Antibody Recognition of Highly and Low-Pathogenic A/H5Nx Influenza Viruses in Sera of Mexican Donors
Abstract
1. Introduction
2. Materials and Methods
2.1. Serum Samples
2.2. Hemagglutination Inhibition (HI) Test
2.3. Viral Propagation
2.4. In Silico Analysis of the H5 Proteins
2.5. Statistical Analysis
3. Results
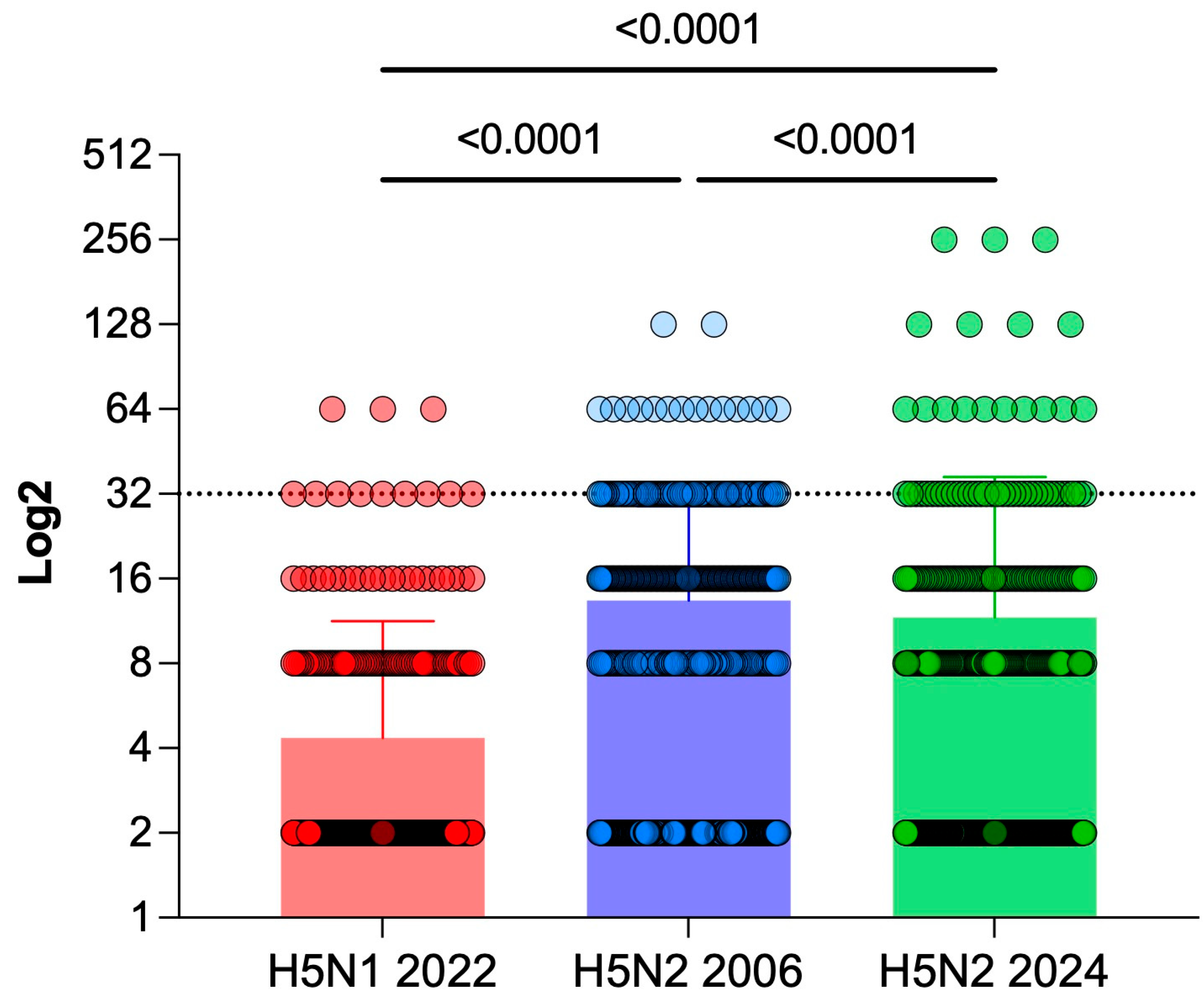
3.1. Serological Results for H5Nx Hemagglutination Inhibition Test
3.2. Age and Sex Association with Antibodies Against H5Nx Viruses
3.3. Predicted Epitopes Recognized by Antibodies in Adults’ Sera
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| HI | Hemagglutinin inhibition |
| RBS | Receptor Binding Site |
References
- ICTV. ICTV Taxonomy Browser. Taxon Details. Available online: https://ictv.global/taxonomy/taxondetails?taxnode_id=202403956&ictv_id=ICTV19710226 (accessed on 3 March 2026).
- Zerbini, F.M.; Simmonds, P.; Adriaenssens Evelien, M.; Lefkowitz Elliot, J.; Oksanen Hanna, M.; Alfenas-Zerbini, P.; Aylward Frank, O.; Freitas-Astúa, J.; Hughes Holly, R.; Łobocka, M.; et al. Virus species names have been standardized; virus names remain unchanged. mSphere 2025, 10, e00020–e00025. [Google Scholar] [CrossRef] [PubMed]
- Peacock, T.P.; Moncla, L.; Dudas, G.; VanInsberghe, D.; Sukhova, K.; Lloyd-Smith, J.O.; Worobey, M.; Lowen, A.C.; Nelson, M.I. The global H5N1 influenza panzootic in mammals. Nature 2025, 637, 304–313. [Google Scholar] [CrossRef]
- World Animal Health Information System Interface. Mexico-High Pathogenicity Avian Influenza Viruses (Poultry) (Inf. with)-Follow up Report 6 [FINAL]. Available online: https://wahis.woah.org/#/in-review/4762?reportId=163107&fromPage=event-dashboard-url (accessed on 20 May 2024).
- Burrough, E.; Magstadt, D.; Petersen, B.; Timmermans, S.; Gauger, P.; Zhang, J.; Siepker, C.; Mainenti, M.; Li, G.; Thompson, A.; et al. Highly Pathogenic Avian Influenza A(H5N1) Clade 2.3.4.4b Virus Infection in Domestic Dairy Cattle and Cats, United States, 2024. Emerg. Infect. Dis. J. 2024, 30, 1335. [Google Scholar] [CrossRef]
- Nuñez, I.A.; Ross, T.M. A review of H5Nx avian influenza viruses. Ther. Adv. Vaccines Immunother. 2019, 7, 2515135518821625. [Google Scholar] [CrossRef] [PubMed]
- Alexakis, L.; Fusaro, A.; Kuiken, T.; Mirinavičiūtė, G.; Ståhl, K.; Staubach, C.; Svartström, O.; Terregino, C.; Willgert, K.; Delacourt, R.; et al. Avian influenza overview March–June 2024. EFSA J. 2024, 22, e8930. [Google Scholar] [CrossRef] [PubMed]
- Vázquez-Pérez, J.A.; Wong-Arámbula, C.; Solís-Hernández, M.; Becerril-Vargas, E.; Barrera-Badillo, G.; Ahumada-Topete, V.H.; Avila-Rios, S.; Pérez-Padilla, R.; Mejía-Nepomuceno, F.; Mendoza-Ramírez, E.; et al. Mexico’s Laboratory-Confirmed Human Case of Infection with the Influenza A(H5N2) Virus. Viruses 2025, 17, 205. [Google Scholar] [CrossRef]
- Apostolopoulos, V.; Sah, R.; Mehta, R.; Diaz, B.; Rodriguez-Morales, A.J. First confirmed human case of H5N2 virus infection in Mexico: An emerging zoonotic concern. Le Infez. Med. 2024, 32, 413–416. [Google Scholar] [CrossRef]
- Vázquez-Pérez, J.A.; Becerril-Vargas, E.; Ramírez-González, J.E.; Solís-Hernández, M.; Davis, C.T.; Garciadiego-Fossas, P.; Villanueva-Reza, M.; Chávez-Morales, H.H.; Mendoza-Ramírez, E.; Olivares-Flores, C.D.; et al. Emergence of a novel reassorted high pathogenicity avian influenza A(H5N2) virus associated with severe pneumonia in a young adult. medRxiv 2025. medRxiv:2025.11.21.25340167. [Google Scholar] [CrossRef]
- Broberg, E.K.; Svartström, O.; Riess, M.; Kraus, A.; Vukovikj, M.; Melidou, A.; Members of the European Reference Laboratory Network for Human Influenza (ERLI-Net). Co-circulation of seasonal influenza A(H1N1)pdm09, A(H3N2) and B/Victoria lineage viruses with further genetic diversification, EU/EEA, 2022/23 influenza season. Euro Surveill. 2024, 29, 2400020. [Google Scholar] [CrossRef]
- World Health Organization. Influenza (Seasonal). Available online: https://www.who.int/news-room/fact-sheets/detail/influenza-(seasonal) (accessed on 3 September 2025).
- Restori, K.H.; Weaver, V.; Patel, D.R.; Merrbach, G.A.; Septer, K.M.; Field, C.J.; Bernabe, M.J.; Kronthal, E.M.; Minns, A.; Lindner, S.E.; et al. Preexisting immunity to the 2009 pandemic H1N1 virus reduces susceptibility to H5N1 infection and disease in ferrets. Sci. Transl. Med. 2025, 17, eadw4856. [Google Scholar] [CrossRef]
- Garretson, T.A.; Liu, J.; Li, S.H.; Scher, G.; Santos, J.J.S.; Hogan, G.; Vieira, M.C.; Furey, C.; Atkinson, R.K.; Ye, N.; et al. Immune history shapes human antibody responses to H5N1 influenza viruses. Nat. Med. 2025, 31, 1454–1458. [Google Scholar] [CrossRef]
- Hernández, J.; Dehesa-Canseco, F.; Vázquez-López, A.B.; Reséndiz-Sandoval, M.; Caire-Juvera, G.; Solís-Hernández, M.; Valenzuela, O.; Gómez-Gil, B.; Mata-Haro, V. Neutralization of Omicron BA.1, BA.5.1.6, BQ.1.3 and XBB1.1 induced by heterologous vaccination Ad5-nCoV and mRNA-1273. Signal Transduct. Target. Ther. 2023, 8, 174. [Google Scholar] [CrossRef]
- Melgoza-González, E.A.; Hinojosa-Trujillo, D.; Reséndiz-Sandoval, M.; Mata-Haro, V.; Hernández-Valenzuela, S.; García-Vega, M.; Bravo-Parra, M.; Arvizu-Flores, A.A.; Valenzuela, O.; Velázquez, E.; et al. Analysis of IgG, IgA and IgM antibodies against SARS-CoV-2 spike protein S1 in convalescent and vaccinated patients with the Pfizer-BioNTech and CanSinoBio vaccines. Transbound. Emerg. Dis. 2022, 69, e734–e745. [Google Scholar] [CrossRef]
- Badillo, B.A.M.; Hernández García, D.L.; García, R.A.M.; Pineda, G.O.; Ramiro, C.J.A.; Castillo, J.C.; Hernández, M.S.; López, R.N.; López, A.G. H5N1 highly pathogenic avian influenza vaccination: Seroresponse of mexican poultry in the 2022–2024. Vaccine X 2025, 26, 100709. [Google Scholar] [CrossRef]
- World Health Organization. WHO Manual on Animal Influenza Diagnosis and Surveillance; World Health Organization: Geneva, Switzerland, 2002. [Google Scholar]
- World Organisation for Animal Health. Avian Influenza (Infection with Avian Influenza Viruses). Available online: https://www.woah.org/es/que-hacemos/normas/codigos-y-manuales/#chapter/?rid=335&volume_no=3&ismanual=true&language=104&standard_type=6&animal_type=7 (accessed on 3 March 2026).
- Possas, C.; Marques, E.T.A.; Oliveira, A.; Schumacher, S.; Siqueira, M.M.; McCauley, J.; Antunes, A.; Homma, A. Highly pathogenic avian influenza: Pandemic preparedness for a scenario of high lethality with no vaccines. Front. Public Health 2025, 13, 1613869. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.; Yang, J.; Jiao, W.; Li, X.; Iqbal, M.; Liao, M.; Dai, M. Clade 2.3.4.4b highly pathogenic avian influenza H5N1 viruses: Knowns, unknowns, and challenges. J. Virol. 2025, 99, e00424–e00425. [Google Scholar] [CrossRef] [PubMed]
- Daniel, K.; Ullrich, L.; Ruchnewitz, D.; Meijers, M.; Halwe, N.J.; Wild, U.; Eberhardt, J.; Schön, J.; Stumpf, R.; Schlotz, M.; et al. Pre-existing neutralizing antibodies against cattle-transmitted influenza A virus H5N1 are detectable in unexposed individuals. Immunity 2026, 59, 494–508.e8. [Google Scholar] [CrossRef]
- Heeringa, M.; Leav, B.; Smolenov, I.; Palladino, G.; Isakov, L.; Matassa, V. Comparability of Titers of Antibodies against Seasonal Influenza Virus Strains as Determined by Hemagglutination Inhibition and Microneutralization Assays. J. Clin. Microbiol. 2020, 58, 10–1128. [Google Scholar] [CrossRef] [PubMed]
- Tsai, C.; Caillet, C.; Hu, H.; Zhou, F.; Ding, H.; Zhang, G.; Zhou, B.; Wang, S.; Lu, S.; Buchy, P.; et al. Measurement of neutralizing antibody responses against H5N1 clades in immunized mice and ferrets using pseudotypes expressing influenza hemagglutinin and neuraminidase. Vaccine 2009, 27, 6777–6790. [Google Scholar] [CrossRef]
- Sanz-Muñoz, I.; Sánchez-Martínez, J.; Rodríguez-Crespo, C.; Concha-Santos Corina, S.; Hernández, M.; Rojo-Rello, S.; Domínguez-Gil, M.; Mostafa, A.; Martinez-Sobrido, L.; Eiros Jose, M.; et al. Are we serologically prepared against an avian influenza pandemic and could seasonal flu vaccines help us? mBio 2024, 16, e03721–e03724. [Google Scholar] [CrossRef]
- de Bruin, A.C.M.; Funk, M.; Spronken, M.I.; Gultyaev, A.P.; Fouchier, R.A.M.; Richard, M. Hemagglutinin Subtype Specificity and Mechanisms of Highly Pathogenic Avian Influenza Virus Genesis. Viruses 2022, 14, 1566. [Google Scholar] [CrossRef]
- Hulse Diane, J.; Webster Robert, G.; Russell Rupert, J.; Perez Daniel, R. Molecular Determinants within the Surface Proteins Involved in the Pathogenicity of H5N1 Influenza Viruses in Chickens. J. Virol. 2004, 78, 9954–9964. [Google Scholar] [CrossRef]
- Zheng, Z.; Teo, S.H.C.; Arularasu, S.C.; Liu, Z.; Mohd-Ismail, N.K.; Mok, C.K.; Ong, C.B.; Chu, J.J.-h.; Tan, Y.-J. Contribution of Fc-dependent cell-mediated activity of a vestigial esterase-targeting antibody against H5N6 virus infection. Emerg. Microbes Infect. 2020, 9, 95–110. [Google Scholar] [CrossRef]
- Motsoeneng, B.M.; Dhar, N.; Nunes, M.C.; Krammer, F.; Madhi, S.A.; Moore, P.L.; Richardson, S.I. Hemagglutinin Stalk-Specific Fc-Mediated Functions Are Associated with Protection Against Influenza Illness After Seasonal Influenza Vaccination. J. Infect. Dis. 2024, 230, 1329–1336. [Google Scholar] [CrossRef]
- Sun, X.; Ma, H.; Wang, X.; Bao, Z.; Tang, S.; Yi, C.; Sun, B. Broadly neutralizing antibodies to combat influenza virus infection. Antivir. Res. 2024, 221, 105785. [Google Scholar] [CrossRef] [PubMed]
- EFSA Panel on Animal Health and Animal Welfare (AHAW); ECDC; Álvarez, J.; Boklund, A.; Dippel, S.; Dórea, F.; Figuerola, J.; Herskin, M.; Michel, V.; Chueca, M.M.; et al. Preparedness, prevention and control related to zoonotic avian influenza. EFSA J. 2025, 23, e9191. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Wang, W.; Wang, Y.; Lai, S.; Yang, J.; Cowling, B.J.; Horby, P.W.; Uyeki, T.M.; Yu, H. Serological evidence of human infections with highly pathogenic avian influenza A(H5N1) virus: A systematic review and meta-analysis. BMC Med. 2020, 18, 377. [Google Scholar] [CrossRef] [PubMed]


| Viral Strain | Sex | Age Group | |||||
|---|---|---|---|---|---|---|---|
| Male | Female | 18–30 | 31–40 | 41–50 | 51–60 | 61–76 | |
| H5N2 2006 | 22.4% (48/214) | 16% (42/262) | 15.2% (12/79) | 12.6% (20/159) | 16.1% (24/149) | 34.2% (27/79) | 70% (7/10) |
| Highest titer | 128 | 128 | 128 | 64 | 128 | 64 | 64 |
| H5N2 2024 | 12.6% (27/214) | 9.2% (24/262) | 3.8% (3/79) | 8.8% (14/159) | 13.4% (20/149) | 15.2% (12/79) | 20% (2/10) |
| Highest titer | 256 | 128 | 256 | 128 | 256 | 128 | 64 |
| H5N1 2022 | 1.9% (4/214) | 3% (8/262) | 6.3% (5/79) | 1.9% (3/159) | 2% (3/149) | 1.3% (1/79) | 0% (0/10) |
| Highest titer | 64 | 64 | 64 | 64 | 64 | 32 | 16 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Cordero-Ortiz, M.; Solís-Hernández, M.; Cayetano-Mondragón, M.; Guzmán, N.C.; Valenzuela, O.; Mata-Haro, V.; Giménez-Lirola, L.G.; Hernández, J. Antibody Recognition of Highly and Low-Pathogenic A/H5Nx Influenza Viruses in Sera of Mexican Donors. Pathogens 2026, 15, 352. https://doi.org/10.3390/pathogens15040352
Cordero-Ortiz M, Solís-Hernández M, Cayetano-Mondragón M, Guzmán NC, Valenzuela O, Mata-Haro V, Giménez-Lirola LG, Hernández J. Antibody Recognition of Highly and Low-Pathogenic A/H5Nx Influenza Viruses in Sera of Mexican Donors. Pathogens. 2026; 15(4):352. https://doi.org/10.3390/pathogens15040352
Chicago/Turabian StyleCordero-Ortiz, Maritza, Mario Solís-Hernández, Marlen Cayetano-Mondragón, Nadia Carrillo Guzmán, Olivia Valenzuela, Verónica Mata-Haro, Luis G. Giménez-Lirola, and Jesús Hernández. 2026. "Antibody Recognition of Highly and Low-Pathogenic A/H5Nx Influenza Viruses in Sera of Mexican Donors" Pathogens 15, no. 4: 352. https://doi.org/10.3390/pathogens15040352
APA StyleCordero-Ortiz, M., Solís-Hernández, M., Cayetano-Mondragón, M., Guzmán, N. C., Valenzuela, O., Mata-Haro, V., Giménez-Lirola, L. G., & Hernández, J. (2026). Antibody Recognition of Highly and Low-Pathogenic A/H5Nx Influenza Viruses in Sera of Mexican Donors. Pathogens, 15(4), 352. https://doi.org/10.3390/pathogens15040352








