Potential Use of Biosensors for the Rapid and Specific Isolation of Listeria monocytogenes from Ready-to-Eat (RTE) Foods
Abstract
1. Introduction
2. Materials and Methods
2.1. Reagents and Chemicals
2.2. Primer Design and Biosensor Probe
2.3. Bacterial Culture
2.4. Spiking of PBS and Salad Samples to Determine the Limit of Detection (LoD) of MNPs and GNPs

2.5. Capture Efficiency of MNPs
2.6. Analytical Specificity of GNP Biosensor for Detecting L. monocytogenes
2.7. Assessment of MNP and GNP Biosensor Combination in Naturally Contaminated Food Samples
2.8. Data Analysis
3. Results
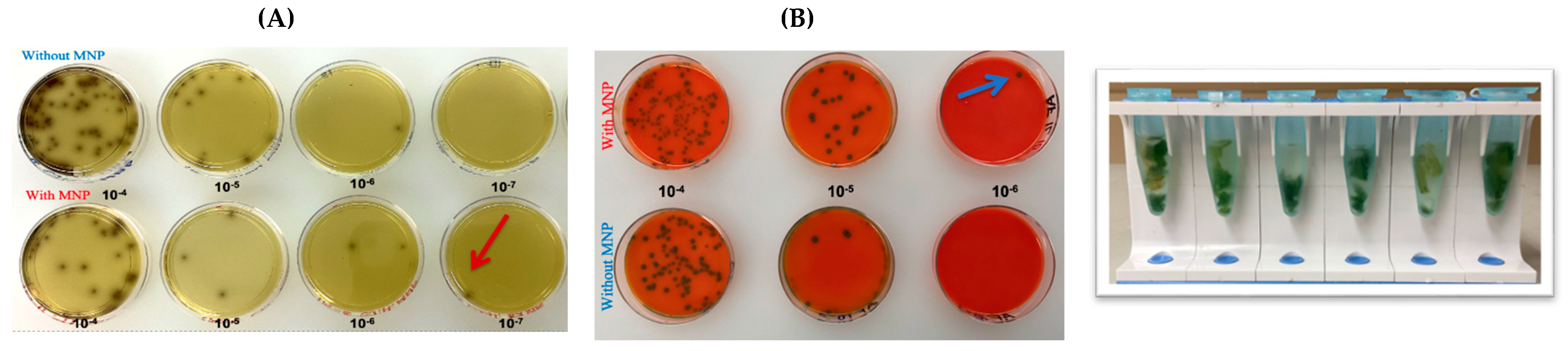
3.1. Spiking of PBS and Salad Samples to Determine the LoD of MNPs and GNPs
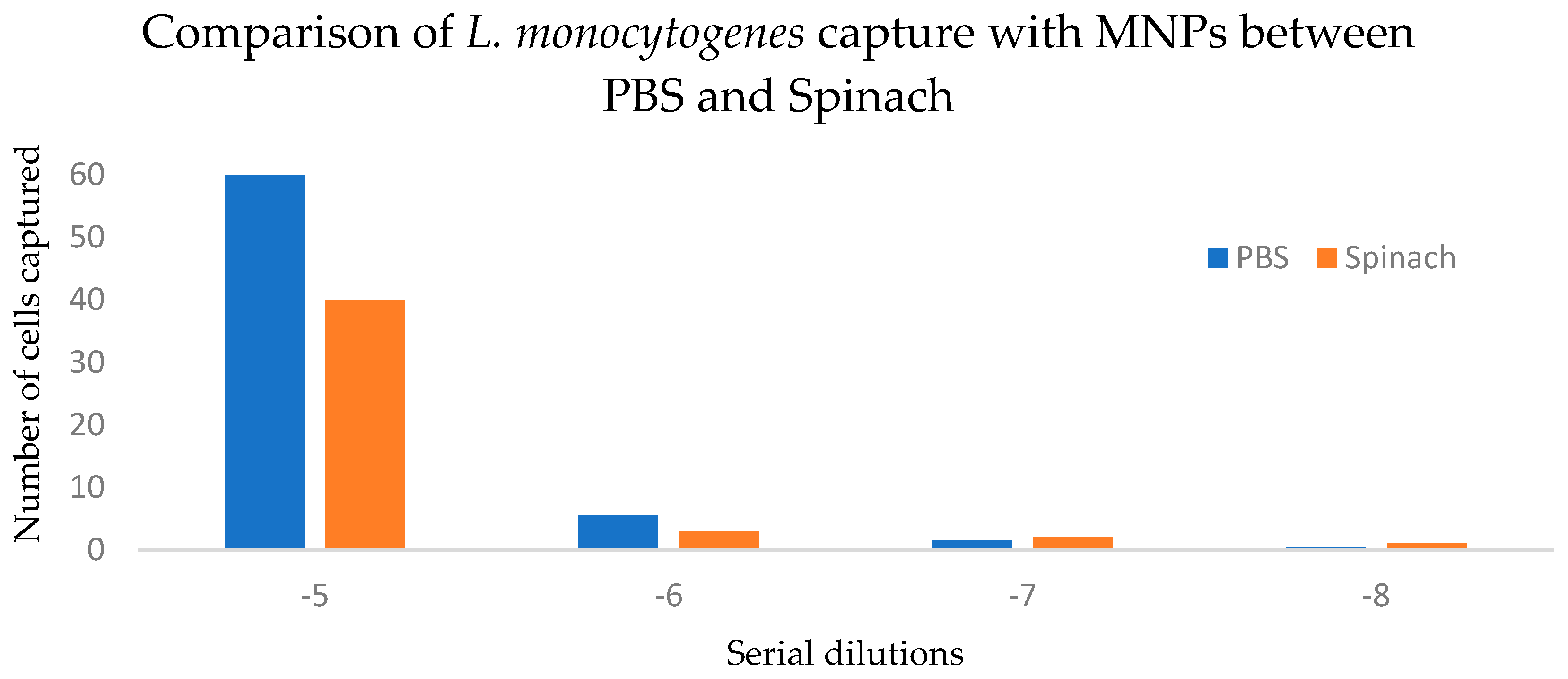
3.2. MNPs Capture Efficiency Results
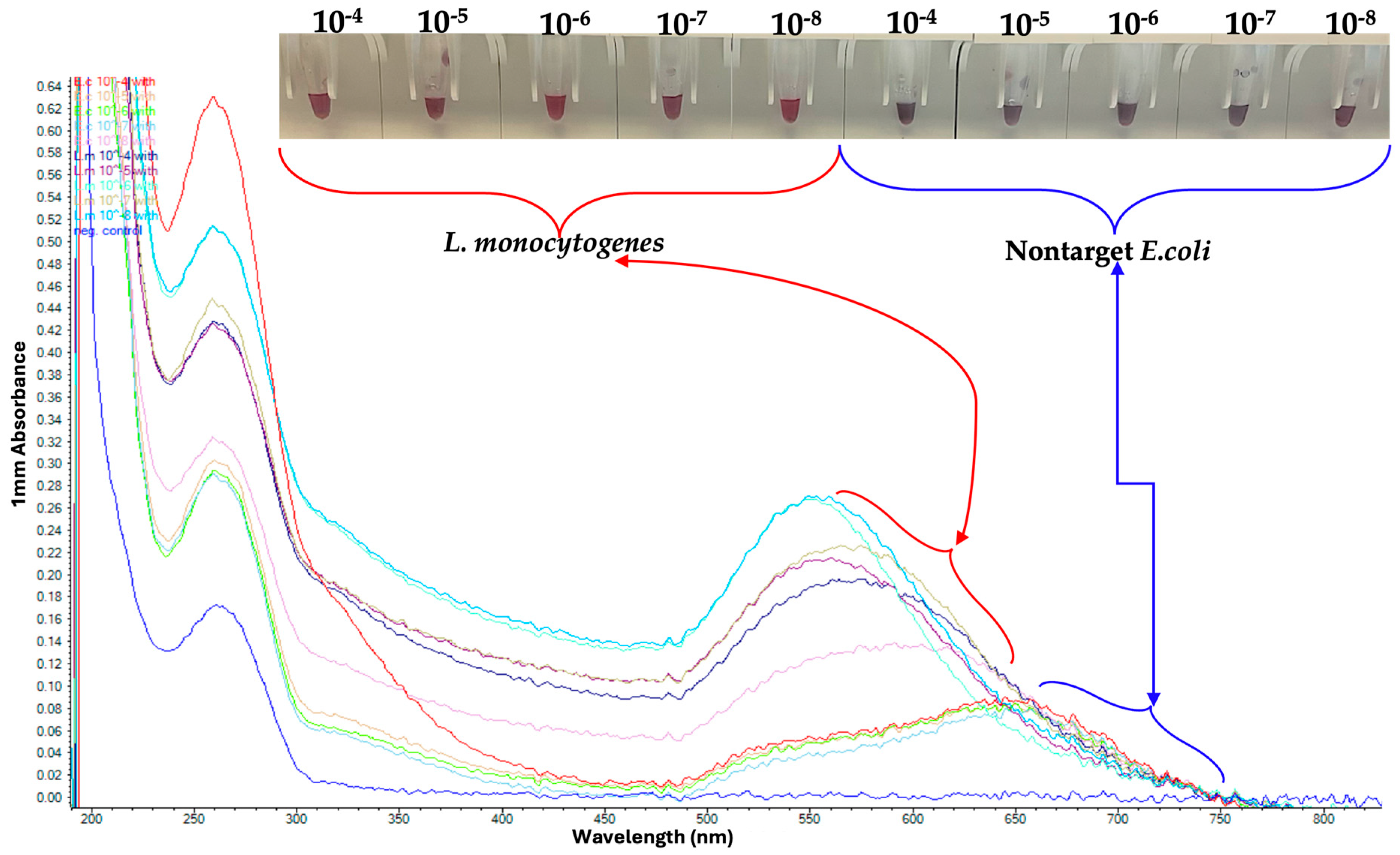
3.3. Analytical Sensitivity of GNP Biosensor for Detecting L. monocytogenes
3.4. Observed Analytical Specificity of GNP Biosensor for Detecting L. monocytogenes
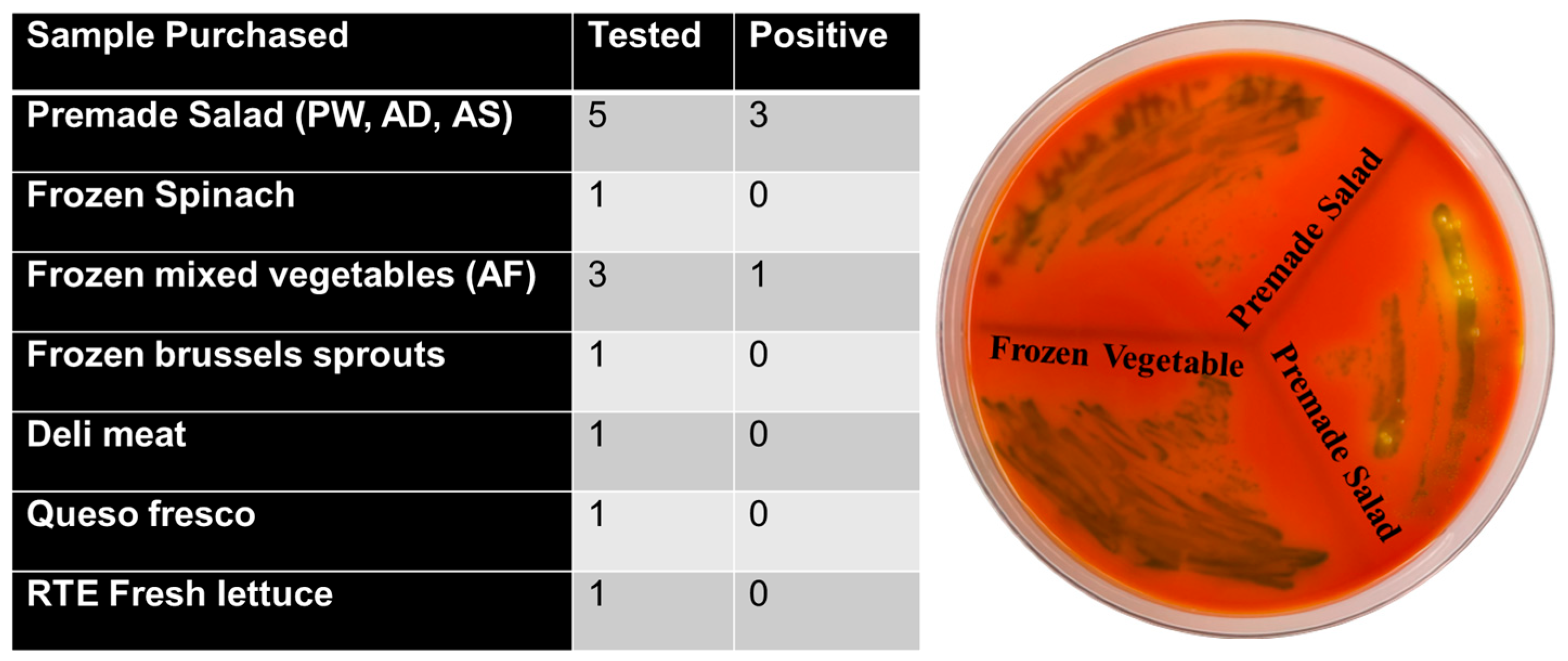
3.5. Assessment of MNP and GNP Biosensors in Naturally Contaminated Food Samples
4. Discussion
| Recognition Molecule/Nanomaterial | Sample Matrix | Detection Time | LOD | Method Type | Ref. |
|---|---|---|---|---|---|
| Glycan-coated MNPs + dextrin-capped GNPs (This study) | RTE foods | 3 h | 1.5 CFU/mL (MNP); 2.5 ng/µL DNA (GNP) | Magnetic capture + plasmonic colorimetric | This study |
| CRISPR/Cas12a (RAA-CRISPR platform) | Pure culture, genomic DNA | 20–30 min | 350 CFU/mL; 5.4 × 10−3 ng/µL | CRISPR fluorescence | [51] |
| High-resolution melting (HRM) qPCR | Roast pork | 3–6 h | 6.2 × 103–6.2 × 104 CFU/mL | mPCR + HRM qPCR | [52] |
| PEG-cefepime MNPs | Lettuce, juices, meat | 110 min | 3.1 × 102 CFU/mL | Antibiotic MNP + colorimetry | [44] |
| Antibody-ZIF-8 GOD@ZIF-8@Ab | Juice | NS | 101 CFU/mL | Colorimetry | [53] |
| Teicoplanin MNPs | PBS, ground beef | NS | 2.6 × 101 CFU/mL | Fluorescence | [54] |
| Ampicillin-MNPs + qPCR | Milk | 2.5 h | 102 CFU/mL | qPCR | [55] |
| Aptamer-linked AuNP + MNP | Milk | NS | 6 CFU/mL | Colorimetric immunoassay | [56] |
| Immunomagnetic beads + SERS | Milk | NS | 12 CFU/mL | SERS | [57] |
| D-amino acid MNPs | Milk, meat | 30 s | 2.17 × 102 CFU/mL | Colorimetric | [58] |
| Antibody–MNP + nitrocellulose | Vegetables | 35 min | 1 × 102 CFU/g | Colorimetry | [59] |
| Fe3O4@silica antibody NPs | Pure culture | 30 min | NS | Magnetic capture | [60] |
| AuNP + Ag nanoclusters + aptamer MNPs | Food | NS | 10 CFU/mL | Colorimetric | [61] |
| Impedance immunosensor + MNPs | Lettuce, milk, beef | 3 h | 104 CFU/mL | Impedance | [62] |
| Aptamer-MNP + AIE fluorescence | Spiked samples | NS | 10 CFU/mL | Fluorescence | [63] |
| Vancomycin-PEG-MNPs + PCR | PBS, lettuce | <4 h | 30 CFU/g | PCR | [64] |
| Fe/Fe3O4 NPs + antibodies (NMR) | Milk powder, lettuce | NS | 3 MPN | NMR | [16] |
| CRISPR/Cas12a electrochemical | Plant samples | 2 h | 0.68 aM; 940 CFU/g | Electrochemical | [36] |
| Mesoporous silica microarray | Food samples | 2 h | 102 CFU/mL | Microarray | [65] |
| Aptamer-MNP + AuNP amplification | Meat, milk | 1.5 h | 10 CFU/mL | Colorimetric | [66] |
| AuNP-SD-PMA-qPCR | Milk | 6 h | 5 × 101 CFU/g | qPCR + viability dye | [67] |
| Aptamer-based MNP system | Food | 18 h | 103 CFU/mL | Fluorescence | [68] |
| Boronate affinity magnetic + fluorescence | Lettuce | 35 min | 2.2 × 101 CFU/mL | Fluorescence | [69] |
| Cefepime-PEG-MNPs + colorimetry | Lettuce, juice, meat | 100 min | 3.1 × 102 CFU/mL | Colorimetric | [44] |
| G-quadruplex DNAzyme colorimetric | Pork | 4 h | 3.1 CFU/mL | Colorimetric | [70] |
| Lateral flow strip (end-on mAb) | Blood, milk, mushrooms | 15 min | 101–104 CFU/mL | Lateral flow | [71] |
| QCM aptasensor + MNPs | Milk, cheese, meats, vegetables | 10 min | 148 CFU/mL | QCM | [72] |
| LAMP electrochemical sensor | Multiple foods | 30 min (post enrichment) | 1 CFU/25g | LAMP + electrochemical | [73] |
| CPA isothermal amplification | Rice flour | 60 min | 104 CFU/mL | CPA | [74] |
| AlphaLISA nucleic acid assay | Milk, juice | NS | 250 attomole | AlphaLISA | [75] |
Limitations and Future Directions
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Centers for Disease Control and Prevention. The Listeria Initiative 2024. Available online: https://www.cdc.gov/listeria/php/surveillance/listeria-initiative.html (accessed on 12 June 2024).
- Byrd-Bredbenner, C.; Berning, J.; Martin-Biggers, J.; Quick, V. Food safety in home kitchens: A synthesis of the literature. Int. J. Environ. Res. Public Health 2013, 10, 4060–4085. [Google Scholar] [CrossRef]
- Thomsen, M.R.; Shiptsova, R.; Hamm, S.J. Sales Responses to Recalls for Listeria monocytogenes: Evidence from Branded Ready--to-Eat Meats. Appl. Econ. Perspect. Policy 2006, 28, 482–493. [Google Scholar] [CrossRef]
- Newell, D.G.; Koopmans, M.; Verhoef, L.; Duizer, E.; Aidara-Kane, A.; Sprong, H.; Opsteegh, M.; Langelaar, M.; Threfall, J.; Scheutz, F.; et al. Food-borne diseases—The challenges of 20 years ago still persist while new ones continue to emerge. Int. J. Food Microbiol. 2010, 139, S3–S15. [Google Scholar] [CrossRef]
- Gahan, C.G.; Hill, C. Listeria monocytogenes: Survival and adaptation in the gastrointestinal tract. Front. Cell. Infect. Microbiol. 2014, 4, 9. [Google Scholar] [CrossRef]
- Välimaa, A.-L.; Tilsala-Timisjärvi, A.; Virtanen, E. Rapid detection and identification methods for Listeria monocytogenes in the food chain—A review. Food Control 2015, 55, 103–114. [Google Scholar] [CrossRef]
- Jordan, K.; McAuliffe, O. Chapter Seven-Listeria monocytogenes in Foods. In Advances in Food and Nutrition Research; Rodríguez-Lázaro, D., Ed.; Academic Press: Amsterdam, The Netherlands, 2018; Volume 86, pp. 181–213. [Google Scholar]
- Ooi, S.T.; Lorber, B. Gastroenteritis due to Listeria monocytogenes. Clin. Infect. Dis. 2005, 40, 1327–1332. [Google Scholar] [CrossRef]
- Chen, Y.; Ross, W.H.; Scott, V.N.; Gombas, D.E. Listeria monocytogenes: Low levels equal low risk. J. Food Prot. 2003, 66, 570–577. [Google Scholar] [CrossRef] [PubMed]
- McLauchlin, J.; Mitchell, R.; Smerdon, W.; Jewell, K. Listeria monocytogenes and listeriosis: A review of hazard characterisation for use in microbiological risk assessment of foods. Int. J. Food Microbiol. 2004, 92, 15–33. [Google Scholar] [CrossRef] [PubMed]
- Willis, C.; McLauchlin, J.; Aird, H.; Amar, C.; Barker, C.; Dallman, T.; Elviss, N.; Lai, S.; Sadler-Reeves, L. Occurrence of Listeria and Escherichia coli in frozen fruit and vegetables collected from retail and catering premises in England 2018–2019. Int. J. Food Microbiol. 2020, 334, 108849. [Google Scholar] [CrossRef]
- U.S. Centers for Disease Control and Prevention. Multistate Outbreak of Listeriosis Linked to Frozen Vegetables. 2016. Available online: https://archive.cdc.gov/www_cdc_gov/listeria/outbreaks/frozen-vegetables-05-16/index.html (accessed on 16 July 2024).
- Madad, A.; Marshall, K.E.; Blessington, T.; Hardy, C.; Salter, M.; Basler, C.; Conrad, A.; Stroika, S.; Luo, Y.; Dwarka, A.; et al. Investigation of a multistate outbreak of Listeria monocytogenes infections linked to frozen vegetables produced at individually quick-frozen vegetable manufacturing facilities. J. Food Prot. 2023, 86, 100117. [Google Scholar] [CrossRef] [PubMed]
- Fay, M.L.; Salazar, J.K.; Stewart, D.S.; Khouja, B.A.; Zhou, X.; Datta, A.R. Survival of Listeria monocytogenes on Frozen Vegetables during Long-term Storage at −18 and −10 °C. J. Food Prot. 2024, 87, 100224. [Google Scholar] [CrossRef] [PubMed]
- Datta, A.R.; Burall, L.S. Diagnosis of Pathogenic Microorganisms Causing Infectious Diseases, 1st ed.; Taylor & Francis Ltd: Oxford, UK, 2024. [Google Scholar]
- Zhao, Y.; Li, Y.; Jiang, K.; Wang, J.; White, W.L.; Yang, S.; Lu, J. Rapid detection of Listeria monocytogenes in food by biofunctionalized magnetic nanoparticle based on nuclear magnetic resonance. Food Control 2017, 71, 110–116. [Google Scholar] [CrossRef]
- Jasson, V.; Jacxsens, L.; Luning, P.; Rajkovic, A.; Uyttendaele, M. Alternative microbial methods: An overview and selection criteria. Food Microbiol. 2010, 27, 710–730. [Google Scholar] [CrossRef]
- Rohde, A.; Hammerl, J.A.; Boone, I.; Jansen, W.; Fohler, S.; Klein, G.; Dieckmann, R.; Danhouk, S.A. Overview of validated alternative methods for the detection of foodborne bacterial pathogens. Trends Food Sci. Technol. 2017, 62, 113–118. [Google Scholar] [CrossRef]
- Janudin, A.A.S.; Lim, Y.C.; Ahmed, M.U. Chapter 8—Strategies and challenges of CRISPR/Cas system in detecting foodborne pathogens. In Biosensors for Foodborne Pathogens Detection; Pal, M.K., Ahmed, M.U., Campbell, K., Eds.; Academic Press: Amsterdam, The Netherlands, 2024; pp. 161–193. [Google Scholar]
- Basak, S.; Venkatram, R.; Singhal, R.S. Recent advances in the application of molecularly imprinted polymers (MIPs) in food analysis. Food Control 2022, 139, 109074. [Google Scholar] [CrossRef]
- Puiu, M.; Bala, C. Microfluidics-integrated biosensing platforms as emergency tools for on-site field detection of foodborne pathogens. TRAC Trends Anal. Chem. 2020, 125, 115831. [Google Scholar] [CrossRef]
- Kendall, H.; Clark, B.; Rhymer, C.; Kuznesof, S.; Hajslova, J.; Tomaniova, M.; Brereton, P.; Frewer, L. A systematic review of consumer perceptions of food fraud and authenticity: A European perspective. Trends Food Sci. Technol. 2019, 94, 79–90. [Google Scholar] [CrossRef]
- Xiao, F.; Li, W.; Xu, H. Advances in magnetic nanoparticles for the separation of foodborne pathogens: Recognition, separation strategy, and application. Compr. Rev. Food Sci. Food Saf. 2022, 21, 4478–4504. [Google Scholar] [CrossRef]
- Augustine, R.; Abraham, A.R.; Kalarikkal, N.; Thomas, S. 9-Monitoring and separation of food-borne pathogens using magnetic nanoparticles. In Novel Approaches of Nanotechnology in Food; Grumezescu, A.M., Ed.; Academic Press: Amsterdam, The Netherlands, 2016; pp. 271–312. [Google Scholar]
- Paniel, N.; Noguer, T. Detection of Salmonella in food matrices, from conventional methods to recent aptamer-sensing technologies. Foods 2019, 8, 371. [Google Scholar] [CrossRef]
- Zhang, Y.; Xu, C.-Q.; Guo, T.; Hong, L. An automated bacterial concentration and recovery system for pre-enrichment required in rapid Escherichia coli detection. Sci. Rep. 2018, 8, 17808. [Google Scholar] [CrossRef]
- Wang, Z.; Cai, R.; Gao, Z.; Yuan, Y.; Yue, T. Immunomagnetic separation: An effective pretreatment technology for isolation and enrichment in food microorganisms detection. Compr. Rev. Food Sci. Food Saf. 2020, 19, 3802–3824. [Google Scholar] [CrossRef] [PubMed]
- Dwivedi, H.P.; Jaykus, L.-A. Detection of pathogens in foods: The current state-of-the-art and future directions. Crit. Rev. Microbiol. 2011, 37, 40–63. [Google Scholar] [CrossRef] [PubMed]
- Sharief, S.A.; Caliskan-Aydogan, O.; Alocilja, E. Carbohydrate-coated magnetic and gold nanoparticles for point-of-use food contamination testing. Biosens. Bioelectron. X 2023, 13, 100322. [Google Scholar] [CrossRef]
- Matta, L.L.; Alocilja, E.C. Carbohydrate ligands on magnetic nanoparticles for centrifuge-free extraction of pathogenic contaminants in pasteurized milk. J. Food Prot. 2018, 81, 1941–1949. [Google Scholar] [CrossRef]
- You, S.-M.; Jeong, K.-B.; Luo, K.; Park, J.-S.; Park, J.-W.; Kim, Y.-R. Paper-based colorimetric detection of pathogenic bacteria in food through magnetic separation and enzyme-mediated signal amplification on paper disc. Anal. Chim. Acta 2021, 1151, 338252. [Google Scholar] [CrossRef]
- Yosief, H.O.; Weiss, A.A.; Iyer, S.S. Capture of Uropathogenic, E. coli by Using Synthetic Glycan Ligands Specific for the Pap-Pilus. ChemBioChem 2013, 14, 251–259. [Google Scholar] [CrossRef]
- El-Boubbou, K.; Gruden, C.; Huang, X. Magnetic Glyco-nanoparticles: A Unique Tool for Rapid Pathogen Detection, Decontamination, and Strain Differentiation. J. Am. Chem. Soc. 2007, 129, 13392–13393. [Google Scholar] [CrossRef]
- Matta, L.L.; Karuppuswami, S.; Chahal, P.; Alocilja, E.C. AuNP-RF sensor: An innovative application of RF technology for sensing pathogens electrically in liquids (SPEL) within the food supply chain. Biosens. Bioelectron. 2018, 111, 152–158. [Google Scholar] [CrossRef]
- Ghazy, A.; Nyarku, R.; Faraj, R.; Bentum, K.; Woube, Y.; Williams, M.; Alocilja, E.; Abebe, W. Gold Nanoparticle-Based Plasmonic Detection of Escherichia coli, Salmonella enterica, Campylobacter jejuni, and Listeria monocytogenes from Bovine Fecal Samples. Microorganisms 2024, 12, 1069. [Google Scholar] [CrossRef]
- Li, F.; Ye, Q.; Chen, M.; Zhou, B.; Zhang, J.; Pang, R.; Xue, L.; Wang, J.; Zeng, H.; Wu, S.; et al. An ultrasensitive CRISPR/Cas12a based electrochemical biosensor for Listeria monocytogenes detection. Biosens. Bioelectron. 2021, 179, 113073. [Google Scholar] [CrossRef]
- Myndrul, V.; Yanovska, A.; Babayevska, N.; Korniienko, V.; Diedkova, K.; Jancelewicz, M.; Pogorielov, M.; Iatsunskyi, I. 1D ZnO–Au nanocomposites as label-free photoluminescence immunosensors for rapid detection of Listeria monocytogenes. Talanta 2024, 271, 125641. [Google Scholar] [CrossRef] [PubMed]
- Karnwal, A.; Kumar Sachan, R.S.; Devgon, I.; Devgon, J.; Pant, G.; Panchpuri, M.; Ahmad, A.; Alshammari, M.B.; Hossain, K.; Kumar, G. Gold Nanoparticles in Nanobiotechnology: From Synthesis to Biosensing Applications. ACS Omega 2024, 9, 29966–29982. [Google Scholar] [CrossRef]
- Arcos Rosero, W.A.; Bueno Barbezan, A.; Daruich de Souza, C.; Chuery Martins Rostelato, M.E. Review of Advances in Coating and Functionalization of Gold Nanoparticles: From Theory to Biomedical Application. Pharmaceutics 2024, 16, 255. [Google Scholar] [CrossRef]
- Dester, E.; Alocilja, E. Current methods for extraction and concentration of foodborne bacteria with glycan-coated magnetic nanoparticles: A review. Biosensors 2022, 12, 112. [Google Scholar] [CrossRef]
- Quintela, I.A.; de Los Reyes, B.G.; Lin, C.-S.; Wu, V.C. Simultaneous colorimetric detection of a variety of Salmonella spp. in food and environmental samples by optical biosensing using oligonucleotide-gold nanoparticles. Front. Microbiol. 2019, 10, 1138. [Google Scholar] [CrossRef]
- Farber, J.M.; Ross, W.H.; Harwig, J. Health risk assessment of Listeria monocytogenes in Canada. Int. J. Food Microbiol. 1996, 30, 145–156. [Google Scholar] [CrossRef] [PubMed]
- Norton, D.M. Polymerase Chain Reaction-Based Methods for Detection of Listeria monocytogenes: Toward Real-Time Screening for Food and Environmental Samples. J. AOAC Int. 2019, 85, 505–515. [Google Scholar] [CrossRef]
- Xiao, F.; Wang, Z.; Li, W.; Qi, W.; Bai, X.; Xu, H. Cefepime-modified magnetic nanoparticles and enzymatic colorimetry for the detection of Listeria monocytogenes in lettuces. Food Chem. 2023, 409, 135296. [Google Scholar] [CrossRef]
- Łepecka, A.; Zielińska, D.; Szymański, P.; Buras, I.; Kołożyn-Krajewska, D. Assessment of the Microbiological Quality of Ready-to-Eat Salads—Are There Any Reasons for Concern about Public Health? Int. J. Environ. Res. Public Health 2022, 19, 1582. [Google Scholar] [CrossRef]
- Ricci, A.; Allende, A.; Bolton, D.; Chemaly, M.; Davies, R.; Fernández Escámez, P.S.; Snary, E.; Threlfall, J.; Arcella, D.; Wagner, M.; et al. Listeria monocytogenes contamination of ready-to-eat foods and the risk for human health in the EU. EFSA J. 2018, 16, e05134. [Google Scholar]
- Osek, J.; Lachtara, B.; Wieczorek, K. Listeria monocytogenes in foods-From culture identification to whole-genome characteristics. Food Sci. Nutr. 2022, 10, 2825–2854. [Google Scholar] [CrossRef]
- Kim, D.H.; Chon, J.W.; Kim, H.; Kim, H.S.; Choi, D.; Kim, Y.J.; Yim, J.-H.; Moon, J.S.; Seo, K.H. Comparison of Culture, Conventional and Real-time PCR Methods for Listeria monocytogenes in Foods. Korean J. Food Sci. Anim. Resour. 2014, 34, 665–673. [Google Scholar] [CrossRef]
- Baetsen-Young, A.M.; Vasher, M.; Matta, L.L.; Colgan, P.; Alocilja, E.C.; Day, B. Direct colorimetric detection of unamplified pathogen DNA by dextrin-capped gold nanoparticles. Biosens. Bioelectron. 2018, 101, 29–36. [Google Scholar] [CrossRef]
- ISO 11290-2:2017; Microbiology of the Food Chain—Horizontal Method for the Detection and Enumeration of Listeria monocytogenes and of Listeria spp.—Part 2: Enumeration Method. ISO: Geneva, Switzerland, 2017; Confirmed 2022.
- Yang, Y.; Kong, X.; Yang, J.; Xue, J.; Niu, B.; Chen, Q. Rapid Nucleic Acid Detection of Listeria monocytogenes Based on RAA-CRISPR Cas12a System. Int. J. Mol. Sci. 2024, 25, 3477. [Google Scholar] [CrossRef]
- Cheng, J.; Wu, S.; Ye, Q.; Gu, Q.; Zhang, Y.; Ye, Q.; Lin, R.; Liang, X.; Liu, Z.; Bai, J.; et al. A novel multiplex PCR based method for the detection of Listeria monocytogenes clonal complex 8. Int. J. Food Microbiol. 2024, 409, 110475. [Google Scholar] [CrossRef]
- Bai, X.; Huang, J.; Li, W.; Song, Y.; Xiao, F.; Xu, Q.; Xu, H. Portable dual-mode biosensor based on smartphone and glucometer for on-site sensitive detection of Listeria monocytogenes. Sci. Total. Environ. 2023, 874, 162450. [Google Scholar] [CrossRef] [PubMed]
- Deng, M.; Chen, G.; Feng, X.; Wang, Z.; Xu, Q.; Xu, H. Detection of Listeria monocytogenes based on teicoplanin functionalized magnetic beads combined with fluorescence assay. Microchem. J. 2021, 171, 106842. [Google Scholar] [CrossRef]
- Bai, X.; Wang, Z.; Li, W.; Xiao, F.; Huang, J.; Xu, Q.; Xu, H. Rapid and accurate detection for Listeria monocytogenes in milk using ampicillin-mediated magnetic separation coupled with quantitative real-time PCR. Microchem. J. 2022, 183, 108063. [Google Scholar] [CrossRef]
- Chen, W.; Cui, L.; Song, Y.; Chen, W.; Su, Y.; Chang, W.; Xu, W. Detection of Listeria monocytogenes Using Luminol-Functionalized AuNF-Labeled Aptamer Recognition and Magnetic Separation. ACS Omega 2021, 6, 26338–26344. [Google Scholar] [CrossRef]
- Yeğenoğlu Akçinar, H.; Aslim, B.; Torul, H.; Güven, B.; Zengin, A.; Suludere, Z.; Boyaci, I.H.; Tamer, U. Immunomagnetic separation and Listeria monocytogenes detection with surface-enhanced Raman scattering. Turk. J. Med. Sci. 2020, 50, 1157–1167. [Google Scholar] [CrossRef]
- Alhogail, S.; Suaifan, G.; Zourob, M. Rapid colorimetric sensing platform for the detection of Listeria monocytogenes foodborne pathogen. Biosens. Bioelectron. 2016, 86, 1061–1066. [Google Scholar] [CrossRef]
- Shim, W.-B.; Lee, C.-W.; Kim, M.-G.; Chung, D.-H. An antibody–magnetic nanoparticle conjugate-based selective filtration method for the rapid colorimetric detection of Listeria monocytogenes. Anal. Methods 2014, 6, 9129–9135. [Google Scholar] [CrossRef]
- Rong, H.; Gao, T.; Zheng, Y.; Li, L.; Xu, D.; Zhang, X.; Hou, Y.; Yan, M. Fe3O4@silica nanoparticles for reliable identification and magnetic separation of Listeria monocytogenes based on molecular-scale physiochemical interactions. J. Mater. Sci. Technol. 2021, 84, 116–123. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, J.; Song, X.; Xu, K.; Chen, H.; Zhao, C.; Li, J. Colorimetric immunoassay for Listeria monocytogenes by using core gold nanoparticles, silver nanoclusters as oxidase mimetics, and aptamer-conjugated magnetic nanoparticles. Microchim. Acta 2018, 185, 360. [Google Scholar] [CrossRef]
- Kanayeva, D.A.; Wang, R.; Rhoads, D.; Erf, G.F.; Slavik, M.F.; Tung, S.; Li, Y. Efficient Separation and Sensitive Detection of Listeria monocytogenes Using an Impedance Immunosensor Based on Magnetic Nanoparticles, a Microfluidic Chip, and an Interdigitated Microelectrode. J. Food Prot. 2012, 75, 1951–1959. [Google Scholar] [CrossRef]
- Guo, Y.; Zhao, C.; Liu, Y.; Nie, H.; Guo, X.; Song, X.; Xu, K.; Li, J.; Wang, J. A novel fluorescence method for the rapid and effective detection of Listeria monocytogenes using aptamer-conjugated magnetic nanoparticles and aggregation-induced emission dots. Analyst 2020, 145, 3857–3863. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.; Li, F.; Li, F.; Xiong, Y.; Xu, H. Vancomycin modified PEGylated-magnetic nanoparticles combined with PCR for efficient enrichment and detection of Listeria monocytogenes. Sens. Actuators B Chem. 2017, 247, 546–555. [Google Scholar] [CrossRef]
- Hormsombut, T.; Mekjinda, N.; Kalasin, S.; Surareungchai, W.; Rijiravanich, P. Mesoporous Silica Nanoparticles-Enhanced Microarray Technology for Highly Sensitive Simultaneous Detection of Multiplex Foodborne Pathogens. ACS Appl. Bio Mater. 2024, 7, 2367–2377. [Google Scholar] [CrossRef] [PubMed]
- Du, J.; Guo, J.; Liu, J.; Huang, Z.; Zhao, D.; Bai, Y. An aptamer magnetic capture based colorimetric method for rapid and sensitive detection of Listeria monocytogenes. J. Food Meas. Charact. 2024, 18, 6319–6330. [Google Scholar] [CrossRef]
- Huang, J.; Pang, X.; Li, X.; Bie, X.; Sun, J.; Lu, Y. Rapid and accurate AuNPs-sodium deoxycholate-propidium monoazide-qPCR technique for simultaneous detection of viable Listeria monocytogenes and Salmonella. Food Control 2024, 166, 110711. [Google Scholar] [CrossRef]
- Bayramoglu, G.; Ozalp, V.C.; Arica, M.Y. Aptamer-based magnetic isolation and specific detection system for Listeria monocytogenes from food samples. Microchem. J. 2024, 203, 110892. [Google Scholar] [CrossRef]
- Xu, Q.; Xiao, F.; Wang, Z.; Ling, Z.; Xu, H. TBA magnetically functionalized materials combined with biomass-derived fluorescent probe for Listeria monocytogenes detection in a sandwich-like strategy. Food Biosci. 2024, 59, 104049. [Google Scholar] [CrossRef]
- Wu, X.; Chen, Q.; Yang, C.; Ning, Q.; Liu, Z. An enhanced visual detection assay for Listeria monocytogenes in food based on isothermal amplified peroxidase-mimicking catalytic beacon. Food Control 2022, 134, 108721. [Google Scholar] [CrossRef]
- Thongmee, P.; Ngernpimai, S.; Srichaiyapol, O.; Mongmonsin, U.; Teerasong, S.; Charoensri, N.; Wongwattanakul, M.; Lulitanond, A.; Kuwatjanakul, W.; Wonglakorn, L.; et al. The Evaluation of a Lateral Flow Strip Based on the Covalently Fixed “End-On” Orientation of an Antibody for Listeria monocytogenes Detection. Anal. Chem. 2024, 96, 8543–8551. [Google Scholar] [CrossRef] [PubMed]
- Beyazit, F.; Arica, M.Y.; Acikgoz-Erkaya, I.; Ozalp, C.; Bayramoglu, G. Quartz crystal microbalance–based aptasensor integrated with magnetic pre-concentration system for detection of Listeria monocytogenes in food samples. Microchim. Acta 2024, 191, 235. [Google Scholar] [CrossRef]
- Rivas-Macho, A.; Eletxigerra, U.; Diez-Ahedo, R.; Merino, S.; Goñi-de-Cerio, F.; Olabarria, G. LAMP based electrochemical sensor for extraction-free detection of Listeria monocytogenes in food samples. Food Control 2024, 163, 110546. [Google Scholar] [CrossRef]
- Liu, J.; Xiang, Z.; Huang, T.; Xu, Z.; Ma, Q.; Yuan, L.; Soteyome, T. Development and verification of crossing priming amplification on rapid detection of virulent and viable L. monocytogenes: In-depth analysis on the target and further application on food screening. LWT 2024, 204, 116422. [Google Scholar] [CrossRef]
- Armstrong, C.M.; Capobianco, J.A.; Nguyen, S.; Guragain, M.; Liu, Y. High-throughput homogenous assay for the direct detection of Listeria monocytogenes DNA. Sci. Rep. 2024, 14, 7026. [Google Scholar] [CrossRef]




| Source | SS | df | MS | % of Total Variation | p-Value | Interpretation |
|---|---|---|---|---|---|---|
| Dilution Level (10−5–10−8) | 3433 | 3 | 1144.3 | 94.4% | 0.0138 | Significant |
| Matrix (PBS vs. Spinach) | 57.8 | 1 | 57.8 | 1.6% | 0.355 | Not significant |
| Residual | 145.6 | 3 | 48.5 | 4.0% | — | — |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Williams, M.; Faraj, R.; Nyarku, R.; Simon, S.; Bentum, K.E.; Ghazy, A.; Woube, Y.; Samuel, T.; Alocija, E.; Abebe, W. Potential Use of Biosensors for the Rapid and Specific Isolation of Listeria monocytogenes from Ready-to-Eat (RTE) Foods. Pathogens 2025, 14, 1280. https://doi.org/10.3390/pathogens14121280
Williams M, Faraj R, Nyarku R, Simon S, Bentum KE, Ghazy A, Woube Y, Samuel T, Alocija E, Abebe W. Potential Use of Biosensors for the Rapid and Specific Isolation of Listeria monocytogenes from Ready-to-Eat (RTE) Foods. Pathogens. 2025; 14(12):1280. https://doi.org/10.3390/pathogens14121280
Chicago/Turabian StyleWilliams, McCoy, Rawah Faraj, Rejoice Nyarku, Savannah Simon, Kingsley E. Bentum, Ahmed Ghazy, Yilkal Woube, Temesgen Samuel, Evangelyn Alocija, and Woubit Abebe. 2025. "Potential Use of Biosensors for the Rapid and Specific Isolation of Listeria monocytogenes from Ready-to-Eat (RTE) Foods" Pathogens 14, no. 12: 1280. https://doi.org/10.3390/pathogens14121280
APA StyleWilliams, M., Faraj, R., Nyarku, R., Simon, S., Bentum, K. E., Ghazy, A., Woube, Y., Samuel, T., Alocija, E., & Abebe, W. (2025). Potential Use of Biosensors for the Rapid and Specific Isolation of Listeria monocytogenes from Ready-to-Eat (RTE) Foods. Pathogens, 14(12), 1280. https://doi.org/10.3390/pathogens14121280








