N-Glycosylation in Piroplasmids: Diversity within Simplicity
Abstract
1. Introduction
2. Results and Discussion
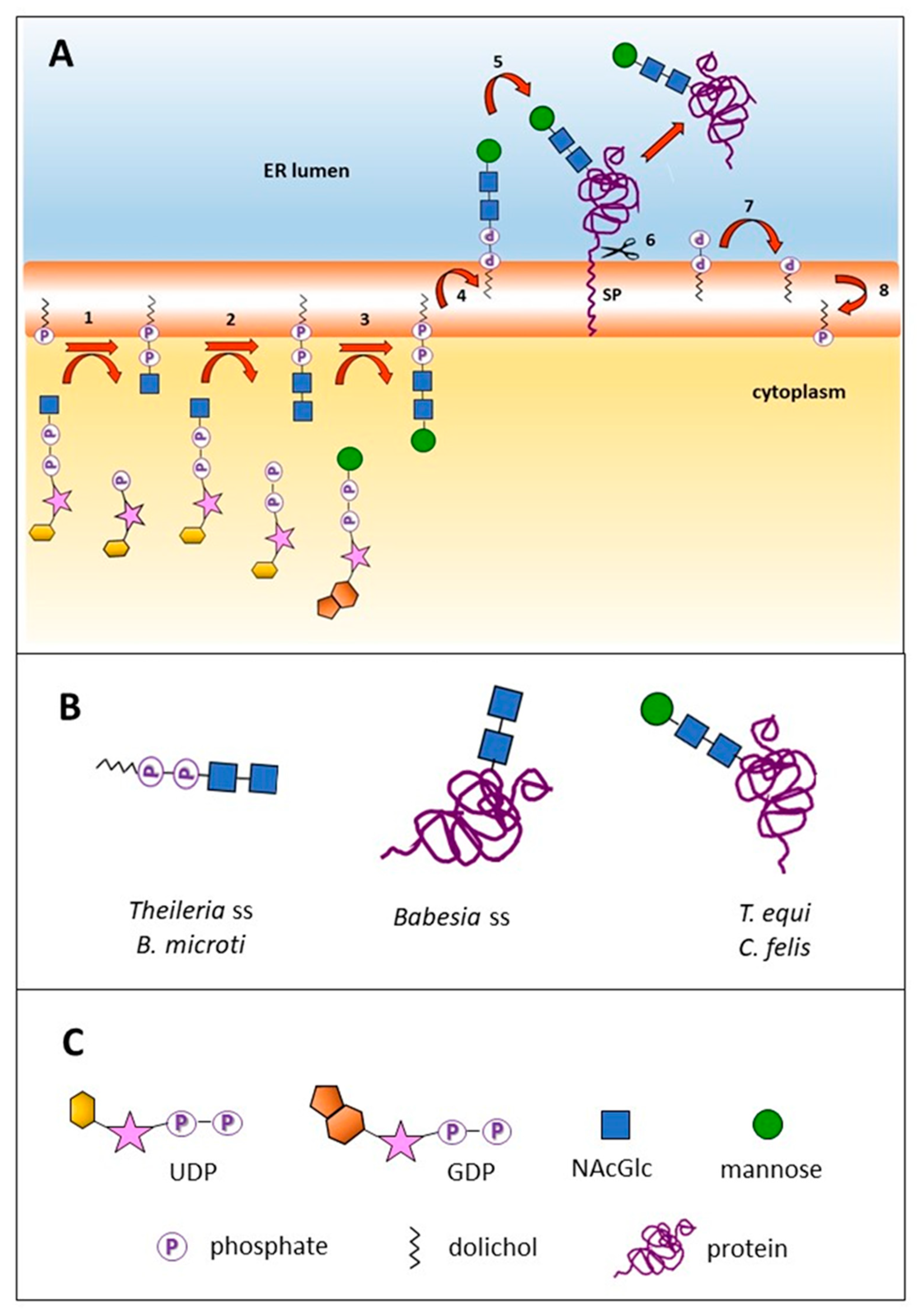
2.1. Three N-Glycosylation Scenarios Are Inferred from the Genomic Analysis of Diverse Piroplasmid Lineages
2.2. Piroplasmid Genomes Encode All Necessary Enzymes to Synthesize N-Glycan Substrates
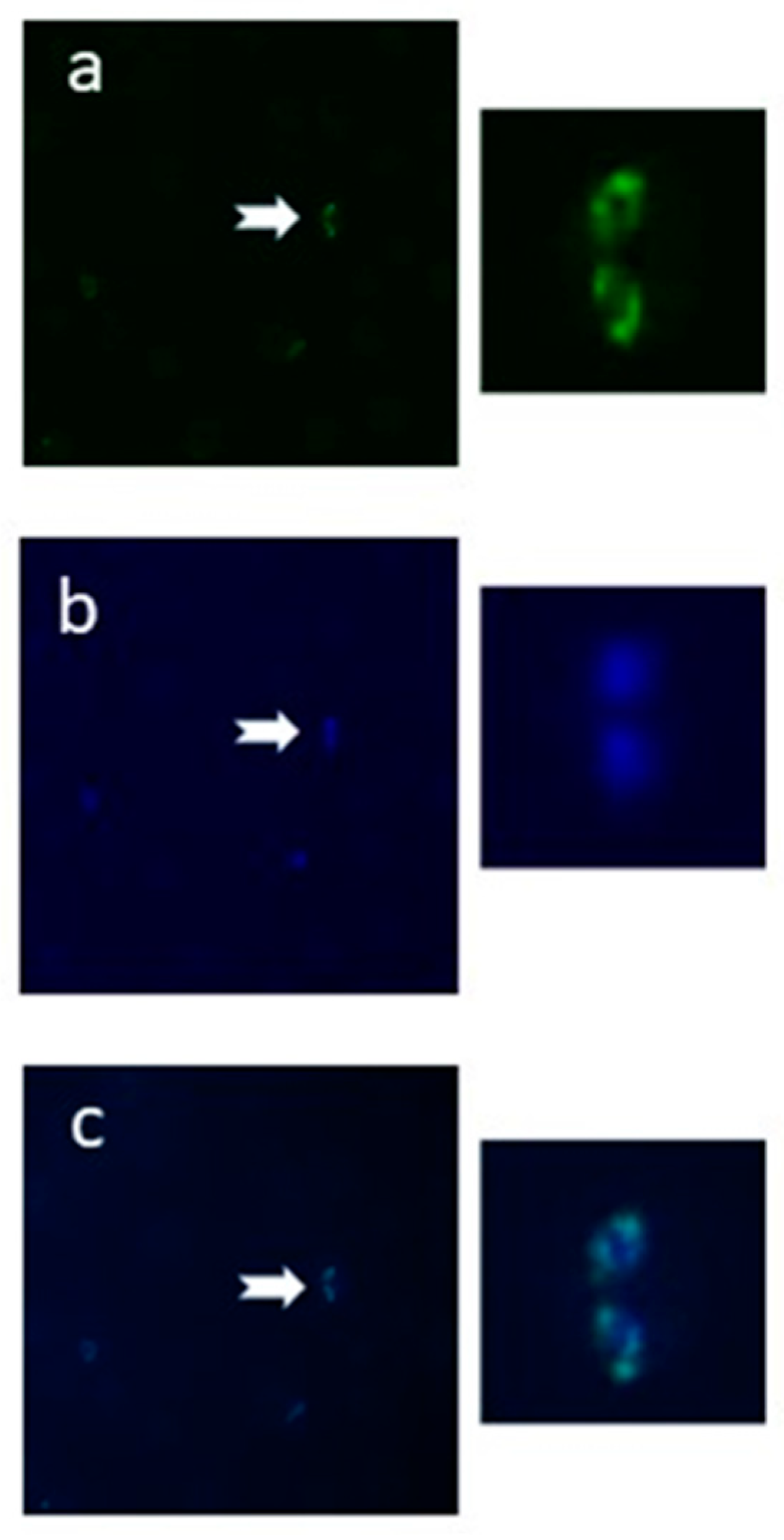
2.3. The N-Glycan N-Acetyl-chitobiose Can Be Detected on the B. bovis mz Surface
2.4. Abundant Proteins of the B. bovis Secretome Are Predicted to Be N-Glycosylated
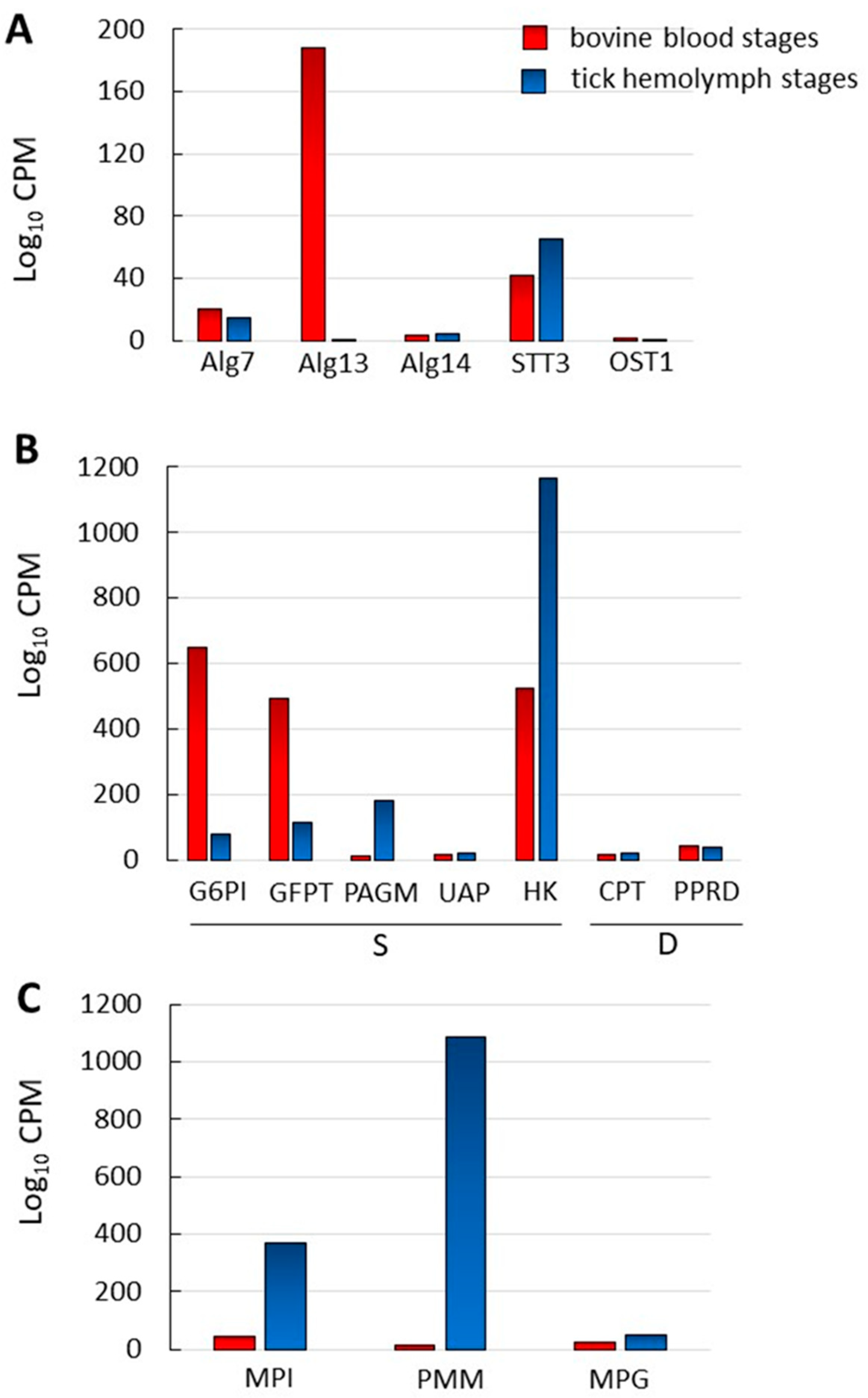
2.5. N-Glycosylation Enzyme Genes Are Transcribed in B. bovis Blood and Tick-Kinete Parasite Stages
2.6. N-Glycosylation Is Important but Not Essential for In Vitro Growth of B. bovis mz
3. Materials and Methods
3.1. Bioinformatic Analysis
3.2. Parasite Cultures
3.3. Detection of N-Glycosylation by Fluorescence Microscopy
3.4. Transcriptomics Analysis
3.5. Inhibition Tests
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Yakubu, R.R.; Weiss, L.M.; Silmon de Monerri, N.C. Posttranslational modifications as key regulators of Apicomplexan biology: Insights from proteome-wide studies. Mol. Microbiol. 2018, 107, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Aebi, M. N-linked protein glycosylation in the ER. Biochim. Biophys. Acta 2013, 1833, 2430–2437. [Google Scholar] [CrossRef] [PubMed]
- Samuelson, J.; Banerjee, S.; Magnelli, P.; Cui, J.; Kelleher, D.J.; Gilmore, R.; Robbins, P.W. The diversity of dolichol-linked precursors to Asn-linked glycans likely results from secondary loss of sets of glycosyltransferases. Proc. Natl. Acad. Sci. USA 2005, 102, 1548–1553. [Google Scholar] [CrossRef] [PubMed]
- Luk, F.C.; Johnson, T.M.; Beckers, C.J. N-linked glycosylation of proteins in the protozoan parasite Toxoplasma gondii. Mol Biochem Parasitol. 2008, 157, 169–178. [Google Scholar] [CrossRef]
- Varki, A. Biological roles of glycans. Glycobiology 2017, 27, 3–49. [Google Scholar] [CrossRef]
- Kimura, E.A.; Couto, A.S.; Peres, V.J.; Casal, O.L.; Katzin, A.M. N-linked glycoproteins are related to schizogony of the intraerythrocytic stage in Plasmodium falciparum. J. Biol. Chem. 1996, 271, 14452–14461. [Google Scholar] [CrossRef]
- Florin-Christensen, M.; Schnittger, L. Piroplasmids and ticks: A long-lasting intimate relationship. Front. Biosci. 2009, 14, 3064–3073. [Google Scholar] [CrossRef]
- Schnittger, L.; Rodriguez, A.E.; Florin-Christensen, M.; Morrison, D.A. Babesia: A world emerging. Infect. Genet. Evol. 2012, 12, 1788–1809. [Google Scholar] [CrossRef]
- Jalovecka, M.; Sojka, D.; Ascencio, M.; Schnittger, L. Babesia Life Cycle—When phylogeny meets biology. Trends Parasitol. 2019, 35, 356–368. [Google Scholar] [CrossRef]
- Sanyal, S.; Menon, A.K. Flipping lipids: Why an’ what’s the reason for? ACS Chem. Biol. 2009, 4, 895–909. [Google Scholar] [CrossRef]
- Bushkin, G.G.; Ratner, D.M.; Cui, J.; Banerjee, S.; Duraisingh, M.T.; Jennings, C.V.; Dvorin, J.D.; Gubbels, M.J.; Robertson, S.D. Suggestive evidence for Darwinian Selection against asparagine-linked glycans of Plasmodium falciparum and Toxoplasma gondii. Eukaryot. Cell 2010, 9, 228–241. [Google Scholar] [CrossRef] [PubMed]
- Kelleher, D.J.; Gilmore, R. An evolving view of the eukaryotic oligosaccharyltransferase. Glycobiology 2006, 16, 47R–62R. [Google Scholar] [CrossRef] [PubMed]
- Tretina, K.; Pelle, R.; Orvis, J.; Gotia, H.T.; Ifeonu, O.O.; Kumari, P.; Palmateer, N.C.; Iqbal, S.B.A.; Fry, L.M.; Nene, V.M.; et al. Re-annotation of the Theileria parva genome refines 53% of the proteome and uncovers essential components of N-glycosylation, aconserved pathway in many organisms. BMC Genom. 2020, 21, 279–291. [Google Scholar] [CrossRef] [PubMed]
- Tai, V.W.; Imperiali, B. Substrate specificity of the glycosyl donor for oligosaccharyl transferase. J. Org. Chem. 2001, 66, 6217–6228. [Google Scholar] [CrossRef] [PubMed]
- Ascencio, M.E.; Florin-Christensen, M.; Mamoun, C.B.; Weir, W.; Shiels, B.; Schnittger, L. Cysteine proteinase C1A paralog profiles correspond with phylogenetic lineages of pathogenic piroplasmids. Vet. Sci. 2018, 5, 41. [Google Scholar] [CrossRef] [PubMed]
- Gallenti, R.; Poklepovich, T.; Florin-Christensen, M.; Schnittger, L. The repertoire of serine rhomboid proteinases of piroplasmids of importance to animal and human health. Int. J. Parasitol. 2020, in press. [Google Scholar]
- Kappmeyer, L.S.; Thiagarajan, M.; Herndon, D.R.; Ramsay, J.D.; Caler, E.; Djikeng, A.; Gillespie, J.J.; Lau, A.O.; Roalson, E.H.; Silva, J.C.; et al. Comparative genomic analysis and phylogenetic position of Theileria equi. BMC Genom. 2012, 13, 603. [Google Scholar] [CrossRef] [PubMed]
- Shaw, M.K. Cell invasion by Theileria sporozoites. Trends Parasitol. 2003, 19, 2–6. [Google Scholar] [CrossRef]
- Sevilla, E.; González, L.M.; Luque, D.; Gray, J.; Montero, E. Kinetics of the invasion and egress processes of Babesia divergens, observed by time-lapse video microscopy. Sci. Rep. 2018, 8, 14116–14126. [Google Scholar] [CrossRef]
- Dacks, J.B.; Doolittle, W.F. Reconstructing/deconstructing review the earliest eukaryotes: How comparative genomics can help. Cell 2001, 107, 419–425. [Google Scholar] [CrossRef]
- Schwarz, F.; Aebi, M. Mechanisms and principles of N-linked protein glycosylation. Curr. Opin. Struct. Biol. 2011, 21, 576–582. [Google Scholar] [CrossRef] [PubMed]
- Bickel, T.; Lehle, L.; Schwarz, M.; Aebi, M.; Jakob, C.A. Biosynthesis of lipid-linked oligosaccharides in Saccharomyces cerevisiae: Alg13p and Alg14p form a complex required for the formation of GlcNAc(2)-PP-dolichol. J. Biol. Chem. 2005, 280, 34500–34506. [Google Scholar] [CrossRef] [PubMed]
- Sanz, S.; Bandini, G.; Ospina, D.; Bernabeu, M.; Mariño, K.; Fernández-Becerra, C.; Izquierdo, L. Biosynthesis of GDP-fucose and other sugar nucleotides in the blood stages of Plasmodium falciparum. J. Biol. Chem. 2013, 288, 16506–16517. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, A.E.; Couto, A.; Echaide, I.; Schnittger, L.; Florin-Christensen, M. Babesia bovis contains an abundant parasite-specific protein-free glycerophosphatidylinositol and the genes predicted for its assembly. Vet. Parasitol. 2010, 167, 227–235. [Google Scholar] [CrossRef] [PubMed]
- Zimbres, F.M.; Valenciano, A.L.; Merino, E.F.; Florentin, A.; Holderman, N.R.; He, G.; Gawarecka, K.; Skorupinska-Tudek, K.; Fernández-Murga, M.L.; Swiezewska, E.; et al. Metabolomics profiling reveals new aspects of dolichol biosynthesis in Plasmodium falciparum. Sci. Rep. 2020, 10, 13264–13281. [Google Scholar] [CrossRef]
- Caballero, M.C.; Pedroni, M.J.; Palmer, G.H.; Suarez, C.E.; Davitt, C.; Lau, A.O. Characterization of acyl carrier protein and LytB in Babesia bovis apicoplast. Mol. Biochem. Parasitol. 2012, 181, 125–133. [Google Scholar] [CrossRef]
- He, L.; He, P.; Luo, X.; Li, M.; Yu, L.; Guo, J.; Zhan, X.; Zhu, G.; Zhao, J. The MEP pathway in Babesia orientalis apicoplast, a potential target for anti-babesiosis drug development. Parasites Vectors 2018, 11, 452–460. [Google Scholar] [CrossRef]
- AbouLaila, M.; Sivakumar, T.; Yokoyama, N.; Igarashi, I. Inhibitory effect of terpene nerolidol on the growth of Babesia parasites. Parasitol. Int. 2010, 59, 278–282. [Google Scholar] [CrossRef]
- Rush, J.S.; Gao, N.; Lehrman, M.A.; Waechter, C.J. Recycling of dolichyl monophosphate to the cytoplasmic leaflet of the endoplasmic reticulum after the cleavage of dolichyl pyrophosphate on the lumenal monolayer. J. Biol. Chem. 2008, 283, 4087–4093. [Google Scholar] [CrossRef]
- D’Alexandri, F.L.; Kimura, E.A.; Peres, V.J.; Katzin, A.M. Protein dolichylation in Plasmodium falciparum. FEBS Lett. 2006, 580, 6343–6348. [Google Scholar] [CrossRef]
- Ruef, B.J.; Dowling, S.C.; Conley, P.G.; Perryman, L.E.; Brown, W.C.; Jasmer, D.P.; Rice-Ficht, A.C. A unique Babesia bovis spherical body protein is conserved among geographic isolates and localizes to the infected erythrocyte membrane. Mol. Biochem. Parasitol. 2000, 105, 1–12. [Google Scholar] [CrossRef]
- Gaffar, F.R.; Yatsuda, A.P.; Franssen, F.F.; de Vries, E. A Babesia bovis merozoite protein with a domain architecture highly similar to the thrombospondin-related anonymous protein (TRAP) present in Plasmodium sporozoites. Mol. Biochem. Parasitol. 2004, 136, 25–34. [Google Scholar] [CrossRef] [PubMed]
- Hope, M.; Riding, G.; Menzies, M.; Colditz, I.; Reverter, A.; Willadsen, P. Potential for recombinant Babesia bovis antigens to protect against a highly virulent isolate. Parasite Immunol. 2005, 27, 439–445. [Google Scholar] [CrossRef] [PubMed]
- Gohil, S.; Kats, L.M.; Seemann, T.; Fernandez, K.M.; Siddiqui, G.; Cooke, B.M. Bioinformatic prediction of the exportome of Babesia bovis and identification of novel proteins in parasite-infected red blood cells. Int. J. Parasitol. 2013, 43, 409–416. [Google Scholar] [CrossRef]
- Ferreri, L.M.; Brayton, K.A.; Sondgeroth, K.S.; Lau, A.O.; Suarez, C.E.; McElwain, T.F. Expression and strain variation of the novel “small open reading frame” (smorf) multigene family in Babesia bovis. Int. J. Parasitol. 2012, 42, 131–138. [Google Scholar] [CrossRef]
- Guha-Niyogi, A.; Sullivan, D.R.; Turco, S.J. Glycoconjugate structures of parasitic protozoa. Glycobiology 2001, 11, 45R–59R. [Google Scholar] [CrossRef]
- Petrescu, A.J.; Milac, A.L.; Petrescu, S.M.; Dwek, R.A.; Wormald, M.R. Statistical analysis of the protein environment of N-glycosylation sites: Implications for occupancy, structure, and folding. Glycobiology 2004, 14, 103–114. [Google Scholar] [CrossRef]
- Jinnelov, A.; Ali, L.; Tinti, M.; Güther, M.L.S.; Ferguson, M.A.J. Single-subunit oligosaccharyltransferases of Trypanosoma brucei display different and predictable peptide acceptor specificities. J. Biol. Chem. 2017, 292, 20328–20341. [Google Scholar] [CrossRef]
- Lehle, L.; Tanner, W. The specific site of tunicamycin inhibition in the formation of dolichol-bound N-acetylglucosamine derivatives. FEBS Lett. 1976, 72, 167–170. [Google Scholar] [CrossRef]
- Yoo, J.; Mashalidis, E.H.; Kuk, A.C.Y.; Yamamoto, K.; Kaeser, B.; Ichikawa, S.; Lee, S.-Y. GlcNAc-1-P-transferase-tunicamycin complex structure reveals basis for inhibition of N-glycosylation. Nat. Struct. Mol. Biol. 2018, 25, 217–224. [Google Scholar] [CrossRef]
- Li, L.; Ren, M.; Xu, Y.; Jin, C.; Zhang, W.; Dong, X. Enhanced glycosylation of an S-layer protein enables a psychrophilic methanogenic archaeon to adapt to elevated temperatures in abundant substrates. FEBS Lett. 2020, 594, 665–677. [Google Scholar] [CrossRef] [PubMed]
- Naik, R.S.; Venkatesan, M.; Gowda, D.C. Plasmodium falciparum: The lethal effects of tunicamycin and mevastatin on the parasite are not mediated by the inhibition of N-linked oligosaccharide biosynthesis. Exp. Parasitol. 2001, 98, 110–114. [Google Scholar] [CrossRef] [PubMed]
- Samuelson, J.; Robbins, P.W. Effects of N-glycan precursor length diversity on quality control of protein folding and on protein glycosylation. In Seminars in Cell & Developmental Biology; Academic Press: Cambridge, MA, USA, 2015; Volume 41, pp. 121–128. [Google Scholar]
- Fauquenoy, S.; Morelle, W.; Hovasse, A.; Bednarczyk, A.; Slomianny, C.; Schaeffer, C.; Van Dorsselaer, A.; Tomavo, S. Proteomics and glycomics analyses of N-glycosylated structures involved in Toxoplasma gondii-host cell interactions. Mol. Cell. Proteom. 2008, 7, 891–910. [Google Scholar] [CrossRef] [PubMed]
- Suarez, C.E.; Bishop, R.P.; Alzan, H.F.; Poole, W.A.; Cooke, B.M. Advances in the application of genetic manipulation methods to apicomplexan parasites. Int. J. Parasitol. 2017, 47, 701–710. [Google Scholar] [CrossRef]
- Hall, N.; Pain, A.; Berriman, M.; Churcher, C.; Harris, B.; Harris, D.; Mungall, K.; Bowman, S.; Atkin, R.; Baker, S.; et al. Sequence of Plasmodium falciparum chromosomes 1,3-9 and 13. Nature 2002, 419, 527–531. [Google Scholar] [CrossRef]
- Brayton, K.A.; Lau, A.O.; Herndon, D.R.; Hannick, L.; Kappmeyer, L.S.; Berens, S.J.; Bidwell, S.L.; Brown, W.C.; Crabtree, J.; Fadrosh, D.; et al. Genome sequence of Babesia bovis and comparative analysis of apicomplexan hemoprotozoa. PLoS Pathog. 2007, 3, 1401–1413. [Google Scholar] [CrossRef]
- Jackson, A.P.; Otto, T.D.; Darby, A.; Ramaprasad, A.; Xia, D.; Echaide, I.E.; Farber, M.; Gahlot, S.; Gamble, J.; Gupta, D.; et al. The evolutionary dynamics of variant antigen genes in Babesia reveal a history of genomic innovation underlying host-parasite interaction. Nucleic Acids Res. 2014, 42, 7113–7131. [Google Scholar] [CrossRef]
- Cuesta, I.; González, L.M.; Estrada, K.; Grande, R.; Zaballos, A.; Lobo, C.A.; Barrera, J.; Sanchez-Flores, A.; Montero, E. High-quality draft genome sequence of Babesia divergens, the etiological agent of cattle and human babesiosis. Genome Announc. 2014, 2, e01194-14. [Google Scholar] [CrossRef]
- Yamagishi, J.; Asada, M.; Hakimi, H.; Tanaka, T.Q.; Sugimoto, C.; Kawazu, S.I. Whole-genome assembly of Babesia ovata and comparative genomics between closely related pathogens. BMC Genom. 2017, 18, 832–841. [Google Scholar] [CrossRef]
- Guan, G.; Korhonen, P.K.; Young, N.D.; Koehler, A.V.; Wang, T.; Li, Y.; Liu, Z.; Luo, J.; Yin, H.; Gasser, R.B. Genomic resources for a unique, low-virulence Babesia taxon from China. Parasites Vectors 2016, 9, 564–572. [Google Scholar] [CrossRef]
- Silva, J.C.; Cornillot, E.; McCracken, C.; Usmani-Brown, S.; Dwivedi, A.; Ifeonu, O.O.; Crabtree, J.; Gotia, H.T.; Virji, A.Z.; Reynes, C.; et al. Genome-wide diversity and gene expression profiling of Babesia microti isolates identify polymorphic genes that mediate host-pathogen interactions. Sci. Rep. 2016, 6, 35284–35299. [Google Scholar] [CrossRef] [PubMed]
- Gardner, M.J.; Bishop, R.; Shah, T.; de Villiers, E.P.; Carlton, J.M.; Hall, N.; Ren, Q.; Paulsen, I.T.; Pain, A.; Berriman, M.; et al. Genome sequence of Theileria parva, a bovine pathogen that transforms lymphocytes. Science 2005, 309, 134–137. [Google Scholar] [CrossRef] [PubMed]
- Pain, A.; Renauld, H.; Berriman, M.; Murphy, L.; Yeats, C.A.; Weir, W.; Kerhornou, A.; Aslett, M.; Bishop, R.; Bouchier, C.; et al. Genome of the host-cell transforming parasite Theileria annulata compared with T. parva. Science 2005, 309, 131–133. [Google Scholar] [CrossRef] [PubMed]
- Hayashida, K.; Hara, Y.; Abe, T.; Yamasaki, C.; Toyoda, A.; Kosuge, T.; Suzuki, Y.; Sato, Y.; Kawashima, S.; Katayama, T.; et al. Comparative genome analysis of three eukaryotic parasites with differing abilities to transform leukocytes reveals key mediators of Theileria-induced leukocyte transformation. mBio 2012, 3, e00204-12. [Google Scholar] [CrossRef] [PubMed]
- Tarigo, J.L.; Scholl, E.H.; McK Bird, D.; Brown, C.C.; Cohn, L.A.; Dean, G.A.; Levy, M.G.; Doolan, D.L.; Trieu, A.; Nordone, S.K.; et al. A novel candidate vaccine for cytauxzoonosis inferred from comparative apicomplexan genomics. PLoS ONE 2013, 8, e71233. [Google Scholar] [CrossRef]
- Anziani, O.S.; Guglielmone, A.A.; Abdala, A.A.; Aguirre, D.H.; Mangold, A.J. Acquired protection in Frisian steers with Babesia bovis attenuated vaccine. Rev. Med. Vet. 1993, 74, 47–49. [Google Scholar]
- Levy, M.G.; Ristic, M. Babesia bovis: Continuous cultivation in a microaerophilous stationary phase culture. Science 1980, 207, 1218–1220. [Google Scholar] [CrossRef]
- Ueti, M.W.; Johnson, W.C.; Kappmeyer, L.S.; Herndon, D.R.; Mousel, M.R.; Reif, K.E.; Taus, N.S.; Ifeonu, O.O.; Silva, J.C.; Suarez, C.E.; et al. Comparative analysis of gene expression between Babesia bovis blood stages and kinetes allowed by improved genome annotation. Int. J. Parasitol. 2020. [Google Scholar] [CrossRef]

| Set | Lineage | Activity | Addition of: | N-Glycan Transfer | ||||
|---|---|---|---|---|---|---|---|---|
| 1st NAcGlc | 2nd NAcGlc | Man | ||||||
| Enzyme Species | Alg7 | Alg13 | Alg14 | Alg1 | STT3 | OST1 | ||
| i | Clade IV | T. equi | XP_004830167 | XP_004829088 | XP_004830080 | XP_004832122 | XP_004832354 | XP_004831427 |
| Clade IIIa | C. felis | CF000673 | CF001791 | CF002639 | CF004094 | CF001242 | CF000008 | |
| ii | Clade VI Babesia s.s. | B. bovis | XP_001609261 | XP_001610083 | XP_001610338 | - | XP_001609551 | XP_001611482 |
| B. bigemina | XP_012769992 | XP_012766341 | - | - | XP_012768251 | XP_012769449 | ||
| B. divergens | Bdiv_013430c | Bdiv_003540c | Bdiv_029110 | - | Bdiv_030100 | Bdiv_036510 | ||
| Babesia sp. Xinjang | XP_028870826 | XP_028872191 | - | - | XP_028869940 | - | ||
| B. ovata | - | BOVATA_021750 | - | BOVATA_014210 | BOVATA_008000 | |||
| iii | Clade I | B. microti | XP_021337988 | - | - | - | - | - |
| Clade V Theileria s.s. | T. annulata | XP_954592 | - | - | - | - | - | |
| T. parva | XP_765645 | XP_765081 | KAF5153350.1 | - | - | - | ||
| T. orientalis | XP_009688943 | XP_009690544 | XP_009689017 | - | - | - | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Florin-Christensen, M.; Rodriguez, A.E.; Suárez, C.E.; Ueti, M.W.; Delgado, F.O.; Echaide, I.; Schnittger, L. N-Glycosylation in Piroplasmids: Diversity within Simplicity. Pathogens 2021, 10, 50. https://doi.org/10.3390/pathogens10010050
Florin-Christensen M, Rodriguez AE, Suárez CE, Ueti MW, Delgado FO, Echaide I, Schnittger L. N-Glycosylation in Piroplasmids: Diversity within Simplicity. Pathogens. 2021; 10(1):50. https://doi.org/10.3390/pathogens10010050
Chicago/Turabian StyleFlorin-Christensen, Monica, Anabel E. Rodriguez, Carlos E. Suárez, Massaro W. Ueti, Fernando O. Delgado, Ignacio Echaide, and Leonhard Schnittger. 2021. "N-Glycosylation in Piroplasmids: Diversity within Simplicity" Pathogens 10, no. 1: 50. https://doi.org/10.3390/pathogens10010050
APA StyleFlorin-Christensen, M., Rodriguez, A. E., Suárez, C. E., Ueti, M. W., Delgado, F. O., Echaide, I., & Schnittger, L. (2021). N-Glycosylation in Piroplasmids: Diversity within Simplicity. Pathogens, 10(1), 50. https://doi.org/10.3390/pathogens10010050









