Torque Teno Virus (TTV) Plasma Load and Immune Reconstitution Post-Transplantation in Patients with Lymphoproliferative Disorders: A Systematic Review
Abstract
1. Introduction
2. Methods
2.1. Eligibility Criteria
2.2. Information Sources and Search Strategy
2.3. Selection and Data Collection Process
2.4. Data Items
2.5. Assessment of Risk of Bias
3. Results
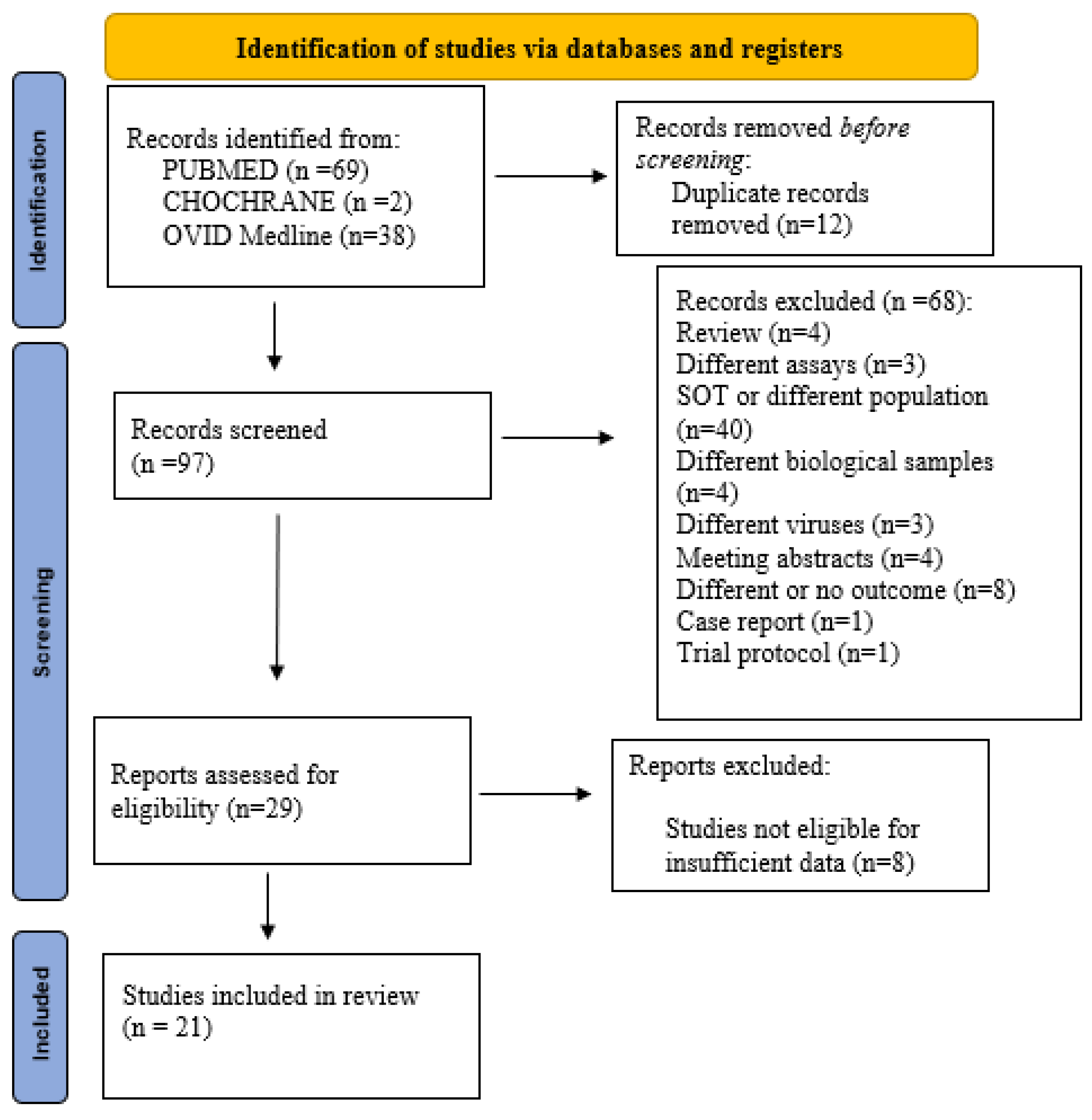
3.1. Study Selection and Search Results
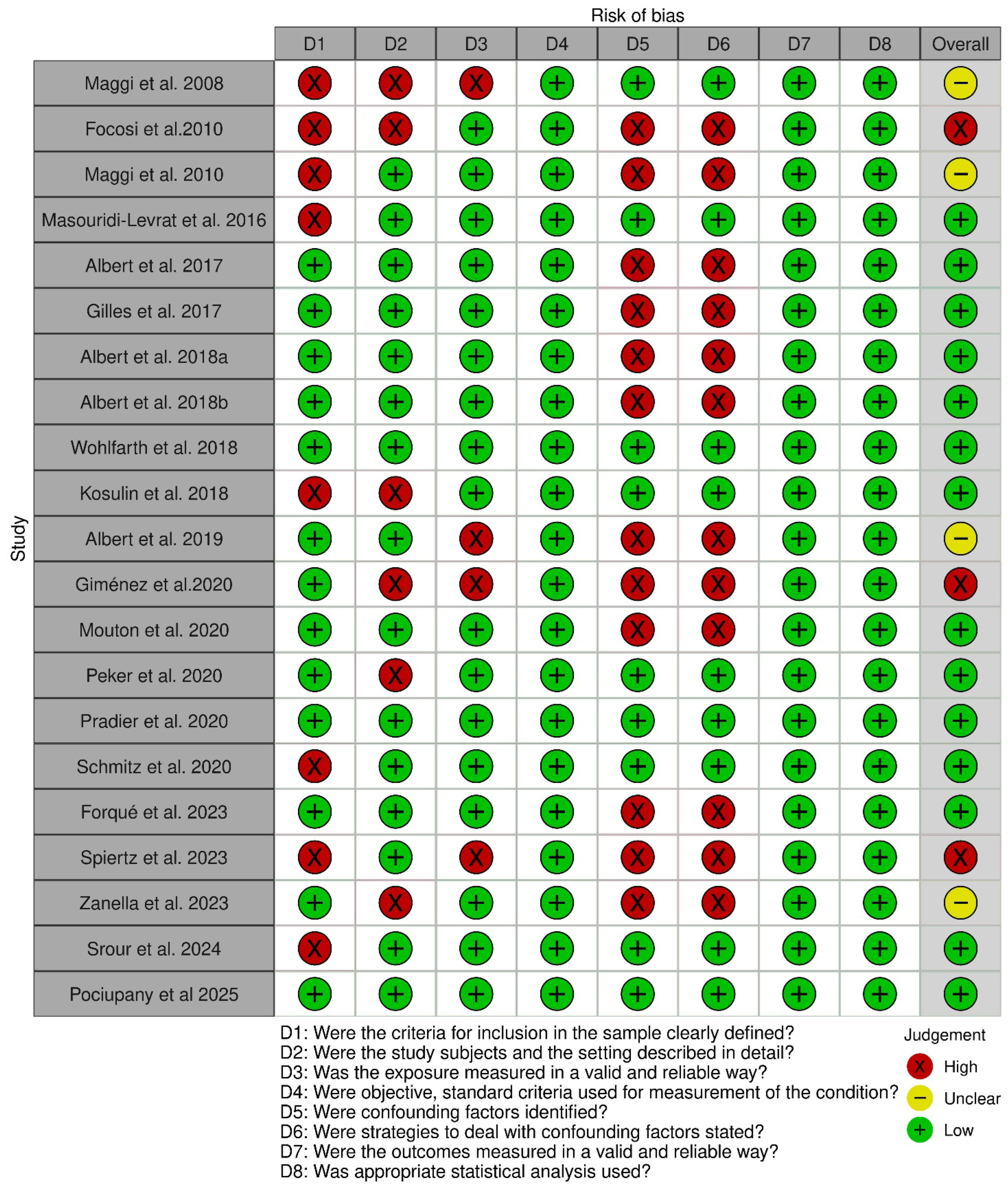
3.2. Quality Assessment of the Articles
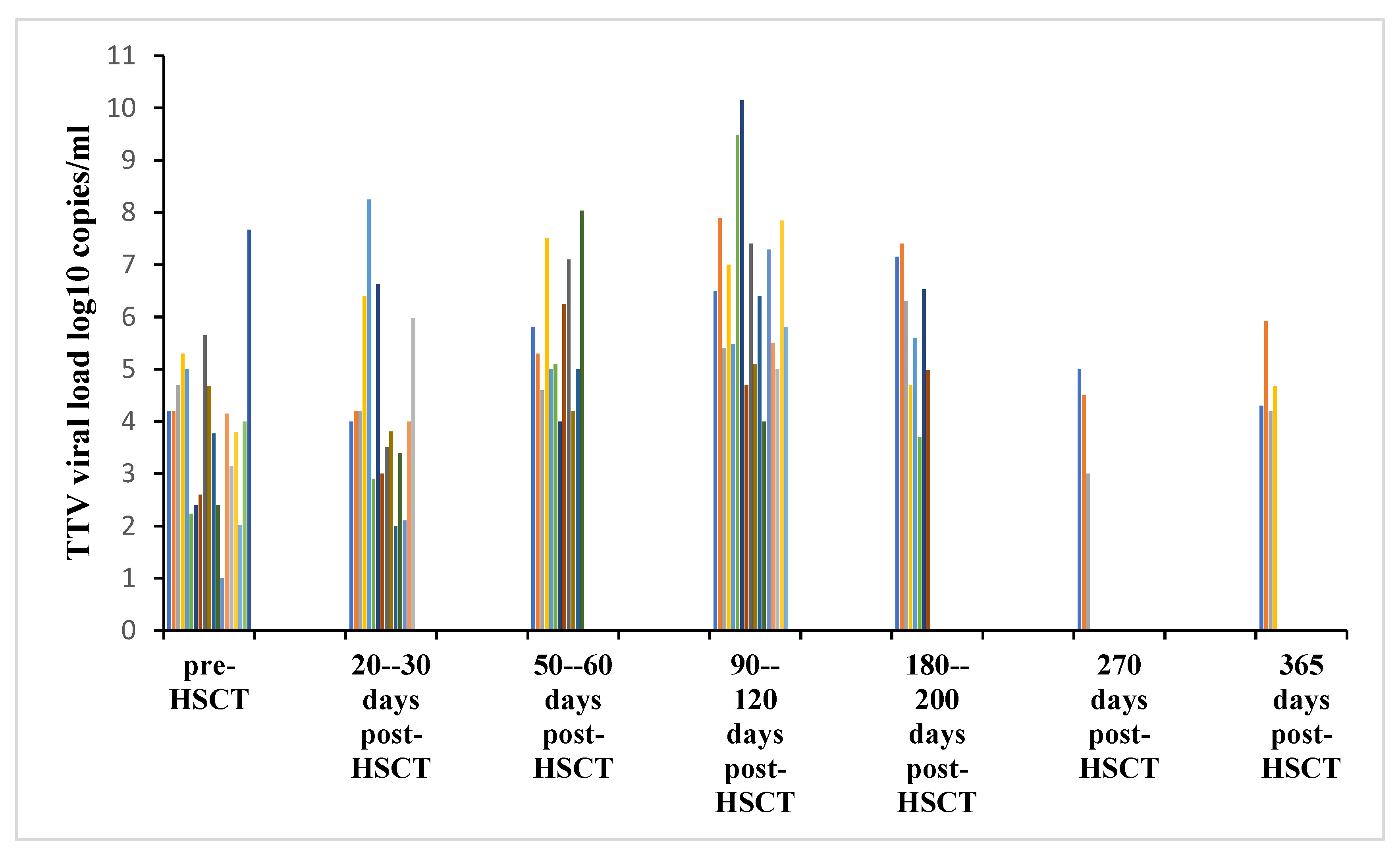
3.3. Dynamics of TTV Viral Load Before and After HSCT
3.4. TTV Viral Load and Immune Reconstitution
3.5. TTV Viral Load, Opportunistic Viral Infections, and GVHD
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Nishizawa, T.; Okamoto, H.; Konishi, K.; Yoshizawa, H.; Miyakawa, Y.; Mayumi, M. A Novel DNA Virus (TTV) Associated with Elevated Transaminase Levels in Posttransfusion Hepatitis of Unknown Etiology. Biochem. Biophys. Res. Commun. 1997, 241, 92–97. [Google Scholar] [CrossRef]
- De Vlaminck, I.; Khush, K.K.; Strehl, C.; Kohli, B.; Luikart, H.; Neff, N.F.; Okamoto, J.; Snyder, T.M.; Cornfield, D.N.; Nicolls, M.R.; et al. Temporal Response of the Human Virome to Immunosuppression and Antiviral Therapy. Cell 2013, 155, 1178–1187. [Google Scholar] [CrossRef] [PubMed]
- Sabbaghian, M.; Gheitasi, A.A.; Shekarchi, A.A.; Tavakoli, A.; Pootahmasebi, V. The mysterious anelloviruses: Investigating its role in human diseases. BMC Microbiol. 2024, 24, 40. [Google Scholar] [CrossRef] [PubMed]
- Okamoto, H. History of discoveries and pathogenicity of TT viruses. Curr. Top. Microbiol. Immunol. 2009, 331, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Varsani, A.; Opriessnig, T.; Celer, V.; Maggi, F.; Okamoto, H.; Blomström, A.L.; Cadar, D.; Harrach, B.; Biagini, P.; Kraberger, S. Taxonomic update for mammalian anelloviruses (family Anelloviridae). Arch. Virol. 2021, 166, 2943–2953. [Google Scholar] [CrossRef]
- Takahashi, K.; Iwasa, Y.; Hijikata, M.; Mishiro, S. Identification of a new human DNA virus (TTV-like mini virus, TLMV) intermediately related to TT virus and chicken anemia virus. Arch. Virol. 2000, 145, 979–993. [Google Scholar] [CrossRef]
- Ninomiya, M.; Nishizawa, T.; Takahashi, M.; Lorenzo, F.R.; Shimosegawa, T.; Okamoto, H. Identification and genomic characterization of a novel human torque teno virus of 3.2 kb. J. Gen. Virol. 2007, 88, 1939–1944. [Google Scholar] [CrossRef]
- Spandole, S.; Cimponeriu, D.; Berca, L.M.; Mihaescu, G. Human anelloviruses: An update of molecular, epidemiological and clinical aspects. Arch. Virol. 2015, 160, 893–908. [Google Scholar] [CrossRef]
- Väisänen, E.; Kuisma, I.; Mäkinen, M.; Ilonen, J.; Veijola, R.; Toppari, J.; Hedman, K.; Söderlund-Venermo, M. Torque teno virus primary infection kinetics in early childhood. Viruses 2022, 14, 1277. [Google Scholar] [CrossRef]
- Tyschik, E.; Rasskazova, A.; Degtyareva, A.V.; Rebrikov, D.V.; Sukhikh, G. Torque teno virus dynamics during the first year of life. Virol. J. 2018, 15, 94–96. [Google Scholar] [CrossRef]
- Goens, M.; Mouton, W.; Oriol, G.; Cheynet, V.; Restier, L.; Bal, A.; Brengel-Pesce, K.; Rozières, A.; Viret, C.; Nancey, S.; et al. Deep characterization of circulating Torque teno virus DNA load in Chron’s disease patients. J. Med. Virol. 2025, 97, e70473. [Google Scholar] [CrossRef] [PubMed]
- Goto, K.; Sugiyama, K.; Ando, T.; Mizutani, F.; Terabe, K.; Tanaka, K.; Nishiyama, M.; Wada, Y. Detection rates of TT virus DNA in serum of umbilical cord blood, breast milk and saliva. Tohoku J. Exp. Med. 2000, 191, 203–207. [Google Scholar] [CrossRef] [PubMed]
- Maillard, A.; Feghoul, L.; Lemiale, V.; Mercier-Delarue, S.; Demoule, A.; Jaber, S.; Klouche, K.; Kouatchet, A.; Argaud, L.; Barbier, F.; et al. Clinical significance of respiratory Torque Teno virus in immunocompromised patients with acute respiratory failure. Clin. Microbiol. Infect. 2025, 31, 2073–2081. [Google Scholar] [CrossRef] [PubMed]
- van Rijn, A.; Wunderink, H.; Sidorov, I.; de Brouwer, C.; Kroes, A.; Putter, H.; de Vries, A.P.; Rotmans, J.I.; Feltkamp, M.C. Torque teno virus loads after kidney transplantation predict allograft rejection but not viral infection. J. Clin. Virol. 2021, 140, 104871. [Google Scholar] [CrossRef]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 statement: An updated guideline for reporting systematic reviews. BMJ 2021, 372, n71. [Google Scholar] [CrossRef]
- Tufanaru, C.; Munn, Z.; Aromataris, E.; Campbell, J.; Hopp, L. Chapter3: Systematic reviews of effectiveness. In JBJ Manual for Evidence Synthesis; Joanna Briggs Institute: North Adelaide, SA, Australia, 2020. [Google Scholar]
- McGuinness, L.A.; Higgins, J.P.T. Risk of bias VISualization (robvis): An R package and Shiny web app for visualization risk of bias assessments. Res. Synth. Methods 2021, 12, 55–61. [Google Scholar] [CrossRef]
- Maggi, F.; Ricci, V.; Bendinelli, M.; Ceccherini-Nelli, L. Changes in CD8+57+ T lymphocyte expansions after autologous hematopoietic stem cell transplantation correlate with changes in Torquetenovirus viremia. Transplantation 2008, 85, 1867–1868. [Google Scholar] [CrossRef]
- Focosi, D.; Maggi, F.; Albani, M.; Macera, L.; Ricci, V.; Gragnani, S.; Di Beo, S.; Ghimenti, M.; Antonelli, G.; Bendinelli, M.; et al. Torquetenovirus viremia kinetics after autologous stem cell transplantation are predictable and may serve as a surrogate marker of functional immune reconstitution. J. Clin. Virol. 2010, 47, 189–192. [Google Scholar] [CrossRef]
- Maggi, F.; Focosi, D.; Albani, M.; Lanini, L.; Vatteroni, M.L.; Petrini, M.; Ceccherini-Nelli, L.; Pistello, M.; Bendinelli, M. Role of hematopoietic cells in the maintenance of chronic human torquetenovirus plasma viremia. J. Virol. 2010, 84, 6891–6893. [Google Scholar] [CrossRef]
- Masouridi-Levrat, S.; Pradier, A.; Simonetta, F.; Kaiser, L.; Chalandon, Y.; Roosnek, E. Torque teno virus in patients undergoing allogeneic hematopoietic stem cell transplantation for hematological malignancies. Bone Marrow Transplant. 2016, 51, 440–442. [Google Scholar] [CrossRef]
- Albert, E.; Solano, C.; Pascual, T.; Torres, I.; Macera, L.; Focosi, D.; Maggi, F.; Giménez, E.; Amat, P.; Navarro, D. Dynamics of Torque Teno virus plasma DNAemia in allogeneic stem cell transplant recipients. J. Clin. Virol. 2017, 94, 22–28. [Google Scholar] [CrossRef] [PubMed]
- Gilles, R.; Herling, M.; Holtick, U.; Heger, E.; Awerkiew, S.; Fish, I.; Höller, K.; Sierra, S.; Knops, E.; Kaiser, R.; et al. Dynamics of Torque Teno virus viremia could predict risk of complications after allogeneic hematopoietic stem cell transplantation. Med. Microbiol. Immunol. 2017, 206, 355–362. [Google Scholar] [CrossRef] [PubMed]
- Albert, E.; Solano, C.; Giménez, E.; Focosi, D.; Pérez, A.; Macera, L.; Piñana, J.L.; Boluda, J.C.H.; Maggi, F.; Navarro, D. The kinetics of torque teno virus plasma DNA load shortly after engraftment predicts the risk of high-level CMV DNAemia in allogeneic hematopietic stem cell transplant recipients. Bone Marrow Transplant. 2018, 53, 180–187. [Google Scholar] [CrossRef] [PubMed]
- Albert, E.; Torres, I.; Talaya, A.; Giménez, E.; Piñana, J.L.; Hernández-Boluda, J.C.; Focosi, D.; Macera, L.; Maggi, F.; Solano, C.; et al. Kinetics of torque teno virus DNA load in saliva and plasma following allogeneic hematopoietic stem cell transplantation. J. Med. Virol. 2018, 90, 1438–1443. [Google Scholar] [CrossRef]
- Wohlfarth, P.; Leiner, M.; Schoergenhofer, C.; Hopfinger, G.; Goerzer, I.; Puchhammer-Stoeckl, E.; Rabitsch, W. Torquetenovirus dynamics and immune marker properties in patients following allogeneic hematopoietic stem cell transplantation: A prospective longitudinal study. Biol. Blood Marrow Transplant. 2018, 24, 185–199. [Google Scholar] [CrossRef]
- Albert, E.; Solano, C.; Giménez, E.; Focosi, D.; Pérez, A.; Macera, L.; Piñana, J.L.; Mateo, E.V.; Hernández-Boluda, J.C.; Maggi, F.; et al. Kinetics of Alphatorquevirus plasma DNAemia at late times after allogeneic hematopoietic stem cell transplantation. Med. Microbiol. Immunol. 2019, 208, 253–258. [Google Scholar] [CrossRef]
- Giménez, E.; Monzó, C.; Albert, E.; Fuentes-Trillo, A.; Seda, E.; Piñana, J.L.; Hernández-Boluda, J.C.; Solano, C.; Chaves, J.; Navarro, D. Diversity and dynamic change of anelloviruses in plasma following allogeneic hematopoietic stem cell transplantation. J. Med. Virol. 2021, 93, 5167–5172. [Google Scholar] [CrossRef]
- Mouton, W.; Conrad, A.; Bal, A.; Boccard, M.; Malcus, C.; Ducastelle-Lepretre, S.; Balsat, M.; Barraco, F.; Larcher, M.V.; Fossard, G.; et al. Torque teno virus viral load as a marker of immune function in allogeneic hematopoietic stem cell transplantation recipients. Viruses 2020, 12, 1292. [Google Scholar] [CrossRef]
- Pradier, A.; Masouridi-Levrat, S.; Bosshard, C.; Dantin, C.; Vu, D.L.; Zanella, M.C.; Boely, E.; Tapparel, C.; Kaiser, L.; Chalandon, Y.; et al. Torque Teno Virus as a Potential Biomarker for Complications and Survival After Allogeneic Hematopoietic Stem Cell Transplantation. Front. Immunol. 2020, 11, 998. [Google Scholar] [CrossRef]
- Schmitz, J.; Kobbe, G.; Kondakci, M.; Schuler, E.; Magorsch, M.; Adams, O. The value of Torque teno virus (TTV) as a marker for the degree of immunosuppression in adult patients after hematopoietic stem cell transplantation (HSCT). Biol. Blood Marrow Transplant. 2020, 26, 643–650. [Google Scholar] [CrossRef]
- Forqué, L.; Albert, E.; Piñana, J.L.; Pérez, A.; Hernani, R.; Solano, C.; Navarro, D.; Giménez, E. Monitoring of plasma Torque teno virus, total Anelloviridae and Human Pegivirus 1 viral load for the prediction of infectious events and acute graft versus host disease in the allogeneic stem cell transplantation setting. J. Med. Virol. 2023, 95, e29107. [Google Scholar] [CrossRef] [PubMed]
- Spiertz, A.; Tsakmaklis, A.; Farowski, F.; Knops, E.; Heger, E.; Wirtz, M.; Kaiser, R.; Holtick, U.; Vehreschild, M.J.G.T.; Di Cristanziano, V. Torque teno virus-DNA load as individual cytomegalovirus risk assessment parameter upon allogeneic hematopoietic stem cell transplantation. Eur. J. Haematol. 2023, 111, 963–969. [Google Scholar] [CrossRef] [PubMed]
- Zanella, M.C.; Vu, D.L.; Hosszu-Fellous, K.; Neofytos, D.; Van Delde, C.; Turin, L.; Poncet, A.; Simonetta, F.; Masouridi-Levrat, S.; Chalandon, Y.; et al. Longitudinal Detection of Twenty DNA and RNA Viruses in Allogeneic Hematopoietic Stem Cell Transplant Recipients Plasma. Viruses 2023, 15, 928. [Google Scholar] [CrossRef] [PubMed]
- Srour, M.; Grenier, C.; Magro, L.; Hober, D.; Yakoub-Agha, I.; Engelmann, I. High Torque teno virus load and outcome of patients undergoing allogeneic hematopoietic cell transplantation. J. Med. Virol. 2024, 96, e29458. [Google Scholar] [CrossRef]
- Pociupany, M.; Tarabella, C.; Snoeck, R.; Dierickx, D.; Andrei, G. Viral Infections in HSCT Recipients with Post-Transplant Lymphoproliferative Disorder: The Role of Torque Teno Virus as a Marker of Immune Functions. Microorganisms 2025, 13, 326. [Google Scholar] [CrossRef]
- Kosulin, K.; Kernbichler, S.; Pichler, H.; Lawitschka, A.; Geyeregger, R.; Witt, V.; Lion, T. Post-transplant Replication of Torque Teno Virus in Granulocytes. Front. Microbiol. 2018, 9, 2956. [Google Scholar] [CrossRef]
- Peker, B.O.; Daloğlu, A.E.; Görzer, I.; Purchhammer-Stöckl, E.; Parkan, O.M.; Akbas, H.; Kintrup, G.T.; Mutlu, D.; Küpesiz, O.A.; Colak, D. Investigation of Torque teno virus (TTV) DNA as an immunological and virological marker in pediatric hematopoietic stem cell transplantation (HSCT) patients. Microb. Pathog. 2020, 149, 104397. [Google Scholar] [CrossRef]
- Medina, J.B.; França Vieira e Silva, F.; Veloso Caixeta, R.A.; de Oliveira Rech, B.; Perez-Jardón, A.; Padin-Iruega, M.E.; Pérez-Sayans, M.; Braz-Silva, P.H.; Ortega, K.L. Torque teno virus as a marker of immune status in immunocompromised patients: A systematic review. Eur. J. Clin. Investig. 2025, 55, e70068. [Google Scholar] [CrossRef]
- Focosi, D.; Antonelli, G.; Pistello, M.; Maggi, F. Torquetenovirus: The human virome from bench to bedside. Clin. Microbiol. Infect. 2016, 22, 589–593. [Google Scholar] [CrossRef]
- Gorzer, I.; Jaksch, P.; Kundi, M.; Seitz, T.; Klepetko, W.; Puchhammer-Stockl, E. Pre-transplant plasma Torque Teno virus load and increase dynamics after lung transplantation. PLoS ONE 2015, 10, e0122975. [Google Scholar] [CrossRef]
- Beland, K.; Dore-Nguyen, M.; Gagne, M.J.; Patey, N.; Brassard, J.; Alvarez, F.; Halac, U. Torque Teno Virus in children who underwent orthotopic liver transplantation: New insights about a common pathogen. J. Infect. Dis. 2014, 209, 247–254. [Google Scholar] [CrossRef]
- Focosi, D.; Macera, L.; Pistello, M.; Maggi, F. Torquetenovirus (TTV) viremia correlates with intensity of maintenance immunosuppression in adult orthotopic liver transplantation. J. Infect. Dis. 2014, 43. 210, 667–668. [Google Scholar] [CrossRef]
- Gorzer, I.; Haloschan, M.; Jaksch, P.; Klepetko, W.; Puchhammer-Stockl, E. Plasma DNA levels of Torque teno virus and immunosuppression after lung transplantation. J. Heart Lung Transpl. 2014, 33, 320–328. [Google Scholar] [CrossRef] [PubMed]
- Schiemann, M.; Puchhammer-Stöckl, E.; Eskandary, F.; Kohlbeck, P.; Rasoul-Rockenschaub, S.; Heilos, A.; Kozakowski, N.; Görzer, I.; Kikić, Ž.; Herkner, H.; et al. Torque teno virus load-inverse association with antibody-mediated rejection after kidney transplantation. Transplantation 2017, 101, 360–367. [Google Scholar] [CrossRef] [PubMed]
- Simonetta, F.; Pradier, A.; Masouridi-Levrat, S.; van Delden, C.; Giostra, E.; Morard, I.; Mueller, N.; Muellhaupt, B.; Valli, P.V.; Semmo, N.; et al. Torque teno virus load and acute rejection after orthotopic liver transplantation. Transplantation 2017, 101, e219–e221. [Google Scholar] [CrossRef] [PubMed]
- Jaksch, P.; Kundi, M.; Görzer, I.; Muraközy, G.; Lambers, C.; Benazzo, A.; Hoetzenecker, K.; Klepetko, W.; Puchhammer-Stöckl, E. Torque teno virus as a novel biomarker targeting the efficacy of immunosuppression after lung transplantation. J. Infect. Dis. 2018, 218, 1922–1928. [Google Scholar] [CrossRef]
- Zanotta, N.; Maximova, N.; Campisciano, G.; Del Savio, R.; Pizzol, A.; Casalicchio, G.; Berton, E.; Comar, M. Up-regulation of the monocyte chemotactic protein-3 in sera from bone marrow transplanted children with torquetenovirus infection. J. Clin. Virol. 2015, 63, 6–11. [Google Scholar] [CrossRef]
- Velardi, E.; Clave, E.; Arruda, L.C.M.; Benini, F.; Locatelli, F.; Toubert, A. The role of the thymus in allogeneic bone marrow transplantation and the recovery of the peripheral T-cell compartment. Semin. Immunopathol. 2021, 43, 101–117. [Google Scholar] [CrossRef]
- Ando, T.; Tachibana, T.; Tanaka, M.; Suzuki, T.; Ishiyama, Y.; Koyama, S.; Ogusa, E.; Numata, A.; Matsumoto, K.; Kanamori, H.; et al. Impact of graft sources on immune reconstitution and survival outcomes following allogeneic stem cell transplantation. Blood Adv. 2020, 4, 408–419. [Google Scholar] [CrossRef]
- Yanir, A.; Schulz, A.; Lawitschka, A.; Nierkens, S.; Eyrich, M. Immune Reconstitution After Allogeneic Haematopoietic Cell Transplantation: From Observational Studies to Targeted Interventions. Front. Pediatr. 2022, 9, 786017. [Google Scholar] [CrossRef]
- Borkosky, S.S.; Whitley, C.; Kopp-Schneider, A.; zur Hausen, H.; de Villiers, E.M. Epstein-Barr virus stimulates torque teno virus replication: A possible relationship to multiple sclerosis. PLoS ONE 2012, 7, 32160. [Google Scholar] [CrossRef]
- Haloschan, M.; Bettesch, R.; Görzer, I.; Weseslindtner, L.; Kundi, M.; Puchhammer-Stöckl, E. TTV DNA plasma load and its association with age, gender, and HCMV IgG serostatus in healthy adults. Age 2014, 36, 9716. [Google Scholar] [CrossRef]
- Alhadid, Y.; Mamroth, M.; Wang, Z.; Chadha, A.; Rumpler, M.; Vlassov, A.; Chikova, A. Development of a novel qPCR test for monitoring immunuppression levels by quantifying torque teno virus (TTV) in whole blood. Hum. Immunol. 2025, 86, 111348. [Google Scholar] [CrossRef]


| Source | Year | Country | Study Design | Patient Number | Male, n (%) | Patient Age, Median (Range) |
Clinical
Endpoint | Donors (n) | Stem Cell Source (n) | Diagnosis (n) | Conditioning | Prophylaxis (n) | Reference |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Maggi et al. | 2008 | Italy | Longitudinal | 19 | NA | NA | IR | NA | PB (15) | MM (19) | MAC | NA | [18] |
| Focosi et al. | 2010 | Italy | Longitudinal | 47 | NA | NA | IR | Related | PB (47) | MM (36), LY (10), AML (1) | MAC (37), HCDT (10) | NA | [19] |
| Maggi et al. | 2010 | Italy | Longitudinal | 4 | 1 (25) | 50 (41–58) | IR | NA | PB (4) | T-ALL (1), B-ALL (1), ALL (1), AML (1) | NA | NA | [20] |
| Masouridi-Levrat et al. | 2016 | Switzerland | Longitudinal | 121 | 67 (55) | 50 (18–70) | IR | MMURD (10), MMRD (7), MSD (29), MUD (37) | PB (121) | AML (58), ALL (15), MDS (12), NHL (10), MPS (6), MM (9), HL (5), CML (39, CLL (1), MDPS (1) | NA | NA | [21] |
| Albert et al. | 2017 | Spain | Longitudinal | 72 | 38 (53) | 54 (18–69) | aGVHD | Related (37); unrelated (35); matched (49); mismatched (23) | PB (68); BM (3); CB (1) | HL (5); NHL (15); CLL (6); ALL (7); AML (19); CML (1); MM (5); MDS (10); others (4) | MAC (13); NMA (59) | Steroids (45), ASCT (30), TKI (16) | [22] |
| Gilles et al. | 2017 | Germany | Longitudinal | 23 | 13 (57) | 57 (33–75) | IR | MRD (5); MUD (8); MMURD (9); haploidentical (1) | PB (23) | AML (11); ALL (1); CLL (1); CMML (1); others (9) | RIC (23) | CSA/MMF (18), CSA/MTX (2), ATG/CSA (1), CSA/TAC (1), MMF/TAC (1) | [23] |
| Albert et al. | 2018a | Spain | Longitudinal | 71 | 42 (59) | 55 (18–70) | OI | Related (39); unrelated (32); matched (50); mismatched (21) | PB (69); BM (1); CB (1) | HL (3); NHL (21); CLL (6); ALL (6); AML (15); MM (5); MDS (7); others (8) | MAC (13), NMA (58) | CSA or MTF/TAC (24), CSA or MMF/TAC (29), regimens including thymoglobulin (4), regimens including cyclophosphamide (14) | [24] |
| Albert et al. | 2018b | Spain | Longitudinal | 38 | 22 (58) | 55 (26–69) | IR | Related (22); unrelated (16); matched (26); mismatched (12) | PB (37), BM (1) | HL (1), NHL (14), ALL (4), CLL (3), AML (6), MM (2), MDS (6), others (2) | MAC (5); RIC (33) | CSA or MTF/TAC (13), CSA or MMF/TAC (18), regimens including cyclophosphamide (7) | [25] |
| Wohlfarth et al. | 2018 | Austria | Longitudinal | 50 | 32 (64) | 49 (37–57) | IR, OI, aGVHD | Unrelated (35), SIB (13), mismatched (18) | PB (45), CB (3), BM (2) | AML (25), ALL (9), MDS (6), NHL (3), others (7) | MAC (26), RIC (16), NMA (8) | CSA + MTX (26), CSA + MMF (24) | [26] |
| Albert et al. | 2019 | Germany | Longitudinal | 33 | 16 (48) | 56 (19–70) | IR | Related (18), unrelated (15) | PB (31), BM (1), CB (1) | Lymphoma (12), leukemia (12), myeloma (4), MDS (3), MF (2) | MAC (6), NMA (27) | CSA + MTX or MMF (19), TAC + SIRO (14) | [27] |
| Gimenez et al. | 2020 | Spain | Longitudinal | 25 | 14 (56) | 54 (24–69) | IR | MRD (7), MUD (8), MMRD (5), Haplo (5) | PB (23), BM (2) | Lymphoma (10), AML (4), ALL (4), MDS (2), CLL (3), MM (2) | NA | NA | [28] |
| Mouton et al. | 2020 | France | Cross-sectional | 41 | 25 (61) | 56 (40–64) | IR, OI | MRD (23), MUD (15), MMURD (3) | PB (28), BM (13) | Myeloid neoplasm and acute leukemia (37), others (4) | MAC (17), RIC (24) | 100% but not reported specifically for each patient | [29] |
| Pradier et al. | 2020 | Switzerland | Longitudinal | 168 | 104 (62) | 51 (39–59) | IR, aGVHD | SIB (71), MUD (75), MMUD (13), Haplo (9) | PB (149), BM (19) | AML (78), ALL (17), MDS (22), MPS (11), lymphoma (12), myeloma (11) | RIC (85), MAC (83) | CSA + MMF (85), CSA + MTX (83) | [30] |
| Schmitz et al. | 2020 | Germany | Retrospective | 123 | 74 (60) | 54 (19–75) | IR, OI, aGVHD | Unrelated (90), related (33), matched (105), mismatched (18) | PB (116), BM (7) | AML (58), ALL (9), MDS (33), NHL (11), others (12) | MAC (48), RIC (75) | MMF + CSA (30), MMF + TAC (83) | [31] |
| Forqué et al. | 2023 | Spain | Retrospective | 75 | 43 (57) | 54 (19–70) | IR,OI, aGVHD | Related (46), unrelated (29), matched (41), mismatched (9), Haplo (25) | PB (75) | AML (28), ALL (1), CML (3), CLL (3), HL (13), MM (1), MDS (4), MF (4), NHL (15), others (3) | MAC (14), RIC (61) | SIRO + MMF + cyclophosphamide (71), TAC + SIRO (2), TAC + CP (2) | [32] |
| Spiertz et al. | 2023 | Germany | Retrospective | 59 | 33 (56) | 52 (19–74) | IR,OI | MUD (36), MRD (13), MMUD (7), MMRD haplo-identical (3) | NA | AML (31), ALL (6), CML (2), CLL (4), MDS (10), others (6) | RIC (54), MAC (5) | CSA + MMF or MTX (59) | [33] |
| Zanella et al. | 2023 | Switzerland | Longitudinal | 109 | 72 (66) | 56 (no range) | IR | MUD (47), MSD (28), Haplo (24), MMURD (10) | PB (96), BM (13) | AML (59), MDS/MDPS (27), ALL (10), MPS (4), lymphoma (4), CLL (2), myeloma (2), CML (1) | RIC (74), MAC (43) | NA | [34] |
| Srour et al. | 2024 | France | Longitudinal | 70 | 41 (59) | 54 (19–73) | IR, OI, aGVHD | MRD (14), MUD (40), MMURD (5), Haplo (11) | PB (37), BM (33) | HL (1), NHL (6), MM (2), ALL (13), AML (27), CML (1), MDS (13), MF (3), others (4) | MAC (36), RIC (34) | NA | [35] |
| Pociupany et al. | 2025 | Belgium | Longitudinal | 48 | 28 (58) | PTLD patients 52 (16–67), not PTLD 53 (0–68) | OI | MUD (27), MRD (15), Haplo (6), CB (1) | PB (40), BM (8) | HL (3), T-cell/NK-cell lymphoma (3), MCL (1), ALL (3), AML (18), MDS (5), MF (1), CML (1), MM (1), others (5), NA (7) | MAC (29), NMA (29) | ATG, CSA + MTX (7), ATG, CSA + MMF (1), CSA + MMF (4), MMF (1), CSA (2), CSA + CP (3), others (3), NA (27) | [36] |
| Kosulin et al. | 2018 | Italy | Longitudinal | 43 | NA | Pediatric patients (age NA) | IR | Unrelated (27), MSD (10), MMRD (6) | NA | HM (45), SCID (7), FAA (1) | MAC (22), RIC (21) | NA | [37] |
| Peker et al. | 2020 | Turkey | Retrospective | 33 | 19 (58) | 7.8 (0.7–18.6) | IR | MUD (19), MRD (11), autolog (2), Haplo (1) | PB (16), BM (16), CB + BM (1) | HLH (2), AML (7), ALL (5), TM (8), FAA (3), JMML (1), WAS (1), SCID (2), CN (2), NB (1) | MAC (21), NMA (12) | CSA + MTX (18), CSA + MMF (4), CSA (6), CSA + MMF + MTX (3), NA (2) | [38] |
| First Author, Year of Publication | Number of Patients Included in Follow Up | Control Group | Quantitative Method Used, Target Gene | TTV DNAemia * Pre-HSCT | TTV DNAemia * Post-HSCT (20–30 Days) | TTV DNAemia * Post-HSCT (50–60 Days) | TTV DNAemia * Post-HSCT (90–120 Days) | TTV DNAemia * Post-HSCT (180–200 Days) | TTV DNAemia * Post-HSCT (270 Days) | TTV DNAemia * Post-HSCT (365 Days) | ALCs in Correlation with TTV Load |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Maggi, 2008 [18] | 19 | No | RT PCR in-house, UTR, detection limit = NR | 4.2 | 4 | 5.8 | 6.5 | NA | NA | NA | Direct correlation (r = 0.049, p = 0.001) |
| Focosi, 2010 [19] | 47 | No | RT-PCR in-house, UTR detection limit = 100 copies/ mL | 4.2 | 4.2 | 5.3 | 7.9 | NA | NA | NA | Direct correlation (r = 0.062, p = 0.001) |
| Maggi, 2010 [20] | 3 | No | RT-PCR in-house, UTR detection limit = 100 copies/ mL | Pz n°1 = 4.7 Pz n°2 = 5.3 Pz n°3 = 5.0 | Pz n°2 = 4.2 | NA | Pz n°1 = 5.4 + 80 d Pz n°3 = 7.0 + 110 d | NA | NA | NA | Direct correlation (r not determined) |
| Masouridi-Levrat, 2016 [21] | 77 | Yes 74 HS | RT-PCR in-house, ORF2 detection limit = 25 copies/mL | HS = 2.23 Pz = 2.39 | NA | NA | 5.48 | NA | NA | NA | Not determined |
| Albert, 2017 [22] | 55 | No | RT-PCR in-house, UTR detection limit = 10 copies/mL | TTV DNA loads ranging from 1.40 to 7.97 in 32 Pz | After 30 days a median increase of 3.34 in 16 Pz | TTV DNA load continued to rise, with a median increase of 4.43 in 22 Pz. | TTV DNA load peaked, with a median increase of 5.02 in 19 Pz | NA | NA | NA | Direct correlation (r = 0.285, p = 0.032) |
| Gilles, 2017 [23] | 23 | Yes 16 HS | RT-PCR in-house, UTR detection limit = 100 copies/mL | HS BL = 5.08 LR Pz = 12 HR Pz = 11 TTV viral load baseline for Pz was not reported | 8 LR Pz = 6.40 11 HR Pz = 9.26 | NA | 10 LR Pz = 9.48 9 HR Pz = 10.15 | 12 LR Pz = 7.15 11 HR Pz = 7.40 | NA | NA | Direct correlation (r not determined) |
| Albert, 2018b [25] | Pre-transplant = 23 FU at +30 = 24 FU at +50 = 32 FU at +90 = 25 | No | RT-PCR in-house, UTR detection limit = 10 copies/mL | 2.6 = 20 Pz pos/23 | 2.9 = 16 Pz pos/24 | 4.6 = 30 Pz pos/32 | 4.7 = 24 Pz pos/25 | NA | NA | NA | Direct correlation (r = 0.317, p = 0.002) |
| Wohlfarth, 2018 [26] | 40 | No | RT-PCR in-house, UTR detection limit = 10 copies/mL | 5.65 | 6.63 | 7.50 | 7.40 | 6.31 | 5.0 | 4.68 | Inverse correlation (r = −0.27, p < 0.01) |
| Mouton, 2020 [29] | 41 | Yes 80 HS | TTV R-GENE® kit (BioMérieux, Marcy-l’Étoile, France ) (bioMérieux) Detection limit = 10 copies/mL | NA | NA | NA | NA | 3.1 = 41 Pz 2.1 = 54/80 HS | NA | NA | No correlation (r = −0.13, p = 0.42) |
| Albert, 2019 [27] | 33 | No | RT PCR in-house, UTR, detection limit = 10 copies/mL | 4.68 | 3 after 20 d 3.5 after 30 d | 5 | 5.1 | 4.7 | 4.5 | NA | Direct correlation until +60 days (r = 0.171, p = 0.031), then inverse correlation (r = −0.263, p = 0.003) |
| Giménez, 2020 [28] | 25 | No | RT-PCR in-house, UTR detection limit = 10 copies/mL | 3.77 | 3.81 | NA | NA | NA | NA | NA | Not determined |
| Pradier, 2020 [30] | 130 | Yes 91 HS | RT-PCR in-house, UTR detection limit = 25 copies/mL | 2.2 = 91 HS 2.4 = 130 Pz | NA | 5.1 = 124 Pz | 6.4 = 115 Pz | 5.6 = 95 Pz | 4.7 = 81 Pz | 4.3 = 64 Pz | Inverse correlation at day 100 (r = −0.271, p = 0.005) |
| Schmitz, 2020 [31] | 123 | No | RT-PCR in-house, UTR detection limit = 100 copies/mL | 1 = 62 Pz | ~2 § | ~4 § | ~4 § | ~3.7 § | ~3 § | NA | No correlation (r = 0.092, p not indicated) |
| Forqué, 2023 [32] | 75 | No | RT PCR in-house, UTR, detection limit = 10 copies/mL | 4.15 = 52 Pz | 3.40 = 64 Pz | 6.24 = 65 Pz | 7.29 = 61 Pz | 6.53 = 33 Pz | NA | NA | Not determined |
| Spiertz, 2023 [33] | 59 | No | RT PCR in-house, UTR, detection limit = 100 copies/mL | 3.14 | NA | 7.10 | NA | NA | NA | 5.92 | Not determined |
| Zanella, 2023 [34] | 109 | No | RT-PCR in-house, ORF2 Detection limit = 25 copies/mL | 3.8 = 42 Pz | NA | 4,2 | 5.51 = 93 Pz | 4.98 = 79 Pz | NA | 4.2 = 48 Pz | Not determined |
| Srour, 2023 [35] | 70 | No | TTV R-GENE® kit (bioMérieux) Detection limit = 10 copies/ mL | 2.02 | 2.1 | 5 | 5.8 after 90 d 5 after 120 d | NA | NA | NA | Not determined |
| Kosulin, 2018 [37] | 43 pediatric patients | No | RT-PCR in-house, UTR detection limit = 10 copies/mL | 4 | 4 | NA | 7.84 | NA | NA | NA | Direct correlation with granulocytes at +30 and +60 days (r = 0.494, p = 0.002) |
| Peker, 2020 [38] | 33 pediatric patients | Yes 38 HS | RT-PCR in-house, UTR detection limit = 100 copies/mL | 5.51 = HS 7.67 = Pz | 5.98 + 20 d | 8.03 | NA | NA | NA | NA | Direct correlation (r = 0.29, p = 0.001) |
| First Author, Year of Publication | TTV Viral Load * Post-HSCT in Patients Without OI | TTV Viral Load * Post-HSCT in Patients with OI | p | TTV Viral Load * Post-HSCT in Patients Without aGVHD | TTV Viral Load * Post-HSCT in Patients with aGVHD | p |
|---|---|---|---|---|---|---|
| Albert, 2017 [22] | Not determined | Not determined | NA | 4 after 60 days | 4.8 after 60 days | 0.09 |
| Gilles, 2017 [23] | 6.40 after 30 days | 9.26 after 30 days | 0.005 | <8.48 after 30 days was related to lower incidence of aGVHD | NA | |
| Albert, 2018a [24] | 4.4 for CMV risk AUCs20–30 7.10 for EBV risk AUCs20–50 | 3.3 for CMV risk AUCs20–30 6.67 for EBV risk AUCs20–50 | 0.123 0.31 | Not determined | Not determined | NA |
| Wohlfarth, 2018 [26] | 4.28 after 365 days | 5.40 after 365 days | 0.04 | 5.76 after 120 days 5.29 after 160 days | 7.59 after 120 days 6.96 after 160 days | 0.04 0.01 |
| Mouton, 2020 [29] | 3.2 for OI after 6 months 3.7 for CMV risk after 6 months | 4.1 for OI after 6 months 4.8 for CMV after 6 months | 0.02 0.02 | Not determined | Not determined | NA |
| Pradier, 2020 [30] | Higher TTV titers at day 100 had higher rates of infection | NA | 5.6 after 100 days | 6.9 after 100 days | 0.013 | |
| Schmitz, 2020 [31] | Virus reactivation > 1000 copies/mL was associated with a higher but not significant TTV viral load | >0.05 | The group of patients without GVHD vs. patients with an aGVHD of grade II or higher revealed no significant difference in TTV viral load | >0.05 | ||
| Forqué, 2023 [32] | 3.14 after 30 days | 3.80 after 30 days | 0.46 | 2.66 after 30 days | 4.53 after 30 days | 0.02 |
| Spiertz, 2023 [33] | 3.03 at early stage of HSCT | 4.40 at early stage of HSCT | NA | Not determined | Not determined | NA |
| Srour, 2023 [35] | No difference in TTV viral load between the two groups of patients | NA | 4.97 after 60 days | 6.18 after 60 days | 0.02 | |
| Pociupany, 2025 [36] | 4.94 after 90 days | 7.56 after 90 days | 0.059 | Not determined | Not determined | NA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Quiros-Roldan, E.; Salvi, M.; Alberti, M.; Tiecco, G.; Biasiotto, G.; Bresciani, R.; Bertoli, D.; Sottini, A.; De Francesco, M.A. Torque Teno Virus (TTV) Plasma Load and Immune Reconstitution Post-Transplantation in Patients with Lymphoproliferative Disorders: A Systematic Review. Pathogens 2026, 15, 105. https://doi.org/10.3390/pathogens15010105
Quiros-Roldan E, Salvi M, Alberti M, Tiecco G, Biasiotto G, Bresciani R, Bertoli D, Sottini A, De Francesco MA. Torque Teno Virus (TTV) Plasma Load and Immune Reconstitution Post-Transplantation in Patients with Lymphoproliferative Disorders: A Systematic Review. Pathogens. 2026; 15(1):105. https://doi.org/10.3390/pathogens15010105
Chicago/Turabian StyleQuiros-Roldan, Eugenia, Martina Salvi, Maria Alberti, Giorgio Tiecco, Giorgio Biasiotto, Roberto Bresciani, Diego Bertoli, Alessandra Sottini, and Maria Antonia De Francesco. 2026. "Torque Teno Virus (TTV) Plasma Load and Immune Reconstitution Post-Transplantation in Patients with Lymphoproliferative Disorders: A Systematic Review" Pathogens 15, no. 1: 105. https://doi.org/10.3390/pathogens15010105
APA StyleQuiros-Roldan, E., Salvi, M., Alberti, M., Tiecco, G., Biasiotto, G., Bresciani, R., Bertoli, D., Sottini, A., & De Francesco, M. A. (2026). Torque Teno Virus (TTV) Plasma Load and Immune Reconstitution Post-Transplantation in Patients with Lymphoproliferative Disorders: A Systematic Review. Pathogens, 15(1), 105. https://doi.org/10.3390/pathogens15010105








