Experimental Infection of Human Volunteers with the Heat-Stable Enterotoxin-Producing Enterotoxigenic Escherichia coli Strain TW11681
Abstract
1. Introduction
2. Material and Methods
2.1. Volunteer Inclusion and Screening
2.2. Setting
2.3. Strain Description
2.4. Preparation of Inocula
2.5. Experimental Infection
2.6. Clinical Evaluation
2.7. Specimen Collection
2.8. Detection of Enterotoxigenic Escherichia coli (ETEC) in Stool Specimens
2.9. Antigen Preparations
2.10. T Cell Assay and Flow Cytometry
2.11. Antibody Assays
2.12. Ethics Approval
3. Results
3.1. Volunteer Characteristics
3.2. Clinical Results
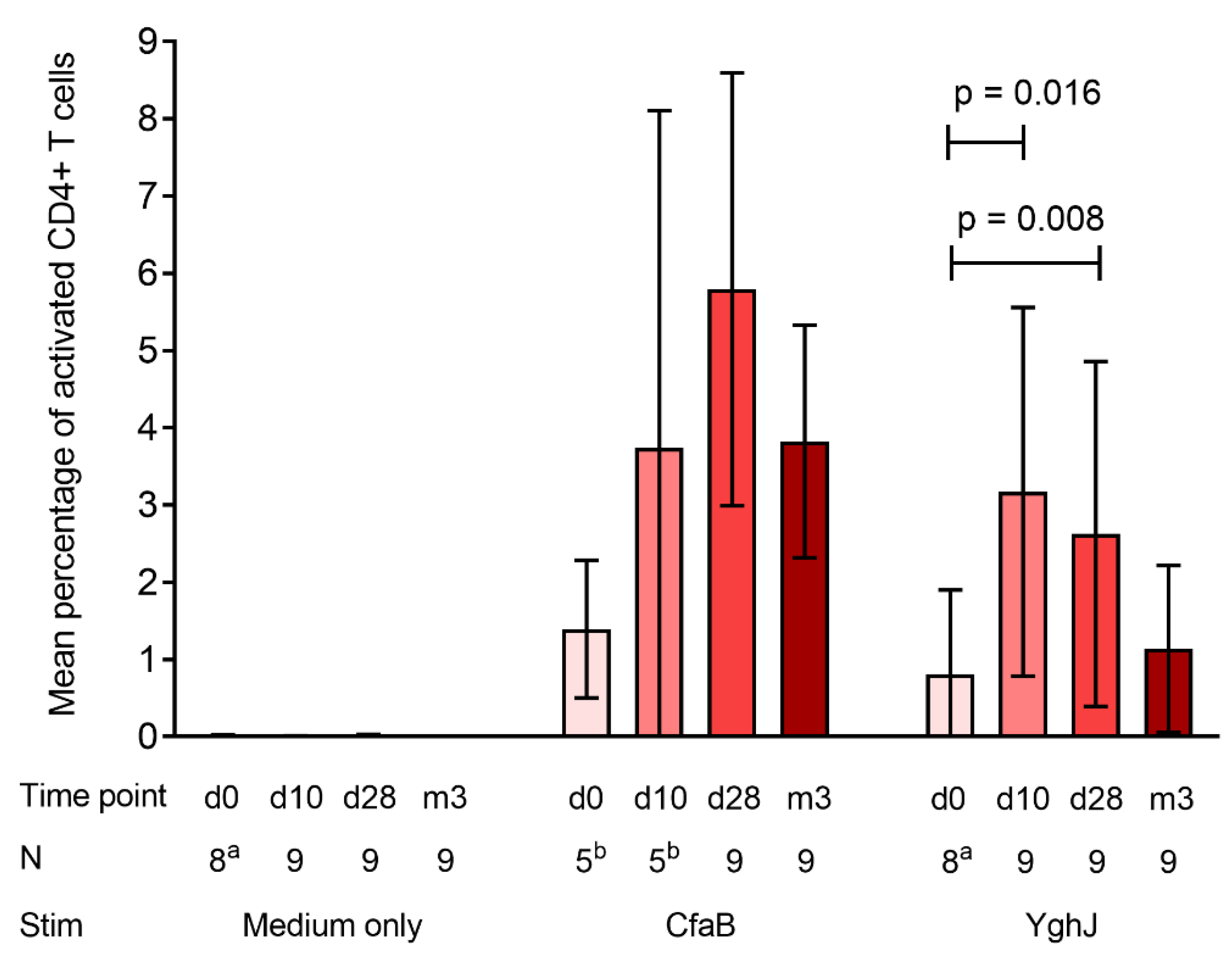
3.3. CD4+ T Cell Responses
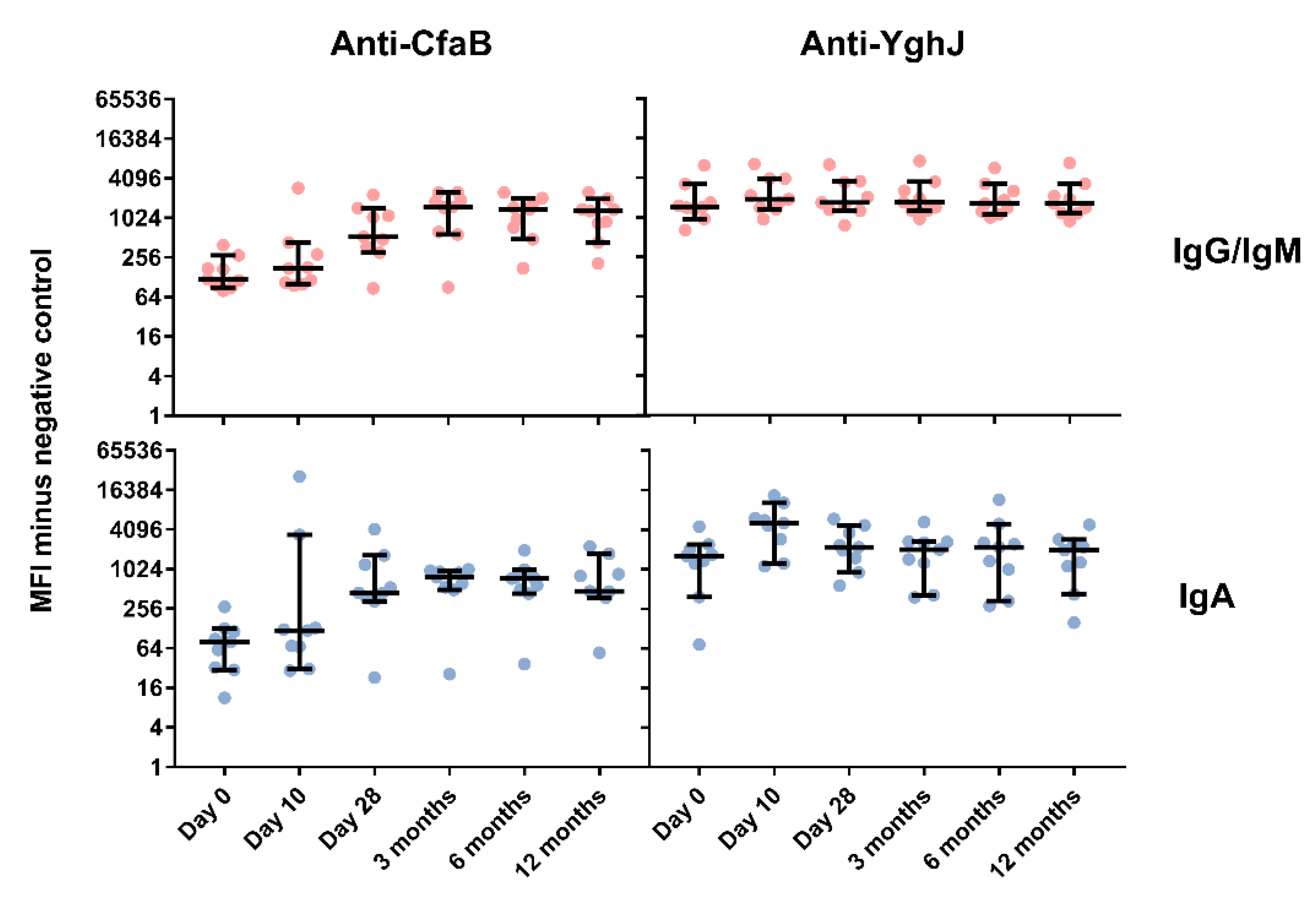
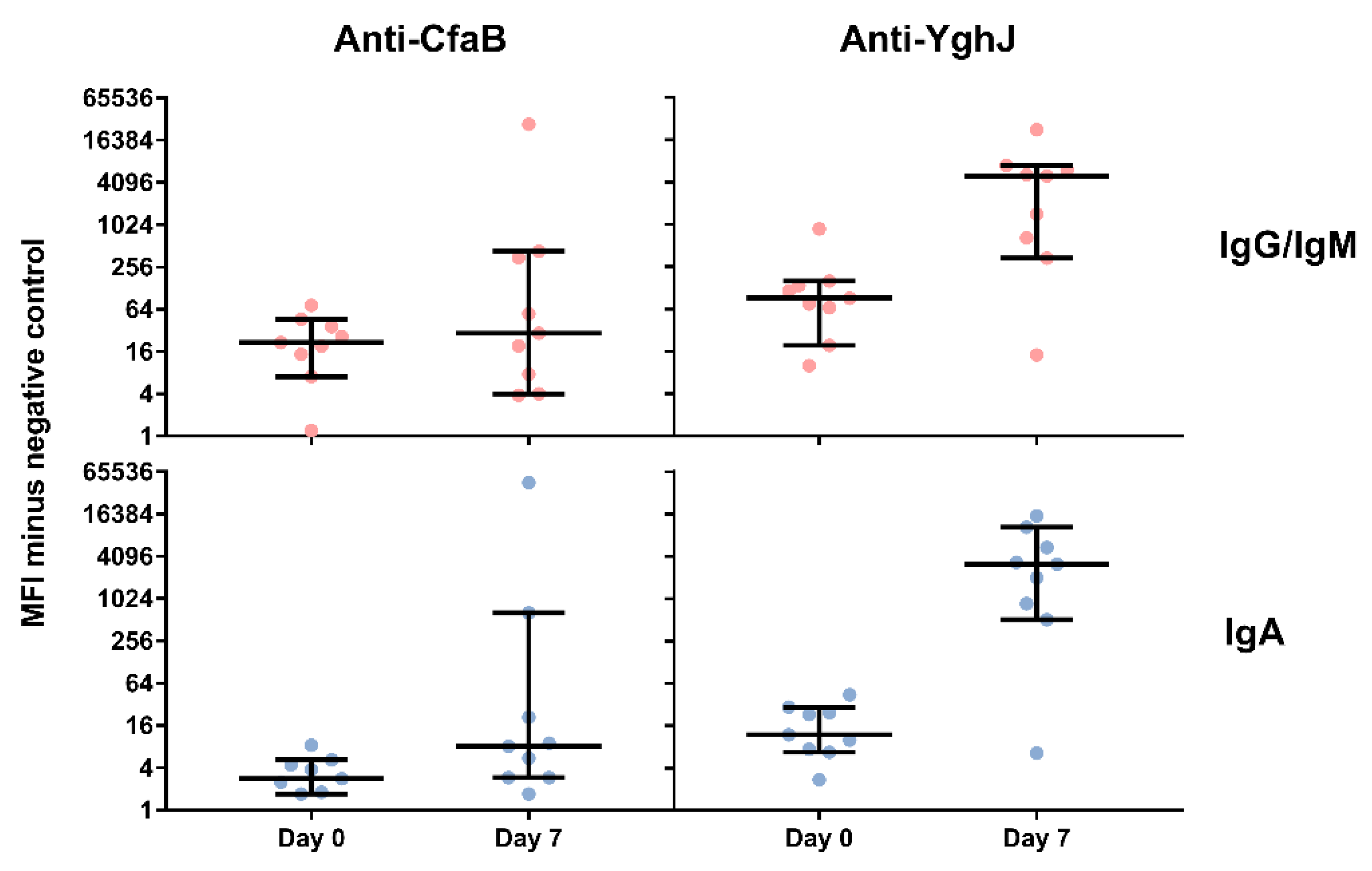
3.4. Serum and Lymphocyte SupernatantAntibody Responses Against TW11681 Antigens
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bourgeois, A.L.; Wierzba, T.F.; Walker, R.I. Status of vaccine research and development for enterotoxigenic Escherichia coli. Vaccine 2016, 34, 2880–2886. [Google Scholar] [CrossRef]
- Kotloff, K.L.; Nataro, J.P.; Blackwelder, W.C.; Nasrin, D.; Farag, T.H.; Panchalingam, S.; Wu, Y.; Sow, S.O.; Sur, D.; Breiman, R.F.; et al. Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): A prospective, case-control study. Lancet 2013, 382, 209–222. [Google Scholar] [CrossRef]
- Porat, N.; Levy, A.; Fraser, D.; Deckelbaum, R.J.; Dagan, R. Prevalence of intestinal infections caused by diarrheagenic Escherichia coli in Bedouin infants and young children in Southern Israel. Pediatr. Infect. Dis. J. 1998, 17, 482–488. [Google Scholar] [CrossRef]
- Steinsland, H.; Valentiner-Branth, P.; Perch, M.; Dias, F.; Fischer, T.K.; Aaby, P.; Molbak, K.; Sommerfelt, H. Enterotoxigenic Escherichia coli infections and diarrhea in a cohort of young children in Guinea-Bissau. J. Infect. Dis. 2002, 186, 1740–1747. [Google Scholar] [CrossRef]
- Clements, M.L.; Levine, M.M.; Black, R.E.; Robins-Browne, R.M.; Cisneros, L.A.; Drusano, G.L.; Lanata, C.F.; Saah, A.J. Lactobacillus prophylaxis for diarrhea due to enterotoxigenic Escherichia coli. Antimicrob. Agents Chemother. 1981, 20, 104–108. [Google Scholar] [CrossRef]
- Levine, M.M.; Caplan, E.S.; Waterman, D.; Cash, R.A.; Hornick, R.B.; Snyder, M.J. Diarrhea caused by Escherichia coli that produce only heat-stable enterotoxin. Infect. Immun. 1977, 17, 78–82. [Google Scholar]
- Levine, M.M.; Rennels, M.B.; Cisneros, L.; Hughes, T.P.; Nalin, D.R.; Young, C.R. Lack of person-to-person transmission of enterotoxigenic Escherichia coli despite close contact. Am. J. Epidemiol. 1980, 111, 347–355. [Google Scholar] [CrossRef]
- Sack, D.A.; Merson, M.H.; Wells, J.G.; Sack, R.B.; Morris, G.K. Diarrhoea associated with heat-stable enterotoxin-producing strains of Escherichia coli. Lancet 1975, 2, 239–241. [Google Scholar] [CrossRef]
- Liu, J.; Platts-Mills, J.A.; Juma, J.; Kabir, F.; Nkeze, J.; Okoi, C.; Operario, D.J.; Uddin, J.; Ahmed, S.; Alonso, P.L.; et al. Use of quantitative molecular diagnostic methods to identify causes of diarrhoea in children: A reanalysis of the GEMS case-control study. Lancet 2016, 388, 1291–1301. [Google Scholar] [CrossRef]
- Luo, Q.; Kumar, P.; Vickers, T.J.; Sheikh, A.; Lewis, W.G.; Rasko, D.A.; Sistrunk, J.; Fleckenstein, J.M. Enterotoxigenic Escherichia coli secretes a highly conserved mucin-degrading metalloprotease to effectively engage intestinal epithelial cells. Infect. Immun. 2014, 82, 509–521. [Google Scholar] [CrossRef]
- Luo, Q.; Qadri, F.; Kansal, R.; Rasko, D.A.; Sheikh, A.; Fleckenstein, J.M. Conservation and immunogenicity of novel antigens in diverse isolates of enterotoxigenic Escherichia coli. PLoS Negl. Trop. Dis. 2015, 9, e0003446. [Google Scholar] [CrossRef]
- Skrede, S.; Steinsland, H.; Sommerfelt, H.; Aase, A.; Brandtzaeg, P.; Langeland, N.; Cox, R.J.; Saevik, M.; Wallevik, M.; Skutlaberg, D.H.; et al. Experimental infection of healthy volunteers with enterotoxigenic Escherichia coli wild-type strain TW10598 in a hospital ward. BMC Infect. Dis. 2014, 14, 482. [Google Scholar] [CrossRef]
- Steinsland, H.; Lacher, D.W.; Sommerfelt, H.; Whittam, T.S. Ancestral lineages of human enterotoxigenic Escherichia coli. J. Clin. Microbiol. 2010, 48, 2916–2924. [Google Scholar] [CrossRef]
- Sahl, J.W.; Steinsland, H.; Redman, J.C.; Angiuoli, S.V.; Nataro, J.P.; Sommerfelt, H.; Rasko, D.A. A comparative genomic analysis of diverse clonal types of enterotoxigenic Escherichia coli reveals pathovar-specific conservation. Infect. Immun. 2011, 79, 950–960. [Google Scholar] [CrossRef]
- Von Mentzer, A.; Connor, T.R.; Wieler, L.H.; Semmler, T.; Iguchi, A.; Thomson, N.R.; Rasko, D.A.; Joffre, E.; Corander, J.; Pickard, D.; et al. Identification of enterotoxigenic Escherichia coli (ETEC) clades with long-term global distribution. Nat. Genet. 2014, 46, 1321–1326. [Google Scholar] [CrossRef]
- Porter, C.K.; Riddle, M.S.; Tribble, D.R.; Louis Bougeois, A.; McKenzie, R.; Isidean, S.D.; Sebeny, P.; Savarino, S.J. A systematic review of experimental infections with enterotoxigenic Escherichia coli (ETEC). Vaccine 2011, 29, 5869–5885. [Google Scholar] [CrossRef]
- Harro, C.; Chakraborty, S.; Feller, A.; DeNearing, B.; Cage, A.; Ram, M.; Lundgren, A.; Svennerholm, A.M.; Bourgeois, A.L.; Walker, R.I.; et al. Refinement of a human challenge model for evaluation of enterotoxigenic Escherichia coli vaccines. Clin. Vaccine Immunol. 2011, 18, 1719–1727. [Google Scholar] [CrossRef]
- Porter, C.K.; Riddle, M.S.; Alcala, A.N.; Sack, D.A.; Harro, C.; Chakraborty, S.; Gutierrez, R.L.; Savarino, S.J.; Darsley, M.; McKenzie, R.; et al. An evidenced-based scale of disease severity following human challenge with enteroxigenic Escherichia coli. PLoS ONE 2016, 11, e0149358. [Google Scholar] [CrossRef]
- Aase, A.; Sommerfelt, H.; Petersen, L.B.; Bolstad, M.; Cox, R.J.; Langeland, N.; Guttormsen, A.B.; Steinsland, H.; Skrede, S.; Brandtzaeg, P. Salivary IgA from the sublingual compartment as a novel noninvasive proxy for intestinal immune induction. Mucosal Immunol. 2016, 9, 884–893. [Google Scholar] [CrossRef]
- Stacy-Phipps, S.; Mecca, J.J.; Weiss, J.B. Multiplex PCR assay and simple preparation method for stool specimens detect enterotoxigenic Escherichia coli DNA during course of infection. J. Clin. Microbiol. 1995, 33, 1054–1059. [Google Scholar]
- Sadler, R.; Bateman, E.A.; Heath, V.; Patel, S.Y.; Schwingshackl, P.P.; Cullinane, A.C.; Ayers, L.; Ferry, B.L. Establishment of a healthy human range for the whole blood "OX40" assay for the detection of antigen-specific CD4+ T cells by flow cytometry. Cytometry B Clin. Cytom. 2014, 86, 350–361. [Google Scholar] [CrossRef]
- Zaunders, J.J.; Munier, M.L.; Seddiki, N.; Pett, S.; Ip, S.; Bailey, M.; Xu, Y.; Brown, K.; Dyer, W.B.; Kim, M.; et al. High levels of human antigen-specific CD4+ T cells in peripheral blood revealed by stimulated coexpression of CD25 and CD134 (OX40). J. Immunol. 2009, 183, 2827–2836. [Google Scholar] [CrossRef]
- Reiss, S.; Baxter, A.E.; Cirelli, K.M.; Dan, J.M.; Morou, A.; Daigneault, A.; Brassard, N.; Silvestri, G.; Routy, J.P.; Havenar-Daughton, C.; et al. Comparative analysis of activation induced marker (AIM) assays for sensitive identification of antigen-specific CD4 T cells. PLoS ONE 2017, 12, e0186998. [Google Scholar] [CrossRef]
- Porter, C.K.; Louis Bourgeois, A.; Frenck, R.W.; Prouty, M.; Maier, N.; Riddle, M.S. Developing and utilizing controlled human models of infection. Vaccine 2017, 35, 6813–6818. [Google Scholar] [CrossRef]
- Vedoy, O.B.; Hanevik, K.; Sakkestad, S.T.; Sommerfelt, H.; Steinsland, H. Proliferation of enterotoxigenic Escherichia coli strain TW11681 in stools of experimentally infected human volunteers. Gut Pathog. 2018, 10, 46. [Google Scholar] [CrossRef]
- Isidean, S.D.; Riddle, M.S.; Savarino, S.J.; Porter, C.K. A systematic review of ETEC epidemiology focusing on colonization factor and toxin expression. Vaccine 2011, 29, 6167–6178. [Google Scholar] [CrossRef]
| Target Dose (CFU) | No. of Volunteers | No. with Diarrhoea | Attack Risk | Median Severity | Mean Incubation Period, Hours (Range) | Mean 24h Maximum Stool Output, Grams (Range) | Mean Whole Episode Stool Output, Grams (Range) | Mean Episode Duration, Hours (Range) | Mean 24h Maximum Stool Output, Count (Range) |
|---|---|---|---|---|---|---|---|---|---|
| 1 × 106 | 3 | 2 | 67% | Mild | 58 (35–80) | 364 (289–389) | 364 (289–389) | 1.3 (0–2.5) | 1.5 (1–2) |
| 1 × 107 | 3 | 0 | 0% | NA | NA | NA | NA | NA | NA |
| 1 × 108 | 3 | 0 | 0% | NA | NA | NA | NA | NA | NA |
| Symptom | Dose (CFU) | Combined (% of Volunteers) | ||
|---|---|---|---|---|
| 1 × 106 | 1 × 107 | 1 × 108 | ||
| Diarrhoea | 2 | 0 | 0 | 2 (22%) |
| Nausea | 1 | 0 | 0 | 1 (11%) |
| Abdominal pain | 2 | 1 | 2 | 5 (56%) |
| Abdominal cramping | 2 | 0 | 2 | 4 (44%) |
| Flatulence | 1 | 1 | 1 | 3 (33%) |
| Decreased appetite | 1 | 1 | 0 | 2 (22%) |
| Bloating | 1 | 0 | 1 | 2 (22%) |
| Headache | 1 | 0 | 0 | 1 (11%) |
| Malaise | 1 | 1 | 0 | 2 (22%) |
| Lightheadedness | 1 | 0 | 0 | 1 (11%) |
| Dose (CFU) | Volunteer ID | Score Components | Disease Severity Score (0–8) | ||
|---|---|---|---|---|---|
| Objective Signs (0–2) | Subjective Symptoms (0–2) | Diarrhoea Score (0–4) | |||
| 1 × 106 | EV10 | 0 | 2 | 1 | 3 |
| 1 × 106 | EV11 | 0 | 1 | 1 | 2 |
| 1 × 106 | EV12 | 0 | 0 | 0 | 0 |
| 1 × 107 | EV13 | 0 | 0 | 0 | 0 |
| 1 × 107 | EV14 | 0 | 0 | 0 | 0 |
| 1 × 107 | EV15 | 0 | 1 | 0 | 1 |
| 1 × 108 | EV16 | 0 | 1 | 0 | 1 |
| 1 × 108 | EV17 | 0 | 0 | 0 | 0 |
| 1 × 108 | EV18 | 0 | 1 | 0 | 1 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Todnem Sakkestad, S.; Steinsland, H.; Skrede, S.; Kleppa, E.; Lillebø, K.; Sævik, M.; Søyland, H.; Rykkje Heien, A.; Gjerde Tellevik, M.; Barry, E.M.; et al. Experimental Infection of Human Volunteers with the Heat-Stable Enterotoxin-Producing Enterotoxigenic Escherichia coli Strain TW11681. Pathogens 2019, 8, 84. https://doi.org/10.3390/pathogens8020084
Todnem Sakkestad S, Steinsland H, Skrede S, Kleppa E, Lillebø K, Sævik M, Søyland H, Rykkje Heien A, Gjerde Tellevik M, Barry EM, et al. Experimental Infection of Human Volunteers with the Heat-Stable Enterotoxin-Producing Enterotoxigenic Escherichia coli Strain TW11681. Pathogens. 2019; 8(2):84. https://doi.org/10.3390/pathogens8020084
Chicago/Turabian StyleTodnem Sakkestad, Sunniva, Hans Steinsland, Steinar Skrede, Elisabeth Kleppa, Kristine Lillebø, Marianne Sævik, Hanne Søyland, Astrid Rykkje Heien, Marit Gjerde Tellevik, Eileen M. Barry, and et al. 2019. "Experimental Infection of Human Volunteers with the Heat-Stable Enterotoxin-Producing Enterotoxigenic Escherichia coli Strain TW11681" Pathogens 8, no. 2: 84. https://doi.org/10.3390/pathogens8020084
APA StyleTodnem Sakkestad, S., Steinsland, H., Skrede, S., Kleppa, E., Lillebø, K., Sævik, M., Søyland, H., Rykkje Heien, A., Gjerde Tellevik, M., Barry, E. M., Sommerfelt, H., & Hanevik, K. (2019). Experimental Infection of Human Volunteers with the Heat-Stable Enterotoxin-Producing Enterotoxigenic Escherichia coli Strain TW11681. Pathogens, 8(2), 84. https://doi.org/10.3390/pathogens8020084





