Arthrobacter woluwensis Bacteremia: A Clinical and Genomic Report
Abstract
1. Introduction
1.1. Specimen Collection and Antibiotic Susceptibility Testing
1.2. Genome Sequencing, Assembly, Annotation, and Phylogenetic Analysis
2. Result
3. Discussion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Funke, G.; Hutson, R.A.; Bernard, K.A.; Pfyffer, G.E.; Wauters, G.; Collins, M.D. Isolation of Arthrobacter spp. from clinical specimens and description of Arthrobacter cumminsii sp. nov. and Arthrobacter woluwensis sp. nov. J. Clin. Microbiol. 1996, 34, 2356–2363. [Google Scholar] [CrossRef]
- Conn, H.J.; Dimmick, I. Soil Bacteria Similar in Morphology to Mycobacterium and Corynebacterium. J. Bacteriol. 1947, 54, 291–303. [Google Scholar] [CrossRef] [PubMed]
- Bernasconi, E.; Valsangiacomo, C.; Peduzzi, R.; Carota, A.; Moccetti, T.; Funke, G. Arthrobacter woluwensis subacute infective endocarditis: Case report and review of the literature. Clin. Infect. Dis. 2004, 38, e27–e31. [Google Scholar] [CrossRef] [PubMed]
- Kim, G.Y.; Suh, J.T.; Choi, S.K.; Lee, H.J. A Case of Bacteremia Caused by Arthrobacter woluwensis. Korean J. Clin. Microbiol. 2007, 10, 160–163. [Google Scholar]
- Park, C.H.; Han, M.S.; Kim, J.K.; Jeong, S.J.; Ku, N.S.; Kim, H.; Kim, S.B.; Chung, H.-S.; Han, S.H.; Choi, J.Y.; et al. Development of Arthrobacter woluwensis Bacteremia in a Patient with Multiple Myeloma: A Case Report and Comprehensive Literature Review. Infect. Chemother. 2012, 44, 205–209. [Google Scholar] [CrossRef]
- Shin, K.S.; Hong, S.B.; Son, B.R. [A Case of Catheter-Related Bacteremia by Arthrobacter woluwensis.]. Korean J. Lab. Med. 2006, 26, 103–106. [Google Scholar] [CrossRef]
- Durand, C.; Kouchit, Y.; Prots, L.; Degand, N.; Dellamonica, P.; Demonchy, E.; Chirio, D. A case of infective endocarditis caused by Arthrobacter woluwensis. Eur. J. Clin. Microbiol. Infect. Dis. 2021. [Google Scholar] [CrossRef]
- Koper, T.E.; El-Sheikh, A.F.; Norton, J.M.; Klotz, M.G. Urease-Encoding Genes in Ammonia-Oxidizing Bacteria. Appl. Environ. Microbiol. 2004, 70, 2342–2348. [Google Scholar] [CrossRef]
- Esteban, J.; Bueno, J.; Perez-Santonja, J.J.; Soriano, F. Endophthalmitis involving an Arthrobacter-like organism following intraocular lens implantation. Clin. Infect. Dis. 1996, 23, 1180–1181. [Google Scholar] [CrossRef][Green Version]
- Shigeta, N.; Ozaki, K.; Hori, K.; Ito, K.; Nakayama, M.; Nakahira, K.; Yanagihara, I. An Arthrobacter spp. bacteremia leading to fetal death and maternal disseminated intravascular coagulation. Fetal Pediatric Pathol. 2013, 31, 25–31. [Google Scholar] [CrossRef]
- Bodaghi, B.; Dauga, C.; Cassoux, N.; Wechsler, B.; Merle-Beral, H.; Poveda, J.D.; Piette, J.C.; LeHoang, P. Whipple’s syndrome (uveitis, B27-negative spondylarthropathy, meningitis, and lymphadenopathy) associated with Arthrobacter sp. infection. Ophthalmology 1998, 105, 1891–1896. [Google Scholar] [CrossRef]
- Bousquet, A.; Soler, C.; MacNab, C.; Le Fleche, A.; Héno, P. Arthrobacter albus infected implantable cardioverter-defibrillator. Med. Mal. Infect. 2016, 46, 59–60. [Google Scholar] [CrossRef] [PubMed]
- Imirzalioglu, C.; Hain, T.; Hossain, H.; Chakraborty, T.; Domann, E. Erythema caused by a localised skin infection with Arthrobacter mysorens. BMC Infect. Dis. 2010, 10, 352. [Google Scholar] [CrossRef] [PubMed]
- Mages, I.S.; Frodl, R.; Bernard, K.A.; Funke, G. Identities of Arthrobacter spp. and Arthrobacter-like bacteria encountered in human clinical specimens. J. Clin. Microbiol. 2008, 46, 2980–2986. [Google Scholar] [CrossRef]
- Yap, D.Y.; Tse, H.; Mok, M.M.; Chan, G.C.; Yip, T.; Lui, S.L.; Lo, W.K.; Chan, T.M. Arthrobacter sanguinis: An uncommon cause of peritonitis in a peritoneal dialysis patient. Nephrology 2015, 20, 868. [Google Scholar] [CrossRef]
- Lai, C.-C.; Cia, C.-T.; Chiang, H.-T.; Kung, Y.-C.; Shi, Z.-Y.; Chuang, Y.-C.; Lee, C.-M.; Ko, W.-C.; Hsueh, P.-R. Implementation of a national bundle care program to reduce central line-associated bloodstream infections in intensive care units in Taiwan. J. Microbiol. Immunol. Infect. 2018, 51, 666–671. [Google Scholar] [CrossRef]
- Hsu, C.L.; Shih, L.Y.; Leu, H.S.; Wu, C.L.; Funke, G. Septicemia due to Arthrobacter species in a neutropenic patient with acute lymphoblastic leukemia. Clin. Infect. Dis. 1998, 27, 1334–1335. [Google Scholar] [CrossRef]
- Friedrich, A.W.; Lukas, R.; Mellmann, A.; Köck, R.; Zhang, W.; Mathys, W.; Bielaszewska, M.; Karch, H. Urease genes in non-O157 Shiga toxin-producing Escherichia coli: Mostly silent but valuable markers for pathogenicity. Clin. Microbiol. Infect. 2006, 12, 483–486. [Google Scholar] [CrossRef]
- Houimel, M.; Mach, J.-P.; Corthésy-Theulaz, I.; Corthésy, B.; Fisch, I. New inhibitors of Helicobacter pylori urease holoenzyme selected from phage-displayed peptide libraries. Eur. J. Biochem. 1999, 262, 774–780. [Google Scholar] [CrossRef] [PubMed]
- Megraud, F.; Lehours, P. Helicobacter pylori detection and antimicrobial susceptibility testing. Clin. Microbiol. Rev. 2007, 20, 280–322. [Google Scholar] [CrossRef] [PubMed]
- Malekshahi, Z.V.; Gargari, S.L.; Rasooli, I.; Ebrahimizadeh, W. Treatment of Helicobacter pylori infection in mice with oral administration of egg yolk-driven anti-UreC immunoglobulin. Microb. Pathog. 2011, 51, 366–372. [Google Scholar] [CrossRef] [PubMed]
- Ardekani, L.S.; Gargari, S.L.; Rasooli, I.; Bazl, M.R.; Mohammadi, M.; Ebrahimizadeh, W.; Bakherad, H.; Zare, H. A novel nanobody against urease activity of Helicobacter pylori. Int. J. Infect. Dis. 2013, 17, e723–e728. [Google Scholar] [CrossRef] [PubMed]


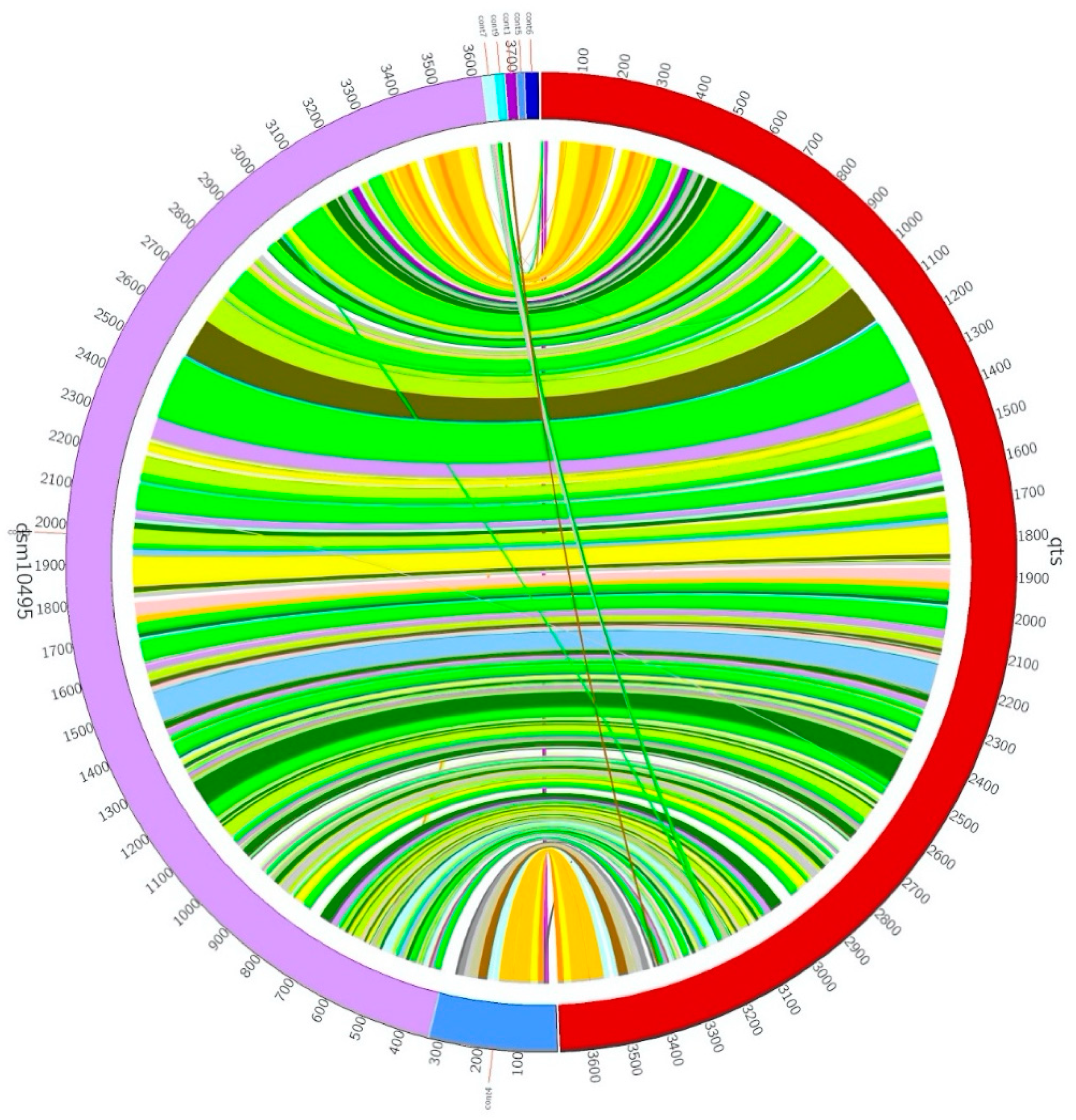
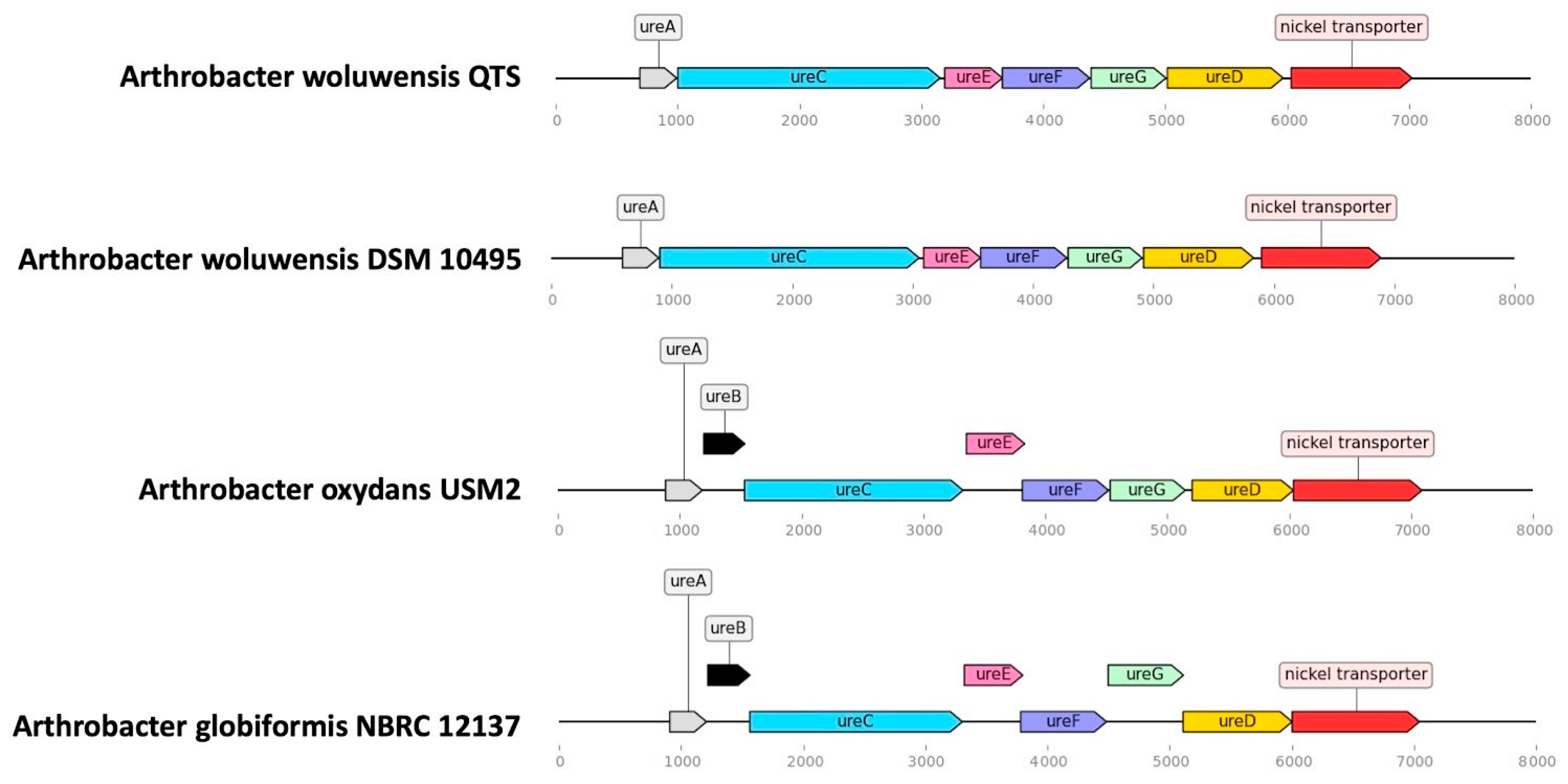
| Strains | Coverage | Genome Size | Genes | Sequencing Technology | Assembly Method | Country | Host | Isolation Source | DNA | Accession Number |
|---|---|---|---|---|---|---|---|---|---|---|
| QTS | 122.0x | 3680971 | 3369 | Oxford Nanopore GridION | Flye v. 2.6 | Taiwan | Homo species | Blood | Circular | CP049819 |
| Case No. | Age/Sex | Underlying Condition | Diagnosis | Specimen | Treatment | Risk Factor | Outcome |
|---|---|---|---|---|---|---|---|
| 1 [1] | 33F | HIV infection, stage C-3 | Bacteremia | Blood | 2 weeks ampicillin | Port-A catheter | Survival |
| 2 [6] | 56M | Metastatic colon cancer | Bacteremia | Blood | Vancomycin, catheter removal | Subclavian catheter | Died 1 |
| 3 [3] | 39M | IVDU, TB | Mitral valve endocarditis | Blood | 6 weeks teicoplanin | IVDU | Survival |
| 4 [4] | 91F | Ischemic stroke | Bacteremia | Blood | 10 days linezolid | Hospital acquired infection | Survival |
| 5 [5] | 76F | Multiple myeloma, HTN, DM | Bacteremia | Blood | 19 days teicoplanin, catheter removal | Chemoport | Survival |
| 6 [7] | 52 | Hepatitis C | Mitral and aortic valve endocarditis | Intraoperative samples and blood | Operation 2, Teicoplanin, TMP/SMX, linezolid 3 | IVDU | Survival |
| Present case | 93M | CAD, HTN, prostate cancer, newly diagnosised gastric cancer | Bacteremia | Blood | 2 weeks ampicillin | CVC, post-OP wound | Survival |
| Case | AMC | AM | CRO | CXM | CE | CIP | CLI | ERY | GM | IPM | PE | RIF | TEC | TET | VA | DAP | LIN |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 [1] | R # | R | R | R | R | R | R | R | R | R | R(4 *) | R | S | S | S(2) | ||
| 2 [6] | R(4) | S(1.5) | |||||||||||||||
| 3 [3] | R | R | R | R | R | I | I | R(4) | S | S | S | S(2) | |||||
| 4 [4] | S | S | S | S | I | S | S | S | |||||||||
| 5 [5] | I | R | I(1) | S(2) | |||||||||||||
| Present case | R | R | R | I(0.75) | S(1) | R |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, S.-Y.; Kao, C.-C.; Hu, Y.-C.; Lai, C.-H.; Jiang, Y.-P.; Mao, Y.-C.; Huang, Y.-T.; Liu, P.-Y. Arthrobacter woluwensis Bacteremia: A Clinical and Genomic Report. Pathogens 2021, 10, 443. https://doi.org/10.3390/pathogens10040443
Li S-Y, Kao C-C, Hu Y-C, Lai C-H, Jiang Y-P, Mao Y-C, Huang Y-T, Liu P-Y. Arthrobacter woluwensis Bacteremia: A Clinical and Genomic Report. Pathogens. 2021; 10(4):443. https://doi.org/10.3390/pathogens10040443
Chicago/Turabian StyleLi, Shu-Yuan, Chin-Chuan Kao, Yu-Cheng Hu, Chung-Hsu Lai, Yi-Ping Jiang, Yan-Chiao Mao, Yao-Ting Huang, and Po-Yu Liu. 2021. "Arthrobacter woluwensis Bacteremia: A Clinical and Genomic Report" Pathogens 10, no. 4: 443. https://doi.org/10.3390/pathogens10040443
APA StyleLi, S.-Y., Kao, C.-C., Hu, Y.-C., Lai, C.-H., Jiang, Y.-P., Mao, Y.-C., Huang, Y.-T., & Liu, P.-Y. (2021). Arthrobacter woluwensis Bacteremia: A Clinical and Genomic Report. Pathogens, 10(4), 443. https://doi.org/10.3390/pathogens10040443







