Abstract
Major advances in sequencing technologies and targeted therapies have accelerated the incorporation of oncology into the era of precision medicine and “biomarker-driven” treatments. However, the impact of this approach on the everyday clinic has yet to be determined. Most precision oncology reports are based on developed countries and usually involve metastatic, hard-to-treat or incurable cancer patients. Moreover, in many cases race and ethnicity in these studies is commonly unreported and real-world evidence in this topic is scarce. Herein, we report data from a total of 202 Chilean advanced stage refractory cancer patients. Retrospectively, we collected patient data from NGS tests and IHC in order to determine the proportion of patients that would benefit from targeted treatments. Overall >20 tumor types were included in our cohort and 37% of patients (n = 74) displayed potentially actionable alterations, including on-label, off-label and immune checkpoint inhibitor recommendations. Our findings were in-line with previous reports such as the cancer genome atlas (TCGA). To our knowledge, this is the first report of its kind in Latin America delivering real-world evidence to estimate the percentage of refractory tumor patients that might benefit from precision oncology. Although this approach is still in its infancy in Chile, we strongly encourage the implementation of mutational tumor boards in our country in order to provide more therapeutic options for advanced stage refractory patients.
1. Introduction
The ultimate goal of precision oncology is to deliver the right cancer treatment to the right patient in a timely manner. In recent years, the rapid development of massively parallel sequencing technologies (also called next generation sequencing; NGS) and the progressive reduction of its associated costs have allowed characterization of the mutational landscape of cancer genomes, identifying “actionable” targets and “driver” mutations responsible for tumor growth and progression. Furthermore, the combined use of NGS gene-panels that expand the number of genes that can be simultaneously analyzed, along with immunohistochemistry (IHC)-based techniques, increases the possibility of finding a matching targeted therapy. Successful examples include anti-HER2 therapy for HER2/neu overexpressing gastric or breast cancer patients [1,2,3,4,5,6], the use of vemurafenib for BRAF-V600E mutant metastatic melanoma patients [7,8,9], or PARP-inhibitors for BRCA-mutant advanced ovarian [10], metastatic breast [11,12] and metastatic pancreatic adenocarcinoma patients [13,14].
Despite major advances in this field, the impact and clinical utility of this approach especially on advanced stage patient survival remains sparse. In most cases, phase II clinical trials demonstrate the effectiveness of targeted therapies. Unfortunately, these studies are commonly focused on a small subset of cancer patients that harbor specific alterations and/or clinical characteristics. In contrast, precision oncology reports on real world data are somewhat scarce. These reports are mainly based on North American, European and Asian countries. Furthermore, race is often unreported in clinical–genomic studies; thereby minorities are usually underrepresented. In fact, NGS studies based in Latin America are extremely scant. Given the geographical heterogeneity observed for some malignancies, precision-oncology real-world evidence studies in specific regions could imply a differential impact of this approach on these geographical areas.
Herein, we retrospectively analyzed the clinical utility of tumor profiling by NGS/IHC in the routine practice across several Chilean tertiary health centers. We also assessed the final decision made by oncologists following the recommendation. To our knowledge, this is the first report of its kind in Latin America.
2. Materials and Methods
2.1. Study Design, Patients and Tumor Molecular Profile
This research was approved by the scientific ethics committee affiliated to the school of medicine of the Pontificia Universidad Catolica de Chile (approval ID#200121002, dated on 30 April 2020). Patients diagnosed with advanced or metastatic solid cancer and who underwent tumor genomic profiling were eligible for inclusion in this study. A total of 202 patients were included. The different platforms used for tumor characterization are included in Supplementary Figure S1. Briefly, in 202 patients the tumor DNA was sequenced through NGS (different number of genes depending on the platform) and, of these, in 117 the result also included the tumor mutational burden (TMB). Furthermore, in 124 patients microsatellite status was determined using DNA sequencing (NGS or Sanger). Finally, PDL1 and HER2 expression was determined using immunohistochemistry (IHC) in 126 and 128 patients, respectively. Commercial companies independently performed all assays (Figure S1). Details of each of the platforms used can be found in Supplementary Table S1. Information on cancer type, platform, and type of sample used is also included in Supplementary Table S2.
2.2. Criteria for Actionability
In order to establish the potential clinical applicability of the genomic alteration found in each patient, we used OncoKB [15], which is an expert-guided precision oncology knowledge database that assigns each variant to different levels of evidence corresponding to its actionability. We selected high levels (1 and 2) of evidence to establish the potential recommendations on label and off label, where level 1 corresponds to a FDA-recognized biomarker predictive of response to an FDA-approved drug in this indication and level 2 corresponds to a standard of care biomarker recommended by the NCCN or other expert panels, predictive of response to an FDA-approved drug in this indication [15]. Furthermore, we established whether patients could potentially benefit from immunotherapy, according to the following criteria: (1) microsatellite instability (MSI), (2) High/Medium expression of PDL1 according to molecular report and (3) High/Medium TMB, TMB-level defined as low ≤ 5 mut/Mb), intermediate > 5 mut/Mb, ≤ 15 mut/Mb), or high > 15 mut/Mb, based on the thresholds described previously [16].
2.3. Comparison of Molecular Alterations to the Cancer Genome Atlas (TCGA) Cohorts
A descriptive comparison of the frequency of molecular alterations between the six main tumor types in our study and matched cohorts of the TCGA was performed. The frequencies were obtained using the bio-portal database [17]. The studies selected for each type of tumor were: colorectal cancer [18], pancreatic cancer [19], sarcoma [20], breast cancer [21,22], ovarian cancer [23], lung cancer [24].
2.4. Oncologists’ Survey
The usefulness of the molecular test in clinical decision was evaluated in a subset of 46 patients through a survey applied to oncologists. The question was: “Has the treatment decision changed based on the results generated by the molecular test?”.
3. Results
A total of 202 advanced stage patients were included in our study. Main cancer types were gastrointestinal (n = 92; 45%) followed by sarcomas/unknown primary (n = 36; 18%), and gynecological/breast (n = 33; 16%). The detailed distribution of tumor types is summarized in Table 1. Overall, 77% of patients had at least one alteration confirmed by NGS.

Table 1.
Distribution of cancer types in this cohort.
First, we sought to determine the mutational landscape of tumors. Our workflow is depicted in Supplementary Figure S1. Briefly, the entire cohort (n = 202) was analyzed by NGS using gene panels at various platforms. Patient subsets were analyzed by IHC to determine HER2 (n = 126) and PDL1 (n = 128) expression. MSI status and TMB were also analyzed by a combination of NGS/IHC or NGS in patient subsets (n = 124 and n = 117, respectively). Mutational landscapes on analyzed tumors are displayed in Figure 1. As pointed out, gastrointestinal cancers were the most prevalent in our cohort. Among these, KRAS (51%) was the most frequently altered gene, mainly due to colorectal and pancreatic cancers followed by TP53, APC, ARID1A, SMAD4 and PIK3CA. Please note the low number of alterations in gallbladder and hepatic tumors. For genitourinary tumors ERBB2 and KIT were the most commonly altered genes, but the number of alterations in these tumors was generally low. As expected, TP53 was the most frequently altered gene on gynecological tumors, followed by ARID1A and PIK3CA. Similarly, BRCA1 and BRCA2 truncations were found in these tumors. As expected PIK3CA mutations and CCND1 amplifications were also recurrent among breast cancer patients; TP53 was frequently altered in lung cancer patients and in those with unknown primary tumors. On the other hand, sarcomas were characterized by a low number of alterations. Other less prevalent tumor types analyzed included CNS and NETs.
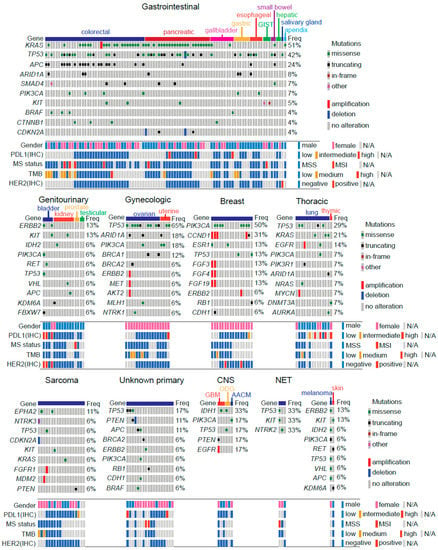
Figure 1.
Mutational landscape of 202 Chilean advanced stage refractory cancer patients. Tumors were grouped as: gastrointestinal, genitourinary, gynecologic, breast, thoracic, sarcoma, unknown primary, central nervous system (CNS), neuroendocrine tumors (NET), melanoma and skin. Abbreviations: GIST: gastrointestinal stromal tumor; MS: microsatellite; TMB: tumor mutational burden; GBM: glioblastoma multiforme; ODG: oligodendroglioma; AACM: anaplastic astrocytoma.
Next, we sought to compare these results with the TCGA database. Figure 2 shows that the frequency of most gene alterations was similar. However, there were some notorious exceptions on well-known relevant genes for certain cancer types, for example, KRAS and APC in colorectal cancer: 55% and 50% in our cohort vs. 35% and 75% in TCGA, respectively; also, BRCA2 and SMAD4 in pancreatic cancer: 8% and 8% in our cohort vs. 2% and 23% in TCGA; EPHA2 on sarcoma: 11% in our cohort vs. 0%; TP53 and ESR1 in breast cancer: 13% and 13% in our cohort vs. 34% and 0% in TCGA. Other examples were TP53 and ARID1A in ovarian cancer, and TP53 and KRAS in lung cancer.
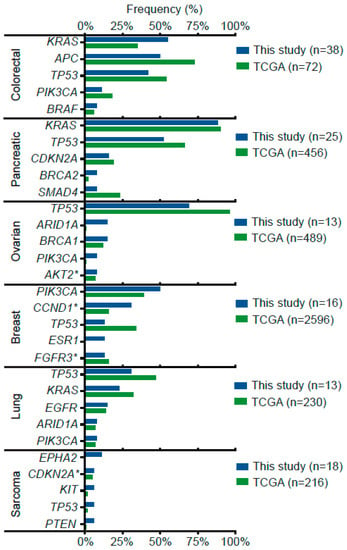
Figure 2.
Most frequent gene alterations (top 5) by cancer type in our cohort versus TCGA cohorts. * Indicates copy number alteration, amplification or deletion.
Our tumor analyses also included immunohistochemistry for certain factors (Supplementary Figure S1). In particular lung and pancreatic cancers displayed medium/high levels of PDL1 expression. Interestingly, PDL1 positivity correlated with HER2+ in bladder cancer patients. Additionally, we determined TMB and microsatellite instability in a subset of patients; in general colorectal cancer patients showed the highest positivity for these markers (Figure 1).
Based on this molecular analysis, 75 out of 202 (37%) patients received informed treatment recommendations that included on-label, off-label and immune checkpoint inhibitor (ICI) recommendations. Overall, 18 out of 202 treatment recommendations (9%) were on-label therapies; within this group, the fulvestrant + alpelisib combination and PARP inhibitors were the most frequently recommended with 39% and 17%, respectively (Figure 3A, left panel). On the other hand, 38 patients (19%) received off-label recommendations. Among these, anti-PI3K, PARP inhibitors and imatinib were the most commonly recommended with 32%, 21% and 18%, respectively (Figure 3A, right panel). Then, ICIs were recommended in 42 out of 202 patients (21%), based on medium/high PDL1 expression, medium/high TMB or MSI status (Figure 3B). Next, we analyzed the percentage of recommendations by cancer type. As shown in Figure 3C the highest percentage of on-label recommendation was observed in breast, followed by ovarian cancer patients. These tumors also displayed the highest rates of overall recommendations (on-label, off-label and ICI). Similarly, lung and colorectal cancer patients had relatively high levels of recommendation. In contrast, pancreatic tumors and sarcomas had the lowest rates of therapy recommendation based on actionable alterations.
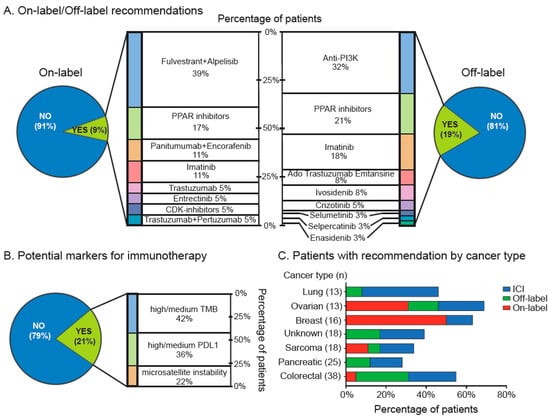
Figure 3.
Clinical actionability for targeted therapies. (A) On-label (left panel) and off-label (right panel) and (B) immune checkpoint inhibitor recommendations based in NGS/IHC profiles. (C) Percentages of recommendation (on-label, off-label or ICI) by cancer type.
Next, we analyzed a small subset of patients (n = 35) from our cohort who had available clinical information. Among these, only four received targeted therapies based on molecular testing: one of them displayed stable disease (to Nivolumab-Ipilimumab), one had a partial response to Olaparib, one had a complete response (to Imatinib) and the other progressed after Regorafenib. The characteristics and outcomes in these patients are summarized in Supplementary Table S3. On the other hand, the clinical characteristics, previous lines of therapy and time between the diagnosis and the molecular test of the patients who did not receive target therapy are detailed in Supplementary Table S4.
Finally, we applied a brief survey to a group of oncologists that made treatment decisions in a subset of 46 patients. The survey asked about the influence of molecular profiling upon treatment choices. Overall, oncologists estimated that molecular testing was useful for clinical management in 50% of the evaluated cases.
4. Discussion
Major advances in massively parallel high-throughput sequencing technologies and the concomitant development of molecularly targeted therapies have shifted the paradigm in oncology from “one size fits all-standard of care” to “biomarker-driven” treatments. Despite this, the true impact of precision oncology in everyday clinical practice remains to be determined. Previous studies have sought to quantify the contribution of these strategies, but several factors must be taken into consideration for such analysis. A German single-center retrospective study analyzed a total of 198 metastatic cancer patients in the context of implementation of a molecular tumor board [25]. Specific treatment recommendations were given to a subset of 104 patients, which included 9.6% off-label targeted therapies (n = 19), 9% combined treatments (n = 18), 18.1% ICI (n = 36) and 6.5% trial inclusions (n = 13). Importantly, in 68% of these cases (n = 71) recommendations were not implemented. A second, similar study retrospectively analyzed 600 incurable solid-tumor patients; 51.7% (n = 310) received therapy recommendations and 15.8% (n = 95) received genomic-driven recommendations; from these 7.09% (n = 22) were on-label and 14.9% (n = 45) were off-label [26]. Surprisingly, and despite the heterogeneity of our cohort, these percentages are in line with our findings which indicated 9% of on-label, 19% off-label and 21% of ICI therapy recommendations (Figure 3). In terms of actionable targets, our cohort displayed a 37% rate of actionability; this is also in line with a recent report of a genomically guided clinical trial coordinated by the NCI that collected a total of 5954 refractory tumor biopsies across >1000 centers [27]. This trial found actionable targets in 37.6% of patients and tested the feasibility of large-scale screenings of molecular alterations by NGS.
Clearly, the approach of our study, which combined NGS and IHC, increases actionability rates in our cohort. Other studies confirm that the addition of IHC or other tests increases the usefulness of the information obtained from NGS studies alone [28]. As noted, a survey applied to oncologists revealed that 50% estimated that molecular testing was useful for clinical management. Similarly, the above mentioned study indicates that 60.4% of the oncologists followed the recommendations of molecular profile reports. Furthermore, a nationally representative survey of oncologists in the US found that 75.6% use NGS tests to guide treatment decisions, including those for advanced refractory cancers, for clinical trial eligibility and for off-label use of therapeutic agents [29].
As explained above, precision oncology reports are usually based on developed countries. Moreover, patient race in these studies goes usually unreported. A systematic review that included a total of >15,000 patients across 231 studies found that only 37% and 17% reported race and ethnicity, respectively [30]. Within this context, Latin American-based reports are extremely rare. To our knowledge, our report is the first of its kind within the region delivering real-world evidence on the mutational landscape in a cohort of 202 Latin American-ancestry advanced stage cancer patients. Our data included >20 tumor types, and besides NGS tests we assessed HER2 and PDL1 expression, MSI status and TMB in specific subsets. As shown in Figure 2, when we analyzed the most frequently altered genes by cancer type in our cohort versus TCGA, we found some coincidences but also some notorious discrepancies. These can be attributed to some of the limitations of our study (listed below) including number of samples, differences in NGS panels (as shown in Supplementary Figure S1 and Supplementary Table S1), tumor stage and previous treatments received by the patients. As an example, the high frequency of ESR1 in our cohort versus TCGA is probably due to the inclusion of a patient with luminal tumors (ER+) who received hormone therapy, and therefore represents a selection bias. Previously, other authors have analyzed the main obstacles to the implementation of precision medicine programs in the region; these are mainly related to high costs and limited (or lack of) access to health coverage [31]. These studies also emphasize the need for locally generated data. Evidently, ethnic and lifestyle influences translate into a wide spectrum of somatic mutations and gene polymorphisms causing differences in biomarkers and treatment response. In this regard, mutational tumor boards inside health institutions are yet to be implemented in Chile and should be encouraged, especially for high incidence/mortality malignancies such as gastric cancers.
Perhaps the major limitation of our study is the lack of data on therapeutic impact. However, we obtained clinical information for a small subset of patients (Supplementary Tables S3 and S4). Interestingly, the average number of therapies received in this subset was 3.1 and the average elapsed time between diagnosis and molecular testing was 34.9 months. This might explain the low number of patients that are effectively treated with targeted therapies. Furthermore, these patients usually have a very limited set of therapeutic alternatives after several lines of treatment. This also explains the negative results, or the lack of benefit reported in several studies. Throughout the literature the debate on this topic is still ongoing; in 2015 a large French open-label phase II trial sought to determine the impact of molecularly targeted therapies on patient survival. This study involved eight centers and enrolled a total of 741 metastatic refractory cancers (any type). From these, 195 patients were randomly assigned to either control (n = 96) or experimental (n = 99) arms [32]. Investigators found no benefit in terms of PFS by the use of molecularly targeted therapies versus the physician’s choice. Furthermore, the study recommended enrollment into clinical trials rather than the off-label use of these agents. Importantly, targeted therapies in this study were restricted to 10 available regimens. Subsequently, in 2017 a single-center single-arm trial in the US enrolled a total of 1035 adult advanced hard-to treat cancer patients. Based on genomic analyses a subset of 199 patients were scheduled to received matched therapy according to actionable targets [33]. The clinical benefit was measured as a ratio of PFS on matched therapy (called PFS2) over PFS on prior therapy (called PFS1), setting a threshold at PFS2/PFS1 = 1.3. Overall, 193 patients were evaluated for PFS and 33% (n = 63) had a PFS2/PFS1 > 1.3. Obviously, these results should be further validated, but they suggest that at least a fraction of hard-to-treat patients could benefit from this approach. More recently, a smaller single-center study compared the outcomes of a group of 22 metastatic cancer patients which received genomically-guided targeted therapies versus 22 matched control patients treated with either chemotherapy or best supportive care [34]. This study demonstrated a significant increase in median OS by the use of targeted therapies. Interestingly, the study also found that patients under targeted therapies had lower weekly costs during their treatment versus controls. Other limitations of our work include the lack of patients’ clinical data such as tumor histology or patient performance status. Furthermore, (1) This was a highly heterogeneous convenience sample, and in many cases the numbers are too small. (2) Data were collected from private health centers. In general, these centers serve higher income patients. On the other hand the Chilean population is highly heterogeneous with a strong European ancestry. Unfortunately this information could not be collected for this study and therefore could represent another bias. Evidently, the frequencies and/or the distribution of cancer types may not reflect the Chilean reality. (3) Finally, as shown by Supplementary Figure S1, a variety of NGS assays were used for this cohort, which further increases the heterogeneity of the sample and may also represent an analysis bias.
5. Conclusions
In summary, our study provides the first Latin American-based precision oncology report delivering real-world evidence. Our data suggest that a 37% of advanced refractory cancer patients displayed actionable targets. Therefore, as suggested by others, our findings indicate that a relatively small but significant percentage of patients could benefit from this approach, especially when NGS tests are combined with IHC. Although the implementation of molecular tumor boards is still pending in Chile, this should be encouraged and incorporated into clinical practice.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/jpm12020195/s1, Supplementary Figure S1: Workflow of the study. Supplementary Table S1: Main characteristic of the platform used for tumor profiling in our patients cohort. Supplementary Table S2: Platforms used for tumor molecular profile according to cancer type, Supplementary Table S3: Clinical characteristics and main outcomes of selected patients on targeted therapy based on molecular testing, Supplementary Table S4: Clinical characteristics of selected patients without targeted therapy based on molecular testing.
Author Contributions
Conceptualization: M.C.-D., M.P.P. and M.G.; methodology, M.C.-D., M.P.P., C.R., L.C., J.M.R., L.I., A.M., P.R., A.F., M.M., M.E.L., J.G., M.P.R.Z., A.B., P.M., J.A.R., B.G.-B. and M.G.; software, M.C.-D. and M.P.P.; formal analysis, M.C.-D., M.P.P. and M.G.; investigation, M.C.-D., M.P.P., C.R., L.C., J.M.R., L.I., A.M., P.R., A.F., M.M., M.E.L., J.G., M.P.R.Z., A.B., P.M., J.A.R., B.G.-B. and M.G.; resources, C.R., L.C., J.M.R., L.I., A.M., P.R., A.F., M.M., M.E.L., J.G., M.P.R.Z., A.B., P.M., J.A.R., B.G.-B. and M.G.; writing—original draft preparation, M.C.-D., M.P.P. and M.G.; writing—review and editing, M.C.-D., M.P.P., C.R., L.C., J.M.R., L.I., A.M., P.R., A.F., M.M., M.E.L., J.G., M.P.R.Z., A.B., P.M., J.A.R., B.G.-B. and M.G.; supervision, J.G., M.P.R.Z., A.B., P.M., J.A.R., B.G.-B. and M.G.; project administration, M.G. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
The study was conducted according to the guidelines of the Declaration of Helsinki, and approved by the Health Sciences and Scientific Ethics Committee at the Universidad Catolica (approval ID# 012321, 20 April 2020).
Informed Consent Statement
All patients signed a written consent form to participate including a consent for publication at the time of enrollment. This procedure was approved by the Health Sciences and Scientific Ethics Committee at the Universidad Catolica (approval ID# 012321, 20 April 2020). Publication of anonymized data was authorized by the Scientific Ethics committee at the Eastern Metropolitan Health Service (ID# CECSSMO040521, 4 May 2021).
Data Availability Statement
The data presented in this study are available on request from the corresponding author.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Bang, Y.-J.; Van Cutsem, E.; Feyereislova, A.; Chung, H.C.; Shen, L.; Sawaki, A.; Lordick, F.; Ohtsu, A.; Omuro, Y.; Satoh, T.; et al. Trastuzumab in combination with chemotherapy versus chemotherapy alone for treatment of HER2-positive advanced gastric or gastro-oesophageal junction cancer (ToGA): A phase 3, open-label, randomised controlled trial. Lancet 2010, 376, 687–697. [Google Scholar] [CrossRef]
- Van Cutsem, E.; Bang, Y.-J.; Feng-Yi, F.; Xu, J.M.; Lee, K.-W.; Jiao, S.-C.; Chong, J.L.; López-Sanchez, R.I.; Price, T.; Gladkov, O.; et al. HER2 screening data from ToGA: Targeting HER2 in gastric and gastroesophageal junction cancer. Gastric Cancer Off. J. Int. Gastric Cancer Assoc. Jpn. Gastric Cancer Assoc. 2015, 18, 476–484. [Google Scholar] [CrossRef] [PubMed]
- Romond, E.H.; Perez, E.A.; Bryant, J.; Suman, V.J.; Geyer, C.E.; Davidson, N.E.; Tan-Chiu, E.; Martino, S.; Paik, S.; Kaufman, P.A.; et al. Trastuzumab plus adjuvant chemotherapy for operable HER2-positive breast cancer. N. Engl. J. Med. 2005, 353, 1673–1684. [Google Scholar] [CrossRef] [Green Version]
- Piccart-Gebhart, M.J.; Procter, M.; Leyland-Jones, B.; Goldhirsch, A.; Untch, M.; Smith, I.; Gianni, L.; Baselga, J.; Bell, R.H.; Jackisch, C.; et al. Trastuzumab after adjuvant chemotherapy in HER2-positive breast cancer. N. Engl. J. Med. 2005, 353, 1659–1672. [Google Scholar] [CrossRef] [Green Version]
- Joensuu, H.; Kellokumpu-Lehtinen, P.-L.; Bono, P.; Alanko, T.; Kataja, V.; Asola, R.; Utriainen, T.; Kokko, R.; Hemminki, A.; Tarkkanen, M.; et al. Adjuvant Docetaxel or Vinorelbine with or without Trastuzumab for Breast Cancer. N. Engl. J. Med. 2006, 354, 809–820. [Google Scholar] [CrossRef] [Green Version]
- Slamon, D.J.; Leyland-Jones, B.; Shak, S.; Fuchs, H.; Paton, V.; Bajamonde, A.; Fleming, T.; Eiermann, W.; Wolter, J.; Pegram, M.; et al. Use of Chemotherapy plus a Monoclonal Antibody against HER2 for Metastatic Breast Cancer That Overexpresses HER2. N. Engl. J. Med. 2001, 344, 783–792. [Google Scholar] [CrossRef]
- Flaherty, K.T.; Puzanov, I.; Kim, K.B.; Ribas, A.; McArthur, G.A.; Sosman, J.A.; O’Dwyer, P.J.; Lee, R.J.; Grippo, J.F.; Nolop, K.; et al. Inhibition of Mutated, Activated BRAF in Metastatic Melanoma. N. Engl. J. Med. 2010, 363, 809–819. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sosman, J.A.; Kim, K.B.; Schuchter, L.; Gonzalez, R.; Pavlick, A.C.; Weber, J.S.; McArthur, G.A.; Hutson, T.E.; Moschos, S.J.; Flaherty, K.T.; et al. Survival in BRAF V600–Mutant Advanced Melanoma Treated with Vemurafenib. N. Engl. J. Med. 2012, 366, 707–714. [Google Scholar] [CrossRef] [Green Version]
- Chapman, P.B.; Hauschild, A.; Robert, C.; Haanen, J.B.; Ascierto, P.; Larkin, J.; Dummer, R.; Garbe, C.; Testori, A.; Maio, M.; et al. Improved Survival with Vemurafenib in Melanoma with BRAF V600E Mutation. N. Engl. J. Med. 2011, 364, 2507–2516. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kaye, S.B.; Lubinski, J.; Matulonis, U.; Ang, J.E.; Gourley, C.; Karlan, B.Y.; Amnon, A.; Bell-McGuinn, K.M.; Chen, L.-M.; Friedlander, M.; et al. Phase II, open-label, randomized, multicenter study comparing the efficacy and safety of olaparib, a poly (ADP-ribose) polymerase inhibitor, and pegylated liposomal doxorubicin in patients with BRCA1 or BRCA2 mutations and recurrent ovarian cancer. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2012, 30, 372–379. [Google Scholar] [CrossRef] [PubMed]
- Robson, M.; Im, S.-A.; Senkus, E.; Xu, B.; Domchek, S.M.; Masuda, N.; Delaloge, S.; Li, W.; Tung, N.; Armstrong, A.; et al. Olaparib for Metastatic Breast Cancer in Patients with a Germline BRCA Mutation. N. Engl. J. Med. 2017, 377, 523–533. [Google Scholar] [CrossRef]
- Litton, J.K.; Rugo, H.S.; Ettl, J.; Hurvitz, S.A.; Gonçalves, A.; Lee, K.-H.; Fehrenbacher, L.; Yerushalmi, R.; Mina, L.A.; Martin, M.; et al. Talazoparib in Patients with Advanced Breast Cancer and a Germline BRCA Mutation. N. Engl. J. Med. 2018, 379, 753–763. [Google Scholar] [CrossRef] [PubMed]
- Golan, T.; Hammel, P.; Reni, M.; Van Cutsem, E.; Macarulla, T.; Hall, M.J.; Park, J.-O.; Hochhauser, D.; Arnold, D.; Oh, D.-Y.; et al. Maintenance Olaparib for Germline BRCA-Mutated Metastatic Pancreatic Cancer. N. Engl. J. Med. 2019, 381, 317–327. [Google Scholar] [CrossRef]
- Kaufman, B.; Shapira-Frommer, R.; Schmutzler, R.K.; Audeh, M.W.; Friedlander, M.; Balmaña, J.; Mitchell, G.; Fried, G.; Stemmer, S.M.; Hubert, A.; et al. Olaparib monotherapy in patients with advanced cancer and a germline BRCA1/2 mutation. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2015, 33, 244–250. [Google Scholar] [CrossRef] [PubMed]
- Chakravarty, D.; Gao, J.; Phillips, S.; Kundra, R.; Zhang, H.; Wang, J.; Rudolph, J.E.; Yaeger, R.; Soumerai, T.; Nissan, M.H.; et al. OncoKB: A Precision Oncology Knowledge Base. JCO Precis. Oncol. 2017, 2017, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Zhou, K.I.; Peterson, B.F.; Serritella, A.; Thomas, J.; Reizine, N.; Moya, S.; Tan, Y.-H.C.; Wang, Y.; Catenacci, D.V.T. Spatial and temporal heterogeneity of PD-L1 expression and tumor mutational burden in gastroesophageal adenocarcinoma at baseline diagnosis and after chemotherapy. Clin. Cancer Res. 2020, 26, 6453–6463. [Google Scholar] [CrossRef]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio Cancer Genomics Portal: An Open Platform for Exploring Multidimensional Cancer Genomics Data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef] [Green Version]
- Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012, 487, 330–337. [CrossRef] [Green Version]
- Bailey, P.; Chang, D.K.; Nones, K.; Johns, A.L.; Patch, A.-M.; Gingras, M.-C.; Miller, D.K.; Christ, A.N.; Bruxner, T.J.C.; Quinn, M.C.; et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 2016, 531, 47–52. [Google Scholar] [CrossRef]
- Barretina, J.; Taylor, B.S.; Banerji, S.; Ramos, A.H.; Lagos-Quintana, M.; DeCarolis, P.L.; Shah, K.; Socci, N.D.; Weir, B.A.; Ho, A.; et al. Subtype-specific genomic alterations define new targets for soft-tissue sarcoma therapy. Nat. Genet. 2010, 42, 715–721. [Google Scholar] [CrossRef] [Green Version]
- The Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 2012, 490, 61–70. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pereira, B.; Chin, S.F.; Rueda, O.M.; Vollan, H.K.; Provenzano, E.; Bardwell, H.A.; Pugh, M.; Jones, L.; Russell, R.; Sammut, S.J.; et al. The somatic mutation profiles of 2433 breast cancers refines their genomic and transcriptomic landscapes. Nat. Commun. 2016, 7, 11479. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- The Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature 2011, 474, 609–615. [Google Scholar] [CrossRef] [PubMed]
- The Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature 2014, 511, 543. [Google Scholar] [CrossRef]
- Hoefflin, R.; Geißler, A.-L.; Fritsch, R.; Claus, R.; Wehrle, J.; Metzger, P.; Reiser, M.; Mehmed, L.; Fauth, L.; Heiland, D.H.; et al. Personalized Clinical Decision Making Through Implementation of a Molecular Tumor Board: A German Single-Center Experience. JCO Precis. Oncol. 2018, 2, 1–16. [Google Scholar] [CrossRef]
- Sadaps, M.; Funchain, P.; Mahdi, H.; Grivas, P.; Pritchard, A.; Klek, S.; Estfan, B.; Abraham, J.; Budd, G.T.; Stevenson, J.P.; et al. Precision Oncology in Solid Tumors: A Longitudinal Tertiary Care Center Experience. JCO Precis. Oncol. 2018, 13, 1–11. [Google Scholar] [CrossRef]
- Flaherty, K.T.; Gray, R.J.; Chen, A.P.; Li, S.; McShane, L.M.; Patton, D.; Hamilton, S.R.; Williams, P.M.; Iafrate, A.J.; Sklar, J.; et al. Molecular Landscape and Actionable Alterations in a Genomically Guided Cancer Clinical Trial: National Cancer Institute Molecular Analysis for Therapy Choice (NCI-MATCH). J. Clin. Oncol. 2020, 38, 3883–3894. [Google Scholar] [CrossRef]
- Laes, J.F.; Aftimos, P.; Barthelemy, P.; Bellmunt, J.; Berchem, G.; Camps, C.; Peñas, R.D.L.; Finzel, A.; García-Foncillas, J.; Hervonen, P.; et al. The clinical impact of using complex molecular profiling strategies in routine oncology practice. Oncotarget 2018, 9, 20282–20293. [Google Scholar] [CrossRef] [Green Version]
- Freedman, A.N.; Klabunde, C.N.; Wiant, K.; Enewold, L.; Gray, S.; Filipski, K.K.; Keating, N.L.; Leonard, D.G.; Lively, T.; McNeel, T.S.; et al. Use of Next-Generation Sequencing Tests to Guide Cancer Treatment: Results from a Nationally Representative Survey of Oncologists in the United States. JCO Precis. Oncol. 2018, 2, 1–13. [Google Scholar] [CrossRef]
- Nugent, A.; Conatser, K.R.; Turner, L.L.; Nugent, J.T.; Sarino, E.M.B.; Ricks-Santi, L.J. Reporting of race in genome and exome sequencing studies of cancer: A scoping review of the literature. Genet. Med. 2019, 21, 2676–2680. [Google Scholar] [CrossRef] [Green Version]
- Calderón-Aparicio, A.; Orue, A. Precision oncology in Latin America: Current situation, challenges and perspectives. Ecancermedicalscience 2019, 13, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Le Tourneau, C.; Delord, J.-P.; Gonçalves, A.; Gavoille, C.; Dubot, C.; Isambert, N.; Campone, M.; Tredan, O.; Massiani, M.-A.; Mauborgne, C.; et al. Molecularly targeted therapy based on tumour molecular profiling versus conventional therapy for advanced cancer (SHIVA): A multicentre, open-label, proof-of-concept, randomised, controlled phase 2 trial. Lancet Oncol. 2015, 16, 1324–1334. [Google Scholar] [CrossRef]
- Massard, C.; Michiels, S.; Ferté, C.; Le Deley, M.-C.; Lacroix, L.; Hollebecque, A.; Verlingue, L.; Ileana, E.; Rosellini, S.; Ammari, S.; et al. High-throughput genomics and clinical outcome in hard-to-treat advanced cancers: Results of the MOSCATO 01 trial. Cancer Discov. 2017, 7, 586–595. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haslem, D.S.; Chakravarty, I.; Fulde, G.; Gilbert, H.; Tudor, B.P.; Lin, K.; Ford, J.M.; Nadauld, L.D. Precision oncology in advanced cancer patients improves overall survival with lower weekly healthcare costs. Oncotarget 2018, 9, 12316–12322. [Google Scholar] [CrossRef] [PubMed] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).