Acanthamoeba castellanii: Non-Steroidal Anti-Inflammatory Drugs Affect Adhesion, Motility, and Encystment, Suggesting a Link with a gp63-like Protein Candidate
Abstract
1. Introduction
2. Materials and Methods
2.1. Drugs
2.2. Amoebic Cultivation Conditions
2.3. Growth Curves
2.4. Adhesion Assay in the Presence of Cyclooxygenase Inhibitors
2.5. Evaluation of Cellular Damage (Cytopathic Effect)
2.6. Brain Tissue Discontinuous Micropatterns (BTDM)
2.7. Monitoring of Trophozoite Migration
2.8. Confocal Microscopy of gp63-like Protein and F-Actin
2.9. Detection of gp63-like Protein by Western Blot
2.10. Encystment Assays and Enrichment of Encysted Forms
2.11. Domain Architecture Analysis of Leishmanolysin (gp63-like Protein)
2.12. Three-Dimensional Modeling of Leishmanolysin (gp63-like Protein) and NSAID Ligands
2.13. Molecular Docking Between Leishmanolysin (gp63-like Protein) and NSAIDs
2.14. Toxicological and Pharmacological Predictions
3. Results
3.1. Non-Steroidal Anti-Inflammatory Drugs (NSAIDs) Do Not Affect the Growth of A. castellanii Trophozoites but Selectively Impair Adhesion
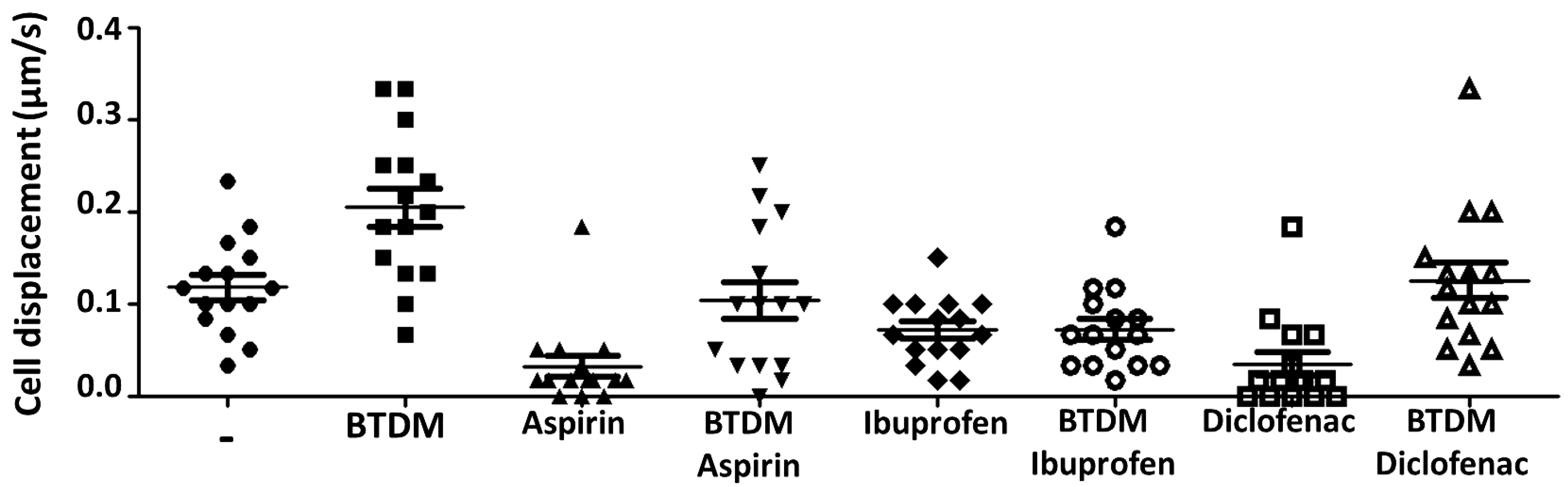
3.2. Brain Tissue Discontinuous Micropatterns (BTDM) Enhance A. castellanii Motility and gp63/Leishmanolysin-like Protein–Actin Association
3.3. NSAIDs Affect Actin Filament Organization in MDCK Cells During A. castellanii–MDCK Cell Interactions
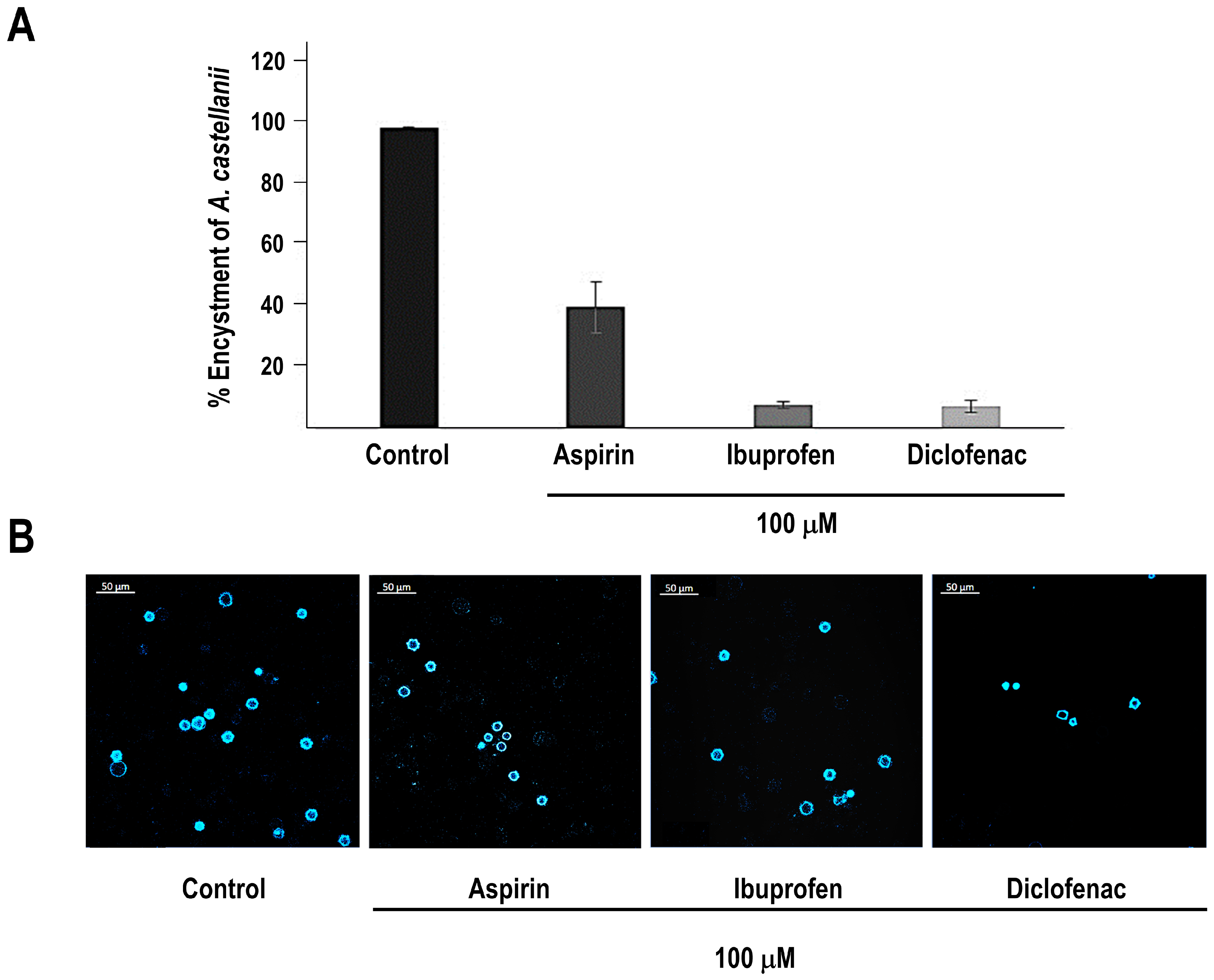
3.4. Cyclooxygenase Inhibitors Differentially Modulate Encystment of A. castellanii
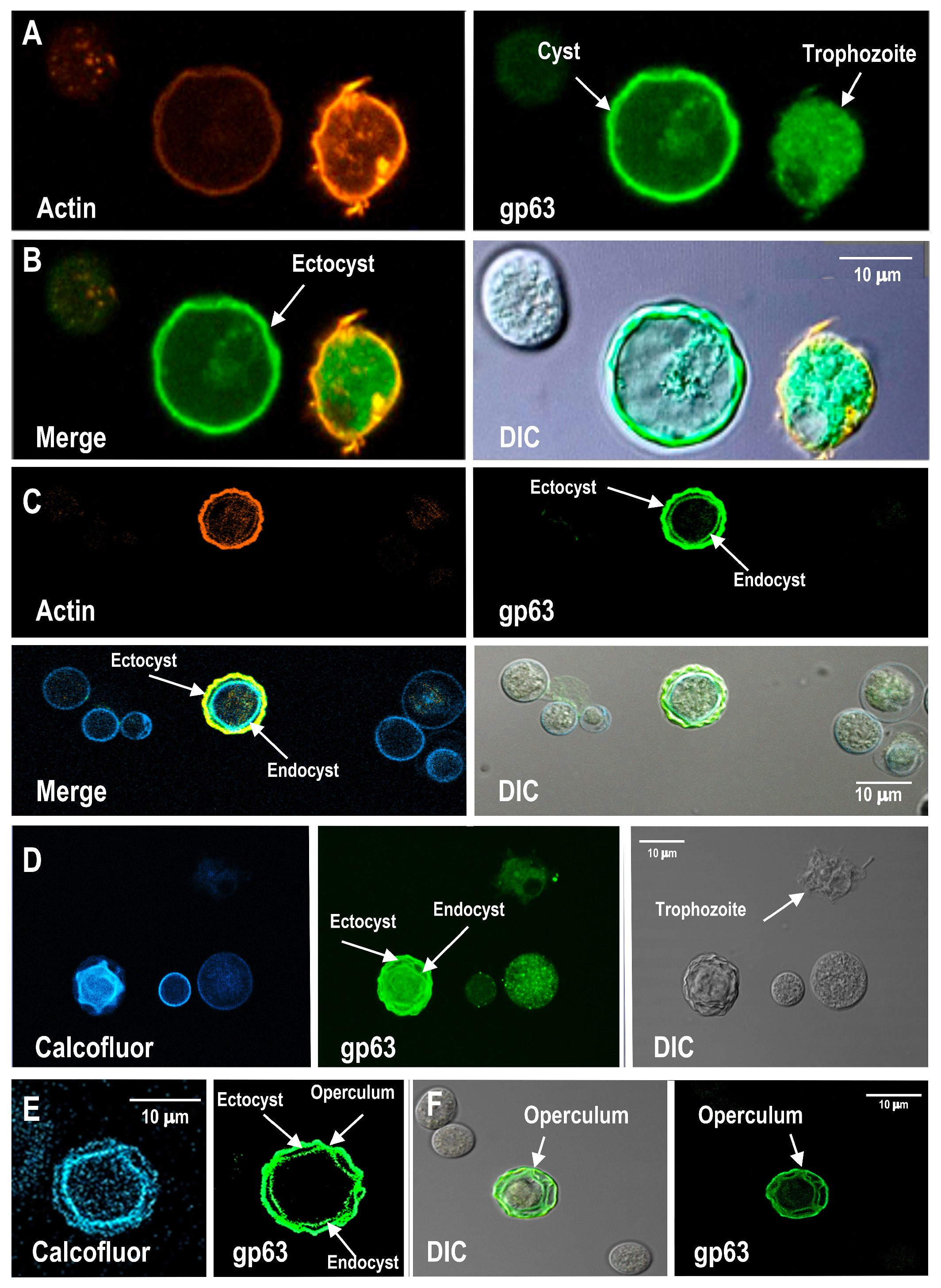
3.5. Distribution of gp63-like Protein in Cyst Wall Structures During Encystment
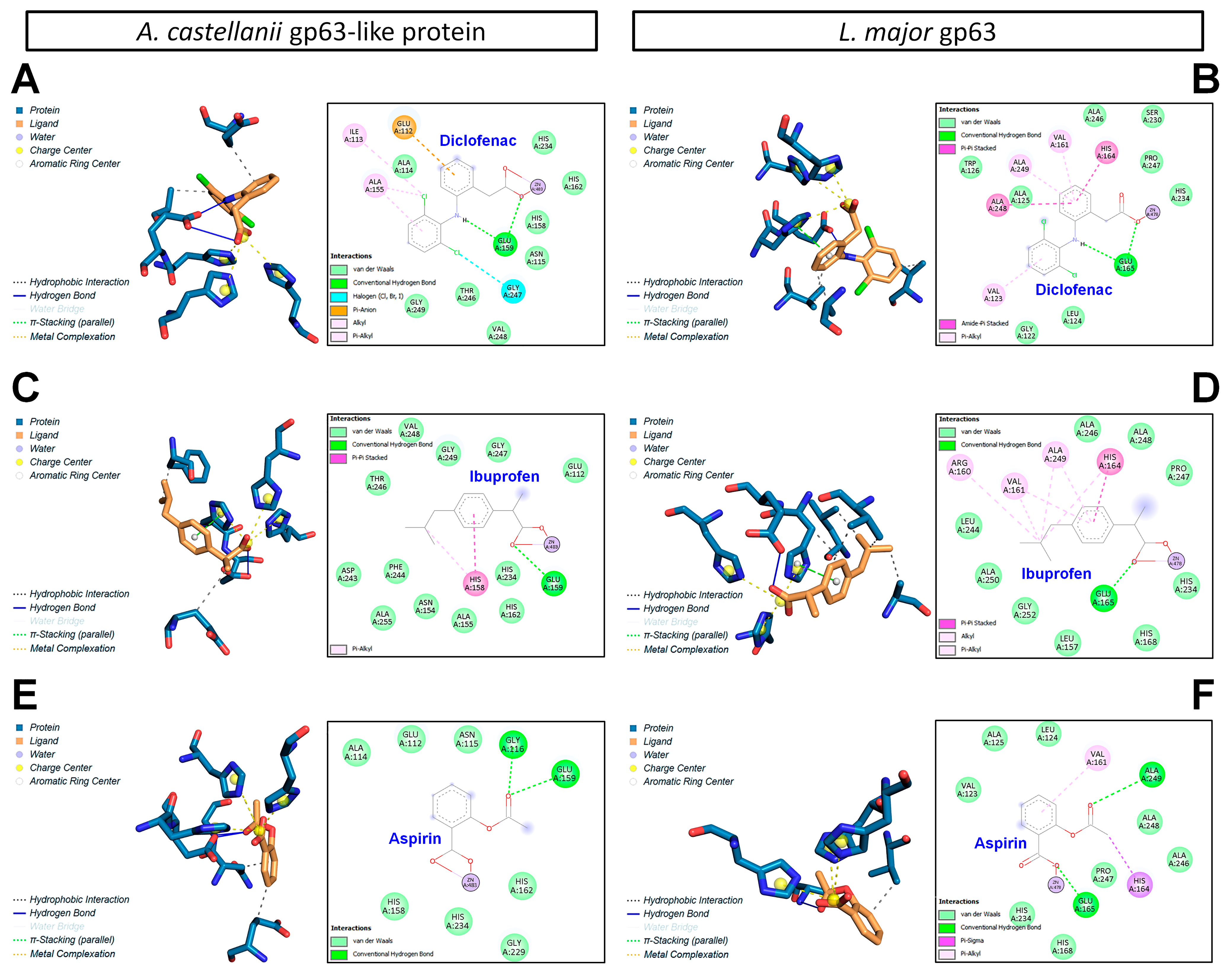
3.6. Structural Comparison of gp63-like Proteins from A. castellanii and L. major
3.7. Molecular Docking Analysis of gp63–NSAID Interactions
3.8. In Silico Assessment of Toxicity and Pharmacological Profiles of NSAIDs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AC50 | Activating concentration 50% |
| BMD | Benchmark dose |
| BTDM | Brain tissue discontinuous micropatterns |
| COX | Cyclooxygenase |
| CSTA | Chemical similarity of toxic analogs |
| DAPI | 4′,6-diamidino-2-fenilindol |
| DIC | Differential interference contrast |
| DMEM | Dulbecco’s modified Eagle medium |
| DMSO | Dimethyl sulfoxide |
| FBS | Fetal bovine serum |
| hERG | Human ether-à-go-go-related gene |
| IC50 | Inhibitory concentration 50% |
| Ki | Inhibition constant |
| MDCK | Madin–Darby canine kidney |
| NSAIDs | Non-steroidal anti-inflammatory drugs |
| pI | Isoelectric point |
| PPARγ | Peroxisome proliferator-activated receptor gamma |
| QED | Quantitative estimation of drug-likeness |
| RMSD | Root-mean-square deviation |
References
- Mungroo, M.R.; Khan, N.A.; Maciver, S.; Siddiqui, R. Opportunistic Free-Living Amoebal Pathogens. Pathog. Glob. Health 2022, 116, 70–84. [Google Scholar] [CrossRef]
- Wang, Y.; Jiang, L.; Zhao, Y.; Ju, X.; Wang, L.; Jin, L.; Fine, R.D.; Li, M. Biological Characteristics and Pathogenicity of Acanthamoeba. Front. Microbiol. 2023, 14, 1147077. [Google Scholar] [CrossRef]
- Haston, J.C.; O’Laughlin, K.; Matteson, K.; Roy, S.; Qvarnstrom, Y.; Ali, I.K.M.; Cope, J.R. The Epidemiology and Clinical Features of Non-Keratitis Acanthamoeba Infections in the United States, 1956–2020. Open Forum Infect. Dis. 2023, 10, ofac682. [Google Scholar] [CrossRef]
- Choaji, M.; Samba-Louaka, A.; Fechtali-Moute, Z.; Aucher, W.; Pomel, S. Encystment and Excystment Processes in Acanthamoeba Castellanii: An Emphasis on Cellulose Involvement. Pathogens 2025, 14, 268. [Google Scholar] [CrossRef]
- Fadaee, M.; Lahouty, M.; Ramzi, F.; Alizadeh, F.; Roshanfar, S.; Kazemi, T.; Spotin, A.; Hatam-Nahavandi, K.; Ahmadpour, E. Extracellular Vesicles in Intestinal Protozoa: Hidden Mediators of Host-Parasite Communication. Gut Pathog. 2025, 17, 69. [Google Scholar] [CrossRef]
- Daugschies, A.; Joachim, A. Eicosanoids in Parasites and Parasitic Infections. Adv. Parasitol. 2000, 46, 181–240. [Google Scholar]
- Dey, I.; Keller, K.; Belley, A.; Chadee, K. Identification and Characterization of a Cyclooxygenase-like Enzyme from Entamoeba histolytica. Proc. Natl. Acad. Sci. USA 2003, 100, 13561–13566. [Google Scholar] [CrossRef]
- Noverr, M.C.; Erb-Downward, J.R.; Huffnagle, G.B. Production of Eicosanoids and Other Oxylipins by Pathogenic Eukaryotic Microbes. Clin. Microbiol. Rev. 2003, 16, 517–533. [Google Scholar] [CrossRef]
- Vallochi, A.L.; Teixeira, L.; Oliveira, K.d.S.; Maya-Monteiro, C.M.; Bozza, P.T. Lipid Droplet, a Key Player in Host-Parasite Interactions. Front. Immunol. 2018, 9, 1022. [Google Scholar] [CrossRef]
- Alves-Ferreira, E.V.C.; Ferreira, T.R.; Walrad, P.; Kaye, P.M.; Cruz, A.K. Leishmania Braziliensis Prostaglandin F2α Synthase Impacts Host Infection. Parasit. Vectors 2020, 13, 9. [Google Scholar] [CrossRef]
- Parreira de Aquino, G.; Mendes Gomes, M.A.; Köpke Salinas, R.; Laranjeira-Silva, M.F. Lipid and Fatty Acid Metabolism in Trypanosomatids. Microbial Cell 2021, 8, 262–275. [Google Scholar] [CrossRef]
- Leroux, M.; Luquain-Costaz, C.; Lawton, P.; Azzouz-Maache, S.; Delton, I. Fatty Acid Composition and Metabolism in Leishmania Parasite Species: Potential Biomarkers or Drug Targets for Leishmaniasis? Int. J. Mol. Sci. 2023, 24, 4702. [Google Scholar] [CrossRef]
- Estrada-Figueroa, L.A.; Díaz-Gandarilla, J.A.; Hernández-Ramírez, V.I.; Arrieta-González, M.M.; Osorio-Trujillo, C.; Rosales-Encina, J.L.; Toledo-Leyva, A.; Talamás-Rohana, P. Leishmania Mexicana Gp63 Is the Enzyme Responsible for Cyclooxygenase (COX) Activity in This Parasitic Protozoa. Biochimie 2018, 151, 73–84. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Ramírez, V.I.; Estrada-Figueroa, L.A.; Medina, Y.; Lizarazo-Taborda, M.R.; Toledo-Leyva, A.; Osorio-Trujillo, C.; Morales-Mora, D.; Talamás-Rohana, P. A Monoclonal Antibody against a Leishmania Mexicana COX-like Enzymatic Activity Also Recognizes Similar Proteins in Different Protozoa of Clinical Importance. Parasitol. Res. 2023, 122, 479–492. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Ramírez, V.I.; Matus-Meza, A.-S.; Oviedo, N.; Magos-Castro, M.A.; Osorio-Trujillo, C.; Salazar-Villatoro, L.; Constantino-Jonapa, L.A.; Talamás-Rohana, P. Exploration of the Binding Site of Arachidonic Acid in Gp63 of Leishmania Mexicana and in Orthologous Proteins in Clinically Important Parasites. Pathogens 2024, 13, 718. [Google Scholar] [CrossRef]
- Sierra-López, F.; Castelan-Ramírez, I.; Hernández-Martínez, D.; Salazar-Villatoro, L.; Segura-Cobos, D.; Flores-Maldonado, C.; Hernández-Ramírez, V.I.; Villamar-Duque, T.E.; Méndez-Cruz, A.R.; Talamás-Rohana, P.; et al. Extracellular Vesicles Secreted by Acanthamoeba culbertsoni Have COX and Proteolytic Activity and Induce Hemolysis. Microorganisms 2023, 11, 2762. [Google Scholar] [CrossRef]
- Yao, C.; Donelson, J.E.; Wilson, M.E. The Major Surface Protease (MSP or GP63) of Leishmania sp. Biosynthesis, Regulation of Expression, and Function. Mol. Biochem. Parasitol. 2003, 132, 1–16. [Google Scholar] [CrossRef]
- Leathers, T.A.; Rogers, C.D. Nonsteroidal Anti-Inflammatory Drugs and Implications for the Cyclooxygenase Pathway in Embryonic Development. Am. J. Physiol.-Cell Physiol. 2023, 324, C532–C539. [Google Scholar] [CrossRef]
- Siddiqui, R.; Lakhundi, S.; Iqbal, J.; Khan, N.A. Effect of Non-Steroidal Anti-Inflammatory Drugs on Biological Properties of Acanthamoeba castellanii Belonging to the T4 Genotype. Exp. Parasitol. 2016, 168, 45–50. [Google Scholar] [CrossRef]
- De Jonckheere, J. Use of an Axenic Medium for Differentiation between Pathogenic and Nonpathogenic Naegleria Fowleri Isolates. Appl. Environ. Microbiol. 1977, 33, 751–757. [Google Scholar] [CrossRef]
- Ramírez-Rico, G.; Martinez-Castillo, M.; Cárdenas-Zúñiga, R.; Coronado-Velázquez, D.; Silva-Olivares, A.; De la Garza, M.; Shibayama, M.; Serrano-Luna, J. Acanthamoeba castellanii Genotype T4: Inhibition of Proteases Activity and Cytopathic Effect by Bovine Apo-Lactoferrin. Microorganisms 2023, 11, 708. [Google Scholar] [CrossRef]
- Sierra-López, F.; Baylón-Pacheco, L.; Espíritu-Gordillo, P.; Lagunes-Guillén, A.; Chávez-Munguía, B.; Rosales-Encina, J.L. Influence of Micropatterned Grill Lines on Entamoeba histolytica Trophozoites Morphology and Migration. Front. Cell. Infect. Microbiol. 2018, 8, 295. [Google Scholar] [CrossRef]
- National Center for Biotechnology Information (NCBI). Bethesda (MD): National Library of Medicine (US), N.C. for B.I. Available online: https://www.ncbi.nlm.nih.gov/ (accessed on 9 October 2025).
- Jones, P.; Binns, D.; Chang, H.-Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-Scale Protein Function Classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef]
- Teufel, F.; Almagro Armenteros, J.J.; Johansen, A.R.; Gíslason, M.H.; Pihl, S.I.; Tsirigos, K.D.; Winther, O.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 6.0 Predicts All Five Types of Signal Peptides Using Protein Language Models. Nat. Biotechnol. 2022, 40, 1023–1025. [Google Scholar] [CrossRef]
- Thumuluri, V.; Almagro Armenteros, J.J.; Johansen, A.R.; Nielsen, H.; Winther, O. DeepLoc 2.0: Multi-Label Subcellular Localization Prediction Using Protein Language Models. Nucleic Acids Res. 2022, 50, W228–W234. [Google Scholar] [CrossRef]
- de Castro, E.; Sigrist, C.J.A.; Gattiker, A.; Bulliard, V.; Langendijk-Genevaux, P.S.; Gasteiger, E.; Bairoch, A.; Hulo, N. ScanProsite: Detection of PROSITE Signature Matches and ProRule-Associated Functional and Structural Residues in Proteins. Nucleic Acids Res. 2006, 34, W362–W365. [Google Scholar] [CrossRef]
- Yang, J.; He, B.-J.; Jang, R.; Zhang, Y.; Shen, H.-B. Accurate Disulfide-Bonding Network Predictions Improve Ab Initio Structure Prediction of Cysteine-Rich Proteins. Bioinformatics 2015, 31, 3773–3781. [Google Scholar] [CrossRef]
- Xie, Y.; Li, H.; Luo, X.; Li, H.; Gao, Q.; Zhang, L.; Teng, Y.; Zhao, Q.; Zuo, Z.; Ren, J. IBS 2.0: An Upgraded Illustrator for the Visualization of Biological Sequences. Nucleic Acids Res. 2022, 50, W420–W426. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An Integrated and Extendable Desktop Software Platform for the Organization and Analysis of Sequence Data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Mirdita, M.; Schütze, K.; Moriwaki, Y.; Heo, L.; Ovchinnikov, S.; Steinegger, M. ColabFold: Making Protein Folding Accessible to All. Nat. Methods 2022, 19, 679–682. [Google Scholar] [CrossRef]
- Varela-Rodríguez, H.; Guzman-Pando, A.; Camarillo-Cisneros, J. Screening and Computational Characterization of Novel Antimicrobial Cathelicidins from Amphibian Transcriptomic Data. Comput. Biol. Chem. 2024, 113, 108276. [Google Scholar] [CrossRef]
- Chen, V.B.; Arendall, W.B.; Headd, J.J.; Keedy, D.A.; Immormino, R.M.; Kapral, G.J.; Murray, L.W.; Richardson, J.S.; Richardson, D.C. MolProbity: All-Atom Structure Validation for Macromolecular Crystallography. Acta Crystallogr. D Biol. Crystallogr. 2010, 66, 12–21. [Google Scholar] [CrossRef]
- Wiederstein, M.; Sippl, M.J. ProSA-Web: Interactive Web Service for the Recognition of Errors in Three-Dimensional Structures of Proteins. Nucleic Acids Res. 2007, 35, W407–W410. [Google Scholar] [CrossRef]
- Yang, J.; Wang, Y.; Zhang, Y. ResQ: An Approach to Unified Estimation of B-Factor and Residue-Specific Error in Protein Structure Prediction. J. Mol. Biol. 2016, 428, 693–701. [Google Scholar] [CrossRef]
- Biasini, M.; Bienert, S.; Waterhouse, A.; Arnold, K.; Studer, G.; Schmidt, T.; Kiefer, F.; Cassarino, T.G.; Bertoni, M.; Bordoli, L.; et al. SWISS-MODEL: Modelling Protein Tertiary and Quaternary Structure Using Evolutionary Information. Nucleic Acids Res. 2014, 42, W252–W258. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Meng, E.C.; Couch, G.S.; Croll, T.I.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Structure Visualization for Researchers, Educators, and Developers. Protein Sci. 2021, 30, 70–82. [Google Scholar] [CrossRef]
- Kim, S.; Chen, J.; Cheng, T.; Gindulyte, A.; He, J.; He, S.; Li, Q.; Shoemaker, B.A.; Thiessen, P.A.; Yu, B.; et al. PubChem 2023 Update. Nucleic Acids Res. 2023, 51, D1373–D1380. [Google Scholar] [CrossRef]
- Hanwell, M.D.; Curtis, D.E.; Lonie, D.C.; Vandermeersch, T.; Zurek, E.; Hutchison, G.R. Avogadro: An Advanced Semantic Chemical Editor, Visualization, and Analysis Platform. J. Cheminform. 2012, 4, 17. [Google Scholar] [CrossRef]
- Pracht, P.; Bohle, F.; Grimme, S. Automated Exploration of the Low-Energy Chemical Space with Fast Quantum Chemical Methods. Phys. Chem. Chem. Phys. 2020, 22, 7169–7192. [Google Scholar] [CrossRef]
- Neese, F. Software Update: The ORCA Program System—Version 5.0. WIREs Comput. Mol. Sci. 2022, 12, e1606. [Google Scholar] [CrossRef]
- Camarillo-Cisneros, J.; Ramirez-Alonso, G.; Arzate-Quintana, C.; Varela-Rodríguez, H.; Guzman-Pando, A. MolGC: Molecular Geometry Comparator Algorithm for Bond Length Mean Absolute Error Computation on Molecules. Mol. Divers. 2024, 28, 1925–1945. [Google Scholar] [CrossRef]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated Docking with Selective Receptor Flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef]
- Dolinsky, T.J.; Czodrowski, P.; Li, H.; Nielsen, J.E.; Jensen, J.H.; Klebe, G.; Baker, N.A. PDB2PQR: Expanding and Upgrading Automated Preparation of Biomolecular Structures for Molecular Simulations. Nucleic Acids Res. 2007, 35, W522–W525. [Google Scholar] [CrossRef]
- Mercado-Camargo, J.; Cervantes-Ceballos, L.; Vivas-Reyes, R.; Pedretti, A.; Serrano-García, M.L.; Gómez-Estrada, H. Homology Modeling of Leishmanolysin (Gp63) from Leishmania Panamensis and Molecular Docking of Flavonoids. ACS Omega 2020, 5, 14741–14749. [Google Scholar] [CrossRef]
- Schake, P.; Bolz, S.N.; Linnemann, K.; Schroeder, M. PLIP 2025: Introducing Protein–Protein Interactions to the Protein–Ligand Interaction Profiler. Nucleic Acids Res. 2025, 53, W463–W465. [Google Scholar] [CrossRef]
- Dassault Systèmes. BIOVIA Discovery Studio Visualizer, Version 21.1; Dassault Systèmes: San Diego, CA, USA. Available online: https://www.3ds.com/products-services/biovia/ (accessed on 22 October 2025).
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A Free Web Tool to Evaluate Pharmacokinetics, Drug-Likeness and Medicinal Chemistry Friendliness of Small Molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef]
- Zoete, V.; Daina, A.; Bovigny, C.; Michielin, O. SwissSimilarity: A Web Tool for Low to Ultra High Throughput Ligand-Based Virtual Screening. J. Chem. Inf. Model. 2016, 56, 1399–1404. [Google Scholar] [CrossRef]
- Bragina, M.E.; Daina, A.; Perez, M.A.S.; Michielin, O.; Zoete, V. The SwissSimilarity 2021 Web Tool: Novel Chemical Libraries and Additional Methods for an Enhanced Ligand-Based Virtual Screening Experience. Int. J. Mol. Sci. 2022, 23, 811. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissTargetPrediction: Updated Data and New Features for Efficient Prediction of Protein Targets of Small Molecules. Nucleic Acids Res. 2019, 47, W357–W364. [Google Scholar] [CrossRef]
- Williams, A.J.; Grulke, C.M.; Edwards, J.; McEachran, A.D.; Mansouri, K.; Baker, N.C.; Patlewicz, G.; Shah, I.; Wambaugh, J.F.; Judson, R.S.; et al. The CompTox Chemistry Dashboard: A Community Data Resource for Environmental Chemistry. J. Cheminform. 2017, 9, 61. [Google Scholar] [CrossRef]
- Keiser, M.J.; Roth, B.L.; Armbruster, B.N.; Ernsberger, P.; Irwin, J.J.; Shoichet, B.K. Relating Protein Pharmacology by Ligand Chemistry. Nat. Biotechnol. 2007, 25, 197–206. [Google Scholar] [CrossRef]
- Cheng, F.; Li, W.; Zhou, Y.; Shen, J.; Wu, Z.; Liu, G.; Lee, P.W.; Tang, Y. AdmetSAR: A Comprehensive Source and Free Tool for Assessment of Chemical ADMET Properties. J. Chem. Inf. Model. 2012, 52, 3099–3105. [Google Scholar] [CrossRef]
- Yang, H.; Lou, C.; Sun, L.; Li, J.; Cai, Y.; Wang, Z.; Li, W.; Liu, G.; Tang, Y. AdmetSAR 2.0: Web-Service for Prediction and Optimization of Chemical ADMET Properties. Bioinformatics 2019, 35, 1067–1069. [Google Scholar] [CrossRef]
- Zdrazil, B.; Felix, E.; Hunter, F.; Manners, E.J.; Blackshaw, J.; Corbett, S.; de Veij, M.; Ioannidis, H.; Lopez, D.M.; Mosquera, J.F.; et al. The ChEMBL Database in 2023: A Drug Discovery Platform Spanning Multiple Bioactivity Data Types and Time Periods. Nucleic Acids Res. 2024, 52, D1180–D1192. [Google Scholar] [CrossRef]
- González-Robles, A.; Castañón, G.; Hernández-Ramírez, V.I.; Salazar-Villatoro, L.; González-Lázaro, M.; Omaña-Molina, M.; Talamás-Rohana, P.; Martínez-Palomo, A. Acanthamoeba castellanii: Identification and Distribution of Actin Cytoskeleton. Exp. Parasitol. 2008, 119, 411–417. [Google Scholar] [CrossRef]
- Ferrera, P.; Zepeda, A.; Arias, C. Nonsteroidal Anti-Inflammatory Drugs Attenuate Amyloid-β Protein-Induced Actin Cytoskeletal Reorganization through Rho Signaling Modulation. Cell. Mol. Neurobiol. 2017, 37, 1311–1318. [Google Scholar] [CrossRef]
- López-Contreras, L.; Hernández-Ramírez, V.I.; Lagunes-Guillén, A.E.; Montaño, S.; Chávez-Munguía, B.; Sánchez-Ramírez, B.; Talamás-Rohana, P. Exploring the Possible Role of Lysine Acetylation on Entamoeba histolytica Virulence: A Focus on the Dynamics of the Actin Cytoskeleton. Biomed Res. Int. 2013, 2013, 757392. [Google Scholar] [CrossRef]
- Łanocha-Arendarczyk, N.; Baranowska-Bosiacka, I.; Kot, K.; Gutowska, I.; Kolasa-Wołosiuk, A.; Chlubek, D.; Kosik-Bogacka, D. Expression and Activity of COX-1 and COX-2 in Acanthamoeba sp.-Infected Lungs According to the Host Immunological Status. Int. J. Mol. Sci. 2018, 19, 121. [Google Scholar] [CrossRef]
- Magistrado-Coxen, P.; Aqeel, Y.; Lopez, A.; Haserick, J.R.; Urbanowicz, B.R.; Costello, C.E.; Samuelson, J. The Most Abundant Cyst Wall Proteins of Acanthamoeba castellanii are Lectins That Bind Cellulose and Localize to Distinct Structures in Developing and Mature Cyst Walls. PLoS Negl. Trop. Dis. 2019, 13, e0007352. [Google Scholar] [CrossRef]
- Brittingham, A.; Morrison, C.J.; McMaster, W.R.; McGwire, B.S.; Chang, K.P.; Mosser, D.M. Role of the Leishmania Surface Protease Gp63 in Complement Fixation, Cell Adhesion, and Resistance to Complement-Mediated Lysis. J. Immunol. 1995, 155, 3102–3111. [Google Scholar] [CrossRef]
- Hallé, M.; Gomez, M.A.; Stuible, M.; Shimizu, H.; McMaster, W.R.; Olivier, M.; Tremblay, M.L. The Leishmania Surface Protease GP63 Cleaves Multiple Intracellular Proteins and Actively Participates in P38 Mitogen-Activated Protein Kinase Inactivation. J. Biol. Chem. 2009, 284, 6893–6908. [Google Scholar] [CrossRef]
- Chen, F.; Zhu, B.; Fang, Y.; Li, Z.; Lei, Z.; Xue, Z.; Shen, T.; Zhou, S.; Chen, X.; Xu, L.; et al. Schistosoma japonicum Leishmanolysin SjLLPi1 Facilitates the Invasion of Cercariae into the Host Skin. PLoS Pathog. 2025, 21, e1013446. [Google Scholar] [CrossRef]
- Schlagenhauf, E.; Etges, R.; Metcalf, P. The Crystal Structure of the Leishmania Major Surface Proteinase Leishmanolysin (Gp63). Structure 1998, 6, 1035–1046. [Google Scholar] [CrossRef]
- Li, D.; Kerns, E.H. Drug-Like Properties: Concepts, Structure Design and Methods from ADME to Toxicity Optimization; Elsevier: Amsterdam, The Netherlands, 2016. [Google Scholar]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and Computational Approaches to Estimate Solubility and Permeability in Drug Discovery and Development Settings. Adv. Drug Deliv. Rev. 2001, 46, 3–26. [Google Scholar] [CrossRef]
- Parepally, J.M.R.; Mandula, H.; Smith, Q.R. Brain Uptake of Nonsteroidal Anti-Inflammatory Drugs: Ibuprofen, Flurbiprofen, and Indomethacin. Pharm. Res. 2006, 23, 873–881. [Google Scholar] [CrossRef]
- Marciano-Cabral, F.; Cabral, G. Acanthamoeba spp. as Agents of Disease in Humans. Clin. Microbiol. Rev. 2003, 16, 273–307. [Google Scholar] [CrossRef]
- Löscher, W.; Potschka, H. Drug Resistance in Brain Diseases and the Role of Drug Efflux Transporters. Nat. Rev. Neurosci. 2005, 6, 591–602. [Google Scholar] [CrossRef]
- Miners, J.O.; Mackenzie, P.I. Drug Glucuronidation in Humans. Pharmacol. Ther. 1991, 51, 347–369. [Google Scholar] [CrossRef]
- Rendic, S.; Guengerich, F.P. Contributions of Human Enzymes in Carcinogen Metabolism. Chem. Res. Toxicol. 2012, 25, 1316–1383. [Google Scholar] [CrossRef]
- Grosser, T.; Smyth, E.; FitzGerald, G. Pharmacotherapy of Inflammation, Fever, Pain, and Gout. Chapter 38. In Goodman & Gilman’s: The Pharmacological Basis of Therapeutics, 13th ed.; Brunton, L.L., Hilal-Dandan, R., Knollmann, B.C., Eds.; McGraw-Hill Education: Columbus, OH, USA; Available online: https://accessmedicine.mhmedical.com/content.aspx?bookid=2189§ionid=172471818 (accessed on 22 October 2025).
- Raies, A.B.; Bajic, V.B. In Silico Toxicology: Computational Methods for the Prediction of Chemical Toxicity. WIREs Comput. Mol. Sci. 2016, 6, 147–172. [Google Scholar] [CrossRef]
- Ekins, S.; Mestres, J.; Testa, B. In Silico Pharmacology for Drug Discovery: Applications to Targets and Beyond. Br. J. Pharmacol. 2007, 152, 21–37. [Google Scholar] [CrossRef]
- van de Waterbeemd, H.; Gifford, E. ADMET in Silico Modelling: Towards Prediction Paradise? Nat. Rev. Drug Discov. 2003, 2, 192–204. [Google Scholar] [CrossRef]




| Compound | Sample Type | Binding Energy (kcal·mol−1) | Inhibition Constant (Ki, 25 °C) |
|---|---|---|---|
| L-Ascorbic acid | Low affinity or low specificity (control) | −4.31 | 689.74 µM |
| Benzoic acid | Moderate affinity (control) | −7.2 | 5.27 µM |
| Aspirin | Sample | −7.85 | 1.76 µM |
| Ibuprofen | Sample | −9.49 | 0.111 µM |
| Diclofenac | Sample | −10.37 | 0.025 µM |
| Amentoflavone | High affinity (control) | −10.73 | 0.014 µM |
| Parameter | Ibuprofen | Aspirin | Diclofenac |
|---|---|---|---|
| Lipinski criteria (%) | 100 | 100 | 100 |
| QED score (%) | 89.5 | 57.1 | 96.5 |
| Permeability (%) | 13.64 | 23.26 | 18.04 |
| Solubility (%) | 51.48 | 34.9 | 63.63 |
| IC50 (theoretical) (μM) | ≈983.75 ± 424 | ≈727.12 ± 137 | ≈135.14 ± 56 |
| AC50 (theoretical) (μM) | 144 | 186 | - |
| BMD (μM) | 50 | 90 | 90 |
| CSTA (%) | Propachlor (23) | Diethyl phthalate (30) | Dichlorobenzamide (25) |
| Main molecular target | Oxidoreductases, transporters | Prostaglandin synthases | Prostaglandins, chemokine |
| Predicted toxicity | Low | Moderate | High |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Hernández-Ramírez, V.I.; Varela-Rodríguez, H.; Varela-Rodríguez, L.; Sierra-López, F.; San Juan-Mora, D.E.; Morales-Mora, J.D.; Falcón-Navarrete, D.; Osorio-Trujillo, C.; Ríos-López, J.; Rodríguez-Mera, I.B.; et al. Acanthamoeba castellanii: Non-Steroidal Anti-Inflammatory Drugs Affect Adhesion, Motility, and Encystment, Suggesting a Link with a gp63-like Protein Candidate. Pathogens 2026, 15, 263. https://doi.org/10.3390/pathogens15030263
Hernández-Ramírez VI, Varela-Rodríguez H, Varela-Rodríguez L, Sierra-López F, San Juan-Mora DE, Morales-Mora JD, Falcón-Navarrete D, Osorio-Trujillo C, Ríos-López J, Rodríguez-Mera IB, et al. Acanthamoeba castellanii: Non-Steroidal Anti-Inflammatory Drugs Affect Adhesion, Motility, and Encystment, Suggesting a Link with a gp63-like Protein Candidate. Pathogens. 2026; 15(3):263. https://doi.org/10.3390/pathogens15030263
Chicago/Turabian StyleHernández-Ramírez, Verónica I., Hugo Varela-Rodríguez, Luis Varela-Rodríguez, Francisco Sierra-López, Daniela Eloísa San Juan-Mora, José Daniel Morales-Mora, Daniela Falcón-Navarrete, Carlos Osorio-Trujillo, Jacqueline Ríos-López, Itzel Berenice Rodríguez-Mera, and et al. 2026. "Acanthamoeba castellanii: Non-Steroidal Anti-Inflammatory Drugs Affect Adhesion, Motility, and Encystment, Suggesting a Link with a gp63-like Protein Candidate" Pathogens 15, no. 3: 263. https://doi.org/10.3390/pathogens15030263
APA StyleHernández-Ramírez, V. I., Varela-Rodríguez, H., Varela-Rodríguez, L., Sierra-López, F., San Juan-Mora, D. E., Morales-Mora, J. D., Falcón-Navarrete, D., Osorio-Trujillo, C., Ríos-López, J., Rodríguez-Mera, I. B., Carrasco-Yépez, M. M., & Talamás-Rohana, P. (2026). Acanthamoeba castellanii: Non-Steroidal Anti-Inflammatory Drugs Affect Adhesion, Motility, and Encystment, Suggesting a Link with a gp63-like Protein Candidate. Pathogens, 15(3), 263. https://doi.org/10.3390/pathogens15030263










