Segmentation-Guided Preprocessing Improves Deep Learning Diagnostic Accuracy and Confidence of Ameloblastoma and Odontogenic Keratocyst in Cone Beam CT Images—A Preliminary Study
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethical Standards
2.2. Data Collection
2.3. Data Set Construction
2.4. Model Development
2.4.1. Model Architecture and Strategy
2.4.2. Mode-Adaptive Patient-Centric Sampling (MAPS)
- Compute the average image count:
- Calculate a patient-specific ratio:
- Obtain the final weight:
2.5. Model Evaluation and Statistical Analysis
- Confusion Matrix
- Accuracy provides the percentage of successful classifications:
- Precision provides the percentage of true positive among all positive ratings:
- Recall provides the percentage of true positive among all true ratings:
- Specificity provides the percentage of true negative among all negative ratings:
- F1-score is calculated as the harmonic mean between recall and precision:
3. Results
3.1. Demographic and Radiographic Characteristics
3.2. Training Process
3.3. Classification Performance
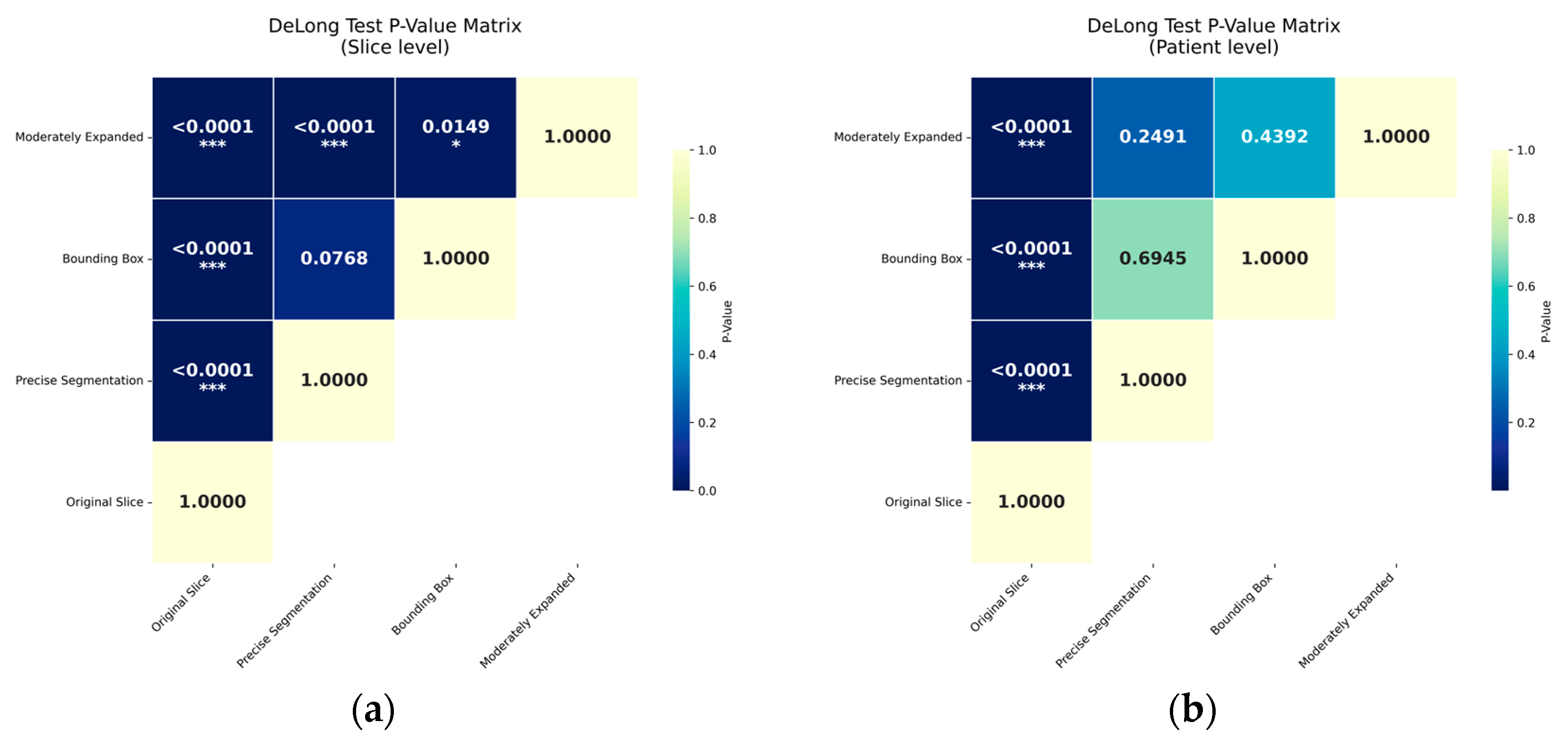
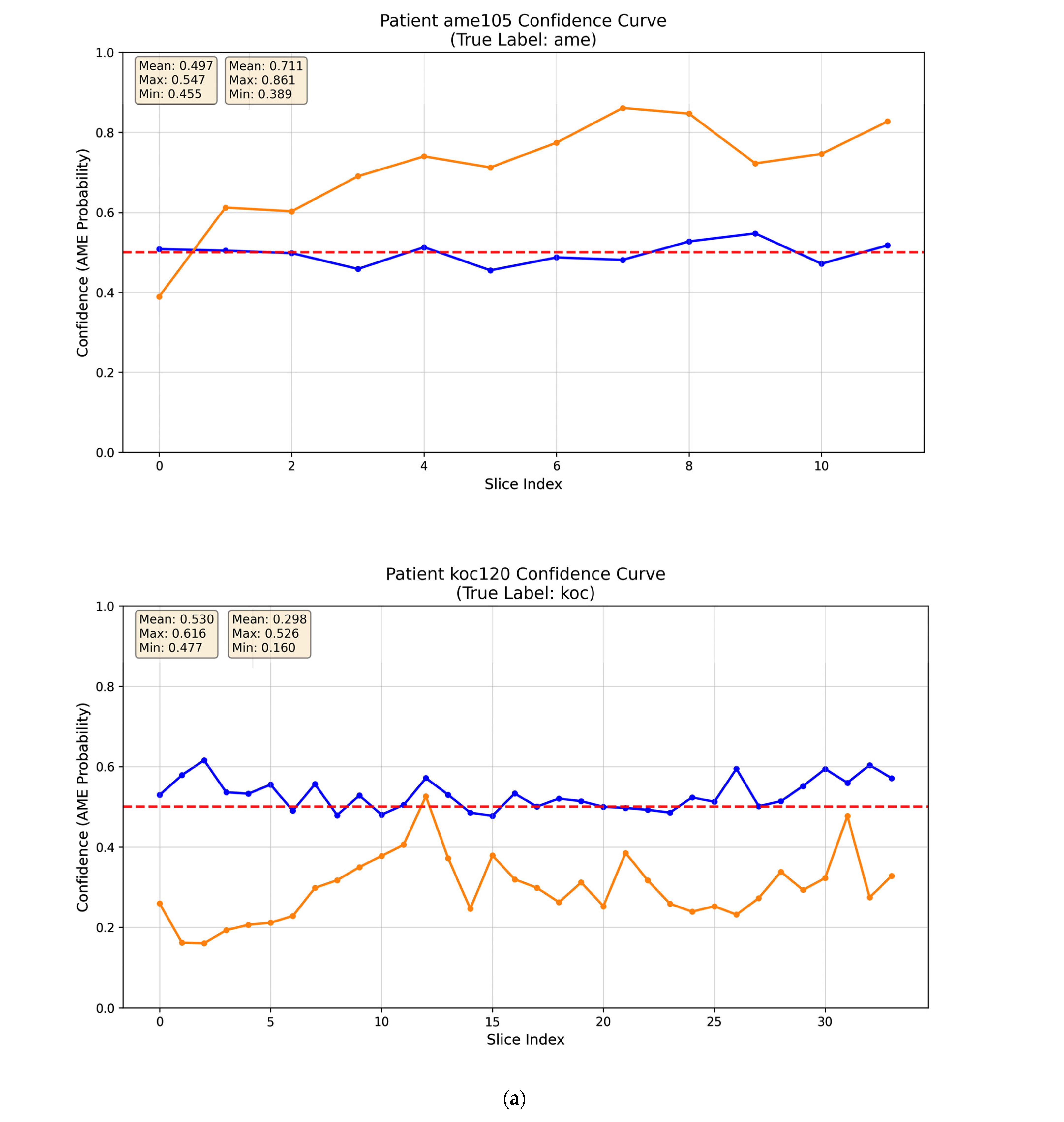
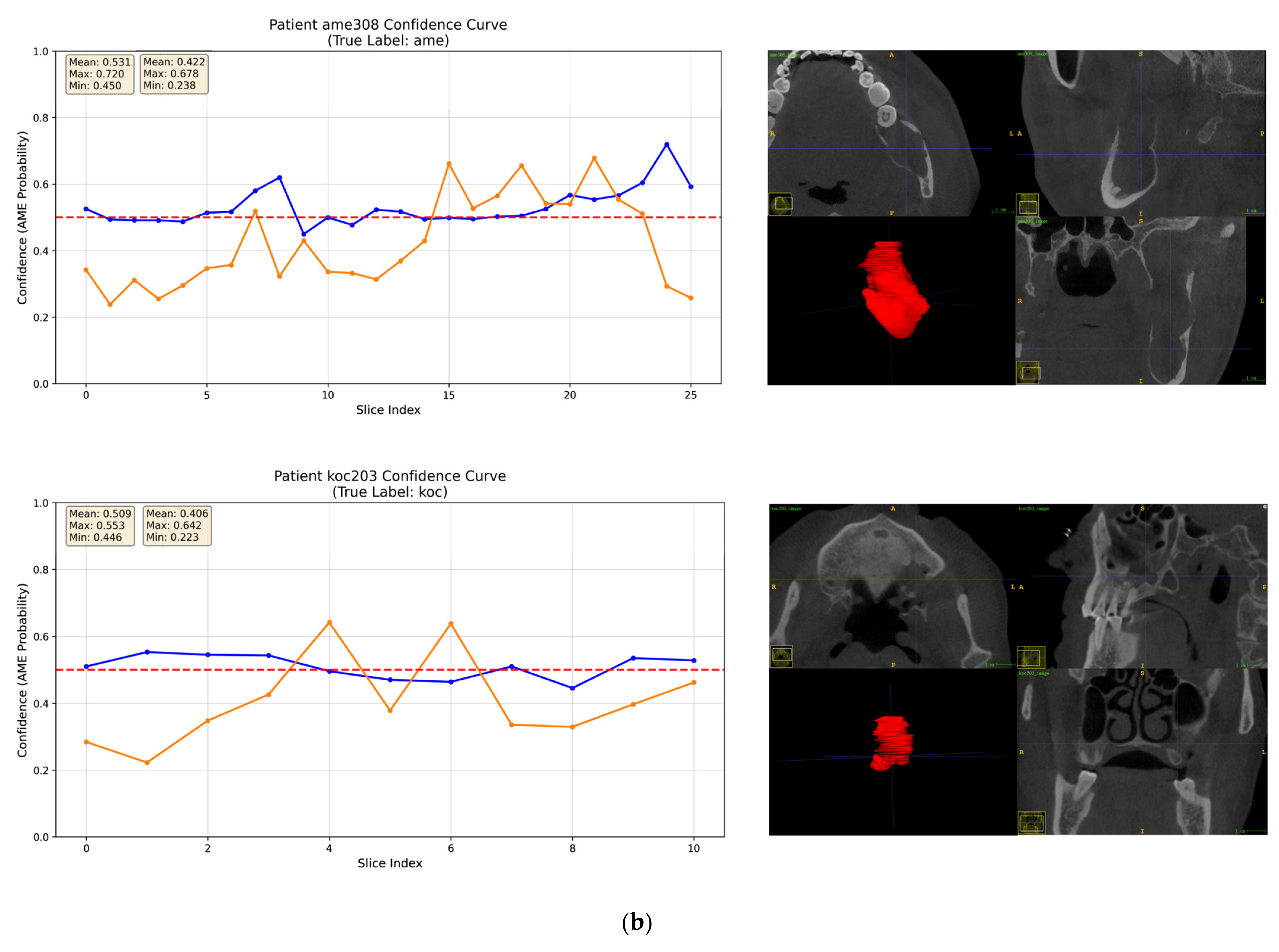
3.4. Interpretability Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hung, K.; Montalvao, C.; Tanaka, R.; Kawai, T.; Bornstein, M.M. The use and performance of artificial intelligence applications in dental and maxillofacial radiology: A systematic review. Dentomaxillofac. Radiol. 2020, 49, 20190107. [Google Scholar] [CrossRef]
- Shi, Y.J.; Li, J.P.; Wang, Y.; Ma, R.H.; Wang, Y.L.; Guo, Y.; Li, G. Deep learning in the diagnosis for cystic lesions of the jaws: A review of recent progress. Dentomaxillofac. Radiol. 2024, 53, 271–280. [Google Scholar] [CrossRef] [PubMed]
- Poedjiastoeti, W.; Suebnukarn, S. Application of Convolutional Neural Network in the Diagnosis of Jaw Tumors. Healthc. Inform. Res. 2018, 24, 236–241. [Google Scholar] [CrossRef]
- Li, M.; Mu, C.C.; Zhang, J.Y.; Li, G. Application of Deep Learning in Differential Diagnosis of Ameloblastoma and Odontogenic Keratocyst Based on Panoramic Radiographs. Zhongguo Yi Xue Ke Xue Yuan Xue Bao 2023, 45, 273–279. [Google Scholar] [CrossRef]
- Liu, Z.J.; Liu, J.N.; Zhou, Z.J.; Zhang, Q.Y.; Wu, H.; Zhai, G.T.; Han, J. Differential diagnosis of ameloblastoma and odontogenic keratocyst by machine learning of panoramic radiographs. Int. J. Comput. Assist. Radiol. Surg. 2021, 16, 415–422. [Google Scholar] [CrossRef]
- Lee, J.H.; Kim, D.H.; Jeong, S.N. Diagnosis of cystic lesions using panoramic and cone beam computed tomographic images based on deep learning neural network. Oral Dis. 2020, 26, 152–158. [Google Scholar] [CrossRef]
- Bispo, M.S.; Pierre Júnior, M.L.G.Q.; Apolinário, A.L., Jr.; Dos Santos, J.N.; Junior, B.C.; Neves, F.S.; Crusoé-Rebello, I. Computer tomographic differential diagnosis of ameloblastoma and odontogenic keratocyst: Classification using a convolutional neural network. Dentomaxillofac. Radiol. 2021, 50, 20210002. [Google Scholar] [CrossRef] [PubMed]
- Chai, Z.K.; Mao, L.; Chen, H.; Sun, T.G.; Shen, X.M.; Liu, J.; Sun, Z.J. Improved Diagnostic Accuracy of Ameloblastoma and Odontogenic Keratocyst on Cone-Beam CT by Artificial Intelligence. Front. Oncol. 2022, 11, 793417. [Google Scholar] [CrossRef]
- Salvi, M.; Acharya, U.R.; Molinari, F.; Meiburger, K.M. The impact of pre- and post-image processing techniques on deep learning frameworks: A comprehensive review for digital pathology image analysis. Comput. Biol. Med. 2021, 128, 104129. [Google Scholar] [CrossRef]
- Li, A.; Winebrake, J.P.; Kovacs, K. Facilitating deep learning through preprocessing of optical coherence tomography images. BMC Ophthalmol. 2023, 23, 158. [Google Scholar] [CrossRef] [PubMed]
- Kwon, O.; Yong, T.H.; Kang, S.R.; Kim, J.E.; Huh, K.H.; Heo, M.S.; Lee, S.S.; Choi, S.C.; Yi, W.J. Automatic diagnosis for cysts and tumors of both jaws on panoramic radiographs using a deep convolution neural network. Dentomaxillofac. Radiol. 2020, 49, 20200185. [Google Scholar] [CrossRef]
- Panyarak, W.; Suttapak, W.; Mahasantipiya, P.; Charuakkra, A.; Boonsong, N.; Wantanajittikul, K.; Iamaroon, A. CrossViT with ECAP: Enhanced deep learning for jaw lesion classification. Int. J. Med. Inform. 2025, 193, 105666. [Google Scholar] [CrossRef] [PubMed]
- Ha, E.G.; Jeon, K.J.; Lee, C.; Kim, H.S.; Han, S.S. Development of deep learning model and evaluation in real clinical practice of lingual mandibular bone depression (Stafne cyst) on panoramic radiographs. Dentomaxillofac. Radiol. 2023, 52, 20220413. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Jo, E.; Kim, H.J.; Cha, I.H.; Jung, Y.S.; Nam, W.; Kim, J.Y.; Kim, J.K.; Kim, Y.H.; Oh, T.G.; et al. Deep Learning for Automated Detection of Cyst and Tumors of the Jaw in Panoramic Radiographs. J. Clin. Med. 2020, 9, 1839. [Google Scholar] [CrossRef]
- Tajima, S.; Okamoto, Y.; Kobayashi, T.; Kiwaki, M.; Sonoda, C.; Tomie, K.; Saito, H.; Ishikawa, Y.; Takayoshi, S. Development of an automatic detection model using artificial intelligence for the detection of cyst-like radiolucent lesions of the jaws on panoramic radiographs with small training datasets. J. Oral Maxillofac. Surg. Med. Pathol. 2022, 34, 553–560. [Google Scholar] [CrossRef]
- Ariji, Y.; Yanashita, Y.; Kutsuna, S.; Muramatsu, C.; Fukuda, M.; Kise, Y.; Nozawa, M.; Kuwada, C.; Fujita, H.; Katsumata, A.; et al. Automatic detection and classification of radiolucent lesions in the mandible on panoramic radiographs using a deep learning object detection technique. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. 2019, 128, 424–430. [Google Scholar] [CrossRef]
- Watanabe, H.; Ariji, Y.; Fukuda, M.; Kuwada, C.; Kise, Y.; Nozawa, M.; Sugita, Y.; Ariji, E. Deep learning object detection of maxillary cyst-like lesions on panoramic radiographs: Preliminary study. Oral Radiol. 2021, 37, 487–493. [Google Scholar] [CrossRef]
- Kise, Y.; Ariji, Y.; Kuwada, C.; Fukuda, M.; Ariji, E. Effect of deep transfer learning with a different kind of lesion on classification performance of pre-trained model: Verification with radiolucent lesions on panoramic radiographs. Imaging Sci. Dent. 2023, 53, 27–34. [Google Scholar] [CrossRef]
- Lee, A.; Kim, M.S.; Han, S.S.; Park, P.; Lee, C.; Yun, J.P. Deep learning neural networks to differentiate Stafne’s bone cavity from pathological radiolucent lesions of the mandible in heterogeneous panoramic radiography. PLoS ONE 2021, 16, e0254997. [Google Scholar] [CrossRef] [PubMed]
- Yu, D.; Hu, J.; Feng, Z.; Song, M.; Zhu, H. Deep learning based diagnosis for cysts and tumors of jaw with massive healthy samples. Sci. Rep. 2022, 12, 1855. [Google Scholar] [CrossRef]
- Altun, O.; Özen, D.Ç.; Duman, Ş.B.; Dedeoğlu, N.; Bayrakdar, İ.Ş.; Eşer, G.; Çelik, Ö.; Sümbüllü, M.A.; Syed, A.Z. Automatic maxillary sinus segmentation and pathology classification on cone-beam computed tomographic images using deep learning. BMC Oral Health 2024, 24, 1208. [Google Scholar] [CrossRef] [PubMed]
- He, H.; Garcia, E.A. Learning from Imbalanced Data. IEEE Trans. Knowl. Data Eng. 2009, 21, 1263–1284. [Google Scholar] [CrossRef]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. In Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 618–626. [Google Scholar] [CrossRef]
- Varoquaux, G.; Cheplygina, V. Machine learning for medical imaging: Methodological failures and recommendations for the future. npj Digit. Med. 2022, 5, 48. [Google Scholar] [CrossRef] [PubMed]





| Predicted OKC | Predicted AME | |
|---|---|---|
| True OKC | True Negative (TN) | False Positive (FP) |
| True AME | True Negative (FN) | True Positive (TP) |
| Characteristic | Groups | p-Value | |
|---|---|---|---|
| AME (n = 64) | OKC (n = 64) | ||
| Age (years), Median (IQR) | 33 (27) | 32 (27) | 0.473 |
| Sex, n (%) | |||
| Male | 32 (50) | 30 (46.9) | 0.724 |
| Female | 32 (50) | 34 (53.1) | |
| Location, n (%) | |||
| Maxilla | 4 (6.3) | 9 (14.1) | 0.143 |
| Mandibular | 60 (93.8) | 55 (85.9) | |
| Locularity, n (%) | |||
| Unilocular | 15 (23.4) | 46 (71.9) | 0.000 * |
| Multilocular | 49 (76.6) | 18 (28.1) | |
| Cortical Integrity, n (%) | |||
| Intact | 46 (71.9) | 31 (48.4) | 0.007 * |
| Disrupted | 18 (28.1) | 33 (51.6) | |
| Impacted Tooth, n (%) | |||
| Present | 17 (26.6) | 30 (46.9) | 0.017 * |
| Absent | 47 (73.4) | 34 (53.1) | |
| Root Resorption, n (%) | |||
| Present | 55 (85.9) | 9 (14.1) | 0.000 * |
| Absent | 11 (17.2) | 53 (82.8) | |
| CBCT Scanner, n (%) | |||
| i-CAT FLX | 50 (78.1) | 49 (76.6) | 0.639 |
| New Tom 1 | 8 (12.5) | 6 (9.4) | |
| New Tom 2 | 6 (9.4) | 9 (14.1) | |
| ROI Extraction Strategy | Accuracy | Precision | Recall | F1-Score |
|---|---|---|---|---|
| Moderately Expanded | 0.7727 ± 0.0483 | 0.8351 ± 0.0896 | 0.7213 ± 0.1428 | 0.7603 ± 0.0761 |
| Bounding box | 0.7595 ± 0.0489 | 0.7630 ± 0.0384 | 0.7955 ± 0.0947 | 0.7747 ± 0.0483 |
| Precise Segmentation | 0.7309 ± 0.0604 | 0.7411 ± 0.0976 | 0.7689 ± 0.1103 | 0.7453 ± 0.0695 |
| Original Slice | 0.5376 ± 0.1004 | 0.5846 ± 0.1329 | 0.6583 ± 0.1519 | 0.5967 ± 0.0622 |
| ROI Extraction Strategy | Accuracy | Precision | Recall | F1-Score |
|---|---|---|---|---|
| Moderately Expanded | 0.8123 ± 0.0385 | 0.8649 ± 0.0950 | 0.7641 ± 0.1018 | 0.8009 ± 0.0441 |
| Bounding box | 0.7975 ± 0.0744 | 0.7967 ± 0.0762 | 0.7974 ± 0.1033 | 0.7948 ± 0.0824 |
| Precise Segmentation | 0.7185 ± 0.0762 | 0.7155 ± 0.0947 | 0.7795 ± 0.1526 | 0.7319 ± 0.0756 |
| Original Slice | 0.4828 ± 0.1595 | 0.5497 ± 0.2298 | 0.6756 ± 0.2014 | 0.5686 ± 0.1101 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Zhang, X.; Yang, Y.; Zhong, C.; Li, J.; Li, G. Segmentation-Guided Preprocessing Improves Deep Learning Diagnostic Accuracy and Confidence of Ameloblastoma and Odontogenic Keratocyst in Cone Beam CT Images—A Preliminary Study. Diagnostics 2026, 16, 416. https://doi.org/10.3390/diagnostics16030416
Zhang X, Yang Y, Zhong C, Li J, Li G. Segmentation-Guided Preprocessing Improves Deep Learning Diagnostic Accuracy and Confidence of Ameloblastoma and Odontogenic Keratocyst in Cone Beam CT Images—A Preliminary Study. Diagnostics. 2026; 16(3):416. https://doi.org/10.3390/diagnostics16030416
Chicago/Turabian StyleZhang, Xinyue, Yuxuan Yang, Chen Zhong, Jupeng Li, and Gang Li. 2026. "Segmentation-Guided Preprocessing Improves Deep Learning Diagnostic Accuracy and Confidence of Ameloblastoma and Odontogenic Keratocyst in Cone Beam CT Images—A Preliminary Study" Diagnostics 16, no. 3: 416. https://doi.org/10.3390/diagnostics16030416
APA StyleZhang, X., Yang, Y., Zhong, C., Li, J., & Li, G. (2026). Segmentation-Guided Preprocessing Improves Deep Learning Diagnostic Accuracy and Confidence of Ameloblastoma and Odontogenic Keratocyst in Cone Beam CT Images—A Preliminary Study. Diagnostics, 16(3), 416. https://doi.org/10.3390/diagnostics16030416






