SBMN: Similarity-Based Memory Network for the Diagnosis of Vertical Root Fracture in Dental Imaging
Abstract
1. Introduction
- We propose a novel classification framework that integrates similarity-based computation with category memory, enabling automatic diagnosis of VRFs.
- To ensure effective categorization of memory, we design a dedicated similarity loss function. By accurately aligning category memory with the corresponding class, the memory can better enhance the features extracted from input images, thereby improving classification—and ultimately diagnostic—accuracy.
- We introduce a bidirectional update mechanism between category memory and input features during training. On the one hand, category memory is continuously refined by assimilating the characteristics of images from its corresponding class. On the other hand, each input image retrieves relevant memory components for fusion and feature enhancement, leading to more robust representation learning.
2. Related Work
2.1. Convolutional Neural Network (CNN)
2.2. Transformer and Similarity
2.3. Memory Networks
3. Materials and Methods
3.1. Materials
3.1.1. Patients and Datasets
3.1.2. Image Preprocessing
3.2. Methods
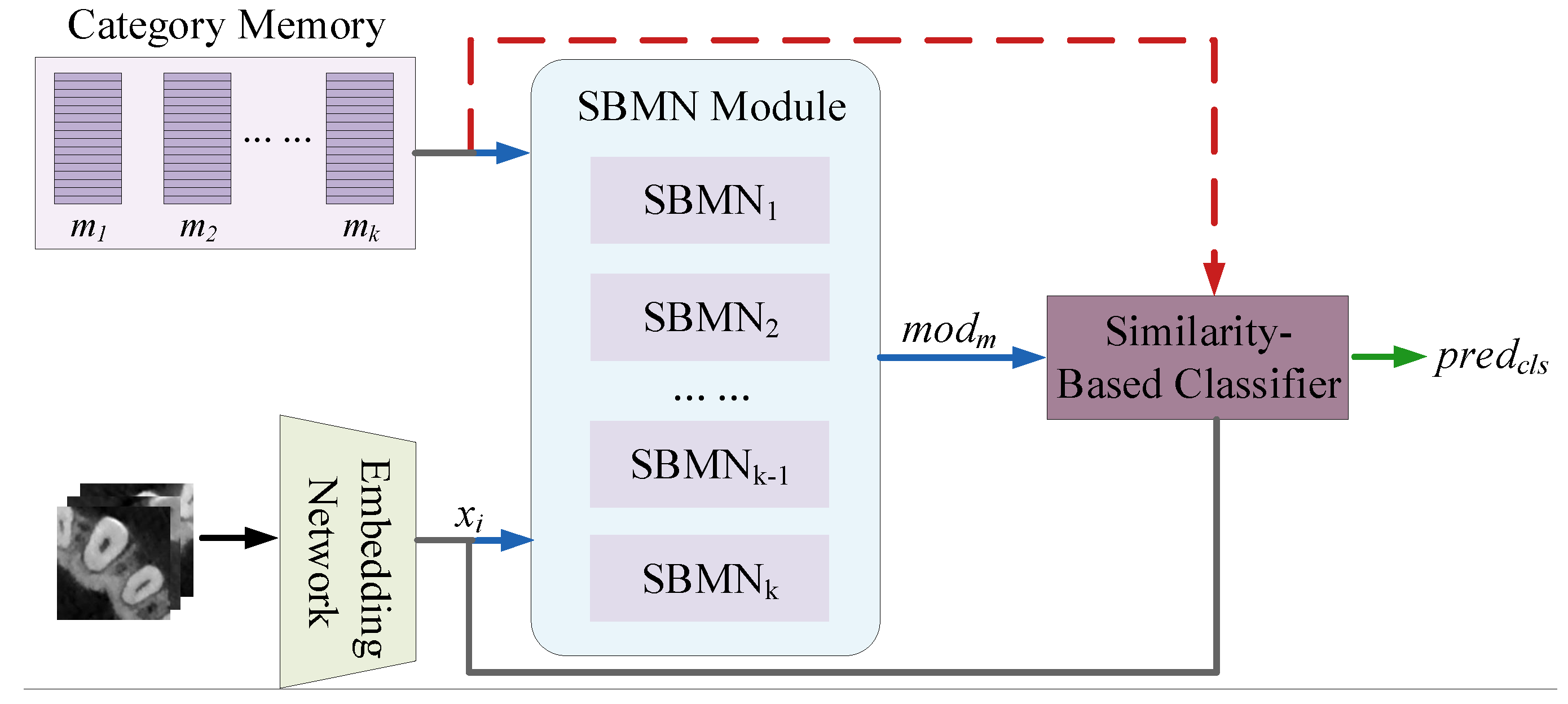
3.2.1. Overview
3.2.2. Network Architecture
Embedding Network
Category Memory
Basic SBMN Module
Similarity-Based Classifier
3.2.3. Training Loss Functions
4. Results
4.1. Evaluation Metrics
4.2. Results of SBMN
4.3. Comparisons with Other Networks
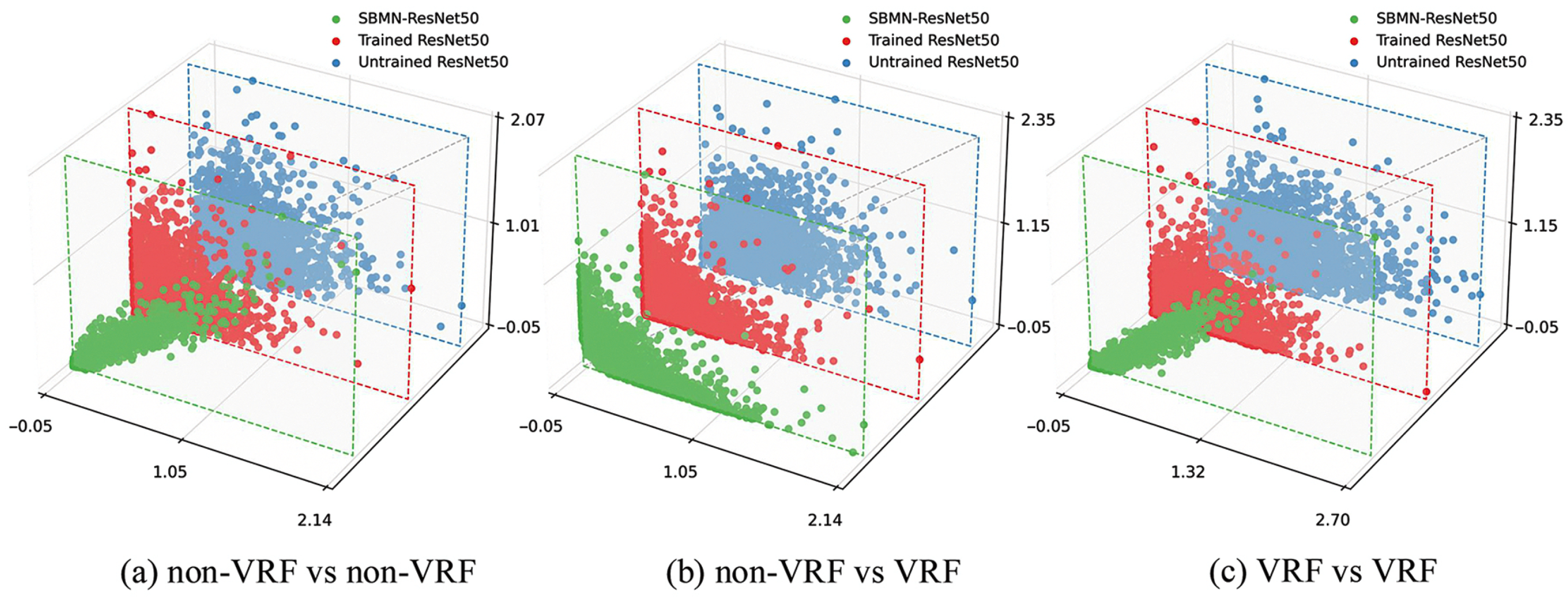
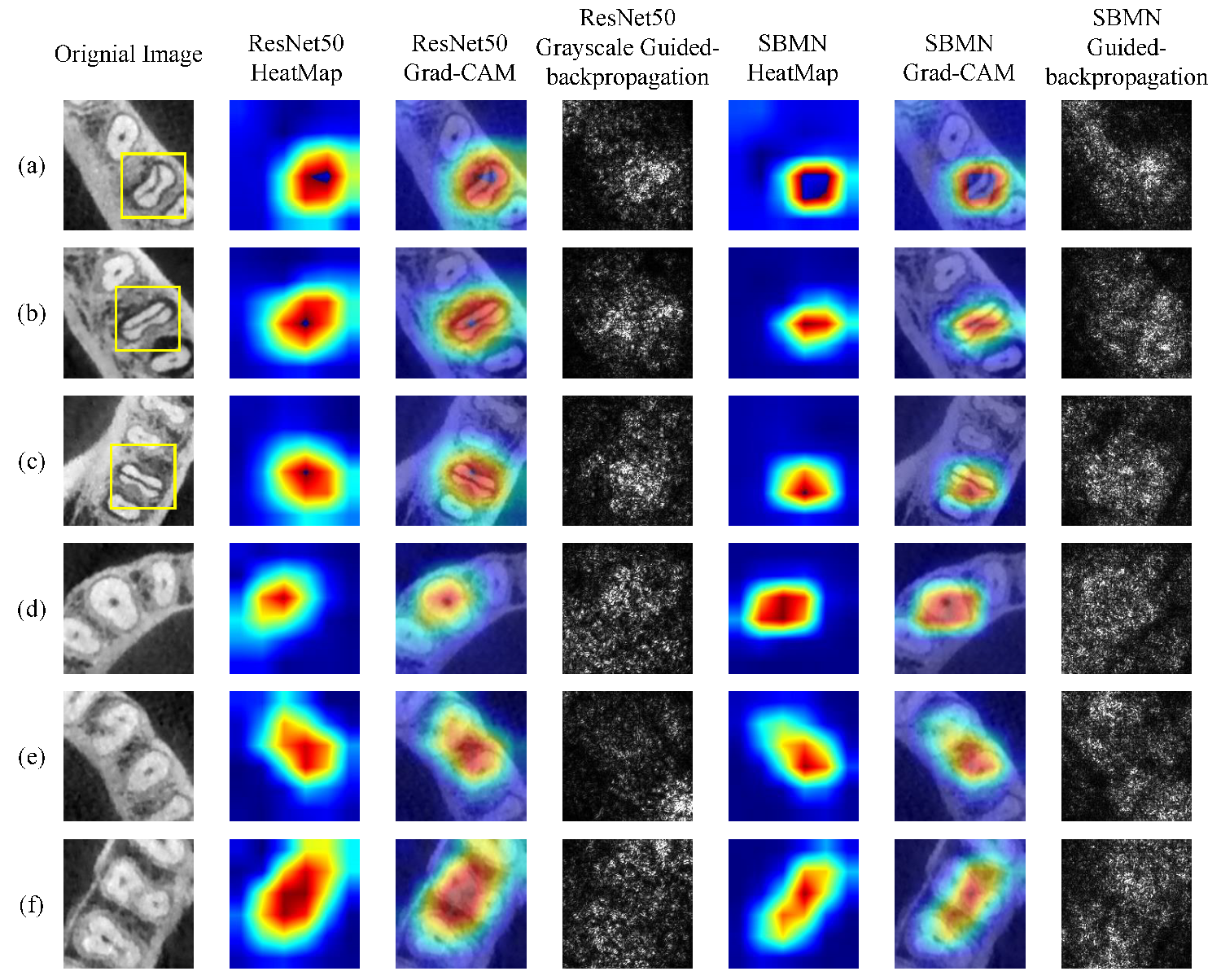
4.4. Visualization
4.5. Ablation Study
5. Discussion
- Introduction of Category Memory: We introduce a category-level memory mechanism that preserves representative patterns of fractured and non-fractured teeth. By retaining stable class-specific information across training samples, the model can better handle limited and heterogeneous medical data, which are common in clinical studies.
- Design of the Basic SBMN Module: This similarity-guided design enables the network to selectively emphasize clinically relevant features while suppressing noise and irrelevant structures frequently observed in CBCT images. By integrating similarity computation with gating operations and a similarity loss function, this module ensures the stability of the Category Memory repository while maintaining discriminative and robust feature representations.
- Proposal of the Similarity-based Classifier: Instead of relying solely on conventional fully connected classifiers, predictions are generated based on similarity between current inputs and stored category representations. This strategy provides a more transparent decision process and improves interpretability, which is desirable for clinical adoption.
- Experimental Validation on Small Medical Datasets: On a small dataset of vertical root fractures (VRF), the SBMN achieved high diagnostic accuracy on both automatically and manually segmented images, indicating that the memory mechanism can effectively capture subtle structural differences associated with fractures. Importantly, competitive performance was maintained under automatic segmentation, which better reflects realistic clinical scenarios where manual annotation is not feasible.
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Rivera, E.M.; Walton, R.E. Longitudinal tooth fractures: Findings that contribute to complex endodontic diagnoses. Endod. Top. 2010, 16, 82–111. [Google Scholar] [CrossRef]
- Huang, C.C.; Lee, B.S. Diagnosis of vertical root fracture in endodontically treated teeth using computed tomography. J. Dent. Sci. 2015, 10, 227–232. [Google Scholar] [CrossRef][Green Version]
- Diagnostic issues dealing with the management of teeth with vertical root fractures: A narrative review. G. Ital. Endod. 2014, 28, 91–96. [CrossRef][Green Version]
- Alsani, A.; Balhaddad, A.A.; Nazir, M.A. Vertical root fracture: A case report and review of the literature. G. Ital. Endod. 2017, 31, 21–28. [Google Scholar] [CrossRef]
- Duak, K.; Kundabala, M.; Bhat, K. Endodontic Miscellany: 1. An unusual vertical root fracture. Endodontology 2004, 16, 23–26. [Google Scholar]
- Silva, E.J.N.L.D.; Santos, G.R.D.; Krebs, R.L.; Coutinho-Filho, T.D.S. Surgical Alternative for Treatment of Vertical Root fracture: A Case Report. Iran. Endod. J. 2012, 7, 40. [Google Scholar]
- Hsiao, L.T.; Ho, J.C.; Huang, C.F.; Hung, W.C.; Chang, C.W. Analysis of clinical associated factors of vertical root fracture cases found in endodontic surgery. J. Dent. Sci. 2020, 15, 200–206. [Google Scholar] [CrossRef] [PubMed]
- Aviad, T.; Tamse, Z.; Fuss, J.; Lustig, J.; Joseph, J. An evaluation of endodontically treated vertically fractured teeth. J. Endod. 1999, 25, 506–508. [Google Scholar] [CrossRef] [PubMed]
- Okaguchi, M.; Kuo, T.; Ho, Y.C. Successful treatment of vertical root fracture through intentional replantation and root fragment bonding with 4-META/MMA-TBB resin. J. Formos. Med. Assoc. 2018, 118, 671–678. [Google Scholar] [CrossRef]
- Wang, P.; Su, L. Clinical observation in 2 representative cases of vertical root fracture in nonendodontically treated teeth. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. Endod. 2009, 107, e39–e42. [Google Scholar] [CrossRef]
- Liao, W.C.; Chen, C.H.; Pan, Y.H.; Chang, M.C.; Jeng, J.H. Vertical Root Fracture in Non-Endodontically and Endodontically Treated Teeth: Current Understanding and Future Challenge. J. Pers. Med. 2021, 11, 1375. [Google Scholar] [CrossRef]
- Hasan, S.; Singh, K.; Salati, N. Cracked tooth syndrome: Overview of literature. Int. J. Appl. Basic Med. Res. 2015, 5, 164–168. [Google Scholar] [CrossRef]
- Taschieri, S.; Tamse, A.; Fabbro, M.D.; Rosano, G.; Tsesis, I. A new surgical technique for preservation of endodontically treated teeth with coronally located vertical root fractures: A prospective case series. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. Endod. 2010, 110, 45–52. [Google Scholar] [CrossRef]
- Kimura, Y.; Tanabe, M.; Amano, Y.; Kinoshita, J.I.; Yamada, Y.; Masuda, Y. Basic study of the use of laser on detection of vertical root fracture. J. Dent. 2009, 37, 909–912. [Google Scholar] [CrossRef]
- Pitts, D.L.; Natkin, E. Diagnosis and treatment of vertical root fractures. J. Endod. 1983, 9, 338–346. [Google Scholar] [CrossRef] [PubMed]
- Corbella, S.; Del Fabbro, M.; Tamse, A.; Rosen, E.; Tsesis, I.; Taschieri, S. Cone beam computed tomography for the diagnosis of vertical root fractures: A systematic review of the literature and meta-analysis. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. 2014, 118, 593–602. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, R.I.; Bahrami, G.; Isidor, F.; Wenzel, A.; Haiter-Neto, F.; Groppo, F.C. Detection of vertical root fractures by cone-beam computerized tomography in endodontically treated teeth with fiber-resin and titanium posts: An in vitro study. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. 2013, 115, E49–E57. [Google Scholar] [CrossRef]
- Kajan, Z.D.; Taromsari, M. Value of cone beam CT in detection of dental root fractures. Dentomaxillofac. Radiol. 2012, 41, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Makeeva, I.M.; Byakova, S.F.; Novozhilova, N.E.; Adzhieva, E.K.; Golubeva, G.I.; Grachev, V.I.; Kasatkina, I.V. Detection of artificially induced vertical root fractures of different widths by CBCT in vitro and in vivo. Int. Endod. J. 2015, 48, 628–635. [Google Scholar] [CrossRef]
- Lecun, Y.; Boser, B.; Denker, J.; Henderson, D.; Howard, R.; Hubbard, W.; Jackel, L. Backpropagation Applied to Handwritten Zip Code Recognition. Neural Comput. 1989, 1, 541–551. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G. ImageNet Classification with Deep Convolutional Neural Networks. In Proceedings of the NIPS; Curran Associates, Inc.: Red Hook, NY, USA, 2012. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR); IEEE: New York, NY, USA, 2016. [Google Scholar] [CrossRef]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention is All You Need. In Proceedings of the 30st Conference on Neural Information Processing Systems (NIPS 2017); NeurIPS Proceedings: New York, NY, USA, 2017. [Google Scholar]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv 2020, arXiv:2010.11929. [Google Scholar]
- Liu, Z.; Lin, Y.; Cao, Y.; Hu, H.; Wei, Y.; Zhang, Z.; Lin, S.; Guo, B. Swin Transformer: Hierarchical Vision Transformer using Shifted Windows. arXiv 2021, arXiv:2103.14030. [Google Scholar] [CrossRef]
- Hubel, D.H.; Wiesel, T.N. Receptive fields, binocular interaction and functional architecture in the cat’s visual cortex. J. Physiol. 1962, 160, 106–154. [Google Scholar] [CrossRef]
- Fukushima, K. Neocognitron: A self-organizing neural network model for a mechanism of pattern recognition unaffected by shift in position. Biol. Cybern. 1980, 36, 193–202. [Google Scholar] [CrossRef] [PubMed]
- Lecun, Y.; Bottou, L. Gradient-based learning applied to document recognition. Proc. IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. Comput. Sci. 2014, abs/1409.1556. [Google Scholar]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Rabinovich, A. Going Deeper with Convolutions. In Proceedings of the 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR); IEEE: New York, NY, USA, 2014. [Google Scholar] [CrossRef]
- Huang, G.; Liu, Z.; Laurens, V.D.M.; Weinberger, K.Q. Densely Connected Convolutional Networks. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR); IEEE: New York, NY, USA, 2016. [Google Scholar] [CrossRef]
- Tan, M.; Le, Q.V. EfficientNet: Rethinking Model Scaling for Convolutional Neural Networks. arXiv 2019, arXiv:1905.11946. [Google Scholar] [CrossRef]
- Zhao, X.; Wang, L.; Zhang, Y.; Han, X.; Deveci, M.; Parmar, M. A review of convolutional neural networks in computer vision. Artif. Intell. Rev. 2024, 57, 43. [Google Scholar] [CrossRef]
- Krichen, M. Convolutional Neural Networks: A Survey. Computers 2023, 12, 151. [Google Scholar] [CrossRef]
- Liu, Y.; Xue, J.; Li, D.; Zhang, W.; Kiang, C.T.; Xu, Z. Image recognition based on lightweight convolutional neural network. Image Vis. Comput. 2024, 146, 105037. [Google Scholar] [CrossRef]
- Liu, Z.; Mao, H.; Wu, C.Y.; Feichtenhofer, C.; Darrell, T.; Xie, S. A ConvNet for the 2020s. arXiv 2022, arXiv:2201.03545. [Google Scholar] [CrossRef]
- Devlin, J.; Chang, M.W.; Lee, K.; Toutanova, K. BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding. arXiv 2019, arXiv:1810.04805. [Google Scholar] [CrossRef]
- Kitaev, N.; Kaiser, U.; Levskaya, A. Reformer: The Efficient Transformer. arXiv 2020, arXiv:2001.04451. [Google Scholar] [CrossRef]
- Wang, S.; Li, B.Z.; Khabsa, M.; Fang, H.; Ma, H. Linformer: Self-Attention with Linear Complexity. arXiv 2020, arXiv:2006.04768. [Google Scholar] [CrossRef]
- Zaheer, M.; Guruganesh, G.; Dubey, A.; Ainslie, J.; Alberti, C.; Ontanon, S.; Pham, P.; Ravula, A.; Wang, Q.; Yang, L. Big Bird: Transformers for Longer Sequences. arXiv 2020, arXiv:2007.14062. [Google Scholar] [CrossRef]
- Ainslie, J.; Ontanon, S.; Alberti, C.; Pham, P.; Ravula, A.; Sanghai, S. ETC: Encoding Long and Structured Data in Transformers. arXiv 2020, arXiv:2004.08483. [Google Scholar] [CrossRef]
- Chen, J.; Lu, Y.; Yu, Q.; Luo, X.; Zhou, Y. TransUNet: Transformers Make Strong Encoders for Medical Image Segmentation. arXiv 2021, arXiv:2102.04306. [Google Scholar] [CrossRef]
- Cheng, B.; Misra, I.; Schwing, A.G.; Kirillov, A.; Girdhar, R. Masked-attention Mask Transformer for Universal Image Segmentation. In Proceedings of the 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR); IEEE: New York, NY, USA, 2022; pp. 1280–1289. [Google Scholar] [CrossRef]
- Fu, J.; Liu, J.; Tian, H.; Li, Y.; Bao, Y.; Fang, Z.; Lu, H. Dual Attention Network for Scene Segmentation. In Proceedings of the 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR); IEEE: New York, NY, USA, 2020. [Google Scholar] [CrossRef]
- Huang, Z.; Wang, X.; Wei, Y.; Huang, L.; Huang, T.S. CCNet: Criss-Cross Attention for Semantic Segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2020, 45, 6896–6908. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Gu, J.; Zhang, Z.; Dai, J.; Wei, Y. Relation Networks for Object Detection. arXiv 2017, arXiv:1711.11575. [Google Scholar] [CrossRef]
- Carion, N.; Massa, F.; Synnaeve, G.; Usunier, N.; Kirillov, A.; Zagoruyko, S. End-to-End Object Detection with Transformers. arXiv 2020, arXiv:2005.12872. [Google Scholar] [CrossRef]
- Zhu, X.; Su, W.; Lu, L.; Li, B.; Dai, J. Deformable DETR: Deformable Transformers for End-to-End Object Detection. arXiv 2020, arXiv:2010.04159. [Google Scholar] [CrossRef]
- Bello, I.; Zoph, B.; Le, Q.; Vaswani, A.; Shlens, J. Attention Augmented Convolutional Networks. In Proceedings of the 2019 IEEE/CVF International Conference on Computer Vision (ICCV); IEEE: New York, NY, USA, 2020. [Google Scholar] [CrossRef]
- Yu, Q.; Xia, Y.; Bai, Y.; Lu, Y.; Shen, W. Glance-and-Gaze Vision Transformer. arXiv 2021, arXiv:2106.02277. [Google Scholar] [CrossRef]
- Khan, A.; Rauf, Z.; Sohail, A.; Khan, A.R.; Asif, H.; Asif, A.; Farooq, U. A survey of the vision transformers and their CNN-transformer based variants. Artif. Intell. Rev. 2023, 56, 2917–2970. [Google Scholar] [CrossRef]
- Maurício, J.; Domingues, I.; Bernardino, J. Comparing Vision Transformers and Convolutional Neural Networks for Image Classification: A Literature Review. Appl. Sci. 2023, 13, 5521. [Google Scholar] [CrossRef]
- Wang, Y.; Deng, Y.; Zheng, Y.; Chattopadhyay, P.; Wang, L. Vision Transformers for Image Classification: A Comparative Survey. Technologies 2025, 13, 32. [Google Scholar] [CrossRef]
- Le, D.P.C.; Wang, D.; Le, V.T. A Comprehensive Survey of Recent Transformers in Image, Video and Diffusion Models. Comput. Mater. Contin. 2024, 80, 37–60. [Google Scholar] [CrossRef]
- Atkeson, C.G.; Schaal, S. Memory-based neural networks for robot learning. Neurocomputing 1995, 9, 243–269. [Google Scholar] [CrossRef]
- Das, S.; Giles, C.L.; Sun, G.Z. Learning Context-free Grammars: Capabilities and Limitations of a Recurrent Neural Network with an External Stack Memory. arXiv 1992. [Google Scholar]
- Graves, A. Generating Sequences with Recurrent Neural Networks. arXiv 2013, arXiv:1308.0850. [Google Scholar] [CrossRef]
- Hochreiter, S.; Schmidhuber, J. Long Short-Term Memory. Neural Comput. 1997, 9, 1735–1780. [Google Scholar] [CrossRef] [PubMed]
- Sukhbaatar, S.; Szlam, A.; Weston, J.; Fergus, R. End-To-End Memory Networks. arXiv 2015, arXiv:1503.08895. [Google Scholar] [CrossRef]
- Weston, J.; Chopra, S.; Bordes, A. Memory networks. arXiv 2014, arXiv:1305.4807. [Google Scholar] [CrossRef]
- Kumar, A.; Irsoy, O.; Ondruska, P.; Iyyer, M.; Bradbury, J.; Gulrajani, I.; Zhong, V.; Paulus, R.; Socher, R. Ask Me Anything: Dynamic Memory Networks for Natural Language Processing. J. Mach. Learn. Res. 2015, arXiv:1506.07285. [Google Scholar] [CrossRef]
- Oh, S.W.; Lee, J.Y.; Xu, N.; Kim, S.J. Video Object Segmentation using Space-Time Memory Networks. arXiv 2019, arXiv:1904.00607. [Google Scholar] [CrossRef]
- Seong, H.; Oh, S.W.; Lee, J.Y.; Lee, S.; Lee, S.; Kim, E. Hierarchical Memory Matching Network for Video Object Segmentation. arXiv 2021, arXiv:2109.11404. [Google Scholar] [CrossRef]
- Liang, S.; Shen, X.; Huang, J.; Hua, X.S. Video Object Segmentation with Dynamic Memory Networks and Adaptive Object Alignment. In Proceedings of the 2021 IEEE/CVF International Conference on Computer Vision (ICCV); IEEE: New York, NY, USA, 2021. [Google Scholar] [CrossRef]
- Vibashan, V.S.; Oza, P.; Sindagi, V.A.; Gupta, V.; Patel, V.M. MeGA-CDA: Memory Guided Attention for Category-Aware Unsupervised Domain Adaptive Object Detection. arXiv 2021, arXiv:2103.04224. [Google Scholar] [CrossRef]
- Chen, Y.; Wang, Y.; Pan, Y.; Yao, T.; Tian, X.; Mei, T. A Style and Semantic Memory Mechanism for Domain Generalization. arXiv 2021, arXiv:2112.07517. [Google Scholar] [CrossRef]
- Hou, J.; Zhang, Y.; Zhong, Q.; Xie, D.; Zhou, H. Divide-and-Assemble: Learning Block-wise Memory for Unsupervised Anomaly Detection. arXiv 2021, arXiv:2107.13118. [Google Scholar] [CrossRef]
- Gong, D.; Liu, L.; Le, V.; Saha, B.; Mansour, M.R.; Venkatesh, S.; Van den Hengel, A. Memorizing Normality to Detect Anomaly: Memory-Augmented Deep Autoencoder for Unsupervised Anomaly Detection. arXiv 2020, arXiv:1904.02639. [Google Scholar] [CrossRef]
- Park, H.; Noh, J.; Ham, B. Learning Memory-guided Normality for Anomaly Detection. arXiv 2020, arXiv:2003.13228. [Google Scholar] [CrossRef]
- Sandler, M.; Zhmoginov, A.; Vladymyrov, M.; Jackson, A. Fine-tuning Image Transformers using Learnable Memory. arXiv 2022, arXiv:2203.15243. [Google Scholar] [CrossRef]
- Wu, D.; Hu, Y.C.; Hsiao, T.Y.; Liu, H. Uniform Memory Retrieval with Larger Capacity for Modern Hopfield Models. arXiv 2024, arXiv:2404.03827. [Google Scholar] [CrossRef]
- Tran, C.D.; Pham, L.H.; Tran, N.N.; Ho, P.N.; Jeon, J.W. Dual Memory Networks Guided Reverse Distillation for Unsupervised Anomaly Detection; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2025; pp. 361–378. [Google Scholar] [CrossRef]
- Lu, H.; Guo, Z.; Zuo, W. Modulated Memory Network for Video Object Segmentation. Mathematics 2024, 12, 863. [Google Scholar] [CrossRef]
- McGrath, C.; Chau, C.W.R.; Molina, G.F. Monitoring oral health remotely: Ethical considerations when using AI among vulnerable populations. Front. Oral Health 2025, 6, 1587630. [Google Scholar] [CrossRef]
- Hu, Z.; Cao, D.; Hu, Y.; Wang, B.; Zhang, Y.; Tang, R.; Zhuang, J.; Gao, A.; Chen, Y.; Lin, Z. Diagnosis of in vivo vertical root fracture using deep learning on cone-beam CT images. BMC Oral Health 2022, 22, 382. [Google Scholar] [CrossRef]
- Wu, Z.; Xiong, Y.; Yu, S.X.; Lin, D. Unsupervised Feature Learning via Non-parametric Instance Discrimination. In Proceedings of the 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition; IEEE: New York, NY, USA, 2018. [Google Scholar] [CrossRef]



| Segmentation | Category | Total (Train/Test) |
|---|---|---|
| Auto Segmentation | VRF | 555 (416/139) |
| Non-VRF | 563 (424/139) | |
| Manual Segmentation | VRF | 276 (208/68) |
| Non-VRF | 276 (208/68) |
| Auto Segmentation | Accuracy | Sensitivity | Specificity | PPV | F1 Score |
| SBMN (ResNet50) | 97.1 | 98.6 | 95.7 | 95.8 | 97.1 |
| SBMN (DenseNet169) | 93.5 | 92.1 | 95.0 | 94.8 | 93.4 |
| SBMN (VGG19) | 94.6 | 93.5 | 95.7 | 95.6 | 94.5 |
| SBMN (Inception_v3) | 78.9 | 91.4 | 66.2 | 73.0 | 81.2 |
| SBMN (ResNeXt) | 94.6 | 92.1 | 97.1 | 97.0 | 94.5 |
| SBMN (ViT) | 92.4 | 93.7 | 91.2 | 91.2 | 92.4 |
| Manual Segmentation | Accuracy | Sensitivity | Specificity | PPV | F1 Score |
| SBMN (ResNet50) | 99.1 | 100.0 | 97.1 | 97.1 | 98.5 |
| SBMN (DenseNet169) | 98.4 | 97.1 | 100.0 | 100.0 | 98.5 |
| SBMN (VGG19) | 98.5 | 100.0 | 97.1 | 97.1 | 98.6 |
| SBMN (Inception_v3) | 80.1 | 80.9 | 79.4 | 79.7 | 80.3 |
| SBMN (ResNeXt) | 97.1 | 100.0 | 94.1 | 94.4 | 97.1 |
| SBMN (ViT) | 99.7 | 100.0 | 99.5 | 99.8 | 99.7 |
| Auto Segmentation | Accuracy | Sensitivity | Specificity | PPV | F1 Score |
| ResNet50 | 91.4 | 92.1 | 90.7 | 90.8 | 91.4 |
| DenseNet169 | 87.1 | 80.6 | 93.5 | 92.6 | 86.6 |
| VGG19 | 87.8 | 89.2 | 86.3 | 86.7 | 87.7 |
| Inception_v3 | 87.1 | 84.9 | 89.2 | 88.7 | 86.8 |
| ResNeXt | 92.4 | 92.8 | 92.1 | 92.1 | 92.5 |
| SCCNN | 93.9 | 95.0 | 92.9 | 93.0 | 94.0 |
| ViT-B | 90.6 | 90.1 | 91.1 | 90.9 | 90.6 |
| Swin-T | 91.5 | 92.8 | 90.2 | 90.4 | 91.5 |
| Mobile-ViT | 87.9 | 86.5 | 89.3 | 88.9 | 87.9 |
| SBMN (ResNet50) | 97.1 | 98.6 | 95.7 | 95.8 | 97.1 |
| Manual Segmentation | Accuracy | Sensitivity | Specificity | PPV | F1 Score |
| ResNet50 | 97.8 | 97.0 | 98.5 | 98.5 | 97.8 |
| DenseNet169 | 96.3 | 94.1 | 98.5 | 98.5 | 96.2 |
| VGG19 | 94.9 | 92.7 | 97.0 | 96.9 | 94.8 |
| Inception_v3 | 87.8 | 86.3 | 89.2 | 88.9 | 87.6 |
| ResNeXt | 97.8 | 97.1 | 98.5 | 98.5 | 97.8 |
| SCCNN | 98.0 | 98.3 | 96.2 | 97.8 | 97.2 |
| ViT-B | 98.2 | 98.1 | 98.3 | 98.2 | 98.2 |
| Swin-T | 98.6 | 98.4 | 98.2 | 98.3 | 98.3 |
| Mobile-ViT | 97.4 | 96.4 | 98.2 | 98.1 | 97.3 |
| SBMN (ResNet50) | 99.1 | 100.0 | 97.1 | 97.1 | 98.5 |
| Similarity Calculation Method | Accuracy | Sensitivity | Specificity | PPV | F1 Score |
|---|---|---|---|---|---|
| Dot Product | 97.1 | 98.6 | 95.7 | 95.8 | 97.1 |
| Cosine Similarity | 97.0 | 97.6 | 98.2 | 96.4 | 97.0 |
| Pearson Correlation | 96.8 | 96.5 | 97.1 | 96.7 | 96.6 |
| Euclidean Distance | 94.7 | 94.9 | 94.6 | 95.1 | 95.0 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Wang, J.; Jin, X.Y.; Zhang, Y.F.; Yuan, J.; Lin, Z.T.; Chen, Y. SBMN: Similarity-Based Memory Network for the Diagnosis of Vertical Root Fracture in Dental Imaging. Diagnostics 2026, 16, 710. https://doi.org/10.3390/diagnostics16050710
Wang J, Jin XY, Zhang YF, Yuan J, Lin ZT, Chen Y. SBMN: Similarity-Based Memory Network for the Diagnosis of Vertical Root Fracture in Dental Imaging. Diagnostics. 2026; 16(5):710. https://doi.org/10.3390/diagnostics16050710
Chicago/Turabian StyleWang, Jie, Xin Yan Jin, Yi Fan Zhang, Jie Yuan, Zi Tong Lin, and Ying Chen. 2026. "SBMN: Similarity-Based Memory Network for the Diagnosis of Vertical Root Fracture in Dental Imaging" Diagnostics 16, no. 5: 710. https://doi.org/10.3390/diagnostics16050710
APA StyleWang, J., Jin, X. Y., Zhang, Y. F., Yuan, J., Lin, Z. T., & Chen, Y. (2026). SBMN: Similarity-Based Memory Network for the Diagnosis of Vertical Root Fracture in Dental Imaging. Diagnostics, 16(5), 710. https://doi.org/10.3390/diagnostics16050710






