Methylation of the NT5E Gene Is Associated with Poor Prognostic Factors in Breast Cancer
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients and Materials
2.2. DNA Extraction and Sodium Bisulfite Treatment
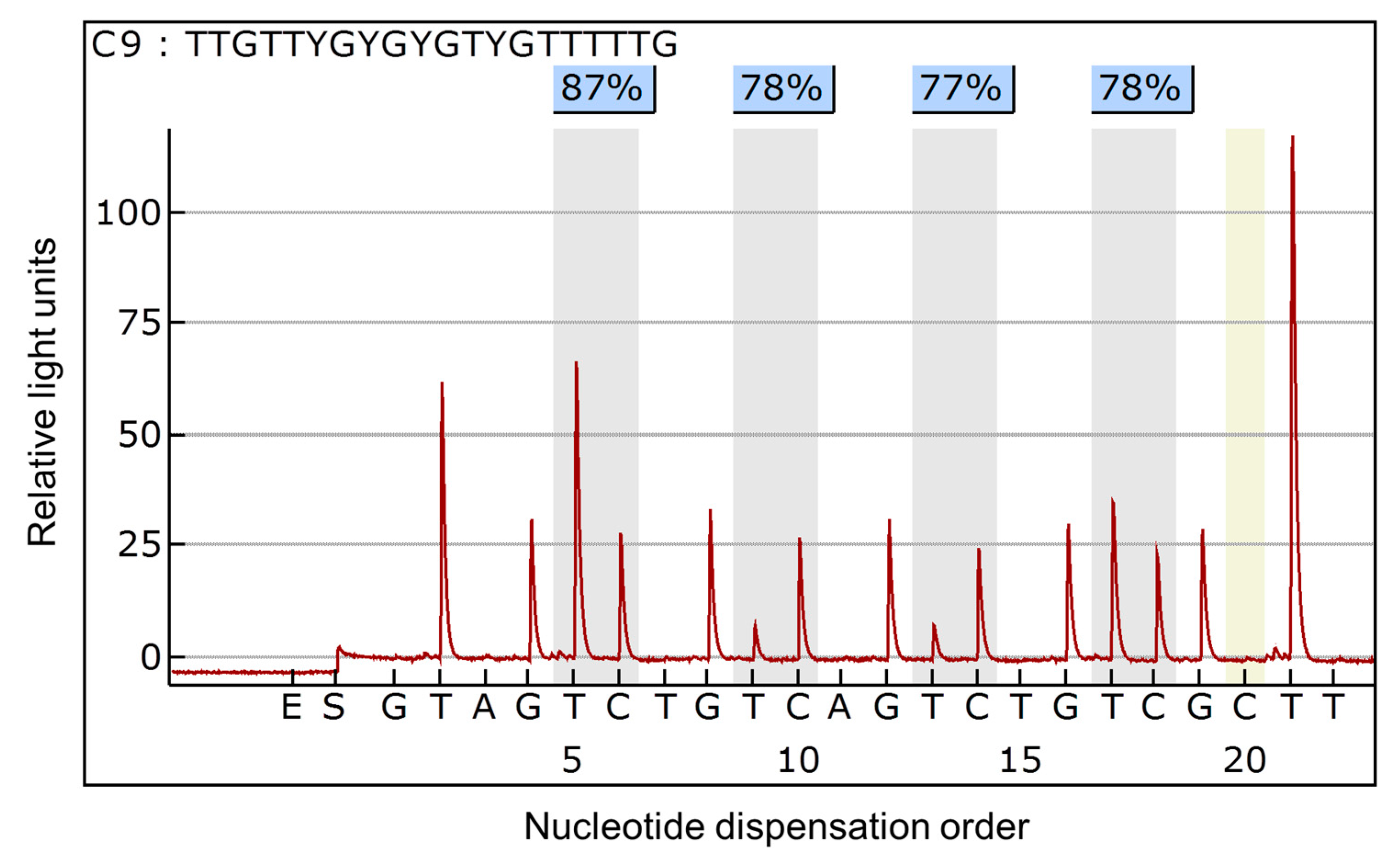
2.3. Methylation Analysis
2.4. Analysis of Inflammatory Markers
2.5. Statistical Analysis
3. Results
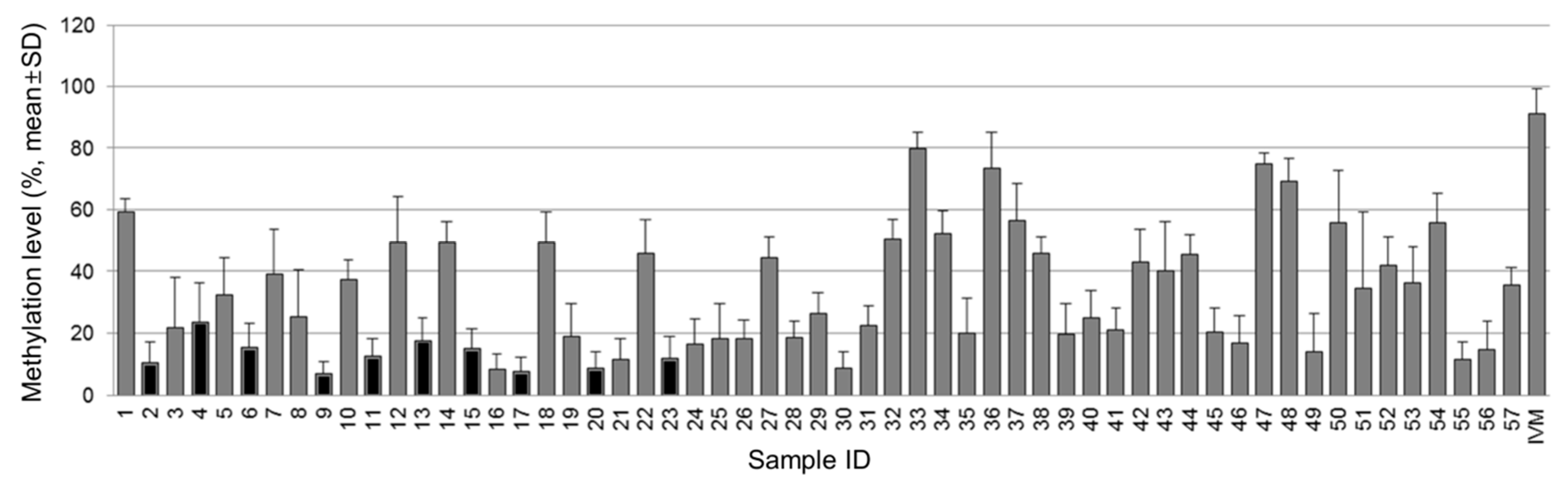
3.1. Methylation Status of the NT5E Gene in Breast Cancer
3.2. Association between the Methylation Levels of the NT5E Gene and Inflammatory Markers in Tumor Tissues
3.3. Association between the Methylation Levels of the NT5E Gene and the Clinicopathologic Characteristics
4. Discussion
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| CD | Cluster of differentiation |
| NT5E | ecto-5′-nucleotidase |
| AMP | adenosine monophosphate |
| HIF | Hypoxia inducible factor |
| DNA | Deoxyribonucleic acid |
| TGF | Transformaing growth factor |
| TNF | Tumor necrosis factor |
| IL | Interleukin |
| NF-κB | Nuclear factor-κB |
| ER | Estrogen receptor |
| Bcl-2 | B-cell lymphoma |
| DFS | Disease-free survival |
| OS | Overall survival |
| HPV | Human papilloma virus |
| PCR | Polymerase chain reaction |
| FFPE | Formalin fixed and paraffin embedded |
| PR | Progesterone receptor |
| HER2 | Human epidermal growth factor receptor 2 |
| TMA | Tissue microarray |
| RNA | Ribonucleic acid |
| RT-PCR | Reverse transcriptase-polymerase chain reaction |
References
- Zimmermann, H.; Zebisch, M.; Sträter, N. Cellular function and molecular structure of ecto-nucleotidases. Purinergic Signal. 2012, 8, 437–502. [Google Scholar] [CrossRef]
- Stagg, J.; Smyth, M.J. Extracellular adenosine triphosphate and adenosine in cancer. Oncogene 2010, 29, 5346–5358. [Google Scholar] [CrossRef] [PubMed]
- Allard, D.; Turcotte, M.; Stagg, J. Targeting A2 adenosine receptors in cancer. Immunol. Cell Biol. 2017, 95, 333–339. [Google Scholar] [CrossRef] [PubMed]
- Antonioli, L.; Blandizzi, C.; Malavasi, F.; Ferrari, D.; Haskó, G. Anti-CD73 immunotherapy: A viable way to reprogram the tumor microenvironment. Oncoimmunology 2016, 5, e1216292. [Google Scholar] [CrossRef] [PubMed]
- Allard, D.; Allard, B.; Gaudreau, P.O.; Chrobak, P.; Stagg, J. CD73-adenosine: A nextgeneration target in immuno-oncology. Immunotherapy 2016, 8, 145–163. [Google Scholar] [CrossRef]
- Allard, B.; Longhi, M.S.; Robson, S.C.; Stagg, J. The ectonucleotidases CD39 and CD73: Novel checkpoint inhibitor targets. Immunol. Rev. 2017, 276, 121–144. [Google Scholar] [CrossRef]
- Allard, D.; Chrobak, P.; Allard, B.; Messaoudi, N.; Stagg, J. Targeting the CD73-adenosine axis in immune-oncology. Immunol. Lett. 2019, 205, 31–39. [Google Scholar] [CrossRef]
- Antonioli, L.; Yegutkin, G.G.; Pacher, P.; Blandizzi, C.; Haskó, G. Anti-CD73 in cancer immunotherapy: Awakening new opportunities. Trends Cancer 2016, 2, 95–109. [Google Scholar] [CrossRef]
- Jiang, T.; Xu, X.; Qiao, M.; Li, X.; Zhao, C.; Zhou, F.; Gao, G.; Wu, F.; Chen, X.; Su, C.; et al. Comprehensive evaluation of NT5E/CD73 expression and its prognostic significance in distinct types of cancers. BMC Cancer 2018, 18, 267. [Google Scholar] [CrossRef]
- Supernat, A.; Markiewicz, A.; Welnicka-Jaskiewicz, M.; Seroczynska, B.; Skokowski, J.; Sejda, A.; Szade, J.; Czapiewski, P.; Biernat, W.; Żaczek, A. CD73 expression as a potential marker of good prognosis in breast carcinoma. Appl. Immunohistochem. Mol. Morphol. 2012, 20, 103–107. [Google Scholar] [CrossRef]
- Sitkovsky, M.V.; Hatfield, S.; Abbott, R.; Belikoff, B.; Lukashev, D.; Ohta, A. Hostile, hypoxia-A2-adenosinergic tumor biology as the next barrier to overcome for tumor immunologists. Cancer Immunol. Res. 2014, 2, 598–605. [Google Scholar] [CrossRef] [PubMed]
- Antonioli, L.; Pacher, P.; Vizi, E.S.; Haskó, G. CD39 and CD73 in immunity and inflammation. Trends Mol. Med. 2013, 19, 355–367. [Google Scholar] [CrossRef] [PubMed]
- Kiss, J.; Yegutkin, G.G.; Koskinen, K.; Savunen, T.; Jalkanen, S.; Salmi, M. IFN-beta protects from vascular leakage via up-regulation of CD73. Eur. J. Immunol. 2007, 37, 3334–3338. [Google Scholar] [CrossRef] [PubMed]
- Kanwal, R.; Gupta, S. Epigenetic modifications in cancer. Clin. Genet. 2012, 81, 303–311. [Google Scholar] [CrossRef]
- Cheng, Y.; He, C.; Wang, M.; Ma, X.; Mo, F.; Yang, S.; Han, J.; Wei, X. Targeting epigenetic regulators for cancer therapy: Mechanisms and advances in clinical trials. Signal Transduct. Targeted Ther. 2019, 4, 62. [Google Scholar] [CrossRef]
- Marzese, D.M.; Hoon, D.S. Emerging technologies for studying DNA methylation for the molecular diagnosis of cancer. Expert Rev. Mol. Diagn. 2015, 15, 647–664. [Google Scholar] [CrossRef]
- de Almeida, B.P.; Apolónio, J.D.; Binnie, A.; Castelo-Branco, P. Roadmap of DNA methylation in breast cancer identifies novel prognostic biomarkers. BMC Cancer 2019, 19, 219. [Google Scholar] [CrossRef]
- Nigro, C.L.; Monteverde, M.; Lee, S.; Lattanzio, L.; Vivenza, D.; Comino, A.; Syed, N.; McHugh, A.; Wang, H.; Proby, C.; et al. NT5E CpG island methylation is a favourable breast cancer biomarker. Br. J. Cancer 2012, 107, 75–83. [Google Scholar] [CrossRef]
- Wang, H.; Lee, S.; Nigro, C.L.; Lattanzio, L.; Merlano, M.; Monteverde, M.; Matin, R.; Purdie, K.; Mladkova, N.; Bergamaschi, D.; et al. NT5E (CD73) is epigenetically regulated in malignant melanoma and associated with metastatic site specificity. Br. J. Cancer 2012, 106, 1446–1452. [Google Scholar] [CrossRef]
- Vogt, T.J.; Genvensleben, H.; Dietrich, J.; Kristiansen, G.; Bootz, F.; Landsberg, J.; Goltz, D.; Dietrich, D. Detailed analysis of adenosine A2a receptor (ADORA2A) and CD73 (5′-nucleodiease, ecto, NT5E) methylation and gene expression in head and neck squamous cell carcinoma patients. Oncoimmunology 2018, 7, e1452579. [Google Scholar] [CrossRef]
- Jeong, Y.J.; Jeong, H.Y.; Bong, J.G.; Park, S.H.; Oh, H.K. Low methylation levels of the SFRP1 gene are associated with the basal-like subtype of breast cancer. Oncol. Rep. 2013, 29, 1946–1954. [Google Scholar] [CrossRef] [PubMed]
- Jeong, Y.J.; Oh, H.K.; Park, S.H.; Bong, J.G. Association between inflammation and cancer cell phenotype in breast cancer. Oncol. Lett. 2018, 15, 2380–2386. [Google Scholar] [CrossRef] [PubMed]
- Clayton, A.; Al-Taei, S.; Webber, J.; Mason, M.D.; Tab, Z. Cancer exosomes express CD39 and CD73, which suppress T cells through adenosine production. J. Immunol. 2011, 187, 676–683. [Google Scholar] [CrossRef] [PubMed]
- Nigro, C.L.; Lattanzio, L.; Monteverde, M.; Syed, N.; Thompson, A.M.; Garrone, O.; Cavicchioli; Tonissi, F.; Comino, A.; McHugh, A.; et al. The frequency of methylation of the NT5E gene in metastatic breast cancer. J. Clin. Oncol. 2011, 29 (Suppl. 15), e11553. [Google Scholar] [CrossRef]
- Hwang, K.T.; Kim, K.; Chang, J.H.; Oh, S.; Kim, Y.A.; Lee, J.Y.; Jung, S.H.; Choi, I.S. BCL2 regulation according to molecular subtype of breast cancer by analysis of The Cancer Genome Atlas Database. Cancer Res. Treat. 2018, 50, 658–669. [Google Scholar] [CrossRef]
- Cheng, Y.Y.; Jin, H.C.; Chan, M.W.Y.; Chu, W.K.; Gryscg, M. Epigenetic biomarkers in cancer. Dis. Markers 2018, 2018, 4987103. [Google Scholar] [CrossRef]
- Kamińska, K.; Nalejska, E.; Kubiak, M.; Wojtysiak, J.; Żołna, Ł.; Kowalewski, J.; Lewandowska, M.A. Prognostic and Predictive Epigenetic Biomarkers in Oncology. Mol. Diagn. Ther. 2019, 23, 83–95. [Google Scholar] [CrossRef]
| Inflammatory Markers | NT5E Gene Methylation | ||
|---|---|---|---|
| Mean Levels (%) | p-Value | ||
| TNF-α | Negative | 35.78 ± 21.28 | 0.996 |
| Positive | 35.84 ± 18.03 | ||
| IL-4 | Negative | 41.63 ± 16.36 | 0.478 |
| Positive | 36.50 ± 21.36 | ||
| NF-κB p50 | Negative | 27.75 ± 18.08 | 0.455 |
| Positive | 34.18 ± 17.53 | ||
| CD4 | Negative | 33.35 ± 17.94 | 0.319 |
| Positive | 39.96 ± 20.17 | ||
| CD8 | Negative | 33.65 ± 15.45 | 0.765 |
| Positive | 36.17 ± 19.57 | ||
| CD68 | Negative | 33.30 ± 14.84 | 0.412 |
| Positive | 37.92 ± 21.81 | ||
| Intratumoral inflammation | Negative | 26.65 ± 16.74 | 0.115 |
| Positive | 31.16 ± 17.07 | ||
| Peritumoral inflammation | Negative | 22.62 ± 15.69 | 0.073 |
| Positive | 37.92 ± 18.99 | ||
| Clinicopathologic Characteristics | NT5E Gene Methylation | ||
|---|---|---|---|
| Mean Levels (%) | p-Value | ||
| Age (years) | <50 | 39.4 ± 17.88 | 0.312 |
| ≥50 | 33.63 ± 19.58 | ||
| Menopausal state | Pre-menopausal | 43.00 ± 19.73 | 0.031 * |
| Post-menopausal | 30.79 ± 16.31 | ||
| Stage | I | 31.77 ± 3.55 | 0.4 |
| II | 36.83 ± 21.02 | ||
| III | 36.76 ± 21.10 | ||
| IV | 59.53 ± 13.89 | ||
| Tumor size (cm) | ≤2 | 30.53 ± 16.60 | 0.024 * |
| >2 | 43.03 ± 19.97 | ||
| Nodal involvement | Negative | 35.10 ± 17.82 | 0.735 |
| Positive | 37.06 ± 21.16 | ||
| Distant metastasis | Negative | 34.80 ± 18.57 | 0.071 |
| Positive | 59.53 ± 13.88 | ||
| Histologic grade | I | 23.42 ± 8.15 | 0.01 * |
| II | 28.62 ± 15.75 | ||
| III | 42.75 ± 19.81 | ||
| Lymphovascular Invasion | Negative | 36.60 ± 16.06 | 0.758 |
| Positive | 34.85 ± 22.71 | ||
| ER status | Negative | 46.09 ± 17.97 | 0.01 * |
| Positive | 31.06 ± 17.70 | ||
| PR status | Negative | 44.25 ± 16.45 | 0.116 |
| Positive | 33.58 ± 19.16 | ||
| HER2 overexpression | Negative | 36.18 ± 21.42 | 0.909 |
| Positive | 36.90 ± 17.17 | ||
| Bcl-2 | Negative | 53.71 ± 19.98 | 0.002 * |
| Positive | 32.19 ± 16.76 | ||
| Molecular subtype | Luminal A | 24.01 ± 13.45 | 0.099 |
| Luminal B | 38.59 ± 20.56 | ||
| HER2 | 47.08 ± 18.21 | ||
| Basal-like | 36.75 ± 16.29 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jeong, Y.J.; Oh, H.K.; Choi, H.R.; Park, S.H. Methylation of the NT5E Gene Is Associated with Poor Prognostic Factors in Breast Cancer. Diagnostics 2020, 10, 939. https://doi.org/10.3390/diagnostics10110939
Jeong YJ, Oh HK, Choi HR, Park SH. Methylation of the NT5E Gene Is Associated with Poor Prognostic Factors in Breast Cancer. Diagnostics. 2020; 10(11):939. https://doi.org/10.3390/diagnostics10110939
Chicago/Turabian StyleJeong, Young Ju, Hoon Kyu Oh, Hye Ryeon Choi, and Sung Hwan Park. 2020. "Methylation of the NT5E Gene Is Associated with Poor Prognostic Factors in Breast Cancer" Diagnostics 10, no. 11: 939. https://doi.org/10.3390/diagnostics10110939
APA StyleJeong, Y. J., Oh, H. K., Choi, H. R., & Park, S. H. (2020). Methylation of the NT5E Gene Is Associated with Poor Prognostic Factors in Breast Cancer. Diagnostics, 10(11), 939. https://doi.org/10.3390/diagnostics10110939





