Comprehensive Gene Panel Analysis of Biliary Tract Cancer Using Next-Generation Sequencing of Endoscopic Transpapillary Brushing/Biopsy/Aspiration Specimens: A Narrative Review
Abstract
1. Introduction
2. Brushing Specimens
2.1. Diagnostic Ability and Accuracy Based on Genetic Analysis
2.2. Detected Mutations/Alterations of Genes
3. Forceps Biopsy Specimens
3.1. Diagnostic Ability and Accuracy Based on Genetic Analysis
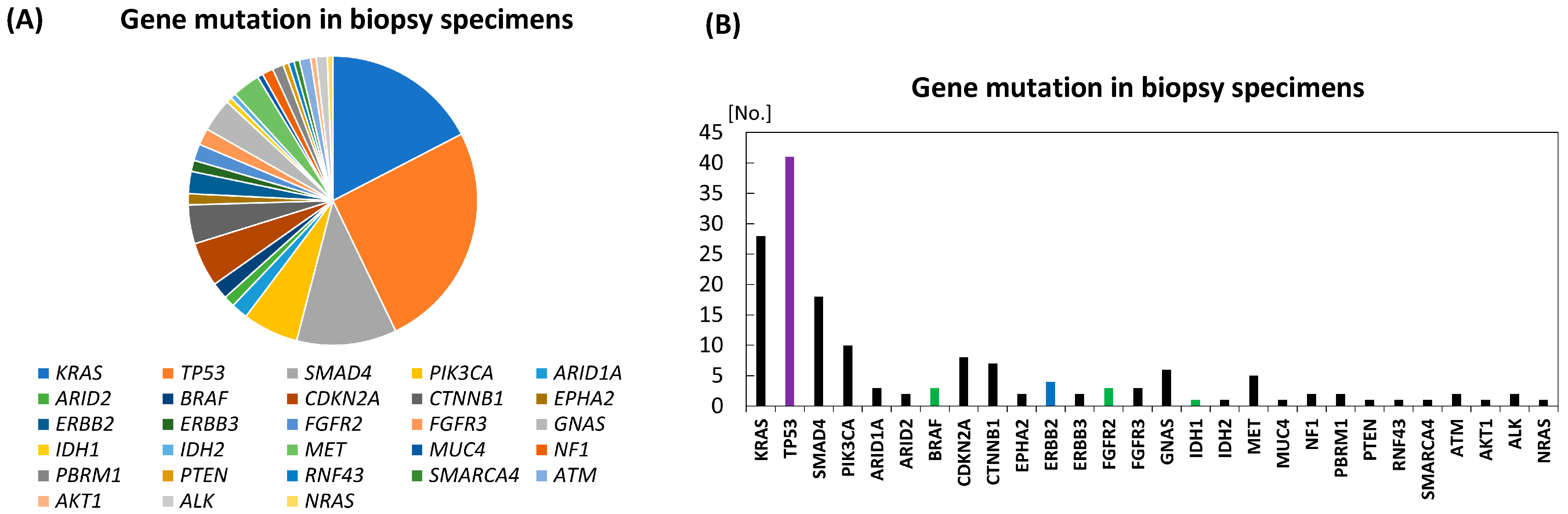
3.2. Detected Mutations/Alterations of Genes
4. Bile (Aspiration) Specimens
4.1. Diagnostic Ability and Accuracy Based on Genetic Analysis
4.2. Detected Mutations/Alterations of Genes
5. Conclusions and Future Perspectives
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- National Cancer Center Japan. Cancer Registry and Statistics; Cancer Information Service. Available online: https://ganjoho.jp/public/qa_links/report/statistics/2025_en.html (accessed on 12 January 2026).
- National Cancer Institute Surveillance. Epidemiology, and End Results (SEER) Program. Available online: https://seer.cancer.gov/ (accessed on 12 January 2026).
- ASGE Standards of Practice Committee; Chandrasekhara, V.; Khashab, M.A.; Muthusamy, V.R.; Acosta, R.D.; Agrawal, D.; Bruining, D.H.; Eloubeidi, M.A.; Fanelli, R.D.; Faulx, A.L.; et al. Adverse events associated with ERCP. Gastrointest. Endosc. 2017, 85, 32–47. [Google Scholar] [CrossRef]
- Dumonceau, J.M.; Kapral, C.; Aabakken, L.; Papanikolaou, I.S.; Tringali, A.; Vanbiervliet, G.; Beyna, T.; Dinis-Ribeiro, M.; Hritz, I.; Mariani, A.; et al. ERCP-related adverse events: European Society of Gastrointestinal Endoscopy (ESGE) Guideline. Endoscopy 2020, 52, 127–149. [Google Scholar] [CrossRef]
- Bishay, K.; Meng, Z.W.; Khan, R.; Gupta, M.; Ruan, Y.; Vaska, M.; Iannuzzi, J.; O’Sullivan, D.E.; Mah, B.; Partridge, A.C.R.; et al. Adverse events associated with endoscopic retrograde cholangiopancreatography: Systematic review and meta-analysis. Gastroenterology 2025, 168, 568–586. [Google Scholar] [CrossRef] [PubMed]
- Ogura, T.; Trong, N.N.; Samanta, J. Advances in Diagnosis and Treatment with Cholangiopancreatoscopy. Dig. Endosc. 2026, 38, e70141. [Google Scholar] [CrossRef]
- Ponchon, T.; Gagnon, P.; Berger, F.; Labadie, M.; Liaras, A.; Chavaillon, A.; Bory, R. Value of endobiliary brush cytology and biopsies for the diagnosis of malignant bile duct stenosis: Results of a prospective study. Gastrointest. Endosc. 1995, 42, 565–572. [Google Scholar] [CrossRef]
- Pugliese, V.; Conio, M.; Nicolò, G.; Saccomanno, S.; Gatteschi, B. Endoscopic retrograde forceps biopsy and brush cytology of biliary strictures: A prospective study. Gastrointest. Endosc. 1995, 42, 520–526. [Google Scholar] [CrossRef]
- Nanda, A.; Brown, J.M.; Berger, S.H.; Lewis, M.M.; Barr Fritcher, E.G.; Gores, G.J.; Keilin, S.A.; Woods, K.E.; Cai, Q.; Willingham, F.F. Triple modality testing by endoscopic retrograde cholangiopancreatography for the diagnosis of cholangiocarcinoma. Ther. Adv. Gastroenterol. 2015, 8, 56–65. [Google Scholar] [CrossRef]
- Naitoh, I.; Nakazawa, T.; Kato, A.; Hayashi, K.; Miyabe, K.; Shimizu, S.; Kondo, H.; Nishi, Y.; Yoshida, M.; Umemura, S.; et al. Predictive factors for positive diagnosis of malignant biliary strictures by transpapillary brush cytology and forceps biopsy. J. Dig. Dis. 2016, 17, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Navaneethan, U.; Njei, B.; Lourdusamy, V.; Konjeti, R.; Vargo, J.J.; Parsi, M.A. Comparative effectiveness of biliary brush cytology and intraductal biopsy for detection of malignant biliary strictures: A systematic review and meta-analysis. Gastrointest. Endosc. 2015, 81, 168–176. [Google Scholar] [CrossRef]
- Weber, A.; von Weyhern, C.; Fend, F.; Schneider, J.; Neu, B.; Meining, A.; Weidenbach, H.; Schmid, R.M.; Prinz, C. Endoscopic transpapillary brush cytology and forceps biopsy in patients with hilar cholangiocarcinoma. World J. Gastroenterol. 2008, 14, 1097–1101. [Google Scholar] [CrossRef] [PubMed]
- Aabakken, L.; Karlsen, T.H.; Albert, J.; Arvanitakis, M.; Chazouilleres, O.; Dumonceau, J.M.; Färkkilä, M.; Fickert, P.; Hirschfield, G.M.; Laghi, A.; et al. Role of endoscopy in primary sclerosing cholangitis: European Society of Gastrointestinal Endoscopy (ESGE) and European Association for the Study of the Liver (EASL) clinical guideline. Endoscopy 2017, 49, 588–608. [Google Scholar] [CrossRef]
- Lee, S.J.; Lee, Y.S.; Lee, M.G.; Lee, S.H.; Shin, E.; Hwang, J.H. Triple-tissue sampling during endoscopic retrograde cholangiopancreatography increases the overall diagnostic sensitivity for cholangiocarcinoma. Gut Liver 2014, 8, 669–673. [Google Scholar] [CrossRef]
- Valle, J.W.; Wasan, H.S.; Palmer, D.H.; Cunningham, D.; Anthoney, A.; Maraveyas, A.; Madhusudan, S.; Iveson, T.; Hughes, S.; Pereira, S.P.; et al. Cisplatin plus gemcitabine versus gemcitabine for biliary tract cancer. N. Engl. J. Med. 2010, 362, 1273–1281. [Google Scholar] [CrossRef]
- Sakai, D.; Kanai, M.; Kobayashi, S.; Eguchi, H.; Baba, H.; Seo, S.; Taketomi, A.; Takayama, T.; Yamaue, H.; Ishioka, C.; et al. Randomized phase III study of gemcitabine, cisplatin plus S-1 (GCS) versus gemcitabine, cisplatin (GC) for advanced biliary tract cancer (KHBO1401-MITSUBA). Ann. Oncol. 2018, 29, viii205. [Google Scholar] [CrossRef]
- Abou-Alfa, G.K.; Sahai, V.; Hollebecque, A.; Vaccaro, G.; Melisi, D.; Al-Rajabi, R.; Paulson, A.S.; Borad, M.J.; Gallinson, D.; Murphy, A.G.; et al. Pemigatinib for previously treated, locally advanced or metastatic cholangiocarcinoma: A multicentre, open-label, phase 2 study. Lancet Oncol. 2020, 21, 671–684. [Google Scholar] [CrossRef] [PubMed]
- Vogel, A.; Sahai, V.; Hollebecque, A.; Vaccaro, G.M.; Melisi, D.; Al Rajabi, R.M.; Paulson, A.S.; Borad, M.J.; Gallinson, D.; Murphy, A.G.; et al. An open-label study of pemigatinib in cholangiocarcinoma: Final results from FIGHT-202. ESMO Open 2024, 9, 103488. [Google Scholar] [CrossRef] [PubMed]
- Oh, D.Y.; Ruth He, A.; Qin, S.; Chen, L.T.; Okusaka, T.; Vogel, A.; Kim, J.W.; Suksombooncharoen, T.; Ah Lee, M.; Kitano, M.; et al. Durvalumab plus gemcitabine and cisplatin in advanced biliary tract cancer. NEJM Evid. 2022, 1, EVIDoa2200015. [Google Scholar] [CrossRef]
- Nakachi, K.; Ikeda, M.; Konishi, M.; Nomura, S.; Katayama, H.; Kataoka, T.; Todaka, A.; Yanagimoto, H.; Morinaga, S.; Kobayashi, S.; et al. Adjuvant S-1 compared with observation in resected biliary tract cancer (JCOG1202, Ascot): A multicentre, open-label, randomised, controlled, phase 3 trial. Lancet 2023, 401, 195–203. [Google Scholar] [CrossRef] [PubMed]
- Primrose, J.N.; Fox, R.P.; Palmer, D.H.; Malik, H.Z.; Prasad, R.; Mirza, D.; Anthony, A.; Corrie, P.; Falk, S.; Finch-Jones, M.; et al. Capecitabine compared with observation in resected biliary tract cancer (BILCAP): A randomised, controlled, multicentre, phase 3 study. Lancet Oncol. 2019, 20, 663–673. [Google Scholar] [CrossRef]
- Sakamoto, Y.; Sereewattanawoot, S.; Suzuki, A. A new era of long-read sequencing for cancer genomics. J. Hum. Genet. 2020, 65, 3–10. [Google Scholar] [CrossRef]
- Alcaide, M.; Cheung, M.; Bushell, K.; Arthur, S.E.; Wong, H.L.; Karasinska, J.; Renouf, D.; Schaeffer, D.F.; McNamara, S.; Tertre, M.C.D.; et al. A novel multiplex droplet digital PCR assay to identify and quantify KRAS mutations in clinical specimens. J. Mol. Diagn. 2019, 21, 214–227. [Google Scholar] [CrossRef] [PubMed]
- Lieb, K.R.; Williams, Z.E.; Sayles, H.; Zaman, M.; Dhir, M. Risk of malignancy for indeterminate cytology of bile duct strictures: A systematic review and meta-analysis. Ann. Surg. Oncol. 2025, 32, 5657–5666. [Google Scholar] [CrossRef]
- Dudley, J.C.; Zheng, Z.; McDonald, T.; Le, L.P.; Dias-Santagata, D.; Borger, D.; Batten, J.; Vernovsky, K.; Sweeney, B.; Arpin, R.N.; et al. Next-generation sequencing and fluorescence in situ hybridization have comparable performance characteristics in the analysis of pancreaticobiliary brushings for malignancy. J. Mol. Diagn. 2016, 18, 124–130. [Google Scholar] [CrossRef]
- Singhi, A.D.; Nikiforova, M.N.; Chennat, J.; Papachristou, G.I.; Khalid, A.; Rabinovitz, M.; Das, R.; Sarkaria, S.; Ayasso, M.S.; Wald, A.I.; et al. Integrating next-generation sequencing to endoscopic retrograde cholangiopancreatography (ERCP)-obtained biliary specimens improves the detection and management of patients with malignant bile duct strictures. Gut 2020, 69, 52–61. [Google Scholar] [CrossRef]
- Rosenbaum, M.W.; Arpin, R.; Limbocker, J.; Casey, B.; Le, L.; Dudley, J.; Iafrate, A.J.; Pitman, M.B. Cytomorphologic characteristics of next-generation sequencing-positive bile duct brushing specimens. J. Am. Soc. Cytopathol. 2020, 9, 520–527. [Google Scholar] [CrossRef]
- Harbhajanka, A.; Michael, C.W.; Janaki, N.; Gokozan, H.N.; Wasman, J.; Bomeisl, P.; Yoest, J.; Sadri, N. Tiny but mighty: Use of next generation sequencing on discarded cytocentrifuged bile duct brushing specimens to increase sensitivity of cytological diagnosis. Mod. Pathol. 2020, 33, 2019–2025. [Google Scholar] [CrossRef] [PubMed]
- Scheid, J.F.; Rosenbaum, M.W.; Przybyszewski, E.M.; Krishnan, K.; Forcione, D.G.; Iafrate, A.J.; Staller, K.D.; Misdraji, J.; Lennerz, J.K.; Pitman, M.B.; et al. Next-generation sequencing in the evaluation of biliary strictures in patients with primary sclerosing cholangitis. Cancer Cytopathol. 2022, 130, 215–230. [Google Scholar] [CrossRef]
- Kamp, E.J.C.A.; Dinjens, W.N.M.; van Velthuysen, M.F.; de Jonge, P.J.F.; Bruno, M.J.; Peppelenbosch, M.P.; de Vries, A.C. Next-generation sequencing mutation analysis on biliary brush cytology for differentiation of benign and malignant strictures in primary sclerosing cholangitis. Gastrointest. Endosc. 2023, 97, 456–465.e6. [Google Scholar] [CrossRef]
- Boyd, S.; Mustamäki, T.; Sjöblom, N.; Nordin, A.; Tenca, A.; Jokelainen, K.; Rantapero, T.; Liuksiala, T.; Lahtinen, L.; Kuopio, T.; et al. NGS of brush cytology samples improves the detection of high-grade dysplasia and cholangiocarcinoma in patients with primary sclerosing cholangitis: A retrospective and prospective study. Hepatol. Commun. 2024, 8, e0415. [Google Scholar] [CrossRef] [PubMed]
- Park, W.; Gwack, J.; Park, J. Implementing massive parallel sequencing into biliary samples obtained through endoscopic retrograde cholangiopancreatography for diagnosing malignant bile duct strictures. Int. J. Mol. Sci. 2024, 25, 9461. [Google Scholar] [CrossRef]
- Arechederra, M.; Rullán, M.; Amat, I.; Oyon, D.; Zabalza, L.; Elizalde, M.; Latasa, M.U.; Mercado, M.R.; Ruiz-Clavijo, D.; Saldaña, C.; et al. Next-generation sequencing of bile cell-free DNA for the early detection of patients with malignant biliary strictures. Gut 2021, 71, 1141–1151. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, H.; Arai, Y.; Totoki, Y.; Shirota, T.; Elzawahry, A.; Kato, M.; Hama, N.; Hosoda, F.; Urushidate, T.; Ohashi, S.; et al. Genomic spectra of biliary tract cancer. Nat. Genet. 2015, 47, 1003–1010. [Google Scholar] [CrossRef]
- Forbes, S.A.; Beare, D.; Boutselakis, H.; Bamford, S.; Bindal, N.; Tate, J.; Cole, C.G.; Ward, S.; Dawson, E.; Ponting, L.; et al. COSMIC: Somatic cancer genetics at high-resolution. Nucleic Acids Res. 2017, 45, D777–D783. [Google Scholar] [CrossRef]
- Macias, R.I.R.; Cardinale, V.; Kendall, T.J.; Avila, M.A.; Guido, M.; Coulouarn, C.; Braconi, C.; Frampton, A.E.; Bridgewater, J.; Overi, D.; et al. Clinical relevance of biomarkers in cholangiocarcinoma: Critical revision and future directions. Gut 2022, 71, 1669–1683. [Google Scholar] [CrossRef] [PubMed]
- Chakravarty, D.; Gao, J.; Phillips, S.M.; Kundra, R.; Zhang, H.; Wang, J.; Rudolph, J.E.; Yaeger, R.; Soumerai, T.; Nissan, M.H.; et al. OncoKB: A precision oncology knowledge base. JCO Precis. Oncol. 2017, 2017, PO.17.00011. [Google Scholar] [CrossRef]
- Mateo, J.; Chakravarty, D.; Dienstmann, R.; Jezdic, S.; Gonzalez-Perez, A.; Lopez-Bigas, N.; Ng, C.K.Y.; Bedard, P.L.; Tortora, G.; Douillard, J.Y.; et al. A framework to rank genomic alterations as targets for cancer precision medicine: The ESMO Scale for Clinical Actionability of molecular Targets (ESCAT). Ann. Oncol. 2018, 29, 1895–1902. [Google Scholar] [CrossRef]
- Chiorean, E.G.; Chiaro, M.D.; Tempero, M.A.; Malafa, M.P.; Benson, A.B.; Cardin, D.B.; Christensen, J.A.; Chung, V.; Czito, B.; Dillhoff, M.; et al. Ampullary adenocarcinoma, version 1.2023, NCCN clinical practice guidelines in oncology. J. Natl. Compr. Canc Netw. 2023, 21, 753–782. [Google Scholar] [CrossRef]
- Benson, A.B.; D’angelica, M.I.; Abrams, T.; Abbott, D.E.; Ahmed, A.; Anaya, D.A.; Anders, R.; Are, C.; Bachini, M.; Binder, D.; et al. NCCN Guidelines® insights: Biliary tract cancers, version 2.2023. J. Natl. Compr. Canc Netw. 2023, 21, 694–704. [Google Scholar] [CrossRef] [PubMed]
- Bankov, K.; Döring, C.; Schneider, M.; Hartmann, S.; Winkelmann, R.; Albert, J.G.; Bechstein, W.O.; Zeuzem, S.; Hansmann, M.L.; Peveling-Oberhag, J.; et al. Sequencing of intraductal biopsies is feasible and potentially impacts clinical management of patients with indeterminate biliary stricture and cholangiocarcinoma. Clin. Transl. Gastroenterol. 2018, 9, 151. [Google Scholar] [CrossRef] [PubMed]
- Wood, H.M.; Conway, C.; Daly, C.; Chalkley, R.; Berri, S.; Senguven, B.; Stead, L.; Ross, L.; Egan, P.; Chengot, P.; et al. The clonal relationships between pre-cancer and cancer revealed by ultra-deep sequencing. J. Pathol. 2015, 237, 296–306. [Google Scholar] [CrossRef]
- Ross-Innes, C.S.; Becq, J.; Warren, A.; Cheetham, R.K.; Northen, H.; O’Donovan, M.; Malhotra, S.; di Pietro, M.; Ivakhno, S.; He, M.; et al. Whole-genome sequencing provides new insights into the clonal architecture of Barrett’s esophagus and esophageal adenocarcinoma. Nat. Genet. 2015, 47, 1038–1046. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, S.; Hijioka, S.; Nagashio, Y.; Yamashige, D.; Agarie, D.; Hagiwara, Y.; Okamoto, K.; Yagi, S.; Komori, Y.; Kuwada, M.; et al. Utility of transpapillary biopsy and endoscopic ultrasound-guided tissue acquisition for comprehensive genome profiling of unresectable biliary tract cancer. Cancers 2024, 16, 2819. [Google Scholar] [CrossRef] [PubMed]
- Vasuri, F.; Albertini, E.; Miranda, L.; Maloberti, T.; Chillotti, S.; Coluccelli, S.; Tallini, G.; D’Errico, A.; Biase, D. Morpho-molecular approach (NGS plus digital PCR) in diagnosis of malignant biliary strictures. Pathologica 2025, 117, 10–17. [Google Scholar] [CrossRef]
- Ishiwatari, H.; Ishikawa, K.; Sato, J.; Niiya, F.; Yasui, H.; Onozawa, Y.; Yamazaki, K.; Todaka, A.; Tsushima, T.; Hamauchi, S.; et al. Optimal number of needle passes in endoscopic ultrasound-guided tissue sampling for genomic profiling: A randomized, noncomparative trial. Gastrointest. Endosc. 2026, 103, 486–493.e2. [Google Scholar] [CrossRef]
- Kinugasa, H.; Nouso, K.; Ako, S.; Dohi, C.; Matsushita, H.; Matsumoto, K.; Kato, H.; Okada, H. Liquid biopsy of bile for the molecular diagnosis of gallbladder cancer. Cancer Biol. Ther. 2018, 19, 934–938. [Google Scholar] [CrossRef]
- Driescher, C.; Fuchs, K.; Haeberle, L.; Goering, W.; Frohn, L.; Opitz, F.V.; Haeussinger, D.; Knoefel, W.T.; Keitel, V.; Esposito, I. Bile-based cell-free DNA analysis is a reliable diagnostic tool in pancreatobiliary cancer. Cancers 2020, 13, 39. [Google Scholar] [CrossRef]
- Nagai, K.; Kuwatani, M.; Hirata, K.; Suda, G.; Hirata, H.; Takishin, Y.; Furukawa, R.; Kishi, K.; Yonemura, H.; Nozawa, S.; et al. Genetic analyses of cell-free DNA in pancreatic juice or bile for diagnosing pancreatic duct and biliary tract strictures. Diagnostics 2022, 12, 2704. [Google Scholar] [CrossRef]
- Miura, Y.; Ohyama, H.; Mikata, R.; Hirotsu, Y.; Amemiya, K.; Mochizuki, H.; Ikeda, J.; Ohtsuka, M.; Kato, N.; Omata, M. The efficacy of bile liquid biopsy in the diagnosis and treatment of biliary tract cancer. J. Hepato-Bil. Pancreat. Sci. 2024, 31, 329–338. [Google Scholar] [CrossRef]
- Ito, S.; Ando, M.; Aoki, S.; Soma, S.; Zhang, J.; Hirano, N.; Kashiwagi, R.; Murakami, K.; Yoshimachi, S.; Sato, H.; et al. Usefulness of multigene liquid biopsy of bile for identifying driver genes of biliary duct cancers. Cancer Sci. 2024, 115, 4054–4063. [Google Scholar] [CrossRef]
- Bardhi, O.; Jones, A.; Ellis, D.; Tielleman, T.; Tavakkoli, A.; Vanderveldt, D.; Goldschmiedt, M.; Singhi, A.; Kubiliun, N.; Sawas, T. Next-generation sequencing improves the detection of malignant biliary strictures and changes management. Gastrointest. Endosc. 2025, 102, 56–63.e1. [Google Scholar] [CrossRef] [PubMed]
- Arechederra, M.; Bik, E.; Rojo, C.; Elurbide, J.; Elizalde, M.; Kruk, B.; Krasnodębski, M.; Pertkiewicz, J.; Kozieł, S.; Grąt, M.; et al. Mutational analysis of bile cell-free DNA in primary sclerosing cholangitis: A pilot study. Liver Int. 2025, 45, e70049. [Google Scholar] [CrossRef] [PubMed]
- Andersson, D.; Kebede, F.T.; Escobar, M.; Österlund, T.; Ståhlberg, A. Principles of digital sequencing using unique molecular identifiers. Mol. Asp. Med. 2024, 96, 101253. [Google Scholar] [CrossRef] [PubMed]
- Dumbrava, E.E.; Shapiro, G.I.; Parikh, A.R.; Johnson, M.L.; Tolcher, A.W.; Thompson, J.A.; El-Khoueiry, A.B.; Vandross, A.L.; Kummar, S.; Shepard, D.R.; et al. Phase 1 study of Rezatapopt, a p53 reactivator, in TP53 Y220C-mutated tumors. N. Engl. J. Med. 2026, 394, 872–883. [Google Scholar] [CrossRef] [PubMed]


| Year | 1st Author | BTC Patient No. Control No. | Panel Gene No. | Detected MAF/VAF | DNA Extraction Success, % (No.) | Amount of DNA Obtained |
|---|---|---|---|---|---|---|
| 2016 | Dudley JC | 11 43 | 39 | ≥5% | 100 (11/11) | ND |
| 2020 | Singhi AD | 41 ‡ 70 | 28 | ≥3% | 100 ‡ | 0.08 to 27.45 ng/μL (mean, 3.17 ng/μL; median, 1.61 ng/μL) ‡ |
| 2020 | Rosenbaum MW | 58 30 | 39 | ≥5% | ND | ND |
| 2020 | Harbhajanka A | 9 51 | 723 | ≥2% (≥0.1%) | 100 | ND |
| 2022 | Scheid JF | 4 56 | Ver. 1, 39 Ver. 2, 116 | ≥5% | 100 (60/60) | ND |
| 2023 | Kamp EJCA | 20 20 | 14 | ≥0.1% | 87 (20/23) | Pursued 50 ng |
| 2024 | Boyd S | 18 8 | 50 | >1–2% | ND | ND |
| 2024 | Park W | 8 || 11 | 161 (Bile, 50) | ≥3% (Bile, ≥0.3%) | 62 (8/13) | ND || (Bile cfDNA ≥ 20 ng, range 1.3 to 20 ng) |
| Alteration No./Gene No. † | Alteration Incidence in Cancer Cases | SN | SP | PPV | NPV | ACC |
| 18/10 | 6/11 (Control: 1/43) | 55 | 98 | 86 | 89 | 89 |
| 47/7 ‡ | 26/41 ‡ (Control: 0/70) | 63 ‡ | 100 ‡ | 100 § (All) | 63 § (All) | 86 § (All) |
| ND | 58/58 | 100 | 73 | 88 | 100 | 91 |
| 15/9 | 8/9 | 89 | 100 | 100 | 98 | 98 |
| 25/11 | 4/4 (PSC control: 6/56) | 100 | 89 | 40 | 100 | 90 |
| 22/7 | 15/20 (PSC control: 4/20) | 75 | 85 | 79 | 76 | 78 |
| 27/7 | 13/18 | 72 | 88 | ND | ND | ND |
| 21/12 || | 8/8 || (Bile: 7/8) | 100 || (Bile 88) | 82 || | 80 || | 100 || | 90 || |
| Year | 1st Author | BTC Patient No. Control No. | Panel Gene No. | Detected MAF/VAF | DNA Extraction Success, % (No.) | Amount of DNA Obtained |
|---|---|---|---|---|---|---|
| 2018 | Bankov K | 16 16 | 41 | 17.7% (range, 4.8–79.9) | 100 (16/16) | All >10 ng |
| 2020 | Singhi AD | 90 ‡ 70 | 28 | ≥3% | 100 ‡ | 0.53 to 35.71 ng/μL (mean, 6.42 ng/μL; median, 4.22 ng/μL) ‡ |
| 2024 | Fukuda S | 350 | 124 or 324 | ND | ND | ND |
| 2025 | Vasuri F | 6 6 | 20 | ND | 92 12/13 | All >10 ng |
| Alteration No./Gene No. † | Alteration Incidence in Cancer Cases | SN | SP | PPV | NPV | ACC |
| 51/20 | 14/16 (Control: /) | 88 | 100 | ND | ND | ND |
| 118/18 ‡ | 64/90 ‡ (Control: 0/70) | 71 ‡ | 100 ‡ | 100 § (All) | 63 § (All) | 86 § (All) |
| ND | ND | ND | ND | ND | ND | ND |
| 11/7 | 5/6 | 83 | 100 | 100 | 86 | 92 |
| Year | 1st Author | BTC Patient No. Control No. | Panel Gene No. | Detected MAF/VAF | DNA Extraction Success, % (No.) | Amount of DNA Obtained |
|---|---|---|---|---|---|---|
| 2018 | Kinugasa H | 24 19 | 49 | ≥5% | 100 (24/24) | ND |
| 2021 | Driescher C | 4 23 | 50 | ≥1% | 100 | ND |
| 2022 | Nagai K | 27 8 | 50 | ≥2% | 100 | 2.4 to 715 ng/μL (after processing) |
| 2022 | Arechederra M | 42 13 | 52 | ≥0.15% | 100 | 886.10 ± 182.3 ng/mL |
| 2024 | Miura Y | 43 29 | 60 | ND | ND | Bile 993.3 ng/mL (IQR, 254.4–3360) |
| 2024 | Ito S | 200 | 7 | >0.005–0.1% | 100 (20/20) | 79.7 to 6165 ng/μL (after processing) |
| 2025 | Bardhi O | 20 § 19 | 28 | ≥3% | 100 (23/23) | ND |
| 2025 | Arechederra M | 4 59 (PSC) | 52 | ≥0.1% | 100 (63/63) | Bile 5.7 μg/mL (range 0.1–70.8 μg/mL) (All cohort) |
| Alteration No./Gene No. † | Alteration Incidence in Cancer Cases | SN | SP | PPV | NPV | ACC |
| 17/4 | 14/24 (Control: 0/19) | 58 | 100 | 100 | 66 | ND |
| 13/5 | 4/4 (Control: 0/23) | 100 | 100 | 100 | 100 | 100 |
| 26/12 | 15/27 | 56 | 75 | 88 | 33 | 60 |
| 116/18 | 42/42 | 100 | 69 | 91 | 100 | 92 |
| 83/34 | 25/43 (PSC control: 6/29) | 47 | 79 | 77 | 50 | 60 |
| 20/7 | 12/20 | 60 | ND | ND | ND | ND |
| ND | ND | 70 § | 96 § | 88 § | 89 § | 88 § |
| 5/5 | 3/4 (PSC control: 20/59) | 75 ‡ | 66 ‡ | 13 ‡ | 98 ‡ | 67 ‡ |
| Mutated Genes | Drug | Drug Class | Approved/Off-Label |
|---|---|---|---|
| FGFR2 | Pemigatinib/ Futibatinib | FGFR inhibitors | Approved |
| IDH1 | Ivosidenib | Inhibitor of isocitrate dehydrogenase 1 | Approved |
| BRAF | Dabrafenib/ Trametinib | BRAF/MEK inhibitor | Approved |
| NTRK | Larotrectinib/ Entrectinib | TRK inhibitors | Approved |
| HER2 | Trastuzumab/ Pertuzumab | Anti-HER2 antibody | Off-label |
| RET | Pralsetinib/ Selpercatinib | Selective inhibitor of RET receptor tyrosine kinase | Off-label |
| NRG-1 | Zenocutuzumab | Anti-HER2/HER3 antibody | Off-label |
| MDM2 | Brigimadlin/ Milademetan | MDM2–p53 antagonist | Off-label |
| BRCA1/2 | Olaparib/ Niraparib | Poly adenosine diphosphate-ribose polymerase inhibitors | Off-label |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Kuwatani, M.; Sakamoto, N. Comprehensive Gene Panel Analysis of Biliary Tract Cancer Using Next-Generation Sequencing of Endoscopic Transpapillary Brushing/Biopsy/Aspiration Specimens: A Narrative Review. Diagnostics 2026, 16, 1516. https://doi.org/10.3390/diagnostics16101516
Kuwatani M, Sakamoto N. Comprehensive Gene Panel Analysis of Biliary Tract Cancer Using Next-Generation Sequencing of Endoscopic Transpapillary Brushing/Biopsy/Aspiration Specimens: A Narrative Review. Diagnostics. 2026; 16(10):1516. https://doi.org/10.3390/diagnostics16101516
Chicago/Turabian StyleKuwatani, Masaki, and Naoya Sakamoto. 2026. "Comprehensive Gene Panel Analysis of Biliary Tract Cancer Using Next-Generation Sequencing of Endoscopic Transpapillary Brushing/Biopsy/Aspiration Specimens: A Narrative Review" Diagnostics 16, no. 10: 1516. https://doi.org/10.3390/diagnostics16101516
APA StyleKuwatani, M., & Sakamoto, N. (2026). Comprehensive Gene Panel Analysis of Biliary Tract Cancer Using Next-Generation Sequencing of Endoscopic Transpapillary Brushing/Biopsy/Aspiration Specimens: A Narrative Review. Diagnostics, 16(10), 1516. https://doi.org/10.3390/diagnostics16101516







