Assessment of the Pathogenicity of Candidatus Rickettsia Colombiensis in a Syrian Hamster Model and Serological Cross-Reactivity Between Spotted Fever Rickettsia Species
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design and Tick Collection
2.2. Isolation and Culture of Candidatus Rickettsia Colombiensis
2.3. Animal Infection Model of Syrian Hamster (Mesocricetus auratus) with Candidatus Rickettsia Colombiensis
2.4. Clinical Monitoring and Histopathology
2.5. Ethical Aspects
2.6. Preparation of Slides and Immunofluorescence Assays (IFA)
2.7. Cross-Reactivity of Syrian Hamsters and Human Serum
2.8. Human Sera for Cross-Reactivity Study of Rickettsia sp. in the Spotted Fever Group
2.9. Molecular Detection Rickettsia by qPCR and PCR Conventional
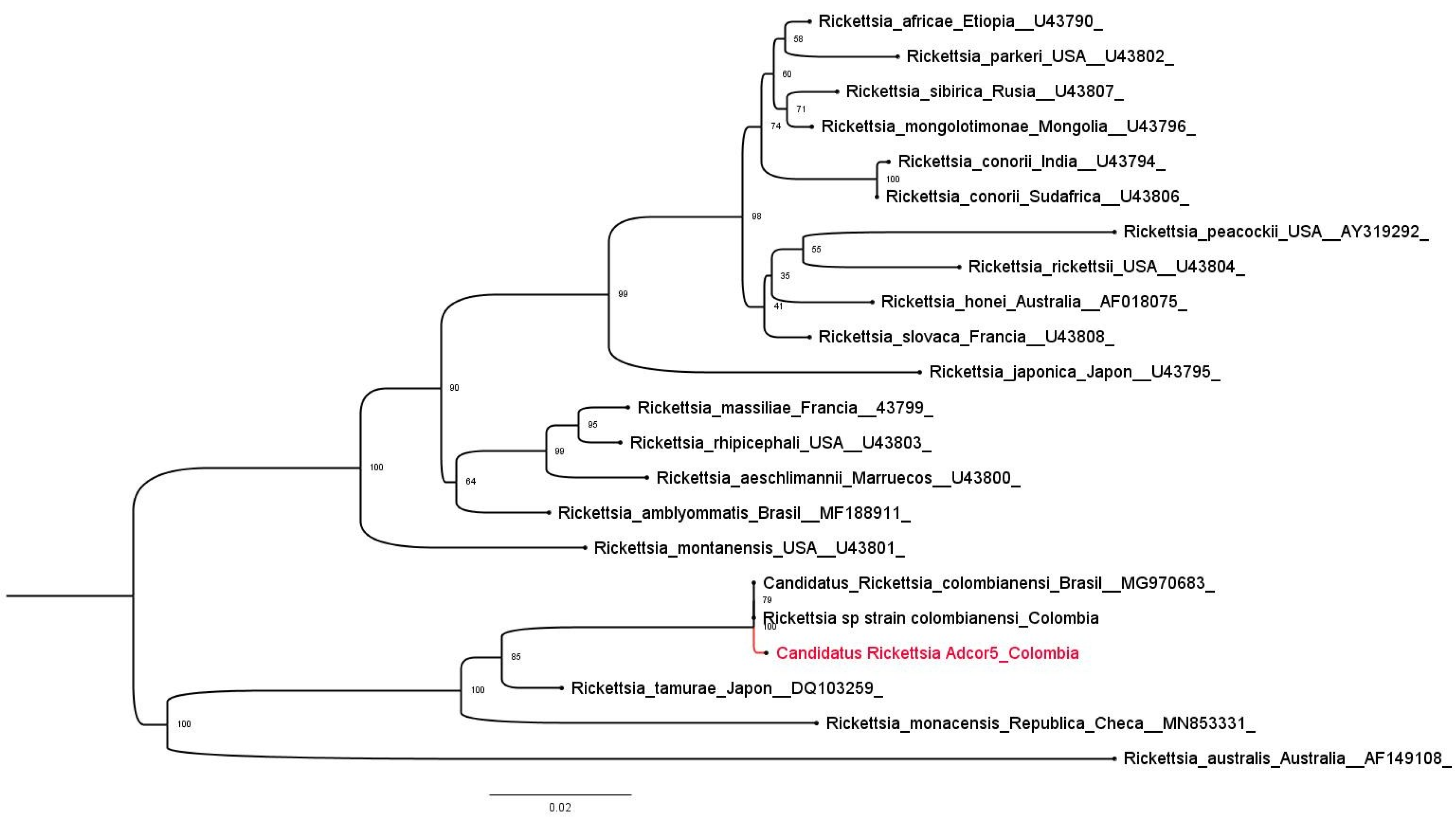
2.10. Phylogenetic Analysis
2.11. Data Analysis
3. Results
3.1. Isolation of Candidatus Rickettsia Colombiensis
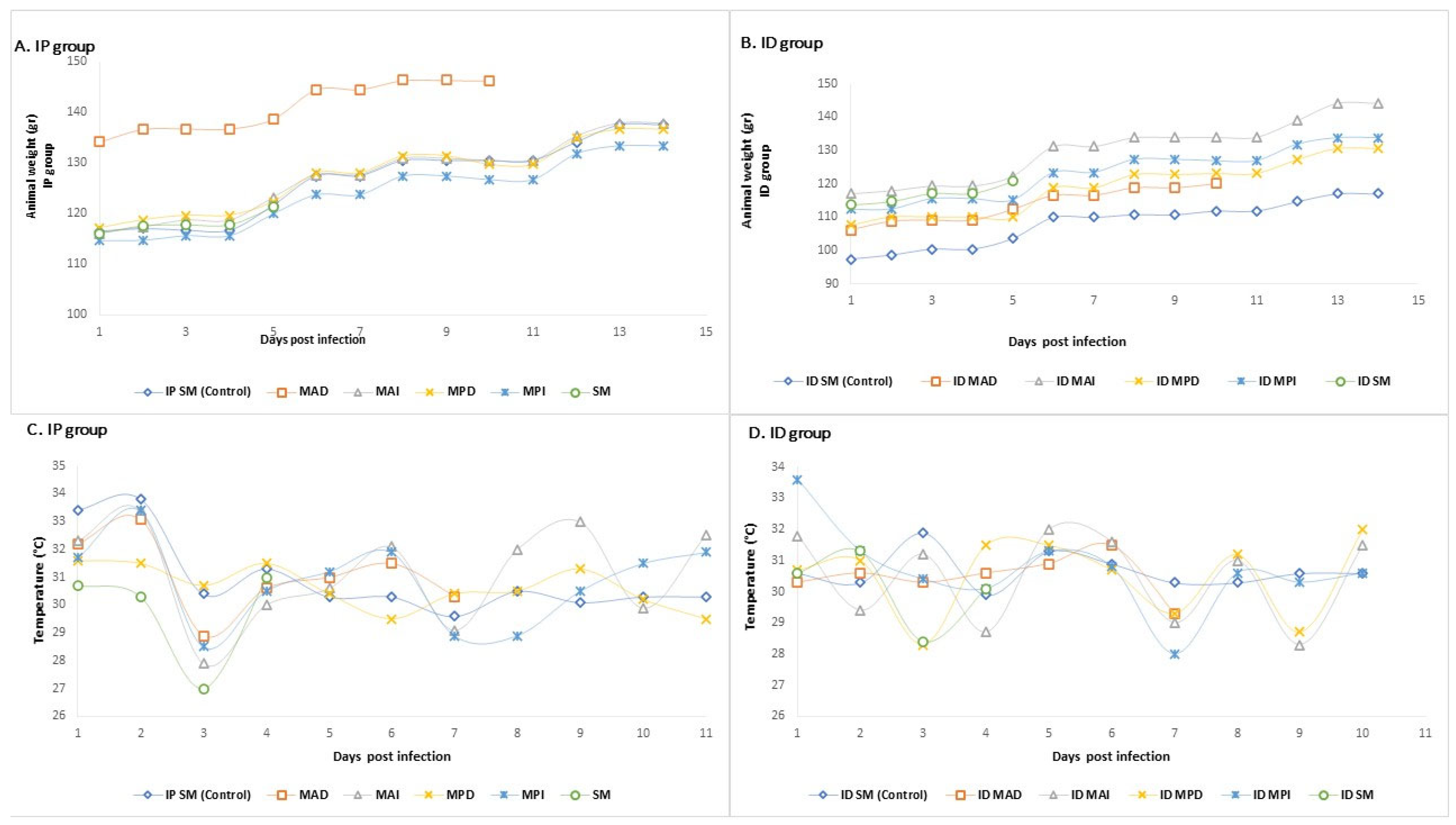
3.2. Evaluation of the Pathogenesis of Ca. R. Colombiensis in an Experimental Syrian Hamster (M. auratus) Model
3.3. Detection of Rickettsiae in Syrian Hamster Tissues
3.4. Indirect Immunofluorescence (IFA) on Syrian Hamster Serum
3.5. Cross-Reactivity in Syrian Hamster and Human Sera
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Ca | Candidatus |
| Ct | Cycle threshold |
| IP | Intraperitoneally |
| ID | Intradermally |
| DNA | Deoxyribonucleic acid |
| qPCR | Quantitative or real-time PCR |
| SFG | Spotted fever group |
| TG | Typhus group |
| TRG | Transitional group |
| AG | Ancestral group |
| BHI | Brain Heart Infusion |
| DMEM | Dulbecco’s Modified Eagle’s medium |
| BFS | Bovine fetal serum |
| IFA | Indirect Immunofluorescence Assay |
| SPG | Sucrose–phosphate–glutamate |
| SPF | Specific pathogen-free |
| AdCor 5 | Amblyomma dissimile (isolated number 5) |
| w | Weight |
| °C | Temperature |
| H&E | Hematoxylin and eosin |
| PBS | Phosphate-buffered saline |
| H + L | Heavy- and light-chain IgG |
| FITC | Fluorescein isothiocyanate |
| gltA | Citrato sintasa (glutamate metabolism) |
| OmpA | Outer membrane protein A |
References
- Parola, P.; Paddock, C.D.; Socolovschi, C.; Labruna, M.B.; Mediannikov, O.; Kernif, T.; Abdad, M.Y.; Stenos, J.; Bitam, I.; Fournier, P.-E.; et al. Update on tick-borne rickettsioses around the world: A geographic approach. Clin. Microbiol. Rev. 2013, 26, 657–702. [Google Scholar] [CrossRef] [PubMed]
- Sahni, A.; Fang, R.; Sahni, S.K.; Walker, D.H. Pathogenesis of rickettsial diseases: Pathogenic and immune mechanisms of an endotheliotropic infection. Annu. Rev. Pathol. Mech. Dis. 2019, 14, 127–152. [Google Scholar] [CrossRef] [PubMed]
- Sit, B.; Lamason, R.L. Pathogenic Rickettsia spp. as emerging models for bacterial biology. J. Bacteriol. 2024, 206, E0040423. [Google Scholar] [CrossRef]
- Helminiak, L.; Mishra, S.; Kim, H.K. Pathogenicity and virulence of Rickettsia. Virulence 2022, 13, 1752–1771. [Google Scholar] [CrossRef]
- Weinert, L.A.; Werren, J.H.; Aebi, A.; Stone, G.N.; Jiggins, F.M. Evolution and diversity of Rickettsia bacteria. BMC Biol. 2009, 7, 6. [Google Scholar] [CrossRef]
- Salje, J. Cells within cells: Rickettsiales and the obligate intracellular bacterial lifestyle. Nat. Rev. Microbiol. 2021, 19, 375–390. [Google Scholar] [CrossRef]
- Parola, P.; Musso, D.; Raoult, D. Rickettsia felis: The next mosquito-borne outbreak? Lancet Infect. Dis. 2016, 16, 1112–1113. [Google Scholar] [CrossRef]
- Gillespie, J.J.; Williams, K.; Shukla, M.; E Snyder, E.; Nordberg, E.K.; Ceraul, S.M.; Dharmanolla, C.; Rainey, D.; Soneja, J.; Shallom, J.M.; et al. Rickettsia phylogenomics: Unwinding the intricacies of obligate intracellular life. PLoS ONE 2008, 3, E2018. [Google Scholar] [CrossRef]
- Snellgrove, A.N.; Krapiunaya, I.; Scott, P.; Levin, M.L. Assessment of the pathogenicity of Rickettsia amblyommatis, Rickettsia bellii, and Rickettsia montanensis in a guinea pig model. Vector-Borne Zoonotic Dis. 2021, 21, 232–241. [Google Scholar] [CrossRef]
- Piotrowski, M.; Rymaszewska, A. Expansion of tick-borne rickettsioses in the world. Microorganisms 2020, 8, 1906. [Google Scholar] [CrossRef]
- Freese, H.M.; Meier-Kolthoff, J.P.; Sardà Carbasse, J.; Afolayan, A.O.; Göker, M. TYGS and LPSN in 2025: A Global Core Biodata Resource for genome-based classification and nomenclature of prokaryotes within DSMZ Digital Diversity. Nucleic Acids Res. 2025, 54, Gkaf1110. [Google Scholar] [CrossRef] [PubMed]
- Portillo, A.; Ibarra, V.; Santibáñez, S.; Pérez-Martínez, L.; Blanco, J.R.; Oteo, J.A. Genetic characterisation of ompA, ompB and gltA genes from Candidatus Rickettsia rioja. Clin. Microbiol. Infect. 2009, 15, 307–308. [Google Scholar] [CrossRef] [PubMed]
- Probert, W.S.; Haw, M.P.; Nichol, A.C.; Glaser, C.A.; Park, S.Y.; Campbell, L.E.; Trivedi, K.K.; Romo, H.; Saunders, M.E.; Kjemtrup, A.M.; et al. Newly Recognized Spotted Fever Group Rickettsia as Cause of Severe Rocky Mountain Spotted Fever–Like Illness, Northern California, USA. Emerg. Infect. Dis. 2024, 30, 1344. [Google Scholar] [CrossRef] [PubMed]
- Tomassone, L.; Portillo, A.; Nováková, M.; De Sousa, R.; Oteo, J.A. Neglected aspects of tick-borne rickettsioses. Parasites Vectors 2018, 11, 263. [Google Scholar] [CrossRef]
- Salje, J.; Weitzel, T.; Newton, P.N.; Varghese, G.M.; Day, N. Rickettsial infections: A blind spot in our view of neglected tropical diseases. PLoS Negl. Trop. Dis. 2021, 15, E0009353. [Google Scholar] [CrossRef]
- Zhang, Y.-Y.; Sun, Y.-Q.; Chen, J.-J.; Teng, A.-Y.; Wang, T.; Li, H.; I Hay, S.; Fang, L.-Q.; Yang, Y.; Liu, W. Mapping the global distribution of spotted fever group rickettsiae: A systematic review with modelling analysis. Lancet Digit. Health 2023, 5, e5–e15. [Google Scholar] [CrossRef]
- Miranda, J.; Portillo, A.; Oteo, J.A.; Mattar, S. Rickettsia sp. strain colombianensi (Rickettsiales: Rickettsiaceae): A new proposed Rickettsia detected in Amblyomma dissimile (Acari: Ixodidae) from iguanas and free-living larvae ticks from vegetation. J. Med. Entomol. 2012, 49, 960–965. [Google Scholar] [CrossRef]
- Oren, A.; Garrity, G.M.; Parker, C.T.; Chuvochina, M.; Trujillo, M.E. Lists of names of prokaryotic Candidatus taxa. Int. J. Syst. Evol. Microbiol. 2020, 70, 3956–4042. [Google Scholar] [CrossRef]
- Miranda, J.; Violet-Lozano, L.; Barrera, S.; Mattar, S.; Monsalve-Buriticá, S.; Rodas, J.; Contreras, V. Candidatus Rickettsia colombianensi in ticks from reptiles in Córdoba, Colombia. Vet. World 2020, 13, 1764. [Google Scholar] [CrossRef]
- Cotes-Perdomo, A.; Cárdenas-Carreño, J.; Hoyos, J.; González, C.; Castro, L.R. Molecular detection of Candidatus Rickettsia colombianensi in ticks (Acari, Ixodidae) collected from herpetofauna in San Juan de Carare, Colombia. Int. J. Parasitol. Parasites Wildl. 2022, 19, 110–114. [Google Scholar] [CrossRef]
- Lerma, L.S.; Cogollo, V.C.; Velilla, S.M.; Gonzalez, I.R.; Atehortua, A.D.M.; Peñuela, D.F.C. First detection of Candidatus Rickettsia colombianensi in the State of Meta, Colombia. Rev. Habanera Cienc. Méd. 2019, 18, 487–499. [Google Scholar]
- Martínez-Sánchez, E.T.; Cardona-Romero, M.; Ortiz-Giraldo, M.; Tobón-Escobar, W.D.; Moreno-López, D.; Ossa-López, P.A.; Pérez-Cárdenas, J.E.; Labruna, M.B.; Martins, T.F.; Castaño-Villa, G.J.; et al. Rickettsia spp. in ticks (Acari: Ixodidae) from wild birds in Caldas, Colombia. Acta Trop. 2021, 213, 105733. [Google Scholar] [CrossRef] [PubMed]
- Santodomingo, A.; Cotes-Perdomo, A.; Foley, J.; Castro, L.R. Rickettsial infection in ticks (Acari: Ixodidae) from reptiles in the Colombian Caribbean. Ticks Tick-Borne Dis. 2018, 9, 623–628. [Google Scholar] [CrossRef] [PubMed]
- Rivera-Páez, F.A.; Martins, T.F.; Ossa-López, P.A.; Sampieri, B.R.; Camargo-Mathias, M.I. Detection of Rickettsia spp. in ticks (Acari: Ixodidae) of domestic animals in Colombia. Ticks Tick-Borne Dis. 2018, 9, 819–823. [Google Scholar] [CrossRef]
- Ogrzewalska, M.; Machado, C.; Rozental, T.; Forneas, D.; Cunha, L.E.; De Lemos, E. Microorganisms in the ticks Amblyomma dissimile Koch 1844 and Amblyomma rotundatum Koch 1844 collected from snakes in Brazil. Med. Vet. Entomol. 2019, 33, 154–161. [Google Scholar] [CrossRef]
- Luz, H.R.; Silva-Santos, E.; Costa-Campos, C.E.; Acosta, I.; Martins, T.F.; Muñoz-Leal, S.; McIntosh, D.; Faccini, J.L.H.; Labruna, M.B. Detection of Rickettsia spp. in ticks parasitizing toads (Rhinella marina) in the northern Brazilian Amazon. Exp. Appl. Acarol. 2018, 75, 309–318. [Google Scholar] [CrossRef]
- Bermúdez, S.; Martínez-Mandiche, J.; Domínguez, L.; Gonzalez, C.; Chavarria, O.; Moreno, A.; Góndola, J.; Correa, N.; Rodríguez, I.; Castillo, B.; et al. Diversity of Rickettsia in ticks collected from wild animals in Panama. Ticks Tick-Borne Dis. 2021, 12, 101723. [Google Scholar] [CrossRef]
- Romero, L.; Costa, F.B.; Labruna, M.B. Ticks and tick-borne Rickettsia in El Salvador. Exp. Appl. Acarol. 2021, 83, 545–554. [Google Scholar] [CrossRef]
- Sánchez-Montes, S.; Isaak-Delgado, A.B.; Guzmán-Cornejo, C.; Rendón-Franco, E.; Muñoz-García, C.I.; Bermúdez, S.; Morales-Diaz, J.; Cruz-Romero, A.; Romero-Salas, D.; Dzul-Rosado, K.; et al. Rickettsia species in ticks that parasitize amphibians and reptiles: Novel report from Mexico and review of the worldwide record. Ticks Tick-Borne Dis. 2019, 10, 987–994. [Google Scholar] [CrossRef]
- Cotes-Perdomo, A.; Santodomingo, A.; Castro, L.R. Hemogregarine and Rickettsial infection in ticks of toads from northeastern Colombia. Int. J. Parasitol. Parasites Wildl. 2018, 7, 237–242. [Google Scholar] [CrossRef]
- Imaoka, K.; Kaneko, S.; Tabara, K.; Kusatake, K.; Morita, E. The First Human Case of Rickettsia tamurae Infection in Japan. Case Rep. Dermatol. 2011, 3, 68–73. [Google Scholar] [CrossRef] [PubMed]
- Jado, I.; Oteo, J.A.; Aldámiz, M.; Gil, H.; Escudero, R.; Ibarra, V.; Portu, J.; Portillo, A.; Lezaun, M.J.; García-Amil, C.; et al. Rickettsia monacensis and human disease, Spain. Emerg. Infect. Dis. 2007, 13, 1405–1407. [Google Scholar] [CrossRef] [PubMed]
- Raoult, D.; Berbis, P.; Roux, V.; Xu, W.; Maurin, M. A new tick-transmitted disease due to Rickettsia slovaca. Lancet 1997, 350, 112–113. [Google Scholar] [CrossRef]
- Paddock, C.D.; Sumner, J.W.; Comer, J.A.; Zaki, S.R.; Goldsmith, C.S.; Goddard, J.; McLellan, S.L.F.; Tamminga, C.L.; Ohl, C.A. Rickettsia parkeri: A newly recognized cause of spotted fever rickettsiosis in the United States. Clin. Infect. Dis. 2004, 38, 805–811. [Google Scholar] [CrossRef]
- Perlman, S.J.; Hunter, M.S.; Zchori-Fein, E. The emerging diversity of Rickettsia. Proc. Biol. Sci. 2006, 273, 2097–2106. [Google Scholar] [CrossRef]
- Oteo, J.A.; Nava, S.; Sousa Rd Mattar, S.; Venzal, J.M.; Abarca, K.; Zavala-Castro, J. Guías Latinoamericanas de la RIICER para el diagnóstico de las rickettsiosis transmitidas por garrapatas. Rev. Chil. Infect. 2014, 31, 54–65. [Google Scholar] [CrossRef]
- Barros-Battesti, D.M.; Arzua, M.; Bechara, G.H. Carrapatos de importância médico-veterinária da região neotropical: Um guia ilustrado para identificação de espécies. In Anonymous Carrapatos De Importância Médico-Veterinária Da Região Neotropical: Um guia Ilustrado Para Identificação De Espécies; Instituto Butantan: São Paulo, Brazil, 2006. [Google Scholar]
- Labruna, M.B.; Whitworth, T.; Horta, M.C.; Bouyer, D.H.; McBride, J.W.; Pinter, A.; Popov, V.; Gennari, S.M.; Walker, D.H. Rickettsia species infecting Amblyomma cooperi ticks from an area in the state of São Paulo, Brazil, where Brazilian spotted fever is endemic. J. Clin. Microbiol. 2004, 42, 90–98. [Google Scholar] [CrossRef]
- Gimenez, D.F. Staining Rickettsiae in Yolk-Sac Cultures. Stain. Technol. 1964, 39, 135–140. [Google Scholar] [CrossRef]
- Kelly, P.J.; Raoult, D.; Mason, P.R. Isolation of spotted fever group rickettsias from triturated ticks using a modification of the centrifugation-shell vial technique. Trans. R. Soc. Trop. Med. Hyg. 1991, 85, 397–398. [Google Scholar] [CrossRef]
- Feng, W.C.; Waner, J.L. Serological cross-reaction and cross-protection in guinea pigs infected with Rickettsia rickettsii and Rickettsia montana. Infect. Immun. 1980, 28, 627–629. [Google Scholar] [CrossRef]
- Stokes, J.V.; Levin, M.L.; Cross, C.E.; Ross, A.L.; Snellgrove, A.N.; Willeford, B.V.; Alugubelly, N.; Varela-Stokes, A.S. Evaluating the clinical and immune responses to spotted fever rickettsioses in the guinea pig-tick-Rickettsia system. Curr. Protoc. 2022, 2, E584. [Google Scholar] [CrossRef]
- Brustolin, J.M.; Krawczak, F.d.S.; Alves, M.E.M.; Weiller, M.A.; de Souza, C.L.; Rosa, F.B.; Cadore, G.C.; Lopes, S.T.D.A.; Labruna, M.B.; Vogel, F.S.F.; et al. Experimental infection in Cavia porcellus by infected Amblyomma ovale nymphs with Rickettsia sp. (Atlantic rainforest strain). Parasitol. Res. 2018, 117, 713–720. [Google Scholar] [CrossRef]
- Soares, J.F.; Soares, H.S.; Barbieri, A.M.; Labruna, M.B. Experimental infection of the tick Amblyomma cajennense, Cayenne tick, with Rickettsia rickettsii, the agent of Rocky Mountain spotted fever. Med. Vet. Entomol. 2012, 26, 139–151. [Google Scholar] [CrossRef] [PubMed]
- Rivas, J.J.; Moreira-Soto, A.; Alvarado, G.; Taylor, L.; Calderón-Arguedas, O.; Hun, L.; Corrales-Aguilar, E.; Morales, J.A.; Troyo, A. Pathogenic potential of a Costa Rican strain of ‘Candidatus Rickettsia amblyommii’in guinea pigs (Cavia porcellus) and protective immunity against Rickettsia rickettsii. Ticks Tick-Borne Dis. 2015, 6, 805–811. [Google Scholar] [CrossRef] [PubMed]
- Horta, M.C.; Labruna, M.B.; Sangioni, L.A.; Vianna, M.C.B.; Gennari, S.M.; Galvão, M.A.M.; Mafra, C.L.; Vidotto, O.; Schumaker, T.T.S.; Walker, D.H. Prevalence of antibodies to spotted fever group rickettsiae in humans and domestic animals in a Brazilian spotted fever-endemic area in the state of São Paulo, Brazil: Serologic evidence for infection by Rickettsia rickettsii and another spotted fever group Rickettsia. Am. J. Trop. Med. Hyg. 2004, 71, 93–97. [Google Scholar] [PubMed]
- Paris, D.H.; Dumler, J.S. State of the art of diagnosis of rickettsial diseases: The use of blood specimens for diagnosis of scrub typhus, spotted fever group rickettsiosis, and murine typhus. Curr. Opin. Infect. Dis. 2016, 29, 433–439. [Google Scholar] [CrossRef]
- Costa, P.S.G.D.; Brigatte, M.E.; Greco, D.B. Antibodies to Rickettsia rickettsii, Rickettsia typhi, Coxiella burnetii, Bartonella henselae, Bartonella quintana, and Ehrlichia chaffeensis among healthy population in Minas Gerais, Brazil. Mem. Do Inst. Oswaldo Cruz 2005, 100, 853–859. [Google Scholar] [CrossRef]
- Hechemy, K.E.; Raoult, D.; Fox, J.; Han, Y.; Elliott, L.B.; Rawlings, J. Cross-reaction of immune sera from patients with rickettsial diseases. J. Med. Microbiol. 1989, 29, 199–202. [Google Scholar] [CrossRef]
- Portillo, A.; de Sousa, R.; Santibáñez, S.; Duarte, A.; Edouard, S.; Fonseca, I.P.; Marques, C.; Novakova, M.; Palomar, A.M.; Santos, M.; et al. Guidelines for the Detection of Rickettsia spp. Vector-Borne Zoonotic Dis. 2017, 17, 23–32. [Google Scholar] [CrossRef]
- Stewart, A.G.; Stewart, A.G. An update on the laboratory diagnosis of Rickettsia spp. Infect. Pathogens 2021, 10, 1319. [Google Scholar] [CrossRef]
- Barrera, S.; Martínez, S.; Tique-Salleg, V.; Miranda, J.; Guzmán, C.; Mattar, S. Seroprevalencia de Hantavirus, Rickettsia y Chikungunya en población indígena del municipio de Tuchín, Córdoba. Infectio 2015, 19, 75–82. [Google Scholar] [CrossRef]
- Hidalgo, M.; Miranda, J.; Heredia, D.; Zambrano, P.; Vesga, J.F.; Lizarazo, D.; Mattar, S.; Valbuena, G. Outbreak of Rocky Mountain spotted fever in Cordoba, Colombia. Mem. Inst. Oswaldo Cruz 2011, 106, 117–118. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Wang, L.; Sakthivel, S.K.; Whitaker, B.; Murray, J.; Kamili, S.; Lynch, B.; Malapati, L.; Burke, S.A.; Harcourt, J.; et al. US CDC real-time reverse transcription PCR panel for detection of severe acute respiratory syndrome coronavirus 2. Emerg. Infect. Dis. 2020, 26, 1654. [Google Scholar] [CrossRef] [PubMed]
- Roux, V.; Fournier, P.E.; Raoult, D. Differentiation of spotted fever group rickettsiae by sequencing and analysis of restriction fragment length polymorphism of PCR-amplified DNA of the gene encoding the protein rOmpA. J. Clin. Microbiol. 1996, 34, 2058–2065. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Trifinopoulos, J.; Nguyen, L.; von Haeseler, A.; Minh, B.Q. W-IQ-TREE: A fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 2016, 44, W232–W235. [Google Scholar] [CrossRef]
- National Toxicology Program. Nonneoplastic Lesion Atlas. Available online: https://ntp.niehs.nih.gov/atlas/nnl (accessed on 13 December 2024).
- Miao, J.; Chard, L.S.; Wang, Z.; Wang, Y. Syrian hamster as an animal model for the study on infectious diseases. Front. Immunol. 2019, 10, 2329. [Google Scholar] [CrossRef]
- Stokes, J.V.; Walker, D.H.; Varela-Stokes, A.S. The guinea pig model for tick-borne spotted fever rickettsioses: A second look. Ticks Tick-Borne Dis. 2020, 11, 101538. [Google Scholar] [CrossRef]
- Feng, H.; Wen, J.; Walker, D.H. Rickettsia australis infection: A murine model of a highly invasive vasculopathic rickettsiosis. Am. J. Pathol. 1993, 142, 1471. [Google Scholar]
- Londoño, A.F.; Mendell, N.L.; Walker, D.H.; Bouyer, D.H. A biosafety level-2 dose-dependent lethal mouse model of spotted fever rickettsiosis: Rickettsia parkeri Atlantic Rainforest strain. PLoS Negl. Negl. Trop. Dis. 2019, 13, E0007054. [Google Scholar] [CrossRef]
- Simser, J.A.; Palmer, A.T.; Fingerle, V.; Wilske, B.; Kurtti, T.J.; Munderloh, U.G. Rickettsia monacensis sp. nov., a spotted fever group Rickettsia, from ticks (Ixodes ricinus) collected in a European city park. Appl. Environ. Microbiol. 2002, 68, 4559–4566. [Google Scholar] [CrossRef]
- McDade, J.E. Evidence supporting the hypothesis that rickettsial virulence factors determine the severity of spotted fever and typhus group infections. Ann. N. Y. Acad. Sci. 1990, 590, 20–26. [Google Scholar] [CrossRef]
- Blanton, L.S.; Mendell, N.L.; Walker, D.H.; Bouyer, D.H. “Rickettsia amblyommii” induces cross protection against lethal Rocky Mountain spotted fever in a guinea pig model. Vector-Borne Zoonotic Dis. 2014, 14, 557–562. [Google Scholar] [CrossRef] [PubMed]
- Burgdorfer, W.; Hayes, S.F.; Thomas, L.A.; Lancaster, J.L. New Spotted Fever Group Rickettsia from the Lone Star Tick, Amblyomma americanum; Rickettsiae and Rickettsial Diseases; Burgdorfer, W., Anacker, R.L., Eds.; Academic Press: Academic Press, NY, USA, 1981. [Google Scholar]
- Philip, R.N.; Casper, E.A.; Anacker, R.L.; Cory, J.; Hayes, S.F.; Burgdorfer, W.; Yunker, C.E. Rickettsia bellii sp. nov.: A tick-borne rickettsia, widely distributed in the United States, that is distinct from the spotted fever and typhus biogroups. Int. J. Syst. Evol. Microbiol. 1983, 33, 94–106. [Google Scholar] [CrossRef]
- Horta, M.C.; Sabatini, G.S.; Moraes-Filho, J.; Ogrzewalska, M.; Canal, R.B.; Pacheco, R.C.; Martins, T.F.; Matushima, E.R.; Labruna, M.B. Experimental infection of the opossum Didelphis aurita by Rickettsia felis, Rickettsia bellii, and Rickettsia parkeri and evaluation of the transmission of the infection to ticks Amblyomma cajennense and Amblyomma dubitatum. Vector-Borne Zoonotic Dis. 2010, 10, 959–967. [Google Scholar] [CrossRef] [PubMed]
- de Sousa, R.; dos Santos, M.L.; Cruz, C.; Almeida, V.; Garrote, A.R.; Ramirez, F.; Seixas, D.; Manata, M.J.; Maltez, F. Rare case of rickettsiosis caused by Rickettsia monacensis, Portugal, 2021. Emerg. Infect. Dis. 2022, 28, 1068. [Google Scholar] [CrossRef]
- Kleba, B.; Clark, T.R.; Lutter, E.I.; Ellison, D.W.; Hackstadt, T. Disruption of the Rickettsia rickettsii Sca2 autotransporter inhibits actin-based motility. Infect. Immun. 2010, 78, 2240–2247. [Google Scholar] [CrossRef]
- Krawczak, F.S.; Binder, L.C.; Gregori, F.; Martins, T.F.; Pádua, G.T.; Sponchiado, J.; Melo, G.L.; Polo, G.; Labruna, M.B. ‘Candidatus Rickettsia andeanae’ and probable exclusion of Rickettsia parkeri in ticks from dogs in a natural area of the Pampa biome in Brazil. Pathogens 2023, 12, 446. [Google Scholar] [CrossRef]
- Vélez, J.C.Q.; Aguirre-Acevedo, D.C.; Rodas, J.D.; Arboleda, M.; Troyo, A.; Aguilar, F.V.; Quintero, L.O.; Arbeláez, C.R. Epidemiological characterization of incident cases of Rickettsia infection in rural areas of Uraba region, Colombia. PLoS Negl. Trop. Dis. 2018, 12, E0006911. [Google Scholar] [CrossRef]
- Delisle, J.; Mendell, N.L.; Stull-Lane, A.; Bloch, K.C.; Bouyer, D.H.; Moncayo, A.C. Human infections by multiple spotted fever group rickettsiae in Tennessee. Am. J. Trop. Med. Hyg. 2016, 94, 1212. [Google Scholar] [CrossRef]
- Vaughn, M.F.; Delisle, J.; Johnson, J.; Daves, G.; Williams, C.; Reber, J.; Mendell, N.L.; Bouyer, D.H.; Nicholson, W.L.; Moncayo, A.C.; et al. Seroepidemiologic study of human infections with spotted fever group rickettsiae in North Carolina. J. Clin. Microbiol. 2014, 52, 3960–3966. [Google Scholar] [CrossRef]
- Biggs, H.M. Diagnosis and management of tickborne rickettsial diseases: Rocky Mountain spotted fever and other spotted fever group rickettsioses, ehrlichioses, and anaplasmosis—United States. MMWR. Recomm. Rep. 2016, 65, 1–44. [Google Scholar] [CrossRef]
- Gage, K.L.; Jerrells, T.R. Demonstration and partial characterization of antigens of Rickettsia rhipicephali that induce cross-reactive cellular and humoral immune responses to Rickettsia rickettsii. Infect. Immun. 1992, 60, 5099–5106. [Google Scholar] [CrossRef]



| Presumptive Infection Rickettsia/(n=) /Country | Rickettsia Cross-Reaction | Samples with Cross-Reaction (%) | Confirmed by Cross-Adsorption and Western Blot | References |
|---|---|---|---|---|
| R. rickettsii/(n = 76) /USA | R. parkeri-R. montanensis-R. amblyommatis-R. rickettsii | 66 | 35 R. amblyommii, 1 R. montanensis, 1 R. parkeri 39 indeterminate 0 R. rickettsii | [65] |
| R. montanensis, R. parkeri, R. amblyommatis | 9 | |||
| R. parkeri, R. amblyommatis | 7 | |||
| R. rickettsii, R. montanensis, R. parkeri | 3 | |||
| R. rickettsii, R. montanensis | 2 | |||
| R. montanensis, R. amblyommatis | 2 | |||
| R. rickettsii, R. montanensis, R. amblyommatis | 1 | |||
| R. rickettsii, R. parkeri, R. amblyommatis | 1 | |||
| R. amblyommatis | 5 | |||
| SFG rickettsiosis/(n = 17) /Colombia | R. rickettsii, R. amblyommatis, R. felis, R. parkeri, R. rhipicephali, R. belli | 14 | ND | [66] |
| R. rickettsii, R. amblyommatis, R. felis, R. belli | 3 | |||
| R. rickettsii/(n = 21) /United States | R. rickettsii R. parkeri R. amblyommatis | 7 | 2 R. parkeri 2 R. rickettsii 1 R. amblyommii 2 indeterminate | [67] |
| R. parkeri y R. amblyommatis | 3 | |||
| R. rickettsii | 1 | |||
| R. parkeri | 4 | |||
| R. amblyommatis | 6 | |||
| R. rickettsii/(n = 30)/Colombia | R. rickettsii, R. parkeri, Ca. R. colombiensis | 30 | ND | Present study |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Miranda, J.; García, A.; Cervera-Acevedo, C.; Santibañez, S.; Portillo, A.; Oteo, J.A.; Mattar, S. Assessment of the Pathogenicity of Candidatus Rickettsia Colombiensis in a Syrian Hamster Model and Serological Cross-Reactivity Between Spotted Fever Rickettsia Species. Pathogens 2026, 15, 146. https://doi.org/10.3390/pathogens15020146
Miranda J, García A, Cervera-Acevedo C, Santibañez S, Portillo A, Oteo JA, Mattar S. Assessment of the Pathogenicity of Candidatus Rickettsia Colombiensis in a Syrian Hamster Model and Serological Cross-Reactivity Between Spotted Fever Rickettsia Species. Pathogens. 2026; 15(2):146. https://doi.org/10.3390/pathogens15020146
Chicago/Turabian StyleMiranda, Jorge, Alejandra García, Cristina Cervera-Acevedo, Sonia Santibañez, Aránzazu Portillo, José A. Oteo, and Salim Mattar. 2026. "Assessment of the Pathogenicity of Candidatus Rickettsia Colombiensis in a Syrian Hamster Model and Serological Cross-Reactivity Between Spotted Fever Rickettsia Species" Pathogens 15, no. 2: 146. https://doi.org/10.3390/pathogens15020146
APA StyleMiranda, J., García, A., Cervera-Acevedo, C., Santibañez, S., Portillo, A., Oteo, J. A., & Mattar, S. (2026). Assessment of the Pathogenicity of Candidatus Rickettsia Colombiensis in a Syrian Hamster Model and Serological Cross-Reactivity Between Spotted Fever Rickettsia Species. Pathogens, 15(2), 146. https://doi.org/10.3390/pathogens15020146










