) induces different mucosal T cell function in nonsmokers and smokers. Clin. Immunol. 2012, 142, 232–236. [Google Scholar] [CrossRef]
- Xu, X.; Wang, H.; Liu, Y.; Wang, Y.; Zeng, L.; Wu, K.; Wang, J.; Ma, F.; Xu, W.; Yin, Y.; et al. Mucosal immunization with the live attenuated vaccine SPY1 induces humoral and Th2-Th17-regulatory T cell cellular immunity and protects against pneumococcal infection. Infect. Immun. 2015, 83, 90–100. [Google Scholar] [CrossRef] [PubMed]
| Memory type | Abbreviation | CD3 | TCRb | CD8 | CD44 | CD62L | CD69 | CD49a | CD103 |
|---|---|---|---|---|---|---|---|---|---|
| Effector | EM | + | + | + | high | low | − | − | − |
| Epithelial TRM | eTRM | + | + | + | high | low | + | + | + |
| Interstitial TRM | iTRM | + | + | + | high | low | + | + | − |
| Epithelial Effector | eeTRM | + | + | + | high | low | + | − | + |
| Marker | Function | Presence on TRM |
|---|---|---|
| CD44 | Binds hyaluronic acids. May serve roles in motility and retention in both peripheral and lymphoid tissues. | +/High |
| CD62L | Ligand for high endothelial venules on SLO. | −/Low |
| CCR7 | Chemokine receptor for S1P1. | −/Low |
| CD49a | Mediates adhesion to type I and IV collagen. | Positive on most TRM |
| CD69 | Early activation marker. S1P1R antagonist. Binds Gal-1. | Positive on most TRM |
| CD103 | Binds E-Cadherin. | Very often present, though can be tissue-dependent. |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Topham, D.J.; Reilly, E.C.; Emo, K.L.; Sportiello, M. Formation and Maintenance of Tissue Resident Memory CD8+ T Cells after Viral Infection. Pathogens 2019, 8, 196. https://doi.org/10.3390/pathogens8040196
Topham DJ, Reilly EC, Emo KL, Sportiello M. Formation and Maintenance of Tissue Resident Memory CD8+ T Cells after Viral Infection. Pathogens. 2019; 8(4):196. https://doi.org/10.3390/pathogens8040196
Chicago/Turabian StyleTopham, David J., Emma C. Reilly, Kris Lambert Emo, and Mike Sportiello. 2019. "Formation and Maintenance of Tissue Resident Memory CD8+ T Cells after Viral Infection" Pathogens 8, no. 4: 196. https://doi.org/10.3390/pathogens8040196
APA StyleTopham, D. J., Reilly, E. C., Emo, K. L., & Sportiello, M. (2019). Formation and Maintenance of Tissue Resident Memory CD8+ T Cells after Viral Infection. Pathogens, 8(4), 196. https://doi.org/10.3390/pathogens8040196





