A Deep Learning Framework for the Detection of Abnormality in Cerebral Blood Flow Velocity Using Transcranial Doppler Ultrasound
Abstract
1. Introduction
1.1. Related Work
1.2. Motivation
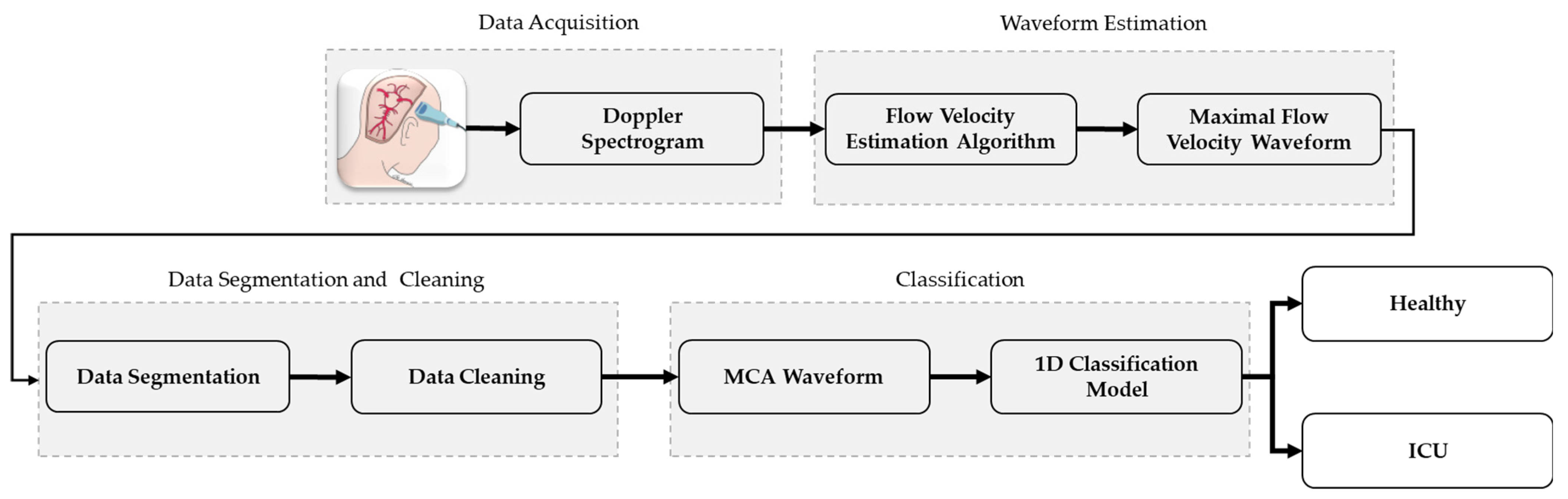
- A novel TCD ultrasound waveform classification system that incorporates cerebral blood flow velocity (CBFV) waveform estimation, data cleaning and segmentation, and classification for classifying data from doppler ultrasounds in both healthy subjects and intensive care unit subjects.
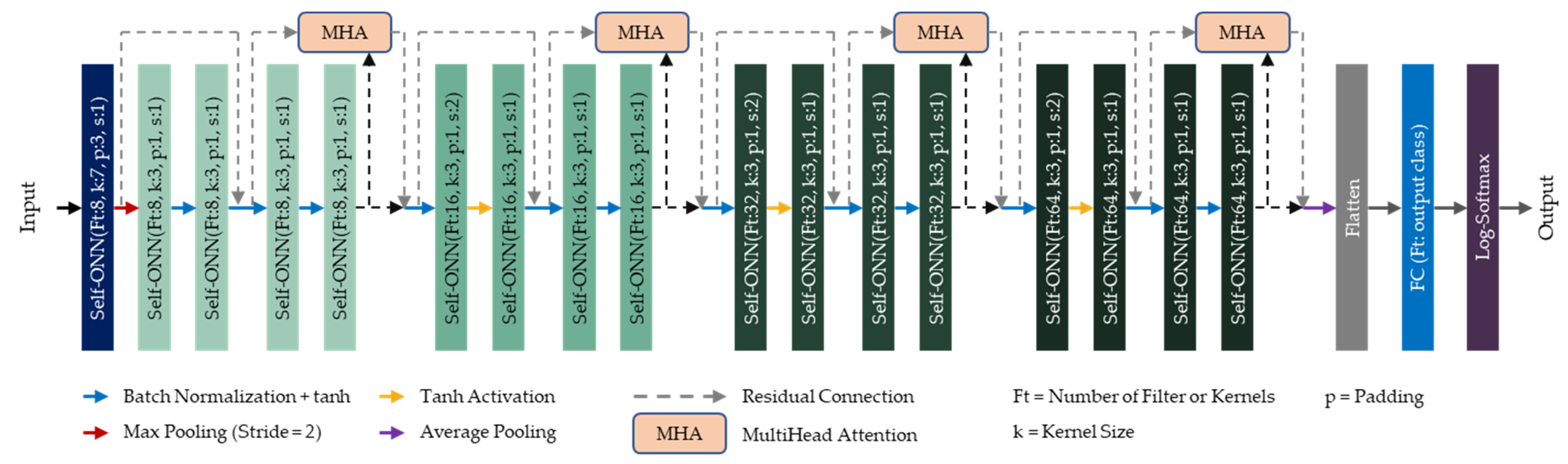
- Two novel SelfONN architectures, (a) a 1D version of ResNet18 and (b) a 1D version of ResNet architecture with a multiheaded attention layer, have been proposed for 1D binary classification, where TCD ultrasound can be used to identify ICU (in this study, “ICU” is an umbrella term used for representing MCA waveforms from hydrocephalus, traumatic brain injury, and intraparenchymal or subarachnoid hemorrhage patients) patients from healthy subjects by analyzing the maximal cerebral blood flow velocity (CBFV) waveforms.
2. Materials and Methods
2.1. Overview of the Dataset
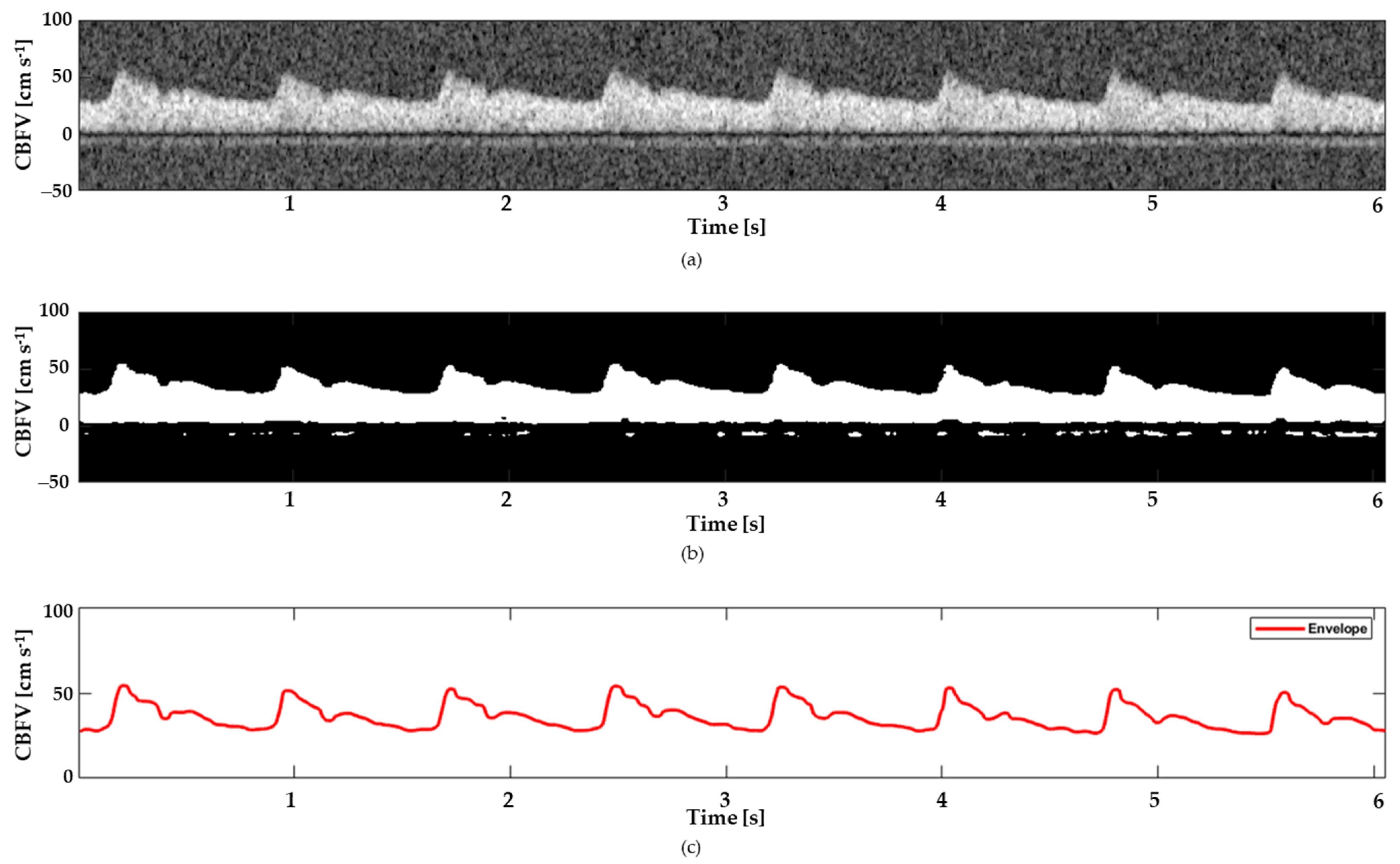
2.2. Signal Extraction
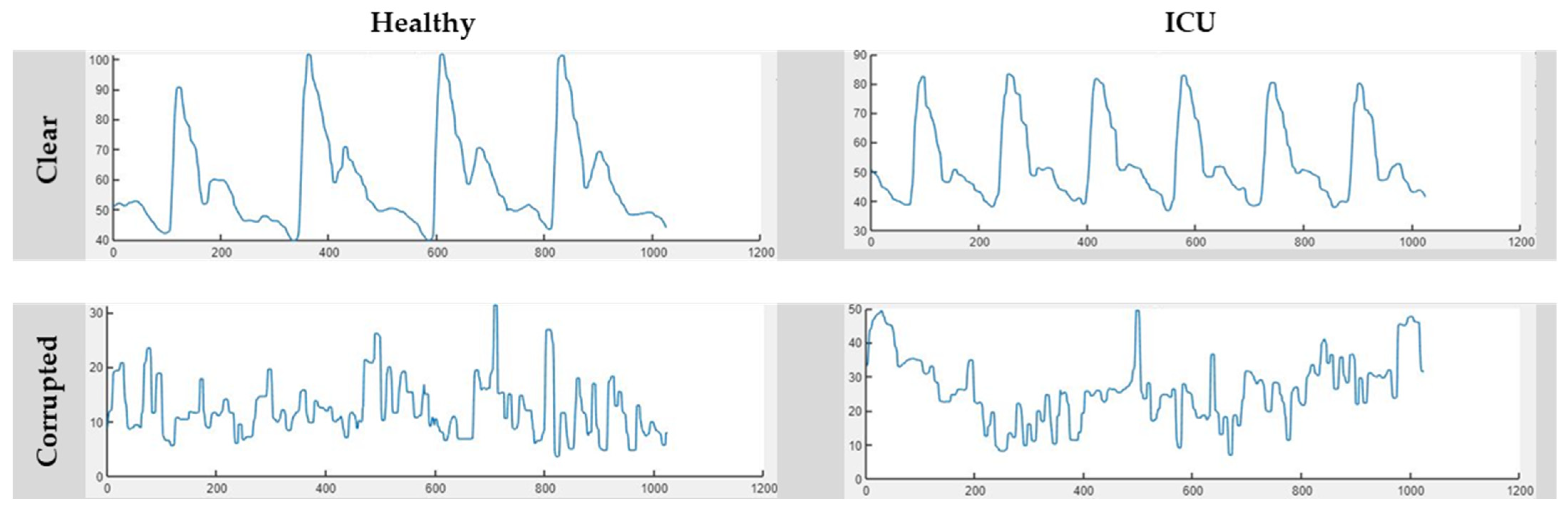
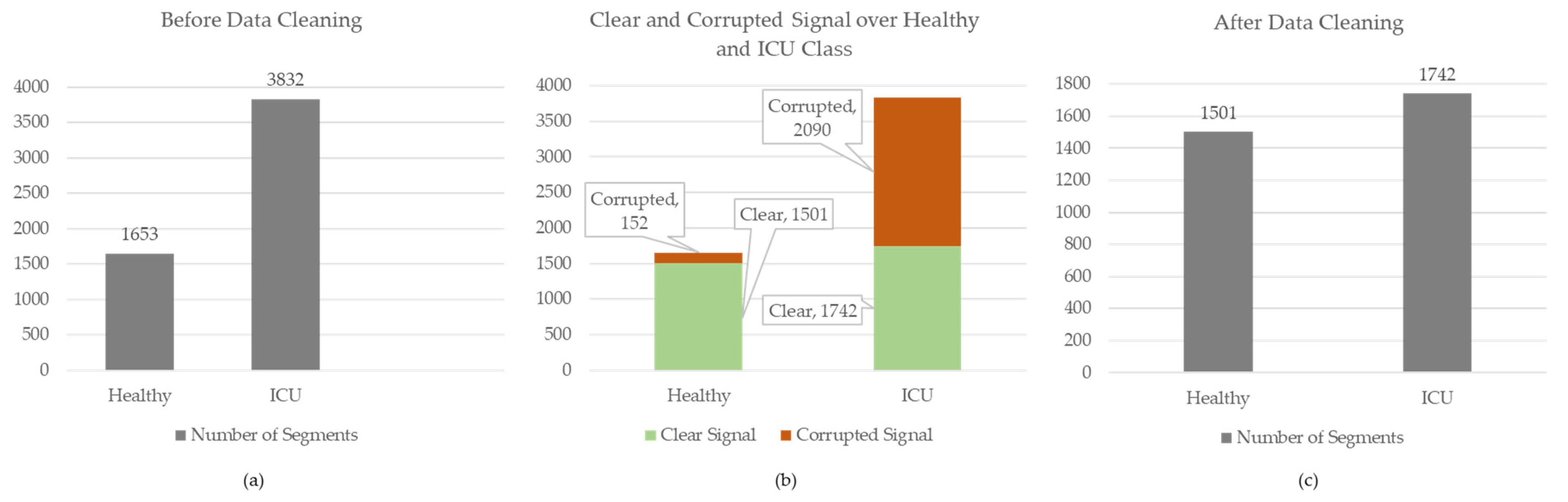
2.3. Manual Data Annotation
2.4. Model Architecture
2.4.1. Self-ONN
2.4.2. Self-ResNet18
2.4.3. Self-ResAttentioNet18
2.5. Experimental Setup
2.6. Performance Metrics
3. Results and Discussion
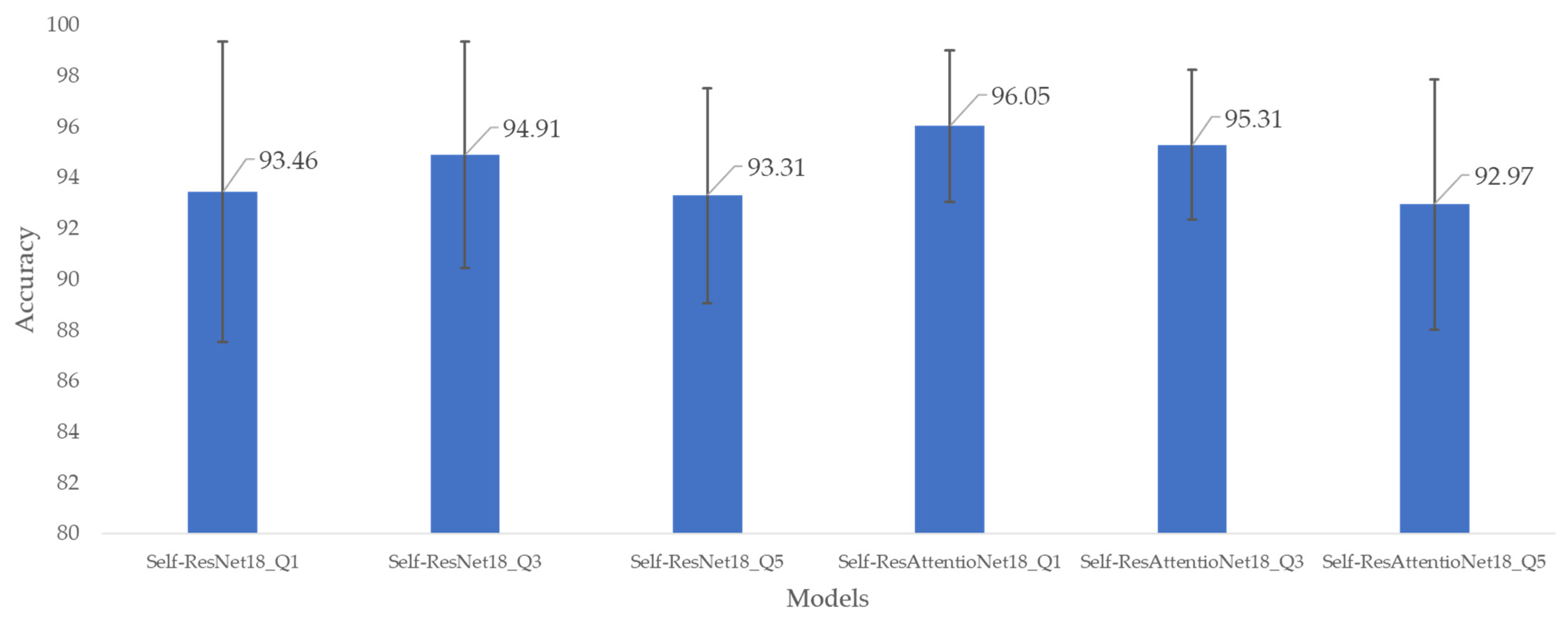
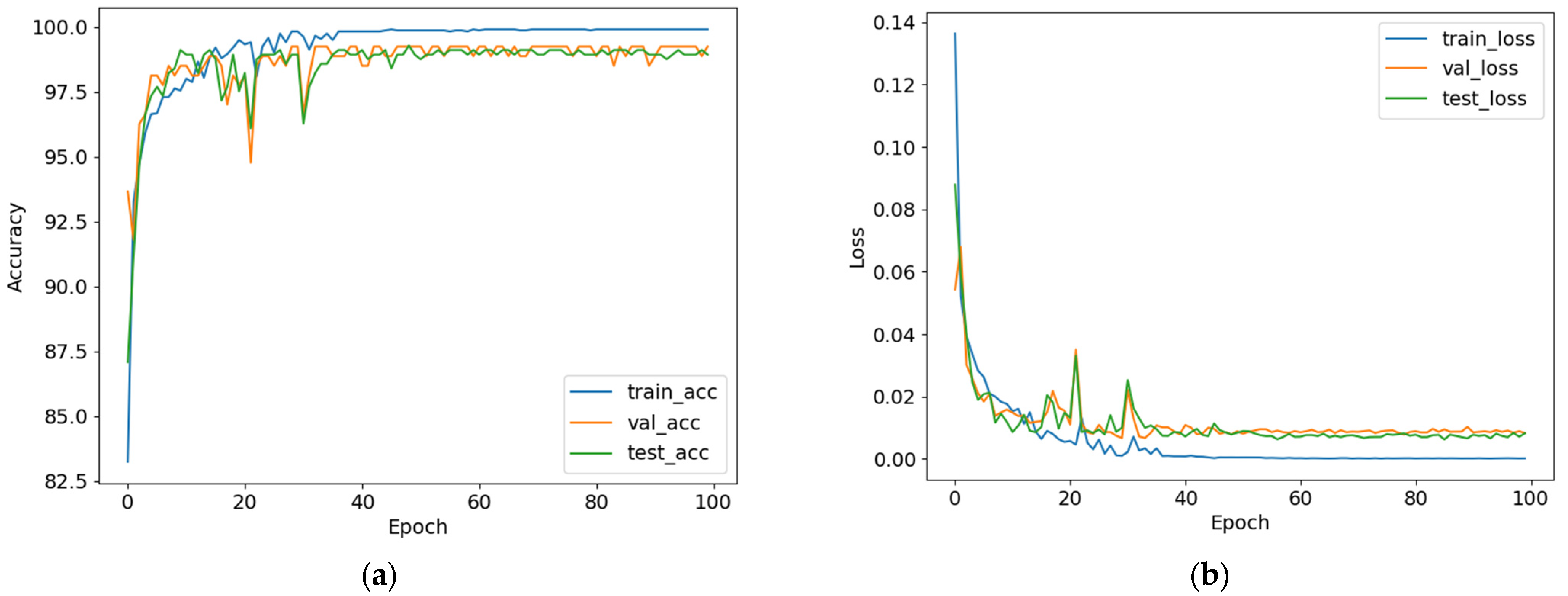
3.1. Accuracy-Based Comparison
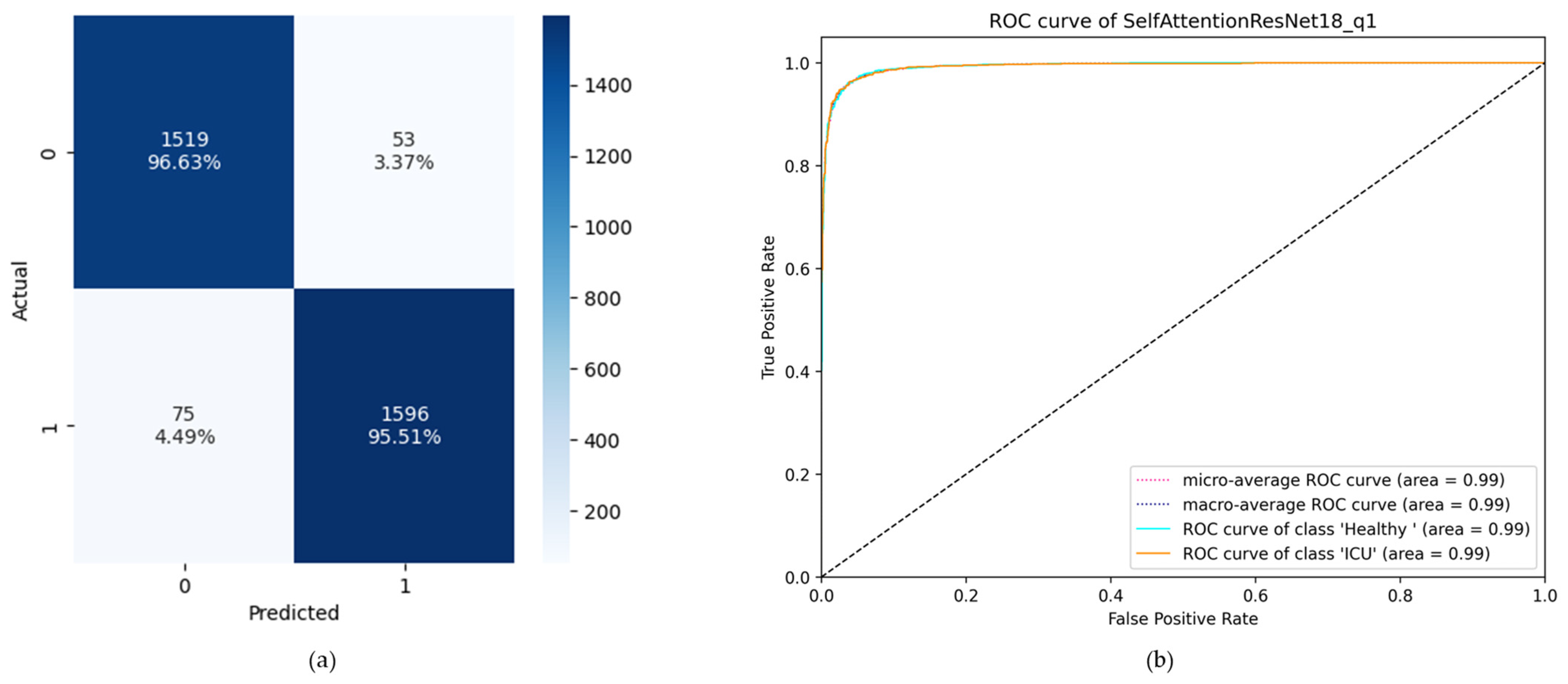
3.2. ROC Curve and Confusion Matrix-Based Comparison
3.3. Comparison with Respect to Previous Literature
4. Conclusions
5. Limitations and Future Work
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Szabo, T.L. Diagnostic Ultrasound Imaging: Inside Out; Academic Press: Cambridge, MA, USA, 2004. [Google Scholar]
- Shung, K.K. Diagnostic Ultrasound: Imaging and Blood Flow Measurements; CRC Press: Boca Raton, FL, USA, 2005. [Google Scholar]
- Birdsill, A.C.; Carlsson, C.M.; Willette, A.A.; Okonkwo, O.C.; Johnson, S.C.; Xu, G.; Oh, J.M.; Gallagher, C.L.; Koscik, R.L.; Jonaitis, E.M.; et al. Low cerebral blood flow is associated with lower memory function in metabolic syndrome. Obesity 2013, 21, 1313–1320. [Google Scholar] [CrossRef] [PubMed]
- Jarrett, C.L.; Shields, K.L.; Broxterman, R.M.; Hydren, J.R.; Park, S.H.; Gifford, J.R.; Richardson, R.S. Imaging transcranial Doppler ultrasound to measure middle cerebral artery blood flow: The importance of measuring vessel diameter. Am. J. Physiol. Integr. Comp. Physiol. 2020, 319, R33–R42. [Google Scholar] [CrossRef] [PubMed]
- Ozturk, E.D.; Tan, C.O. Human cerebrovascular function in health and disease: Insights from integrative approaches. J. Physiol. Anthropol. 2018, 37, 1–11. [Google Scholar] [CrossRef]
- Christou, I.; Felberg, R.A.; Demchuk, A.M.; Grotta, J.C.; Burgin, W.; Malkoff, M.; Alexandrov, A.V. A broad diagnostic battery for bedside transcranial Doppler to detect flow changes with internal carotid artery stenosis or occlusion. J. Neuroimaging 2001, 11, 236–242. [Google Scholar] [CrossRef]
- Thorpe, S.G.; Thibeault, C.M.; Wilk, S.J.; O’brien, M.; Canac, N.; Ranjbaran, M.; Devlin, C.; Devlin, T.; Hamilton, R.B. Velocity curvature index: A novel diagnostic biomarker for large vessel occlusion. Transl. Stroke Res. 2019, 10, 475–484. [Google Scholar] [CrossRef] [PubMed]
- Thorpe, S.G.; Thibeault, C.M.; Canac, N.; Jalaleddini, K.; Dorn, A.; Wilk, S.J.; Devlin, T.; Scalzo, F.; Hamilton, R.B. Toward automated classification of pathological transcranial Doppler waveform morphology via spectral clustering. PLoS ONE 2020, 15, e0228642. [Google Scholar] [CrossRef] [PubMed]
- Wadehn, F.; Heldt, T. Adaptive Maximal Blood Flow Velocity Estimation From Transcranial Doppler Echos. IEEE J. Transl. Eng. Health Med. 2020, 8, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Aaslid, R.; Markwalder, T.-M.; Nornes, H. Noninvasive transcranial Doppler ultrasound recording of flow velocity in basal cerebral arteries. J. Neurosurg. 1982, 57, 769–774. [Google Scholar] [CrossRef] [PubMed]
- Übeyli, E.D.; Güler, İ. Improving medical diagnostic accuracy of ultrasound Doppler signals by combining neural network models. Comput. Biol. Med. 2005, 35, 533–554. [Google Scholar] [CrossRef]
- Seddik, A.F.; Shawky, D.M. A low-cost screening method for the detection of the carotid artery diseases. Knowl.-Based Syst. 2013, 52, 236–245. [Google Scholar] [CrossRef]
- Übeyli, E.D.; Güler, İ. Wavelet-based neural network analysis of internal carotid arterial Doppler signals. J. Med. Syst. 2006, 30, 221–229. [Google Scholar] [CrossRef] [PubMed]
- Samiappan, D.; Chakrapani, V. Classification of carotid artery abnormalities in ultrasound images using an artificial neural classifier. Int. Arab J. Inf. Technol. 2016, 13, 756–762. [Google Scholar]
- Uğuz, H. Detection of carotid artery disease by using learning vector quantization neural network. J. Med. Syst. 2012, 36, 533–540. [Google Scholar] [CrossRef] [PubMed]
- Mei, Y.J.; Hu, R.T.; Lin, J.; Xu, H.Y.; Wu, L.Y.; Li, H.P.; Ye, Z.M.; Qin, C. Diagnosis of Middle Cerebral Artery Stenosis Using Transcranial Doppler Images Based on Convolutional Neural Network. World Neurosurg. 2022, 161, e118–e125. [Google Scholar] [CrossRef]
- Seera, M.; Lim, C.P.; Tan, K.S.; Liew, W.S. Classification of transcranial Doppler signals using individual and ensemble recurrent neural networks. Neurocomputing 2017, 249, 337–344. [Google Scholar] [CrossRef]
- Li, Z.; He, W. Stenosis diagnosis based on peripheral arterial and artificial neural network. Netw. Model. Anal. Health Inform. Bioinform. 2021, 10, 1–21. [Google Scholar] [CrossRef]
- Sharma, S.; Lubrica, R.J.; Song, M.; Vandse, R.; Boling, W.; Pillai, P. The role of transcranial Doppler in cerebral vasospasm: A literature review. In Subarachnoid Hemorrhage: Neurological Care and Protection; Springer: Berlin/Heidelberg, Germany, 2020; pp. 201–205. [Google Scholar]
- Podder, K.K.; Chowdhury, M.E.; Al-Maadeed, S.; Nisha, N.N.; Mahmud, S.; Hamadelneil, F.; Almkhlef, T.; Aljofairi, H.; Mushtak, A.; Khandakar, A.; et al. Deep learning-based middle cerebral artery blood flow abnormality detection using flow velocity waveform derived from transcranial Doppler ultrasound. Biomed. Signal Process. Control. 2023, 85, 104882. [Google Scholar] [CrossRef]
- Hossain, S.; Mahmud, S.; Khandakar, A.; Al-Emadi, N.; Chowdhury, F.A.; Bin Mahbub, Z.; Reaz, M.B.I.; Chowdhury, M.E.H. MultiResUNet3+: A Full-Scale Connected Multi-Residual UNet Model to Denoise Electrooculogram and Electromyogram Artifacts from Corrupted Electroencephalogram Signals. Bioengineering 2023, 10, 579. [Google Scholar] [CrossRef]
- Mahmud, S.; Hossain, S.; Chowdhury, M.E.H.; Reaz, M.B.I. MLMRS-Net: Electroencephalography (EEG) motion artifacts removal using a multi-layer multi-resolution spatially pooled 1D signal reconstruction network. Neural Comput. Appl. 2023, 35, 8371–8388. [Google Scholar] [CrossRef]
- Alam, J.; Hassan, M.; Khan, A.; Chaudhry, A. Robust fuzzy RBF network based image segmentation and intelligent decision making system for carotid artery ultrasound images. Neurocomputing 2015, 151, 745–755. [Google Scholar] [CrossRef]
- Shuzan, N.I.; Chowdhury, M.E.; Reaz, M.B.I.; Khandakar, A.; Abir, F.F.; Faisal, A.A.; Ali, S.H.M.; Bakar, A.A.A.; Chowdhury, M.H.; Mahbub, Z.B.; et al. Machine learning-based classification of healthy and impaired gaits using 3D-GRF signals. Biomed. Signal Process. Control. 2023, 81, 104448. [Google Scholar] [CrossRef]
- Rahman, A.; Chowdhury, M.E.H.; Khandakar, A.; Kiranyaz, S.; Zaman, K.S.; Reaz, M.B.I.; Islam, M.T.; Ezeddin, M.; Kadir, M.A. Multimodal EEG and keystroke dynamics based biometric system using machine learning algorithms. IEEE Access 2021, 9, 94625–94643. [Google Scholar] [CrossRef]
- Abir, F.F.; Chowdhury, M.E.; Tapotee, M.I.; Mushtak, A.; Khandakar, A.; Mahmud, S.; Hasan, A. PCovNet+: A CNN-VAE anomaly detection framework with LSTM embeddings for smartwatch-based COVID-19 detection. Eng. Appl. Artif. Intell. 2023, 122, 106130. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, M.H.; Shuzan, N.I.; Chowdhury, M.E.H.; Reaz, M.B.I.; Mahmud, S.; Al Emadi, N.; Ayari, M.A.; Ali, S.H.M.; Bakar, A.A.A.; Rahman, S.M.; et al. Lightweight End-to-End Deep Learning Solution for Estimating the Respiration Rate from Photoplethysmogram Signal. Bioengineering 2022, 9, 558. [Google Scholar] [CrossRef]
- Faisal, A.A.; Chowdhury, M.E.H.; Bin Mahbub, Z.; Pedersen, S.; Ahmed, M.U.; Khandakar, A.; Alhatou, M.; Nabil, M.; Ara, I.; Bhuiyan, E.H.; et al. NDDNet: A deep learning model for predicting neurodegenerative diseases from gait pattern. Appl. Intell. 2023, 1–13. [Google Scholar] [CrossRef]
- Campadelli, P.; Casiraghi, E.; Pratissoli, S. A segmentation framework for abdominal organs from CT scans. Artif. Intell. Med. 2010, 50, 3–11. [Google Scholar] [CrossRef]
- Hassan, M.; Chaudhry, A.; Khan, A.; Kim, J.Y. Carotid artery image segmentation using modified spatial fuzzy c-means and ensemble clustering. Comput. Methods Programs Biomed. 2012, 108, 1261–1276. [Google Scholar] [CrossRef]
- Wright, I.A.; Gough, N.A.; Rakebrandt, F.; Wahab, M.; Woodcock, J.P. Neural network analysis of Doppler ultrasound blood flow signals: A pilot study. Ultrasound Med. Biol. 1997, 23, 683–690. [Google Scholar] [CrossRef]
- Ceylan, M.; Ceylan, R.; Dirgenali, F.; Kara, S.; Özbay, Y. Classification of carotid artery Doppler signals in the early phase of atherosclerosis using complex-valued artificial neural network. Comput. Biol. Med. 2007, 37, 28–36. [Google Scholar] [CrossRef]
- Ceylan, M.; Ceylan, R.; Özbay, Y.; Kara, S. Application of complex discrete wavelet transform in classification of Doppler signals using complex-valued artificial neural network. Artif. Intell. Med. 2008, 44, 65–76. [Google Scholar] [CrossRef]
- Jaishankar, R.; Fanelli, A.; Filippidis, A.; Vu, T.; Holsapple, J.; Heldt, T. A spectral approach to model-based noninvasive intracranial pressure estimation. IEEE J. Biomed. Health Informatics 2019, 24, 2398–2406. [Google Scholar] [CrossRef] [PubMed]
- Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. 1979, 9, 62–66. [Google Scholar] [CrossRef]
- Li, Z.; Liu, F.; Yang, W.; Peng, S.; Zhou, J. A survey of convolutional neural networks: Analysis, applications, and prospects. IEEE Trans. Neural Networks Learn. Syst. 2021, 33, 6999–7019. [Google Scholar] [CrossRef] [PubMed]
- Kiranyaz, S.; Ince, T.; Iosifidis, A.; Gabbouj, M. Operational neural networks. Neural Comput. Appl. 2020, 32, 6645–6668. [Google Scholar] [CrossRef]
- Kiranyaz, S.; Malik, J.; Ben Abdallah, H.; Ince, T.; Iosifidis, A.; Gabbouj, M. Self-organized operational neural networks with generative neurons. Neural Netw. 2021, 140, 294–308. [Google Scholar] [CrossRef]
- Malik, J.; Devecioglu, O.C.; Kiranyaz, S.; Ince, T.; Gabbouj, M. Real-time patient-specific ecg classification by 1d self-operational neural networks. IEEE Trans. Biomed. Eng. 2021, 69, 1788–1801. [Google Scholar] [CrossRef]
- Malik, J.; Kiranyaz, S.; Gabbouj, M. Self-organized operational neural networks for severe image restoration problems. Neural Networks 2021, 135, 201–211. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Linear. PyTorch 1.13 Documentation. 2022. Available online: https://pytorch.org/docs/stable/generated/torch.nn.Linear.html (accessed on 19 January 2023).
- Logsoftmax. PyTorch 1.13 Documentation. 2022. Available online: https://pytorch.org/docs/stable/generated/torch.nn.LogSoftmax.html (accessed on 19 January 2023).
- Kohavi, R. A study of cross-validation and bootstrap for accuracy estimation and model selection. In Proceedings of the Appears in the International Joint Conference on Articial Intelligence (IJCAI), Montreal, QC, Canada, 20–25 August 1995; pp. 1137–1145. [Google Scholar]
- Elzaafarany, K.; Aly, M.H.; Kumar, G.; Nakhmani, A. Cerebral artery vasospasm detection using transcranial doppler signal analysis. J. Ultrasound Med. 2019, 38, 2191–2202. [Google Scholar] [CrossRef]
- Hsu, K.-C.; Lin, C.-H.; Johnson, K.R.; Liu, C.-H.; Chang, T.-Y.; Huang, K.-L.; Fann, Y.-C.; Lee, T.-H. Autodetect extracranial and intracranial artery stenosis by machine learning using ultrasound. Comput. Biol. Med. 2020, 116, 103569. [Google Scholar] [CrossRef]
- Hang, S.; Miao, Y.; Chen, J.; Zhang, X.; Han, L.; Huang, Z.; Pei, N.; Liu, H.; An, C. CCS-Net: Cascade Detection Network with the Convolution Kernel Switch Block and Statistics Optimal Anchors Block in Hypopharyngeal Cancer MRI. IEEE J. Biomed. Health Inform. 2022, 27, 433–444. [Google Scholar]
- Zhang, S.; Miao, Y.; Chen, J.; Zhang, X.; Han, L.; Ran, D.; Huang, Z.; Pei, N.; Liu, H.; An, C. Twist-Net: A multi-modality transfer learning network with the hybrid bilateral encoder for hypopharyngeal cancer segmentation. Comput. Biol. Med. 2023, 154, 106555. [Google Scholar] [CrossRef] [PubMed]

| Healthy | ICU | |
|---|---|---|
| Number of subjects | 6 | 12 |
| Diagnosis | Healthy | Intraparenchymal hemorrhage, hydrocephalus, and traumatic brain injury (TBI) |
| Age group | 25–45 years | 23–74 years |
| Number of recordings | 16 | 30 |
| The total duration of all recordings | 2 h | 2 h 40 min |
| Targeted arteries | MCA | MCA, ICA |
| Data of targeted artery used in this study | MCA | MCA |
| Ultrasound system | Philips CX-50 with a 1.75 MHz transducer (S5-1) | |
| Field of view | 90° | |
| US probe placement | Temporal region and the M1 segment of the MCA | |
| Blow flow direction for MCA | Towards the probe | |
| Blood flow velocity for MCA | Positive | |
| Fold Num | Session ID | |||
|---|---|---|---|---|
| Train (Healthy) | Test (Healthy) | Train (ICU) | Test (ICU) | |
| 1 | Rest | “14–16” | Rest | “1, 2, 5” |
| 2 | Rest | “10–12” | Rest | “24–26” |
| 3 | Rest | “7–10” | Rest | “21–23” |
| 4 | Rest | “4–6” | Rest | “4, 5, 20” |
| 5 | Rest | “1–3” | Rest | “27–29” |
| Training Parameters | Value |
|---|---|
| Number of folds | 5 |
| Batch size | 4 |
| Number of epochs | 100 |
| Learning rate | 0.0001 |
| Epoch patience | 7 |
| Learning factor | 0.2 |
| Loss type | SoftM_MSE |
| Optimization function | Adam |
| Model Name | Overall Accuracy | Precision | Recall | F1 Score | Specificity |
|---|---|---|---|---|---|
| Self-ResNet18_Q1 | 93.46 | 93.48 | 93.46 | 93.46 | 93.39 |
| Self-ResNet18_Q3 | 94.91 | 94.92 | 94.91 | 94.91 | 94.88 |
| Self-ResNet18_Q5 | 93.31 | 93.32 | 93.31 | 93.3 | 93.26 |
| Self- ResAttentioNet18_Q1 | 96.05 | 96.06 | 96.05 | 96.06 | 96.09 |
| Self- ResAttentioNet18_Q3 | 95.31 | 95.34 | 95.31 | 95.31 | 95.37 |
| Self- ResAttentioNet18_Q5 | 92.97 | 93.01 | 92.97 | 92.97 | 93.03 |
| Ref. | Data | Classes | Classification Model | Accuracy | Recall | Specificity |
|---|---|---|---|---|---|---|
| [45] | TCD signal from MCA | Vasospasm & normal | Decision tree | 89.17 | 87.5% | 89.74% |
| [18] | Peripheral pulse wave | Five degrees of stenosis | ANN | 88.7% | - | - |
| [46] | Cerebral ultrasound | Stenosis & normal | SVM | 80.8% to 81.9% | 70.9% to 73.1% | 90.7% to 90.8% |
| [17] | TCD from basal arteries | Stenosis & non-stenosis | RNN | 71.1% to 75.89% | 74.6% to 75.53% | 71.15% to 74.89% |
| Ours | TCD signal from MCA | Healthy & ICU | Self- ResAttentioNet18_Q1 | 96.05% | 96.05% | 96.09% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nisha, N.N.; Podder, K.K.; Chowdhury, M.E.H.; Rabbani, M.; Wadud, M.S.I.; Al-Maadeed, S.; Mahmud, S.; Khandakar, A.; Zughaier, S.M. A Deep Learning Framework for the Detection of Abnormality in Cerebral Blood Flow Velocity Using Transcranial Doppler Ultrasound. Diagnostics 2023, 13, 2000. https://doi.org/10.3390/diagnostics13122000
Nisha NN, Podder KK, Chowdhury MEH, Rabbani M, Wadud MSI, Al-Maadeed S, Mahmud S, Khandakar A, Zughaier SM. A Deep Learning Framework for the Detection of Abnormality in Cerebral Blood Flow Velocity Using Transcranial Doppler Ultrasound. Diagnostics. 2023; 13(12):2000. https://doi.org/10.3390/diagnostics13122000
Chicago/Turabian StyleNisha, Naima Nasrin, Kanchon Kanti Podder, Muhammad E. H. Chowdhury, Mamun Rabbani, Md. Sharjis Ibne Wadud, Somaya Al-Maadeed, Sakib Mahmud, Amith Khandakar, and Susu M. Zughaier. 2023. "A Deep Learning Framework for the Detection of Abnormality in Cerebral Blood Flow Velocity Using Transcranial Doppler Ultrasound" Diagnostics 13, no. 12: 2000. https://doi.org/10.3390/diagnostics13122000
APA StyleNisha, N. N., Podder, K. K., Chowdhury, M. E. H., Rabbani, M., Wadud, M. S. I., Al-Maadeed, S., Mahmud, S., Khandakar, A., & Zughaier, S. M. (2023). A Deep Learning Framework for the Detection of Abnormality in Cerebral Blood Flow Velocity Using Transcranial Doppler Ultrasound. Diagnostics, 13(12), 2000. https://doi.org/10.3390/diagnostics13122000









