Decoding Uncertainty Quantification for Oncology—An Illustration Using Radiomics
Abstract
1. Introduction
What Is Prediction Accuracy and Prediction Uncertainty?
| Somekey terms and their explanations Machine Learning (ML)—A subset of AI involving algorithms that learn patterns from data and improve performance on a task without being explicitly programmed for each scenario. ML methods are commonly used in risk prediction, image analysis, and clinical outcome modelling. Features: The variables that contribute towards the model’s output. Feature space represents a multi-dimensional space where each dimension corresponds to a specific feature of the given data. Every patient represents a unique data point in feature space. The dimensionality of the feature space corresponds to the number of features in the model. Example: Imagine plotting patients on a graph where one axis is age and the other is blood pressure; age and blood pressure are features, the 2D graph is the feature space, each patient is a data point on it, and adding more features like heart rate or cholesterol would add more axes (dimensions) to this space. Distance function: In the feature space, the notion of distance allows us to quantify how similar or different our features are collectively in a dataset. The distance between the points helps us identify clusters of similar or different classes. Decision boundary is typically an equation (a line, curve, or surface) that uses several features together (e.g., age + blood sugar + BMI), and the model decides based on which side of that surface a data point lies. For complex ML models (e.g., decision trees, random forests, deep nets), the boundary can be an irregular, wiggly surface, but conceptually it is still “the place where the prediction flips from one class to another”. The algorithm learns to draw this boundary from the data and is not random. |
2. Materials and Methods
2.1. Illustrative Example
2.2. Background Information About the Example Model
2.3. Uncertainty Quantification
3. Results
Distances in Feature Space for the Training Dataset
4. Discussion
| Anecdotalexample of how this model could be used for a new patient in observational mode. A 67-year-old male was evaluated for complaints of lower urinary tract symptoms for the past 3 months in June 2025. On evaluation, his Prostate Specific Antigen (PSA) was evaluated, and biopsy was reported as acinar adenocarcinoma (GS 4+3). A PSMA PET scan done in August 2025 showed locally advanced carcinoma of the prostate and a well-defined soft tissue density lesion measuring 19 mm × 28.5 mm in the anterior mediastinum abutting the ascending thoracic aorta with no fat/calcified components (incidental finding). Based on these incidental findings, the thoracic lesion, which could be a thymic epithelial tumor (TET), was evaluated using the Radiomics8 risk model. | ||||||
| Output from our prediction model with UQ | ||||||
| Model Predicted Risk Class | Prob. of HR Class (%) | Dist. to Nearest LR | Dist. to Nearest HR | LR UQ Score | HR UQ Score | Final Risk Class |
| LR | 10 | 0.21 | 0.99 | 0.04 | 4.76 | LR |
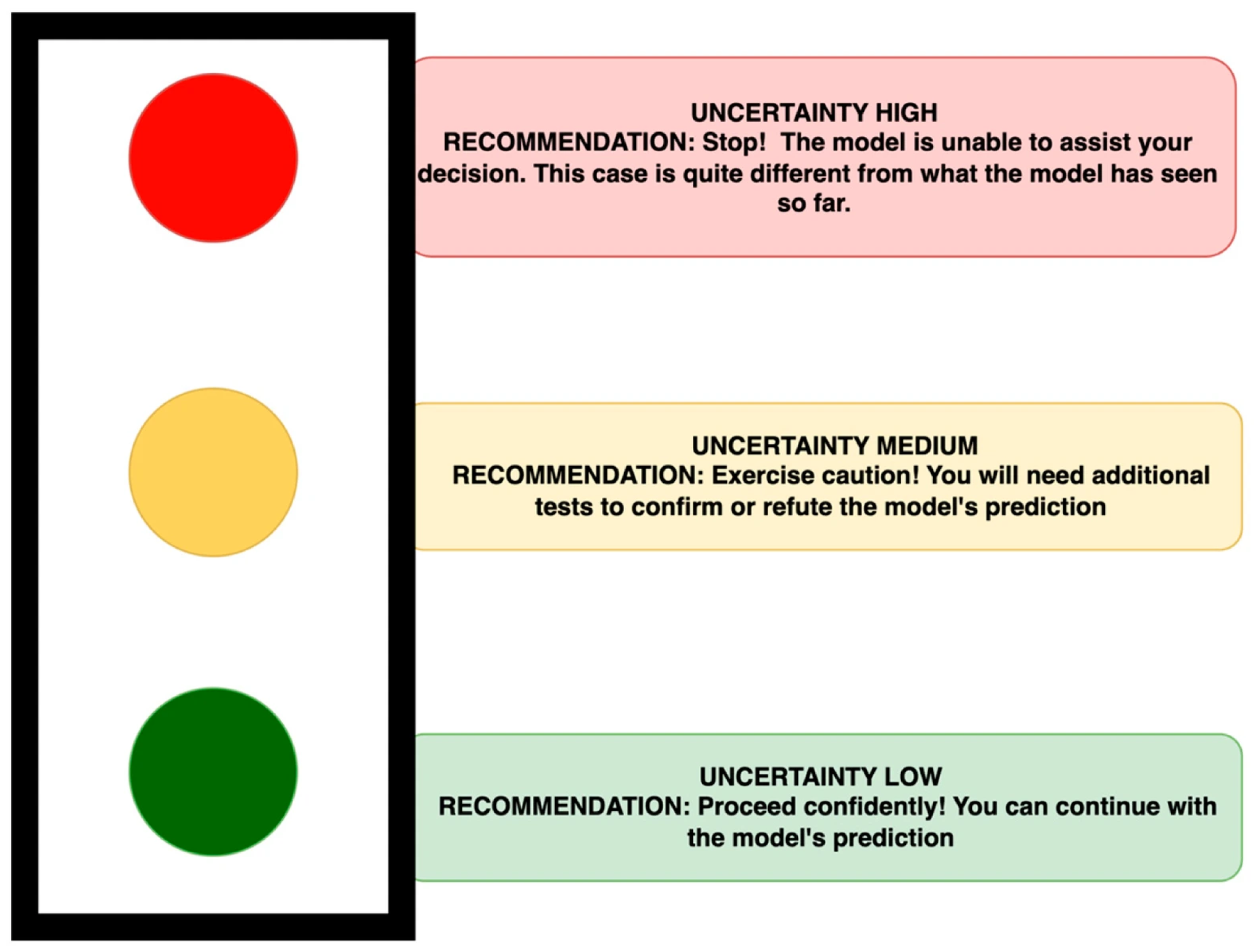
| The model predicted this patient to be low risk, with a probability of being high risk of 10%. Based on the distance metric, the patient’s features were ‘closer (0.21)’ to LR examples than the HR (0.99) examples in the training cohort. The final lower UQ score for LR (0.04) indicates that this is most likely to be LR and hence was labelled so in the final risk class. Implementing the TLP (Figure 3) by using a simple threshold (>75% = uncertainty high; 50–75% = uncertainty medium; <50% = uncertainty low), this patient’s distance (LR = 0.21) when compared with Table 1 was categorized in the amber category = uncertainty medium—exercise caution. The clinicians received the predicted risk and the degree of uncertainty in the classification in a two-step report. This aligned the multi-disciplinary team’s decision to go ahead with prostate cancer treatment and to take a ‘wait and watch’ strategy regarding the TET, given the patient’s age. | ||||||
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| RM | Risk Model |
| TET | Thymic Epithelial Tumour |
| UQ | Uncertainty Quantification |
| WHO | World Health Organization |
| HR | High Risk |
| LR | Low Risk |
| IQR | Interquartile Range |
| AI | Artificial Intelligence |
Appendix A

References
- Kompa, B.; Snoek, K.; Beam, A.L. Second opinion needed: Communicating uncertainty in medical machine learning. npj Digit. Med. 2021, 4, 4. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Tyralis, H.; Papacharalampous, G. A review of predictive uncertainty estimation with machine learning. Artif. Intell. Rev. 2024, 57, 94. [Google Scholar] [CrossRef]
- Singh, Y.; Andersen, J.B.; Hathaway, Q.; Venkatesh, S.K.; Gores, G.J.; Erickson, B. Deep learning-based uncertainty quantification for quality assurance in hepatobiliary imaging-based techniques. Oncotarget 2025, 16, 249–255. [Google Scholar] [CrossRef] [PubMed]
- Varghese, A.J.; Pathinathan, M.; Sasidharan, B.K.; Praveenraj, C.; Kuchipudi, R.B.; Kodiatte, T.; Mathew, M.; Isiah, R.; Pavamani, S.; Irodi, A.; et al. Are quantitative radiomics features comparable to semantic radiology features for pre-operative risk classification of thymic epithelial tumours? medRxiv 2025. [Google Scholar] [CrossRef]
- van den Goorbergh, R.; van Smeden, M.; Timmerman, D.; Van Calster, B. The harm of class imbalance corrections for risk prediction models: Illustration and simulation using logistic regression. J. Am. Med. Inform. Assoc. JAMIA 2022, 29, 1525–1534. [Google Scholar] [CrossRef] [PubMed]
- Russell, S.J.; Norvig, P. Artificial Intelligence: A Modern Approach; Pearson: London, UK, 2016. [Google Scholar]
- Kwee, T.C.; Cheng, G.; Lam, M.G.E.H.; Basu, S.; Alavi, A. SUVmax of 2.5 should not be embraced as a magic threshold for separating benign from malignant lesions. Eur. J. Nucl. Med. Mol. Imaging 2013, 40, 1475–1477. [Google Scholar] [CrossRef] [PubMed]
- Jones, C.K.; Wang, G.; Yedavalli, V.; Sair, H. Direct quantification of epistemic and aleatoric uncertainty in 3D U-net segmentation. J. Med. Imaging 2022, 9, 034002. [Google Scholar] [CrossRef] [PubMed]
- Froicu, E.-M.; Creangă-Murariu, I.; Afrăsânie, V.-A.; Gafton, B.; Alexa-Stratulat, T.; Miron, L.; Pușcașu, D.M.; Poroch, V.; Bacoanu, G.; Radu, I.; et al. Artificial Intelligence and Decision-Making in Oncology: A Review of Ethical, Legal, and Informed Consent Challenges. Curr. Oncol. Rep. 2025, 27, 1002–1012. [Google Scholar] [CrossRef] [PubMed]
- van Daalen, F.; Brecheisen, R.; Wee, L.; Dekker, A.; Bermejo, I. Multinomial Classification Certainty: A new uncertainty metric for multinomial outcome prediction. Prog. Artif. Intell. 2025. [Google Scholar] [CrossRef]



| Training Dataset | Metric/From–To | LR | HR |
|---|---|---|---|
| Within-group distance distribution | Min | 0.04 | 0.02 |
| 25% | 0.13 | 0.09 | |
| 50% (Median) | 0.20 | 0.19 | |
| 75% | 0.34 | 0.38 | |
| Max | 2.45 | 2.87 | |
| Mean intra-/inter-group distances | LR → LR (intra-LR) | 1.69 | |
| LR → HR (inter-group) | 5.37 | ||
| HR → LR (inter-group) | 1.78 | ||
| HR → HR (intra-HR) | 1.12 |
| E.g. # | Predicted Risk Class | Prob. of HR Class (%) | Dist. to Nearest LR | Dist. to Nearest HR | LR UQ Score | HR UQ Score | Final Risk Class | True Risk Class (Pathology) |
|---|---|---|---|---|---|---|---|---|
| 1 | HR | 48 | 0.19 | 0.28 | 0.13 | 0.42 | LR | LR |
| 2 | HR | 70 | 0.19 | 0.34 | 0.11 | 0.60 | LR | HR |
| 3 | HR | 98 | 3.83 | 0.51 | 28.48 | 0.07 | HR | LR |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
van Daalen, F.; Sasidharan, B.K.; Praveenraj, C.; Varghese, A.J.; Dekker, A.; Wee, L.; Fijten, R.; Irodi, A.; Thomas, H.M.T. Decoding Uncertainty Quantification for Oncology—An Illustration Using Radiomics. Diagnostics 2026, 16, 700. https://doi.org/10.3390/diagnostics16050700
van Daalen F, Sasidharan BK, Praveenraj C, Varghese AJ, Dekker A, Wee L, Fijten R, Irodi A, Thomas HMT. Decoding Uncertainty Quantification for Oncology—An Illustration Using Radiomics. Diagnostics. 2026; 16(5):700. https://doi.org/10.3390/diagnostics16050700
Chicago/Turabian Stylevan Daalen, Florian, Balu Krishna Sasidharan, C. Praveenraj, Amal Joseph Varghese, Andre Dekker, Leonard Wee, Rianne Fijten, Aparna Irodi, and Hannah Mary T. Thomas. 2026. "Decoding Uncertainty Quantification for Oncology—An Illustration Using Radiomics" Diagnostics 16, no. 5: 700. https://doi.org/10.3390/diagnostics16050700
APA Stylevan Daalen, F., Sasidharan, B. K., Praveenraj, C., Varghese, A. J., Dekker, A., Wee, L., Fijten, R., Irodi, A., & Thomas, H. M. T. (2026). Decoding Uncertainty Quantification for Oncology—An Illustration Using Radiomics. Diagnostics, 16(5), 700. https://doi.org/10.3390/diagnostics16050700





