Phenotype-First Diagnostic Framework for Tracking Fluoroquinolone Resistance in Escherichia coli
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design and Oversight
2.2. Isolates and Inclusion Criteria
2.3. Species Identification (Conventional Methods Followed by MALDI-TOF MS)
2.4. Antimicrobial Susceptibility Testing (AST)
2.5. ESBL/AmpC Phenotypes, Multidrug Resistance (MDR), and MAR Index
2.6. Selection of Founders for Experimental Evolution
2.7. Experimental Evolution Under Ciprofloxacin
2.8. Post-Selection Stability and Compensatory Adaptation
2.9. Growth Kinetics (Fitness) Assays
2.10. Biofilm Quantification
2.11. Outcomes and Definitions
2.12. Data Handling and Statistical Analysis
3. Results
3.1. Isolate Recovery, Identification, and Quality Control
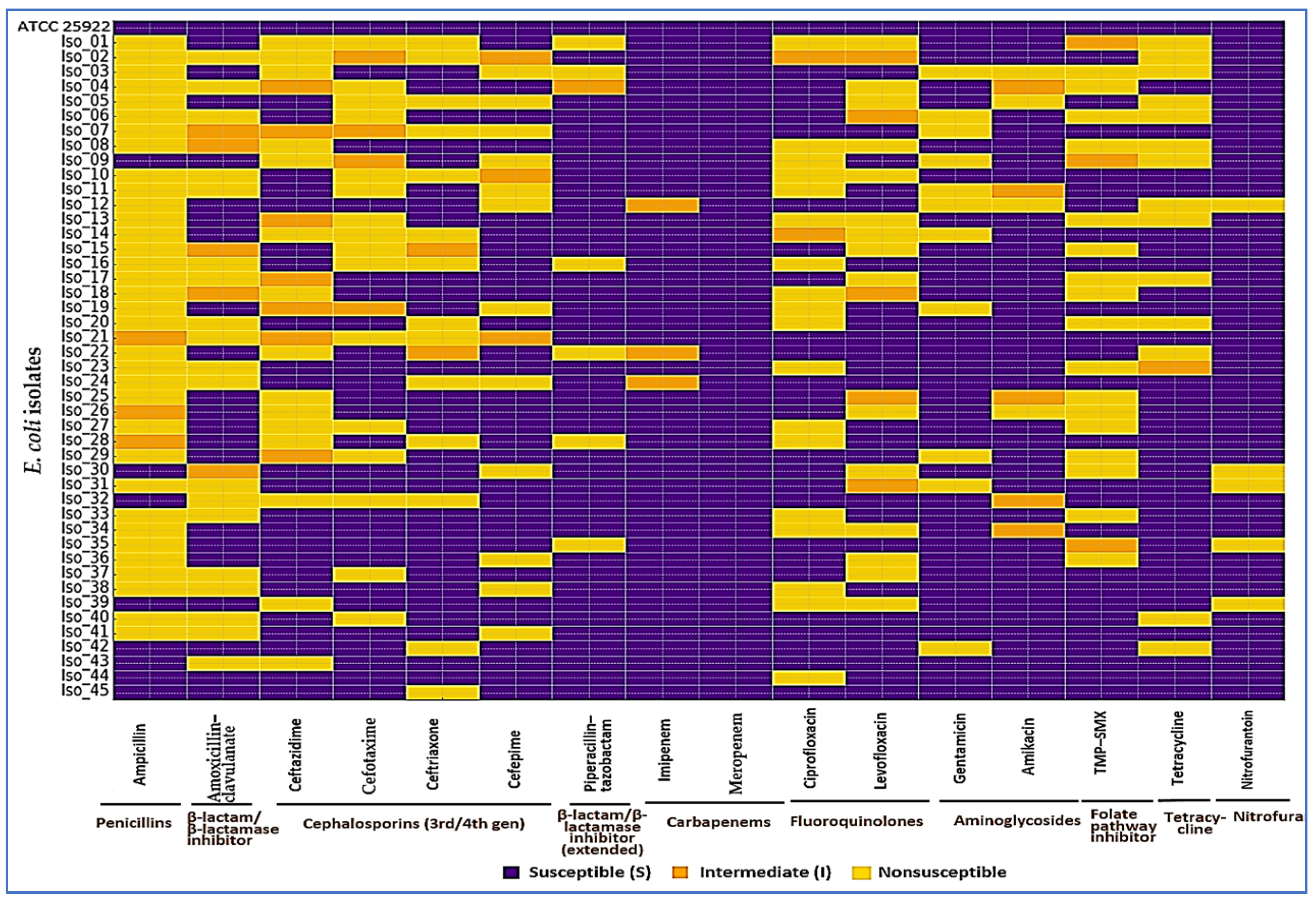
3.2. Baseline Susceptibility, ESBL/AmpC, MDR, and MAR (Arm 1)
3.3. Founder Set and Baseline Characteristics (Arm 2 Entry)
3.4. Evolution Under Ciprofloxacin: Trajectories and Time to High-Level Resistance
3.5. Collateral Resistance and MAR Dynamics
3.6. Growth Fitness and Biofilm Metrics
4. Discussion
5. Strengths and Limitations of the Study
6. Future Directions
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tamma, P.D.; Heil, E.L.; Justo, J.A.; Mathers, A.J.; Satlin, M.J.; Bonomo, R.A. Infectious Diseases Society of America 2024 guidance on the treatment of antimicrobial-resistant gram-negative infections. Clin. Infect. Dis. 2024, ciae403. [Google Scholar] [CrossRef] [PubMed]
- Ajulo, S.; Awosile, B. Global antimicrobial resistance and use surveillance system (GLASS 2022): Investigating the relationship between antimicrobial resistance and antimicrobial consumption data across the participating countries. PLoS ONE 2024, 19, e0297921. [Google Scholar] [CrossRef] [PubMed]
- Cuevas, O.; Oteo, J.; Lazaro, E.; Aracil, B.; De Abajo, F.; Garcia-Cobos, S.; Ortega, A.; Campos, J.; Group, S.E.-N.S.; Fontanals, D. Significant ecological impact on the progression of fluoroquinolone resistance in Escherichia coli with increased community use of moxifloxacin, levofloxacin and amoxicillin/clavulanic acid. J. Antimicrob. Chemother. 2011, 66, 664–669. [Google Scholar] [CrossRef] [PubMed]
- WHO. WHO Bacterial Priority Pathogens List, 2024: Bacterial Pathogens of Public Health Importance to Guide Research, Development and Strategies to Prevent and Control Antimicrobial Resistance; World Health Organization: Geneva, Switzerland, 2024; Available online: https://www.who.int/publications/i/item/9789240093461 (accessed on 15 September 2025).
- Stapleton, A.E.; Wagenlehner, F.M.; Mulgirigama, A.; Twynholm, M. Escherichia coli resistance to fluoroquinolones in community-acquired uncomplicated urinary tract infection in women: A systematic review. Antimicrob. Agents Chemother. 2020, 64, e00862-20. [Google Scholar] [CrossRef]
- WHO. Global Antimicrobial Resistance and Use Surveillance System (GLASS) Report: 2022; World Health Organization: Geneva, Switzerland, 2022; Available online: https://www.who.int/pubLications/i/item/9789240062702 (accessed on 15 September 2025).
- Ruiz-Lievano, A.P.; Cervantes-Flores, F.; Nava-Torres, A.; Carbajal-Morales, P.J.; Villaseñor-Garcia, L.F.; Zavala-Cerna, M.G. Fluoroquinolone resistance in Escherichia coli causing community-acquired urinary tract infections: A systematic review. Microorganisms 2024, 12, 2320. [Google Scholar] [CrossRef]
- Hooper, D.C.; Jacoby, G.A. Mechanisms of drug resistance: Quinolone resistance. Ann. N. Y. Acad. Sci. 2015, 1354, 12–31. [Google Scholar] [CrossRef]
- Strahilevitz, J.; Jacoby, G.A.; Hooper, D.C.; Robicsek, A. Plasmid-mediated quinolone resistance: A multifaceted threat. Clin. Microbiol. Rev. 2009, 22, 664–689. [Google Scholar] [CrossRef]
- Andersson, D.I.; Hughes, D. Antibiotic resistance and its cost: Is it possible to reverse resistance? Nat. Rev. Microbiol. 2010, 8, 260–271. [Google Scholar] [CrossRef]
- Byrd, B.A.; Zenick, B.; Rocha-Granados, M.C.; Englander, H.E.; Hare, P.J.; LaGree, T.J.; DeMarco, A.M.; Mok, W.W. The AcrAB-TolC efflux pump impacts persistence and resistance development in stationary-phase Escherichia coli following delafloxacin treatment. Antimicrob. Agents Chemother. 2021, 65, 5806–5816. [Google Scholar] [CrossRef]
- Nesse, L.L.; Osland, A.M.; Asal, B.; Mo, S.S. Evolution of antimicrobial resistance in E. coli biofilm treated with high doses of ciprofloxacin. Front. Microbiol. 2023, 14, 1246895. [Google Scholar] [CrossRef]
- Wilson, C.; Lukowicz, R.; Merchant, S.; Valquier-Flynn, H.; Caballero, J.; Sandoval, J.; Okuom, M.; Huber, C.; Brooks, T.D.; Wilson, E. Quantitative and qualitative assessment methods for biofilm growth: A mini-review. Res. Rev. J. Eng. Technol. 2017, 6, 1–7. [Google Scholar]
- Pitout, J.D.; Peirano, G.; Chen, L.; DeVinney, R.; Matsumura, Y. Escherichia coli ST1193: Following in the footsteps of E. coli ST131. Antimicrob. Agents Chemother. 2022, 66, e00511-22. [Google Scholar] [CrossRef]
- Reid, C.J.; McKinnon, J.; Djordjevic, S.P. Clonal ST131-H 22 Escherichia coli strains from a healthy pig and a human urinary tract infection carry highly similar resistance and virulence plasmids. Microb. Genom. 2019, 5, e000295. [Google Scholar] [PubMed]
- Muloi, D.M.; Jauneikaite, E.; Anjum, M.F.; Essack, S.Y.; Singleton, D.A.; Kasudi, M.R.; Wade, M.J.; Egyir, B.; Nunn, J.G.; Midega, J.T. Exploiting genomics for antimicrobial resistance surveillance at One Health interfaces. Lancet Microbe 2023, 4, e1056–e1062. [Google Scholar] [CrossRef] [PubMed]
- Sundermann, A.J.; Rosa, R.; Harris, P.N.; Snitkin, E.; Javaid, W.; Moore, N.M.; Hayden, M.K.; Allen, K.; Rodino, K.; Peacock, S.J. Pathogen genomics in healthcare: Overcoming barriers to proactive surveillance. Antimicrob. Agents Chemother. 2025, 69, e01479-24. [Google Scholar] [CrossRef]
- Teichmann, L.; Luitwieler, S.; Bengtsson-Palme, J.; Ter Kuile, B. Fluoroquinolone-specific resistance trajectories in E. coli and their dependence on the SOS-response. BMC Microbiol. 2025, 25, 37. [Google Scholar] [CrossRef] [PubMed]
- Dash, J.; Naykodi, A.; Mohakud, N.K.; Deb, S. MALDI TOF-MS for microbial identification and diagnosis. In Evolving Landscape of Molecular Diagnostics; Elsevier: Amsterdam, The Netherlands, 2024; pp. 43–57. [Google Scholar] [CrossRef]
- Patel, R. MALDI-TOF MS for the diagnosis of infectious diseases. Clin. Chem. 2015, 61, 100–111. [Google Scholar] [CrossRef]
- Elbehiry, A.; Abalkhail, A. Spectral Precision: Recent Advances in Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry for Pathogen Detection and Resistance Profiling. Microorganisms 2025, 13, 1473. [Google Scholar] [CrossRef]
- Elbehiry, A.; Marzouk, E.; Moussa, I.M.; Alenzi, A.; Al-Maary, K.S.; Mubarak, A.S.; Alshammari, H.D.; Al-Sarar, D.; Alsubki, R.A.; Hemeg, H.A. Multidrug-resistant Escherichia coli in Raw Milk: Molecular Characterization and the potential impact of camel’s Urine as an Antibacterial Agent. Saudi J. Biol. Sci. 2021, 28, 2091–2097. [Google Scholar] [CrossRef]
- Papadomanolaki, A.; Siopi, M.; Karakosta, P.; Vourli, S.; Pournaras, S. Comparative Evaluation of Vitek 2 and Etest versus Broth microdilution for ceftazidime/avibactam and ceftolozane/tazobactam susceptibility testing of Enterobacterales and Pseudomonas aeruginosa. Antibiotics 2022, 11, 865. [Google Scholar] [CrossRef]
- Weinstein, M.P.; Limbago, B.; Patel, J.; Mathers, A.; Campeau, S.; Mazzulli, T.; Eliopoulos, G.; Patel, R.; Galas, M.; Richter, S. M100 performance standards for antimicrobial susceptibility testing. Clin. Lab. Stand. Inst. 2018, 27, 210–214. [Google Scholar]
- Wiegand, I.; Hilpert, K.; Hancock, R.E. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 2008, 3, 163–175. [Google Scholar] [CrossRef] [PubMed]
- O’Toole, G.A. Microtiter Dish Biofilm Formation Assay. J. Vis. Exp. 2011, 47, e2437. [Google Scholar] [CrossRef] [PubMed]
- ASM. MacConkey Agar Plates—Protocols. Available online: https://asm.org/protocols/macconkey-agar-plates-protocols (accessed on 24 September 2025).
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing (M100); Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2025; Available online: https://clsi.org/shop/standards/m100/? (accessed on 24 September 2025).
- Kim, T.Y.; Baek, J.Y.; Suh, E.; Lee, J.-K.; Yu, H.-J.; Ko, J.-H.; Huh, H.J.; Lee, N.Y. Evaluation of the VITEK 2 AST-N439 card for susceptibility testing of novel β-lactam/β-lactamase inhibitor combinations and colistin in carbapenem-non-susceptible gram-negative bacilli. Microbiol. Spectr. 2025, 13, e00166-25. [Google Scholar] [CrossRef] [PubMed]
- Coudron, P.E. Inhibitor-based methods for detection of plasmid-mediated AmpC β-lactamases in Klebsiella spp., Escherichia coli, and Proteus mirabilis. J. Clin. Microbiol. 2005, 43, 4163–4167. [Google Scholar] [CrossRef]
- Magiorakos, A.-P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.; Giske, C.; Harbarth, S.; Hindler, J.; Kahlmeter, G.; Olsson-Liljequist, B. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef]
- Krumperman, P.H. Multiple antibiotic resistance indexing of Escherichia coli to identify high-risk sources of fecal contamination of foods. Appl. Environ. Microbiol. 1983, 46, 165–170. [Google Scholar] [CrossRef]
- WHO. Antimicrobial Resistance. Available online: https://www.who.int/news-room/fact-sheets/detail/antimicrobial-resistance (accessed on 29 September 2025).
- Naghavi, M.; Vollset, S.E.; Ikuta, K.S.; Swetschinski, L.R.; Gray, A.P.; Wool, E.E.; Aguilar, G.R.; Mestrovic, T.; Smith, G.; Han, C. Global burden of bacterial antimicrobial resistance 1990–2021: A systematic analysis with forecasts to 2050. Lancet 2024, 404, 1199–1226. [Google Scholar] [CrossRef]
- Sati, H.; Carrara, E.; Savoldi, A.; Hansen, P.; Garlasco, J.; Campagnaro, E.; Boccia, S.; Castillo-Polo, J.A.; Magrini, E.; Garcia-Vello, P. The WHO Bacterial Priority Pathogens List 2024: A prioritisation study to guide research, development, and public health strategies against antimicrobial resistance. Lancet Infect. Dis. 2025, 25, 1033–1043. [Google Scholar] [CrossRef]
- Calderaro, A.; Chezzi, C. MALDI-TOF MS: A reliable tool in the real life of the clinical microbiology laboratory. Microorganisms 2024, 12, 322. [Google Scholar] [CrossRef]
- Elbehiry, A.; Aldubaib, M.; Abalkhail, A.; Marzouk, E.; Albeloushi, A.; Moussa, I.; Ibrahem, M.; Albazie, H.; Alqarni, A.; Anagreyyah, S. How MALDI-TOF mass spectrometry technology contributes to microbial infection control in healthcare settings. Vaccines 2022, 10, 1881. [Google Scholar] [CrossRef] [PubMed]
- Anorue, M.; Ejikeugwu, C.; Iroha, C.S.; David, E.E.; Nwabueze, E.F.; Iroha, I.R. Extended spectrum beta-lactamase producing Escherichia coli encoding aminoglycoside and fluoroquinolone resistant genes in urinary tract infection patients in a tertiary hospital in Nigeria. BMC Infect. Dis. 2025, 25, 1171. [Google Scholar] [CrossRef] [PubMed]
- Lawal, S.T.; Usman, F.A.; Adams, Z.A.; Ogunbayo, O.S.; Ekwedigwe, C.M.; Jimoh, R.O.; Oladeru, F.O.; Osho, O.; Essiet, U.U.; Ajayi, A. Genetic Determinants of Carbapenem and Fluoroquinolone Resistance in Escherichia coli Isolates of Clinical Origin. Infect. Chemother. 2025, 57, 102. [Google Scholar] [CrossRef] [PubMed]
- Topa, A.-E.; Ionescu, C.; Pinzaru, A.; Mocanu, E.; Iancu, A.M.; Dumea, E.; Nitu, B.F.; Panculescu, F.G.; Cambrea, S.C. Challenges in the Treatment of Urinary Tract Infections: Antibiotic Resistance Profiles of Escherichia coli Strains Isolated from Young and Elderly Patients in a Southeastern Romanian Hospital. Biomedicines 2025, 13, 1066. [Google Scholar] [CrossRef]
- Alós, J.-I.; Serrano, M.-G.; Gómez-Garcés, J.-L.; Perianes, J. Antibiotic resistance of Escherichia coli from community-acquired urinary tract infections in relation to demographic and clinical data. Clin. Microbiol. Infect. 2005, 11, 199–203. [Google Scholar] [CrossRef]
- García-Meniño, I.; García, V.; Lumbreras-Iglesias, P.; Fernández, J.; Mora, A. Fluoroquinolone resistance in complicated urinary tract infections: Association with the increased occurrence and diversity of Escherichia coli of clonal complex 131, together with ST1193. Front. Cell. Infect. Microbiol. 2024, 14, 1351618. [Google Scholar] [CrossRef]
- Zlamal, J.E.; Leyn, S.A.; Iyer, M.; Elane, M.L.; Wong, N.A.; Wamsley, J.W.; Vercruysse, M.; Garcia-Alcalde, F.; Osterman, A.L. Shared and unique evolutionary trajectories to ciprofloxacin resistance in gram-negative bacterial pathogens. mBio 2021, 12, e0098721. [Google Scholar] [CrossRef]
- de Lastours, V.; El Meouche, I.; Chau, F.; Beghain, J.; Chevret, D.; Aubert-Frambourg, A.; Clermont, O.; Royer, G.; Bouvet, O.; Denamur, E. Evolution of fluoroquinolone-resistant Escherichia coli in the gut after ciprofloxacin treatment. Int. J. Med. Microbiol. 2022, 312, 151548. [Google Scholar] [CrossRef]
- Garoff, L.; Pietsch, F.; Huseby, D.L.; Lilja, T.; Brandis, G.; Hughes, D. Population bottlenecks strongly influence the evolutionary trajectory to fluoroquinolone resistance in Escherichia coli. Mol. Biol. Evol. 2020, 37, 1637–1646. [Google Scholar] [CrossRef]
- Toprak, E. Evolutionary paths to strong antibiotic resistance under dynamically sustained drug stress. Nat. Genet. 2011, 44, 101–105. [Google Scholar] [CrossRef]
- Huseby, D.L.; Pietsch, F.; Brandis, G.; Garoff, L.; Tegehall, A.; Hughes, D. Mutation supply and relative fitness shape the genotypes of ciprofloxacin-resistant Escherichia coli. Mol. Biol. Evol. 2017, 34, 1029–1039. [Google Scholar] [CrossRef] [PubMed]
- Jacoby, G.A.; Strahilevitz, J.; Hooper, D.C. Plasmid-Mediated Quinolone Resistance. In Plasmids: Biology and Impact in Biotechnology and Discovery; Wiley: Hoboken, NJ, USA, 2015; pp. 475–503. [Google Scholar]
- Blair, J.M.; Richmond, G.E.; Piddock, L.J. Multidrug efflux pumps in Gram-negative bacteria and their role in antibiotic resistance. Future Microbiol. 2014, 9, 1165–1177. [Google Scholar] [CrossRef] [PubMed]
- Webber, M.A.; Piddock, L.J. The importance of efflux pumps in bacterial antibiotic resistance. J. Antimicrob. Chemother. 2003, 51, 9–11. [Google Scholar] [CrossRef] [PubMed]
- Chetri, S. The culmination of multidrug-resistant efflux pumps vs. meager antibiotic arsenal era: Urgent need for an improved new generation of EPIs. Front. Microbiol. 2023, 14, 1149418. [Google Scholar] [CrossRef]
- Melnyk, A.H.; Wong, A.; Kassen, R. The fitness costs of antibiotic resistance mutations. Evol. Appl. 2015, 8, 273–283. [Google Scholar] [CrossRef]
- IDSA. IDSA 2024 Guidance on the Treatment of Antimicrobial Resistant Gram-Negative Infections. Available online: https://www.idsociety.org/practice-guideline/amr-guidance/ (accessed on 29 September 2025).
- Lázár, V.; Pal Singh, G.; Spohn, R.; Nagy, I.; Horváth, B.; Hrtyan, M.; Busa-Fekete, R.; Bogos, B.; Méhi, O.; Csörgő, B. Bacterial evolution of antibiotic hypersensitivity. Mol. Syst. Biol. 2013, 9, 700. [Google Scholar] [CrossRef]
- Imamovic, L.; Sommer, M.O. Use of collateral sensitivity networks to design drug cycling protocols that avoid resistance development. Sci. Transl. Med. 2013, 5, 204ra132. [Google Scholar] [CrossRef]
- Podnecky, N.L.; Fredheim, E.G.; Kloos, J.; Sørum, V.; Primicerio, R.; Roberts, A.P.; Rozen, D.E.; Samuelsen, Ø.; Johnsen, P.J. Conserved collateral antibiotic susceptibility networks in diverse clinical strains of Escherichia coli. Nat. Commun. 2018, 9, 3673. [Google Scholar] [CrossRef]
- Nichol, D.; Rutter, J.; Bryant, C.; Hujer, A.M.; Lek, S.; Adams, M.D.; Jeavons, P.; Anderson, A.R.; Bonomo, R.A.; Scott, J.G. Antibiotic collateral sensitivity is contingent on the repeatability of evolution. Nat. Commun. 2019, 10, 334. [Google Scholar] [CrossRef]
- Sørum, V.; Øynes, E.L.; Møller, A.S.; Harms, K.; Samuelsen, Ø.; Podnecky, N.L.; Johnsen, P.J. Evolutionary instability of collateral susceptibility networks in ciprofloxacin-resistant clinical Escherichia coli strains. MBio 2022, 13, e00441-22. [Google Scholar] [CrossRef]
- Liu, D.Y.; Phillips, L.; Wilson, D.M.; Fulton, K.M.; Twine, S.M.; Wong, A.; Linington, R.G. Collateral sensitivity profiling in drug-resistant Escherichia coli identifies natural products suppressing cephalosporin resistance. Nat. Commun. 2023, 14, 1976. [Google Scholar] [CrossRef] [PubMed]
- Shehata, A.A.; Abd-Elfatah, E.B.; Elsheik, H.E.; Salman, M.B.; Khater, A.S.I.; El-Emam, M.M.A. Epidemiological Features, Biochemical Indices, Antibiogram Susceptibility Profile and Biofilm Factor Genes of Klebsiella pneumoniae Isolated from Bovine Clinical Mastitis Cases. Pak. Vet. J. 2024, 44, 141–147. [Google Scholar]
- Aldred, K.J.; Kerns, R.J.; Osheroff, N. Mechanism of quinolone action and resistance. Biochemistry 2014, 53, 1565–1574. [Google Scholar] [CrossRef] [PubMed]
- Chubiz, L.M. The mar, sox, and rob systems. EcoSal Plus 2023, 11, eesp00102022. [Google Scholar] [CrossRef]
- Agnello, M.; Finkel, S.E.; Wong-Beringer, A. Fitness cost of fluoroquinolone resistance in clinical isolates of Pseudomonas aeruginosa differs by type III secretion genotype. Front. Microbiol. 2016, 7, 1591. [Google Scholar] [CrossRef]
- Bhatnagar, K.; Wong, A. The mutational landscape of quinolone resistance in Escherichia coli. PLoS ONE 2019, 14, e0224650. [Google Scholar] [CrossRef]
- Vanacker, M.; Lenuzza, N.; Rasigade, J.-P. The fitness cost of horizontally transferred and mutational antimicrobial resistance in Escherichia coli. Front. Microbiol. 2023, 14, 1186920. [Google Scholar] [CrossRef]
- FDA. FDA Drug Safety Communication: FDA Updates Warnings for Oral and Injectable Fluoroquinolone Antibiotics Due to Disabling Side Effects. Available online: https://www.fda.gov/drugs/drug-safety-and-availability/fda-drug-safety-communication-fda-updates-warnings-oral-and-injectable-fluoroquinolone-antibiotics (accessed on 23 September 2025).
- WHO. WHO Updates List of Drug-Resistant Bacteria Most Threatening to Human Health. 2024. Available online: https://www.who.int/news/item/17-05-2024-who-updates-list-of-drug-resistant-bacteria-most-threatening-to-human-health (accessed on 26 September 2025).
- Schuster, S.; Bohnert, J.A.; Vavra, M.; Rossen, J.W.; Kern, W.V. Proof of an outer membrane target of the efflux inhibitor Phe-Arg-β-naphthylamide from random mutagenesis. Molecules 2019, 24, 470. [Google Scholar] [CrossRef]





| Antimicrobial Class | Agent(s) Tested | Nonsusceptible | |
|---|---|---|---|
| Number of Isolates | Percentage | ||
| Penicillins | Ampicillin | 34/45 | 75.6 |
| β-lactam/β-lactamase inhibitor | Amoxicillin–clavulanate | 19/45 | 42.2 |
| Cephalosporins (3rd/4th gen) | Cefotaxime | 16/45 | 35.6 |
| Ceftazidime | 15/45 | 33.3 | |
| Ceftriaxone | 14/45 | 31.1 | |
| Cefepime | 12/45 | 26.7 | |
| β-lactam/β-lactamase inhibitor (extended) | Piperacillin–tazobactam | 6/45 | 13.3 |
| Carbapenems | Imipenem | 0/45 | 0.0 |
| Meropenem | 0/45 | 0.0 | |
| Fluoroquinolones | Ciprofloxacin | 18/45 | 40.0 |
| Levofloxacin | 15/45 | 33.3 | |
| Aminoglycosides | Gentamicin | 11/45 | 24.4 |
| Amikacin | 4/45 | 8.9 | |
| Folate pathway inhibitor | TMP–SMX | 17/45 | 37.8 |
| Tetracycline class | Tetracycline | 14/45 | 31.1 |
| Nitrofuran | Nitrofurantoin | 5/45 | 11.1 |
| Resistance phenotypes | ESBL confirmed | 13/45 | 28.9 |
| AmpC screen-positive (cefoxitin) | 11/45 | 24.4 | |
| — inhibitor-based AmpC confirmed | 7/45 | 15.6 | |
| Composite indices | MDR prevalence (≥1 agent in ≥3 classes) | 21/45 | 46.7 |
| MAR index, median (IQR) [value: 0.29 (0.14–0.50)] | — | — | |
| Frequent co-resistance partners | Third-generation cephalosporins; TMP–SMX; tetracyclines | — | — |
| Founder Group | No. | Ciprofloxacin MIC, Median (IQR) (µg/mL) | ESBL Positive | AmpC Confirmed | MDR | MAR Index, Median (IQR) | |||
|---|---|---|---|---|---|---|---|---|---|
| No. | % | No. | % | No. | % | ||||
| FQ-S | 5 | 0.06 (0.06–0.125) | 2 | 40% | 1 | 20% | 2 | 40% | 0.18 (0.10–0.29) |
| LLR | 5 | 1 (0.5–2) | 3 | 60% | 1 | 20% | 3 | 60% | 0.40 (0.25–0.55) |
| Overall | 10 | 0.25 (0.06–2) | 5 | 50% | 2 | 20% | 5 | 50% | 0.29 (0.14–0.50) |
| Outcome/Metric | Overall 10 Founders; 20 Lineages | FQ-S Lineages 5 Founders; 10 Lineages | LLR Lineages 5 Founders; 10 Lineages | Statistics/Notes |
|---|---|---|---|---|
| Reached HLR, n/N (%) | 14/20 (70%) | 7/10 (70%) | 7/10 (70%) | Kaplan–Meier cumulative incidence at passage 15 ≈ 68% |
| Passages to HLR, median (IQR) | NE 1 | 11 (9–14) | 7 (6–9) | Log-rank p = 0.018 (LLR earlier than FQ-S) |
| Lineage extinction before HLR, n/N (%) | 3/20 (15%) | 3/10 (30%) | 0/10 (0%) | Failures at 2–4× founder MIC |
| Endpoint ciprofloxacin MIC (μg/mL), median (IQR) | 8 (4–16) | NR 2 | NR 2 | Median among surviving lineages at endpoint |
| MIC fold-increase vs. founder, median (IQR) | 32× (8–128×) | NR 2 | NR 2 | Median across lineages |
| Mixed-effects model: passage effect (β on log2(MIC)) | 0.22 per passage | NE 1 | NE 1 | 95% CI 0.18–0.26; p < 0.001 |
| Mixed-effects model: founder effect (LLR vs. FQ-S) | NE 1 | NE 1 | NE 1 | Δβ = 0.06 (95% CI 0.02–0.10); p = 0.004 |
| Category | Metric | Group | Founders Median (IQR) [n] | Endpoints Median (IQR) [n] | Delta Endpoint–Founder Median (IQR) | Notes/p-Value |
|---|---|---|---|---|---|---|
| Drug-free growth | µmax, h−1 | FQ-S | 1.30 (1.22–1.36) [n = 5] | 1.18 (1.12–1.24) [n = 10] | −0.12 (−0.20 to −0.05) | paired Wilcoxon p = 0.020 |
| LLR | 1.25 (1.18–1.32) [n = 5] | 1.20 (1.12–1.26) [n = 10] | −0.05 (−0.12 to 0.00) | paired Wilcoxon p = 0.090 | ||
| Overall | 1.28 (1.22–1.34) [n = 10] | 1.19 (1.12–1.25) [n = 20] | −0.08 (−0.16 to −0.02) | paired Wilcoxon p = 0.030 | ||
| Lag time (h) | FQ-S | 0.25 (0.22–0.28) [n = 5] | 0.32 (0.28–0.36) [n = 10] | +0.07 (+0.04 to +0.10) | paired Wilcoxon p = 0.010 | |
| LLR | 0.24 (0.21–0.27) [n = 5] | 0.29 (0.26–0.33) [n = 10] | +0.05 (+0.02 to +0.08) | paired Wilcoxon p = 0.060 | ||
| Overall | 0.24 (0.22–0.27) [n = 10] | 0.30 (0.27–0.34) [n = 20] | +0.06 (+0.03 to +0.09) | paired Wilcoxon p = 0.020 | ||
| Carrying capacity (OD600) | FQ-S | 0.92 (0.88–0.96) [n = 5] | 0.90 (0.86–0.95) [n = 10] | −0.02 (−0.05 to +0.01) | paired Wilcoxon p = 0.280 | |
| LLR | 0.94 (0.90–0.98) [n = 5] | 0.93 (0.89–0.97) [n = 10] | −0.01 (−0.04 to +0.02) | paired Wilcoxon p = 0.410 | ||
| Overall | 0.93 (0.89–0.97) [n = 10] | 0.92 (0.88–0.96) [n = 20] | −0.01 (−0.04 to +0.01) | paired Wilcoxon p = 0.190 | ||
| AUC, AU | FQ-S | 18.5 (17.8–19.2) [n = 5] | 17.4 (16.5–18.2) [n = 10] | −1.1 (−1.8 to −0.4) | paired Wilcoxon p = 0.020 | |
| LLR | 18.8 (18.0–19.5) [n = 5] | 18.4 (17.6–19.0) [n = 10] | −0.4 (−1.2 to +0.3) | paired Wilcoxon p = 0.180 | ||
| Overall | 18.7 (17.9–19.3) [n = 10] | 17.9 (17.1–18.7) [n = 20] | −0.8 (−1.5 to −0.2) | paired Wilcoxon p = 0.030 | ||
| Sub-inhibitory ciprofloxacin | AUC, AU | FQ-S | 12.0 (11.3–12.8) [n = 5] | 12.9 (12.1–13.7) [n = 10] | +0.9 (+0.2 to +1.6) | paired Wilcoxon p = 0.040 |
| LLR | 13.4 (12.6–14.1) [n = 5] | 14.0 (13.3–14.7) [n = 10] | +0.6 (0.0 to +1.3) | paired Wilcoxon p = 0.070 | ||
| Overall | 12.7 (12.0–13.6) [n = 10] | 13.5 (12.7–14.3) [n = 20] | +0.7 (+0.1 to +1.4) | paired Wilcoxon p = 0.030 | ||
| Biofilm phenotype | Crystal violet biomass (OD590) | FQ-S | 0.42 (0.38–0.46) [n = 5] | 0.44 (0.39–0.48) [n = 10] | +0.02 (−0.01 to +0.04) | group effect (endpoint) FQ-S vs. LLR: p = 0.62 |
| LLR | 0.43 (0.40–0.47) [n = 5] | 0.45 (0.41–0.49) [n = 10] | +0.02 (−0.01 to +0.05) | group effect (endpoint) FQ-S vs. LLR: p = 0.62 | ||
| Overall | 0.42 (0.39–0.47) [n = 10] | 0.45 (0.40–0.49) [n = 20] | +0.02 (−0.01 to +0.04) | no systematic shift across lineages |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marzouk, E.; Almuzaini, A.M. Phenotype-First Diagnostic Framework for Tracking Fluoroquinolone Resistance in Escherichia coli. Diagnostics 2025, 15, 2831. https://doi.org/10.3390/diagnostics15222831
Marzouk E, Almuzaini AM. Phenotype-First Diagnostic Framework for Tracking Fluoroquinolone Resistance in Escherichia coli. Diagnostics. 2025; 15(22):2831. https://doi.org/10.3390/diagnostics15222831
Chicago/Turabian StyleMarzouk, Eman, and Abdulaziz M. Almuzaini. 2025. "Phenotype-First Diagnostic Framework for Tracking Fluoroquinolone Resistance in Escherichia coli" Diagnostics 15, no. 22: 2831. https://doi.org/10.3390/diagnostics15222831
APA StyleMarzouk, E., & Almuzaini, A. M. (2025). Phenotype-First Diagnostic Framework for Tracking Fluoroquinolone Resistance in Escherichia coli. Diagnostics, 15(22), 2831. https://doi.org/10.3390/diagnostics15222831






