1. Introduction
Members of the bacterial genus
Aeromonas are natural inhabitants of aquatic environments, including freshwater, estuarine, and marine water bodies. Several species are pathogenic for fish, causing ulcerative, hemorrhagic and even septicemic diseases with relevant economic losses [
1]. Among those diseases, furunculosis is commonly associated with skin lesions which rapidly progress to a septicemic infection with high mortality, typically caused by members of the
Aeromonas salmonicida clade [
2]. Classical furunculosis is produced by brown-pigmented, psychrophilic, Gram-negative bacteria, grouped into the subspecies
A. salmonicida subsp.
salmonicida which mainly affects wild and cultured salmonids. By contrast, non-pigmented atypical subspecies are able to infect a broader number of hosts including carp (
Cyprinus carpio), goldfish (
Carassius auratus), flounder (
Platichthys flesus), Siberian sturgeons (
Acipenser baerii), and captive snakehead fish (
Channa argus) [
2,
3,
4,
5], with the exception of
A. salmonicida subsp.
pectinolytica, a mesophilic subspecies that was only isolated from environmental samples [
6].
Aeromonas piscicola is closely related to
A. salmonicida. This bacterium has first been isolated from cultured salmonids in Spain that exhibited signs of furunculosis-like disease. Phenotypically,
A. piscicola comprises mesophilic, motile microorganisms that do not produce brown pigment, and which can be distinguished from mesophilic
A. salmonicida by its inability to ferment lactose and L-arabinose. Conventional 16S rRNA sequencing, however, cannot differentiate between them [
7]. More recently and along with other pathogenic members of the Aeromonadaceae family,
A. piscicola has been associated with Red Skin Disease, a condition affecting wild salmonid populations in Europe [
8]. In the context of increasingly challenging environmental conditions driven by climate change, it is important to draw attention to mesophilic bacteria, particularly pathogens affecting fish species reared in freshwater, which is especially susceptible to warming.
Chile, as the world’s second-largest salmonid producer, faces several sanitary challenges in both stages of the production cycle. In freshwater, the incidence of furunculosis in Atlantic salmon (
Salmo salar), the most relevant reared salmonid species in the Chilean industry, has increased in recent years, from almost marginal to 7.8% of the total amount of secondary mortality (i.e., caused by infectious diseases). Still, a decline was noted in the last report [
9]. Atypical
A. salmonicida has long been recognized as responsible for furunculosis [
10], but in the face of increasing incidence, some authors have suggested its re-emergence [
11]. The discovery of an
A. salmonicida strain associated with mild disease with limited mortality, which may turn more deadly when stress and suboptimal rearing conditions occur, added a new layer of complexity to the epidemiological scenario [
12].
Using our improved PCR scheme based on vapA-fstA genes for typing A. salmonicida isolates from our strain collection, we observed that some A. salmonicida-like isolates did not amplify in the molecular assay. In the present work, we unveiled the identity and phenotypic characteristics of those isolates by combining sequencing and microbiological approaches. Our results strongly support the identification of A. piscicola as the causative agent of some recent furunculosis-like outbreaks, which is herein reported for the first time in Chile. We further characterized its virulence potential in Atlantic salmon and present evidence on the scarce serological response elicited by the current furunculosis vaccine in comparison with the immune response directed against A. salmonicida.
2. Materials and Methods
2.1. Biological Material and Culture
Tissue samples from suspected cases of furunculosis were collected during diagnostic routine at ADL Diagnostic’s laboratory in Puerto Montt, Chile. Samples, maintained at −20 °C in ethanol, were subjected to a DNA purification protocol (E.Z.N.A. tissue DNA purification kit, Omega Bio-tek, Norcross, GA, USA). Bacterial strains stored at −80 °C in the ADL strain collection were recovered and cultured on trypticase soy agar (TSA; BD) plates at 18 °C for 48 h. Some isolates were grown in trypticase soy broth (TSB; BD). For molecular and sequencing analysis, a single colony of each isolate was processed for DNA purification using a commercial kit (GeneJET genomic DNA purification kit, Thermo Scientific, Waltham, MA, USA). Metadata of bacterial strains are shown in
Table 1.
2.2. Bacterial Growth Kinetics, Counts and Antibiotic Susceptibility Testing
Bacterial isolates were grown on TSA plates at 18 °C for 48 h. Then, a loopful of each isolate was transferred to sterile saline to prepare bacterial suspensions, which were then adjusted to OD600 = 1.00. Next, 96-well microplates were inoculated with 100 µL of these preparations diluted in TSB at 1:100. Bacterial growth kinetics of three A. piscicola and two A. salmonicida isolates were evaluated in an automatic spectrophotometer EPOCH 2 (BioTek, Winooski, VT, USA) at incubation temperatures of 10, 23, and 37 °C. Reads were taken hourly for 48 h. To assess the effect of salinity on growth, TSB was supplemented with 0, 0.5, and 2% of NaCl. For bacterial counts, suspensions were 10-fold serially diluted in saline, plated in triplicate onto TSA, maintained at 18 °C for 48 h, and counted.
For antibiotic susceptibility assays, a saline suspension of bacteria freshly cultured on TSA was prepared and likewise adjusted to OD
600 = 1.00. Minimal inhibitory concentration (MIC) testing was performed using a 2-fold serial microdilution method in Müller-Hinton (M-H) broth, with concentrations ranging from 0.03 to 64 µg × mL
−1. Antimicrobial agents florfenicol and oxytetracycline were purchased from Sigma-Aldrich Co., (St. Louis, MO, USA) and were prepared following the CLSI recommendations for broth dilution susceptibility testing of bacteria isolated from aquatic animals, guidelines VET04-A2 and VET03/VET04-S2 [
13,
14]. Standard stock solutions were prepared by dissolving 10 mg of each antibiotic in 500 µL of 95% ethanol and 1 M NaOH for florfenicol or 100% methanol for oxytetracycline, and the final volume was adjusted to 10 mL with sterilized Milli-Q water. Stock solutions were stored at −80 °C. Challenge trays included positive control wells containing bacterial suspensions in M-H broth without antibiotic, and negative controls consisting of uninoculated broth. Each test was carried out in triplicate, considering incubation for 48 h at 18 °C. The MIC was defined as the lowest concentration of the antibiotic that prevented bacterial growth.
2.3. PCR Assays and Whole Genome Sequencing
Specific TaqMan qPCR assays for
vapA-fstA genes were performed for the identification of
A. salmonicida according to our previous work [
12]. The same validated AgPath-ID One-Step RT-PCR chemistry used in our routine diagnostic workflow was maintained for these assays, although DNA was the nucleic acid template in this study. PCR conditions were as follows: For every primer/probe set, 300 nM of primer and 200 nM of probe were combined and mixed with 3 µL of nucleic acid template, enzyme and master mix using AgPath-ID™ One-Step RT-PCR and nuclease-free water as per the manufacturer’s instructions (Applied Biosystems™, Carlsbad, CA, USA). Furthermore, we considered 10 min at 45 °C for reverse transcription and 10 min at 95 °C for reverse transcriptase inactivation, followed by 45 cycles of 5 s at 95 °C and 30 s at 60 °C for annealing-extension. Reactions were run in a Quant Studio 3 PCR machine (Applied Biosystems™). A similar amplification protocol was followed for examining tissue samples derived from the bioassay, employing a primer set for
Aeromonas spp. described elsewhere [
15]. Sequences of primers and probe set are listed in
Table S1.
Whole genome sequencing was conducted at Codebreaker Bioscience (Santiago, Chile). Libraries were created using the standard protocol of Nextera Flex (Illumina™, San Diego, CA, USA). All standard reagents from the Nextera DNA Flex library prep kit were used to prepare each standard Flex library, in accordance with the manufacturer’s instructions. A Qubit high-sensitivity (HS) dsDNA kit (Thermo Fisher Scientific, Waltham, MA, USA) and the S2 Cartridge from BIOtic (Pinneberg, Germany) with the Qsep1 Machine were used to assess the concentration of eluted libraries and the library size. For sequencing, a 600-cycle kit configured for 2 × 300 bp paired-end reads was used on an Illumina™ Nextseq1000™ platform.
2.4. Bioinformatic Analysis
A custom Nextflow pipeline was applied to analyze raw FASTQ files
https://github.com/gene2dis/mgap (accessed on 24 February 2026). That pipeline included quality control checks and cleaning of the raw reads with FastP (v.0.23.2) [
16], assembly with Spades (v.3.15.5) [
17] and contig annotation with Bakta (v.1.7.0) [
18]. Finally, we employed BLASTN v2.15.0 to screen public databases for homologous sequences of contigs of interest [
19]. Furthermore, the clinker tool described by Gilchrist and Chooi [
20] allowed us to visualize groups of homologous biosynthesis gene clusters. Average nucleotide identities (ANI) were calculated with the ANI calculator tool hosted at
https://www.ezbiocloud.net/ (accessed on 25 February 2026) [
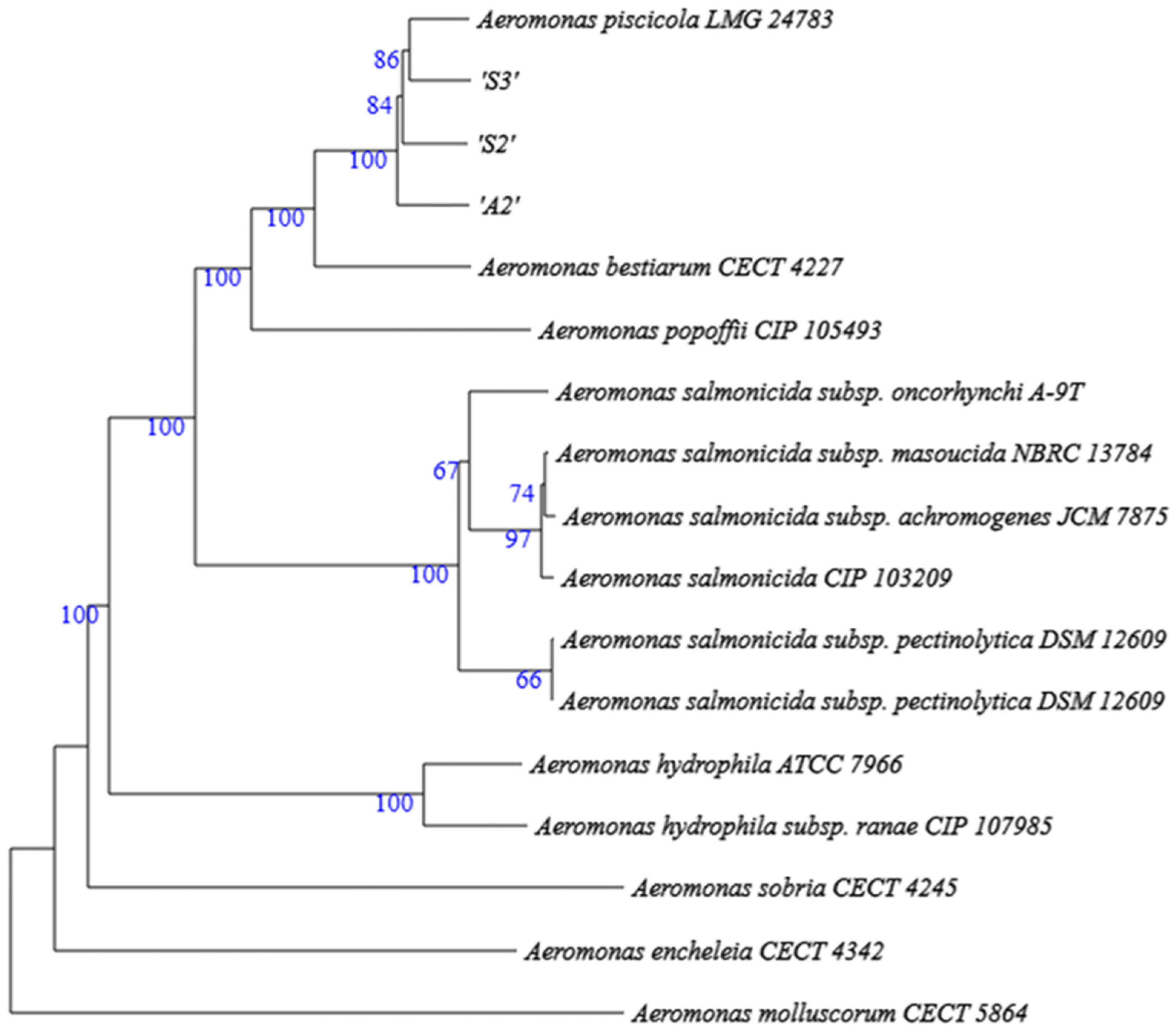
21]. Phylogenomic analysis was conducted using the Type Strain Genome Server [
22] to compare A2, S2, and S3 strains with sequences of reference type strains from the DSMZ database including
A. salmonicida CIP 103209,
A. salmonicida subsp.
oncorhynchi A-9,
A. salmonicida subsp.
masoucida NBRC 13784,
A. salmonicida subsp.
achromogenes JCM 7875,
A. salmonicida subsp.
pectinolytica DSM 12609 (two assemblies),
A. hydrophila ATCC 7966,
A.
hydrophila subsp.
ranae CIP 107985,
A. piscicola LMG 24783,
A. sobria CECT 4245,
A. molluscorum CECT 5864,
A. bestiarum CECT 4247,
A. encheleia CECT 4342, and
A. popoffii CIP 105493.
The functional genome characterization of A1, A2, A4, S2, and S3 was conducted using customized Python v.3.12.13-based bioinformatics pipelines run in Google Colab (
https://colab.google/, accessed on 24 February 2026) to predict virulence factors (VF) and antibiotic resistance genes. Virulence-associated genes were inferred by screening the genomes against the Virulence Factor Database (VFDB) v5 [
23]. This database contains curated coding sequences (CDS) of genes categorized into toxins, secretion systems, adhesion factors, and immune evasion mechanisms. Reciprocal Best Hit (RBH) analysis was performed to validate VF identification. Briefly, the VFDB protein dataset was indexed to create a DIAMOND reference database. Each *.faa file was subjected to a DIAMOND BLASTP v.2.0.14 search against the prepared VFDB. This step identified initial best hits from the query genome to VFDB entries. Hits from the forward search were filtered using stringent criteria: minimum 60% protein sequence identity (MIN_PIDENT), minimum 90% query and subject coverage (MIN_COV), maximum E-value of 1 × 10
−50 (MAX_EVALUE), and a minimum alignment length of 100 amino acids (MIN_ALN). A subset of the VFDB was dynamically generated, containing only the VF entries that passed the strict filtering from the forward search. Concurrently, a DIAMOND database was created from the original query genome’s proteome. A reverse DIAMOND BLASTP search was then performed, querying the VFDB subset against the query genome database. The results from the strictly filtered forward and reverse searches were merged. Only hits that were RBH (i.e., protein A from genome is the best hit for VF B, and VF B is the best hit for protein A) were retained as putative VF orthologs. The identified RBH pairs were annotated with their corresponding VFDB header, gene symbol (extracted from the header when available), VFDB Group ID, and a robust VF_key for consistent identification across genomes. A sequence with a known number of VF was used as calibrator [
24].
Protein FASTA files from bacterial genomes were then processed to identify antimicrobial resistance (AMR) genes using the AMRFinderPlus tool v.4.2.7 [
25], with results compiled into a Pandas data frame. For comparative validation, AMRFinderPlus outputs were cross-referenced with the RGI v.6.0.7 of Comprehensive Antibiotic Resistance Database (CARD) prediction dataset [
26].
2.5. Purification and Analysis of Bacterial Membrane Antigens
Outer membrane protein (OMP) fractions were obtained following a modified, Sarkosyl-based extraction strategy [
27]. Briefly, isolates grown on TSA were suspended in saline and subsequently centrifuged at 10,000×
g for 10 min. Pellets were resuspended in lysis buffer containing 300 mM NaCl, 10 mM HEPES, 2 mM PMSF, and 8 M urea, before sonication on ice for 1 min at 70% potency (300 V/T, Biologics Inc., Cary, NC, USA). Crude extracts thus obtained were centrifuged at 10,000×
g for 10 min at 4 °C. They were resuspended in 1 mL of 0.2% Sarkosyl and incubated overnight with agitation at room temperature (RT). These samples were centrifuged at 10,000×
g for 1 h at 4 °C. Pellets were then washed twice with 10 mM Tris-HCl (pH 7.4) and recovered by centrifugation at 10,000×
g for 10 min at 4 °C. Lastly, OMP fractions were resuspended in buffer containing 10 mM Tris-HCl (pH 7.4) and 8 M urea. Proteins were stored at −20 °C until use.
We adapted the method described by Yi and Hackett [
28] in order to isolate and purify lipopolysaccharide (LPS). Bacteria grown on TSA were suspended in 300 µL of Trizol reagent and incubated for 15 min at RT. Next, 100 µL of chloroform was added. Samples were mixed thoroughly and incubated for another 10 min, maintaining RT. Centrifugation of the water phase at 12,000×
g for 10 min was followed by treatment with 500 µL of 0.375 M MgCl
2 in cold ethanol (−20 °C). Pellets were recovered by a final centrifugation step at 12,000×
g for 15 min at 4 °C. Discontinuous SDS-PAGE electrophoresis was performed to separate membrane antigens. We used 5% and 15% polyacrylamide stacking/solving gels which were subsequently stained with Coomassie blue dye to assess proteins; silver stains were used for LPS analysis, following a protocol described elsewhere [
29]. We ran semi-dry Western blots (Bio-Rad, Dreieich, Germany), revealing signals with a polyclonal antibody obtained from rabbit serum after immunization with a
A. salmonicida VapA+ isolate and the corresponding HRP-conjugated secondary antibody.
2.6. Virulence In Vivo Testing
Salmo salar with a mean weight of 28 g were intraperitoneally challenged with either one of three infectious doses of representative
A. piscicola field isolates (~10
6, ~10
7, or ~10
8 cfu/fish of A2 or S2, respectively; for exact counts, see
Table S2). Bacterial suspensions were freshly prepared as described in
Section 2.2, including a wash step with saline before final resuspension. The assay was conducted in a single tank containing 500 L of filtered, UV-treated freshwater. We worked with a total of 140 fish: 20 healthy control fish (14.3%) which received an intraperitoneal injection of 0.1 mL saline and 120 fish (85.7%) to be injected with bacterial inoculum. All experimentally challenged and control fish were maintained in that same tank. The 120 fish to be challenged were divided into six subgroups and marked accordingly by Visible Implant Elastomer (VIE tagging), and each subgroup was treated with
A. piscicola A2 or S2 strains at a determined infectious dose. Fish biomass density was kept below 30 kg/m
3. The experiment was carried out at 10.7 ± 0.1 °C and 4.0 ppt salinity, and it was ended 30 days post-inoculation (dpi). Mortality was registered daily, and pooled samples of anterior kidney, liver, and spleen were collected from each deceased fish. These samples were stored in tubes with RNAlater (Life Technologies, Carlsbad, CA, USA) and kept at −80 °C until further use. At 30 dpi, all survivor fish, including controls, were sacrificed to obtain internal organ samples for PCR analysis. Pathogen re-isolation was not performed during the challenge; infection status was assessed by qPCR on pooled internal-organ samples.
2.7. Serum Immune Response
In preparation for a field study intended to shed light on the immune response to A. piscicola, we collected serum samples from S. salar on seven fish farms in Southern Chile where a commercial pentavalent vaccine against atypical furunculosis, Infectious Pancreatic Necrosis (IPN), Infectious Salmon Anemia (ISA), Salmonid Rickettsial Syndrome (SRS), and vibriosis caused by Vibrio ordalii was to be applied. Serum samples were taken from five fish per farm at different time points immediately before and after vaccination: First, unvaccinated S. salar pre-smolts were euthanized applying an overdose of benzocaine. These fish were bled to obtain serum samples, which were subsequently centrifuged at 1000 rpm for 3 min. Further fish were sampled during the freshwater stage at ~300 and ~600 thermal units (TU) time points post vaccination. Immune responses were further monitored post sea transfer with additional samples being collected at ~900 and ~1200 TU post vaccination. All serum samples were stored at −80 °C until further analysis.
To assess immune responses, 96-well microplates were coated with 5 µg of bacterial protein purified from the representative isolates A. piscicola S2 (vapA−) or A. salmonicida A4 (vapA+). Enzyme-linked immunosorbent assays (ELISA) were performed in these pretreated microplates, using 5% skim milk as the blocking agent and PBS containing 0.05% Tween (PBS-T) as the washing buffer. Fish serum was diluted 1:100 in skim milk and added to the wells. Loaded microplates were incubated for 1 h at RT, washed with PBS-T, incubated with a monoclonal HRP-conjugated antibody anti-salmon IgM, and washed again. Signals were developed using the chromogenic substrate 3,3′,5,5′-tetramethylbenzidine and measured as absorbance at 450 nm with a spectrophotometer.
2.8. Statistical Analysis and Plots
Plots were generated using the ggplot2 package on R (version 4.0.5). The Kaplan–Meier method was used to analyze cumulative mortality percentages in the in vivo assay, and the differences were evaluated using log-rank. Statistical tests were also performed with R.
4. Discussion
This study provides evidence that
A. piscicola is associated with recent furunculosis-like outbreaks in Chile and that isolates can cause mortality in Atlantic salmon under experimental conditions, a fact that adds complexity to the epidemiological scenario of the disease. The detection of
A. piscicola in Chile raises concerns regarding its potential role as a pathogen in aquatic systems. Recent reports from other countries have associated
A. piscicola with infections in
S. salar, causing symptoms such as hemorrhagic septicemia and skin ulcerations [
8]. The bacterium’s adaptation to the prevailing environmental conditions in Chile highlights its resilience and potential for dissemination. In fact, our evidence suggests that
A. piscicola isolates, as well as
A. salmonicida vapA-absent variants, are more versatile than the long-known, highly virulent
A. salmonicida vapA+ strain: They are well able to replicate in a wide range of temperature and salinities. Our findings underscore the need for surveillance programs to monitor the presence of
A. piscicola in aquaculture facilities and natural water bodies. Current diagnostic strategies are only focused on the detection of
A. salmonicida, based on the assumption that only one species (and its variants) is responsible for furunculosis in reared salmon in Chile. However, this work expands our knowledge, highlights the virulence potential of
A. piscicola strains, and thus points out the need to update diagnostic procedures. Ongoing research is addressing this issue; we are developing specific molecular assays to better distinguish
A. salmonicida variants as well as non-
salmonicida isolates such as
A. piscicola.
A striking finding of this study is the extensive repertoire of VF identified in
A. piscicola genomes. With approximately 220 predicted virulence genes, these isolates possess a pathogenic arsenal roughly one-third larger than the reference
A. salmonicida subsp.
oncorhynchi A-9. Notably, T3SS genes have been identified in all
A. piscicola isolates, whereas they are characteristically absent from the
A. salmonicida subsp.
onchorynchi A-9 reference genome. At the same time, this observation should not be generalized to all representatives of the
A. salmonicida species, because T3SS determinants have been reported in other
A. salmonicida genomes [
32], consistent with our observation in the genome of Chilean
A. salmonicida vapA+ strain A4. Therefore, the present comparison should be interpreted as reference-genome-based rather than species-wide, and a more comprehensive comparison against multiple typical and atypical
A. salmonicida genomes will be required to determine whether the differences observed here are species-level or lineage-specific. Virulence is a complex biological phenomenon that is not determined solely by the number of VF, and hence the total number of VF identified in
A. piscicola genomes cannot support the notion that
A. piscicola isolates described here are more virulent than their
A. salmonicida counterparts.
Unlike classical A. salmonicida, the A. piscicola isolates described herein were mesophilic, vapA-negative, and antigenically distinct in their OMP/LPS profiles. Furthermore, structural and genomic differences in the outer membrane composition provide a plausible explanation for the observed variations in virulence among the A. piscicola strains themselves. The highly virulent A2 isolate (causing >80% mortality) was shown to possess a complete LPS structure featuring both the lipid A-core and a high-molecular-weight O-antigen. Conversely, the less virulent S2 isolate exhibited a truncated LPS profile lacking the polymeric O-antigen fraction. This phenotypic divergence is strongly supported by our genomic analysis, which revealed a unique ~10 kb O-antigen biosynthesis gene cluster in the A2 genome that is absent in the A. piscicola type strain LMG 24783. The presence of this novel O-antigen likely enhances serum resistance and host colonization, making A2 significantly more lethal than the S2 variant. This feature may also explain the early mortality observed in fish inoculated with what appears to be a bacterium carrying a more biologically active LPS, although the precise contribution of this locus to virulence require functional validation.
These two strains also differ with regard to antimicrobial susceptibility: While the majority of the mesophilic A. piscicola isolates were confirmed to be susceptible to the primary antibiotics used in Chilean salmon farming, the A2 strain demonstrated pronounced resistance to oxytetracycline (MIC > 64 µg/mL). Our bioinformatic analysis successfully linked this phenotype to the presence of the tet(E) MFS efflux pump gene. The circulation of such resistance markers within a virulent, environmentally versatile strain poses a severe risk to standard therapeutic interventions in freshwater aquaculture, emphasizing the critical need for continuous AMR monitoring.
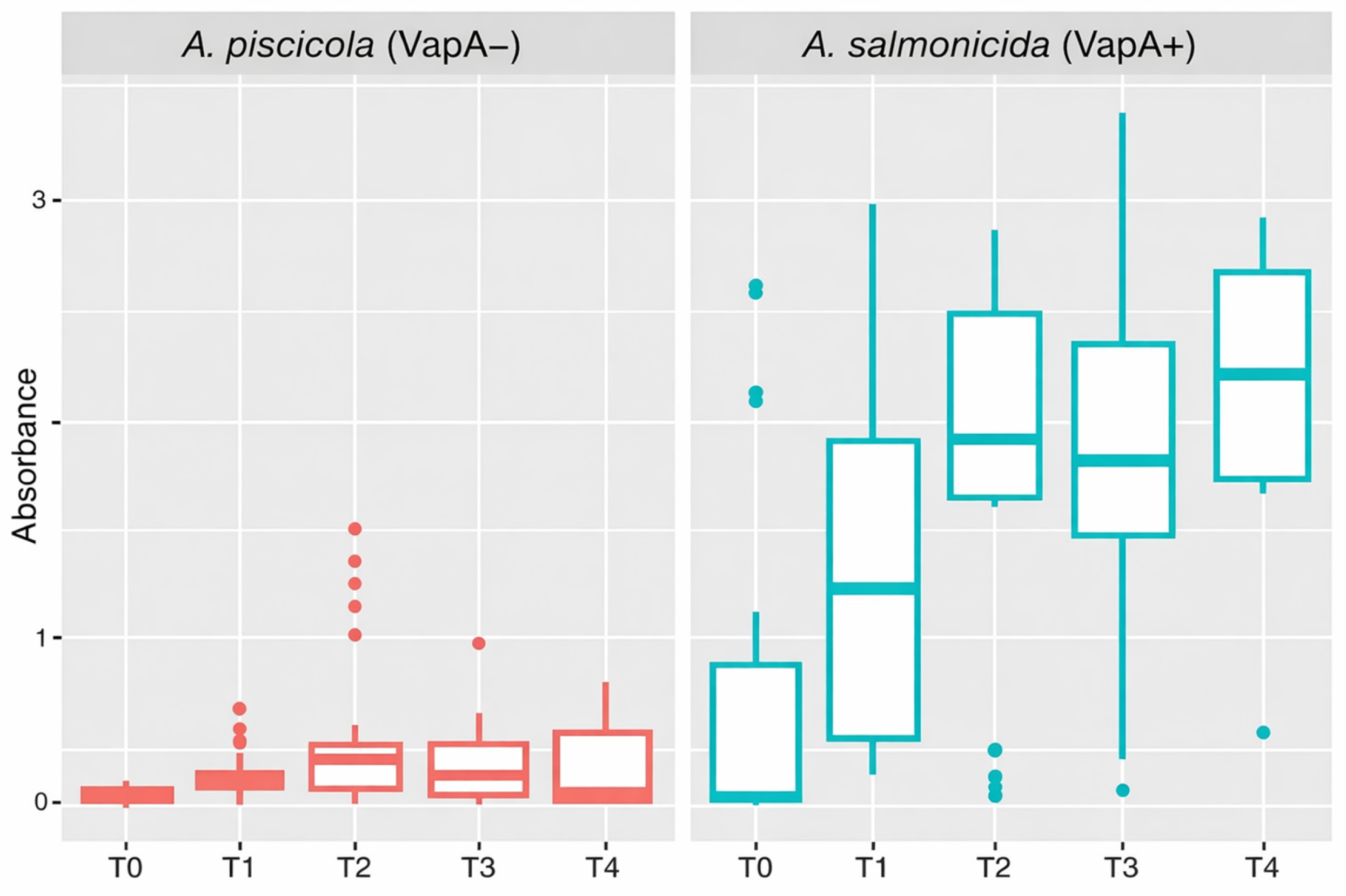
From an immunological and preventative standpoint, our results reveal a significant vulnerability in current prophylactic strategies. The ELISA data clearly demonstrate that the commercial pentavalent vaccine, which relies on homologous
A. salmonicida antigens (such as the A-layer protein VapA), elicits a robust IgM response against classical
A. salmonicida but fails to trigger a comparable cross-reactive response against
A. piscicola in the field. This immune evasion is almost certainly due to the heterologous nature of the outer membrane of
A. piscicola, which lacks the VapA protein and, in the case of the A2 isolate, expresses a highly divergent, unrecognized O-antigen, as confirmed by our Western blot analysis (
Figure 4D). Consequently, vaccinated Atlantic salmon remain immunologically naïve to
A. piscicola, facilitating the outbreaks observed in recent years.
In conclusion, the emergence of virulent, mesophilic A. piscicola strains capable of circumventing current commercial vaccines represents a paradigm shift in the management of furunculosis-like diseases in Chile. Future efforts must prioritize the development of multivalent or autogenous vaccines that incorporate A. piscicola-specific antigens, alongside the implementation of multiplexed diagnostic tools capable of accurately differentiating between these closely related, yet functionally distinct, Aeromonas species.