Recent Advances towards the Understanding of Secondary Acute Myeloid Leukemia Progression
Abstract
1. Introduction
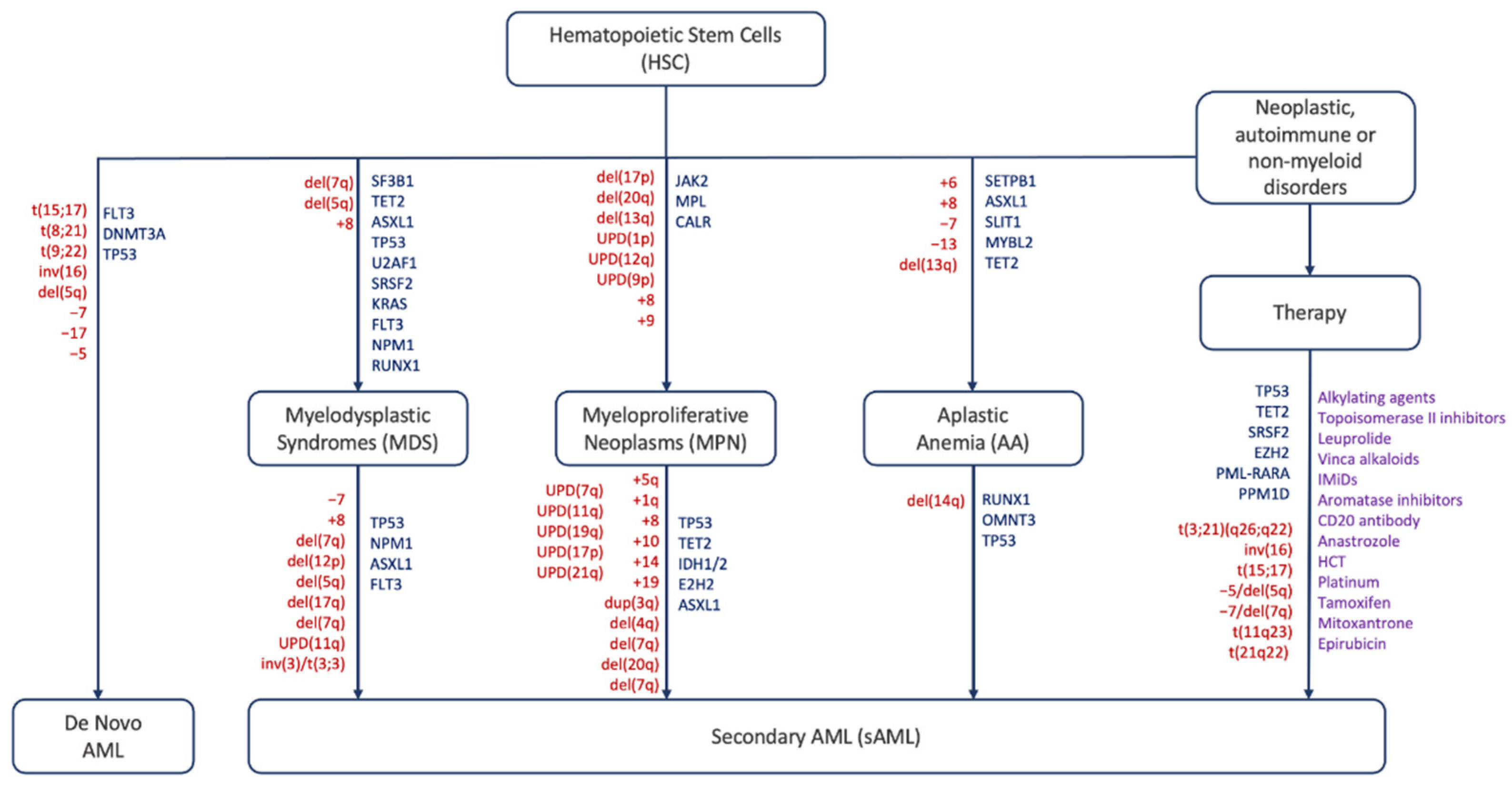
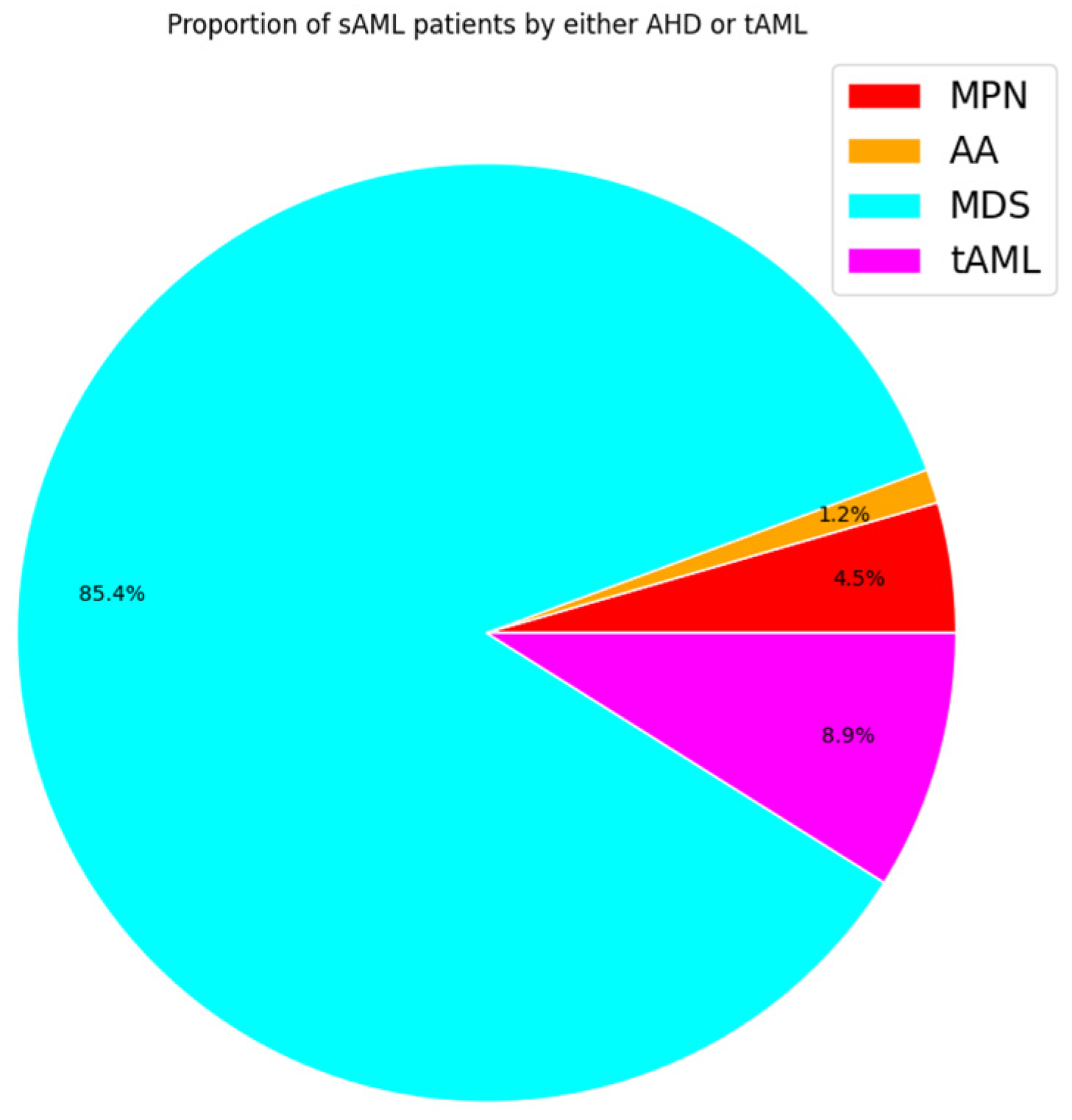
2. Pathophysiology of sAML
3. Cytogenetics and Mutational Landscape of sAML
| Category of Genes | Examples | Citations |
|---|---|---|
| Spliceosome | SRSF2, U2AF1, SF3B1 | [86,87] |
| DNA Methylation | DMNT3A, TET 1/2, IDH 1/2, | [88,89] |
| Activated Signaling | CALR, JAK2, PTPN11, TpoR, KRAS, FLT3, NRAS | [90,91] |
| Transcription Factors | RUNX1, NFE2, TP53 | [92,93,94,95,96] |
| Chromatin Modification | EZH2, ASXL1, NPM1 | [89,97] |
| Type of Chromosomal Abnormality | Examples | Citations |
|---|---|---|
| Deletions | del(7q), del(5q), del(17p) | [106,113] |
| Duplications | dup(1q), dup(3q), dup(11q), dup(17q) | [114,115,116] |
| Translocations | t(1;11)(q21;p15), t(10;11)(q22;q23), t(8;21) | [65,117,118] |
| Inversions | inv(3)/t(3;3) | [24] |
| Monosomy | −7 | [87,119] |
| Trisomy | +8, +19, +21 | [59,87,120] |
| Uniparental disomy | UPD(9p), UPD(1p), UPD (17p) | [75,92,121] |
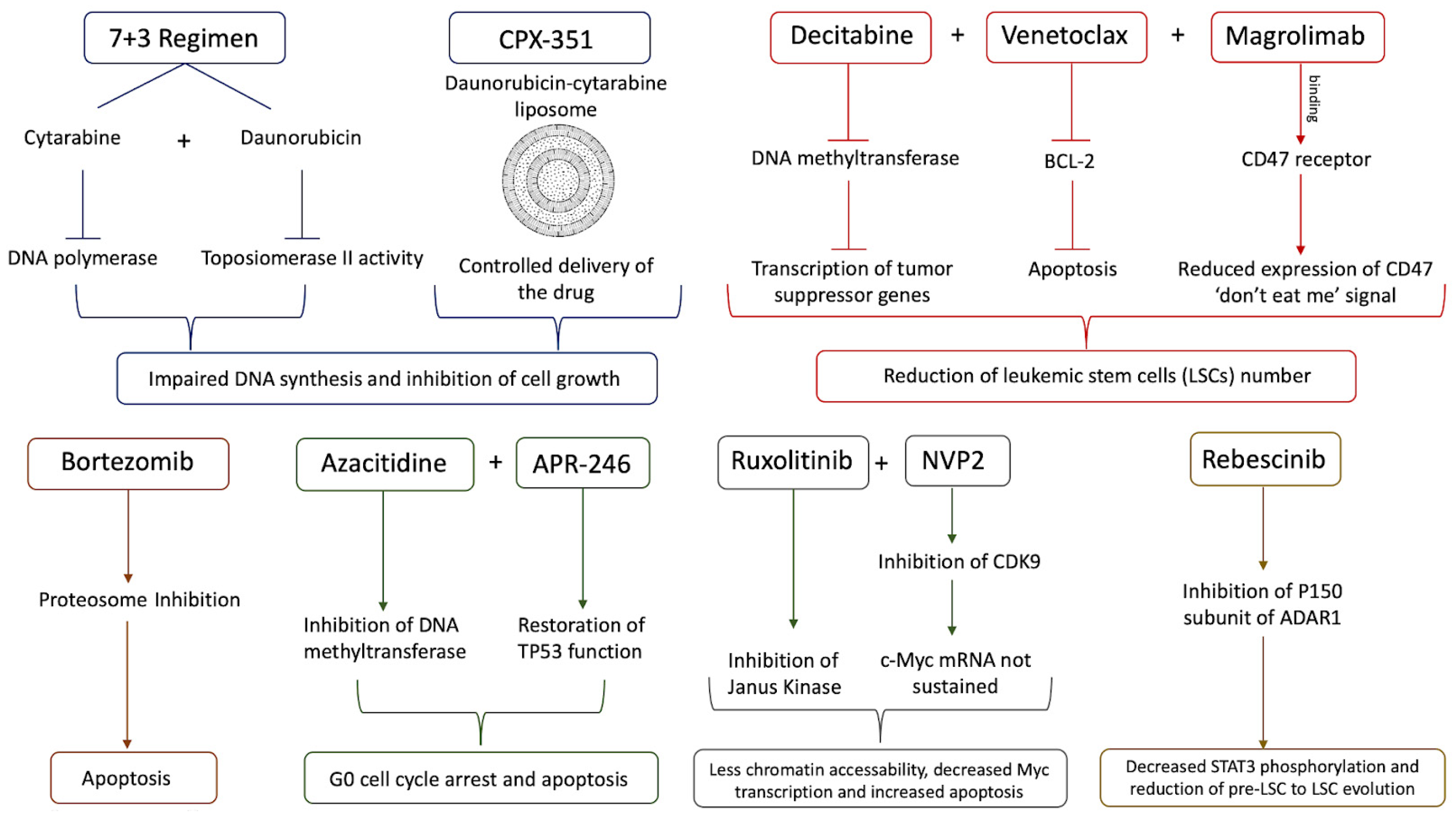
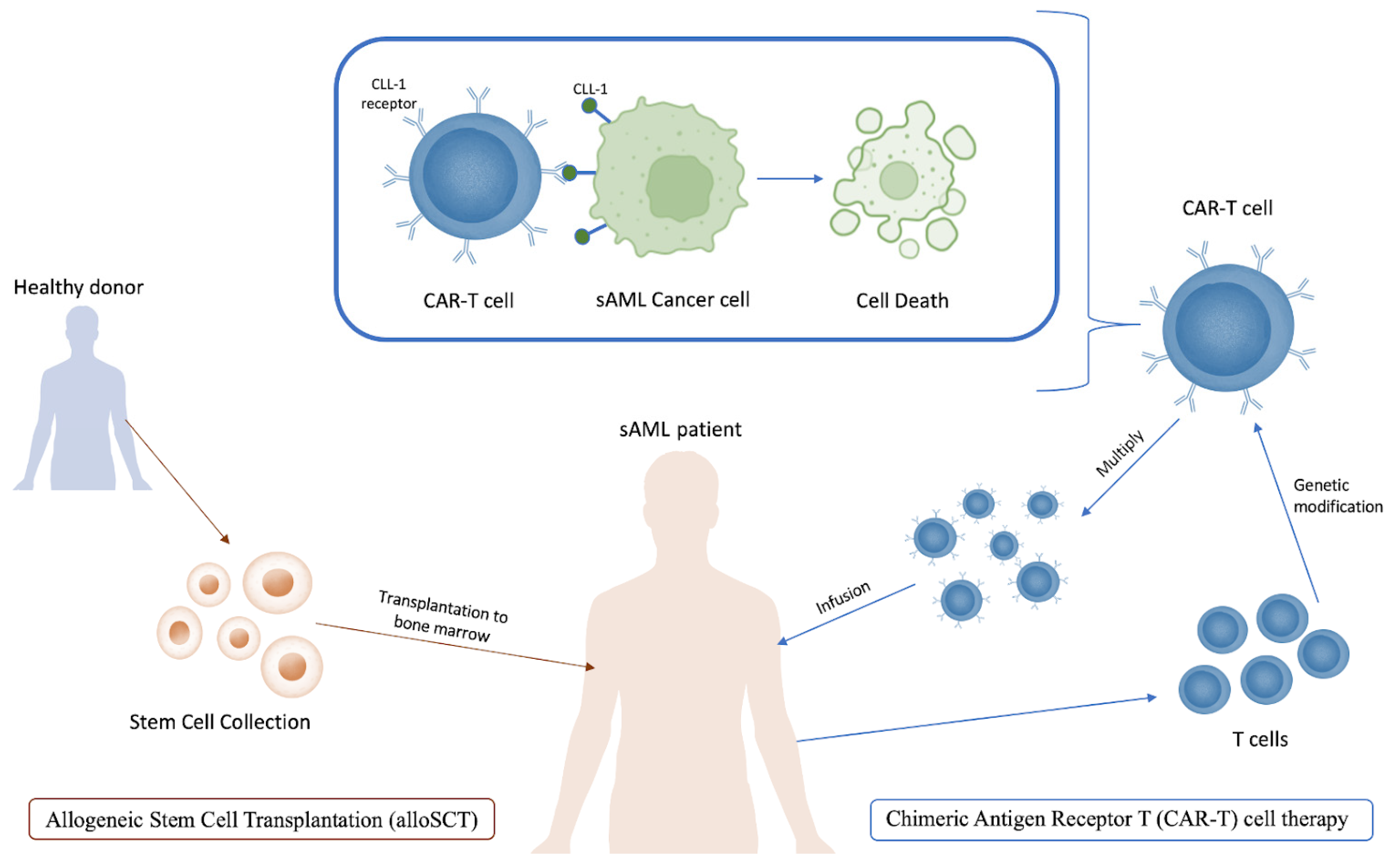
4. Treatment of sAML
5. Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Estey, E.; Döhner, H. Acute myeloid leukaemia. Lancet 2006, 368, 1894–1907. [Google Scholar] [CrossRef]
- Higgins, A.; Shah, M.V. Genetic and genomic landscape of secondary and therapy-related acute myeloid leukemia. Genes 2020, 11, 749. [Google Scholar] [CrossRef]
- Pedersen-Bjergaard, J.; Andersen, M.K.; Andersen, M.T.; Christiansen, D.H. Genetics of therapy-related myelodysplasia and acute myeloid leukemia. Leukemia 2008, 22, 240–248. [Google Scholar] [CrossRef]
- McMullin, M.F.; Anderson, L.A. Aetiology of Myeloproliferative Neoplasms. Cancers 2020, 12, 1810. [Google Scholar] [CrossRef]
- Shahin, O.A.; Chifotides, H.T.; Bose, P.; Masarova, L.; Verstovsek, S. Accelerated Phase of Myeloproliferative Neoplasms. Acta Haematol. 2021, 144, 484–499. [Google Scholar] [CrossRef]
- Adès, L.; Itzykson, R.; Fenaux, P. Myelodysplastic syndromes. Lancet 2014, 383, 2239–2252. [Google Scholar] [CrossRef] [PubMed]
- Dan, C.; Chi, J.; Wang, L. Molecular mechanisms of the progression of myelodysplastic syndrome to secondary acute myeloid leukaemia and implication for therapy. Ann. Med. 2015, 47, 209–217. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Babushok, D.V. Secondary myelodysplastic syndrome and leukemia in acquired aplastic anemia and paroxysmal nocturnal hemoglobinuria. Blood 2020, 136, 36–49. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Liu, H. Pathogenesis of aplastic anemia. Hematology 2019, 24, 559–566. [Google Scholar] [CrossRef]
- Cook, M.R.; Karp, J.E.; Lai, C. The spectrum of genetic mutations in myelodysplastic syndrome: Should we update prognostication? eJHaem 2022, 3, 301–313. [Google Scholar] [CrossRef] [PubMed]
- Heuser, M.; Gabdoulline, R.; Löffeld, P.; Dobbernack, V.; Kreimeyer, H.; Pankratz, M.; Flintrop, M.; Liebich, A.; Klesse, S.; Panagiota, V.; et al. Individual outcome prediction for myelodysplastic syndrome (MDS) and secondary acute myeloid leukemia from MDS after allogeneic hematopoietic cell transplantation. Ann. Hematol. 2017, 96, 1361–1372. [Google Scholar] [CrossRef]
- Hussein, K.; Van Dyke, D.L.; Tefferi, A. Conventional cytogenetics in myelofibrosis: Literature review and discussion. Eur. J. Haematol. 2009, 82, 329–338. [Google Scholar] [CrossRef]
- Rubin, C.M.; Larson, R.A.; Anastasi, J.; Winter, J.N.; Thangavelu, M.; Vardiman, J.W.; Rowley, J.D.; Le Beau, M.M. t(3;21)(q26;q22): A recurring chromosomal abnormality in therapy- related myelodysplastic syndrome and acute myeloid leukemia. Blood 1990, 76, 2594–2598. [Google Scholar] [CrossRef]
- Andersen, M.K.; Larson, R.A.; Mauritzson, N.; Schnittger, S.; Jhanwar, S.C.; Pedersen-Bjergaard, J. Balanced chromosome abnormalities inv(16) and t(15;17) in therapy-related myelodysplastic syndromes and acute leukemia: Report from an International Workshop †. Genes Chromosom. Cancer 2002, 33, 395–400. [Google Scholar] [CrossRef]
- Kotsiafti, A.; Giannakas, K.; Christoforou, P.; Liapis, K. Progress toward Better Treatment of Therapy-Related AML. Cancers 2023, 15, 1658. [Google Scholar] [CrossRef] [PubMed]
- Hosono, N. Genetic abnormalities and pathophysiology of MDS. Int. J. Clin. Oncol. 2019, 24, 885–892. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Wang, Y.; Wang, H.; Chen, Z.; Lou, J.; Xu, H.; Wang, H.; Qian, W.; Meng, H.; Lin, M.; et al. Cytogenetic profile of de novo acute myeloid leukemia: A study based on 1432 patients in a single institution of China. Leukemia 2009, 23, 1801–1806. [Google Scholar] [CrossRef] [PubMed]
- Heuser, M.; Schlarmann, C.; Dobbernack, V.; Panagiota, V.; Wiehlmann, L.; Walter, C.; Beier, F.; Ziegler, P.; Yun, H.; Kade, S.; et al. Genetic characterization of acquired aplastic anemia by targeted sequencing. Haematologica 2014, 99, e165. [Google Scholar] [CrossRef] [PubMed]
- Milosevic, J.D.; Puda, A.; Malcovati, L.; Berg, T.; Hofbauer, M.; Stukalov, A.; Klampfl, T.; Harutyunyan, A.S.; Gisslinger, H.; Gisslinger, B.; et al. Clinical significance of genetic aberrations in secondary acute myeloid leukemia. Am. J. Hematol. 2012, 87, 1010–1016. [Google Scholar] [CrossRef]
- Maciejewski, J.P.; Risitano, A.; Sloand, E.M.; Nunez, O.; Young, N.S. Distinct clinical outcomes for cytogenetic abnormalities evolving from aplastic anemia. Blood 2002, 99, 3129–3135. [Google Scholar] [CrossRef] [PubMed]
- Strickland, S.A.; Vey, N. Diagnosis and treatment of therapy-related acute myeloid leukemia. Crit. Rev. Oncol. Hematol. 2022, 171, 103607. [Google Scholar] [CrossRef]
- Keung, Y.-K.; Pettenati, M.J.; Cruz, J.M.; Powell, B.L.; Woodruff, R.D.; Buss, D.H. Bone Marrow Cytogenetic Abnormalities of Aplastic Anemia. J. Hematol. 2001, 66, 167–171. [Google Scholar] [CrossRef]
- Van Gelder, M.; Schetelig, J.; Volin, L.; Maertens, J.; Socié, G.; Petersen, E.; Thomssen, H.; Biezen, A.; Brand, R.; Witte, T.M.; et al. Monosomal Karyotype Predicts Poor Outcome for MDS/sAML Patients with Chromosome 7 Abnormalities After Allogeneic Stem Cell Transplantation for MDS/sAML. A Study of the MDS Subcommittee of the Chronic Leukemia Working Party of the European Group for Blood and Marrow Transplantation (EBMT). Blood 2009, 114, 293. [Google Scholar] [CrossRef]
- Birdwell, C.; Fiskus, W.; Kadia, T.M.; Di Nardo, C.D.; Mill, C.P.; Bhalla, K.N. EVI1 dysregulation: Impact on biology and therapy of myeloid malignancies. Blood Cancer J. 2021, 11, 64. [Google Scholar] [CrossRef]
- Marcellino, B.K.; Hoffman, R.; Tripodi, J.; Lu, M.; Kosiorek, H.; Mascarenhas, J.; Rampal, R.K.; Dueck, A.; Najfeld, V. Advanced forms of MPNs are accompanied by chromosomal abnormalities that lead to dysregulation of TP53. Blood Adv. 2018, 2, 3581. [Google Scholar] [CrossRef]
- Thoennissen, N.H.; Kawamata, N.; Lasho, T.L.; Weiss, T.; Nowak, D.; Kato, M.; Takita, J.; Sanada, M.; Haferlach, T.; Mesa, R.A.; et al. Genomic Changes Associated with Leukemic Transformation of Myeloproliferative Disorders. Blood 2008, 112, 3371. [Google Scholar] [CrossRef]
- Venton, G.; Courtier, F.; Charbonnier, A.; D’incan, E.; Saillard, C.; Mohty, B.; Mozziconacci, M.J.; Birnbaum, D.; Murati, A.; Vey, N.; et al. Impact of gene mutations on treatment response and prognosis of acute myeloid leukemia secondary to myeloproliferative neoplasms. Am. J. Hematol. 2018, 93, 330–338. [Google Scholar] [CrossRef]
- Dunbar, A.J.; Rampal, R.K.; Levine, R. Leukemia secondary to myeloproliferative neoplasms. Blood 2020, 136, 61–70. [Google Scholar] [CrossRef]
- Schwind, S.; Jentzsch, M.; Kubasch, A.S.; Metzeler, K.H.; Platzbecker, U. Myelodysplastic syndromes: Biological and therapeutic consequences of the evolving molecular aberrations landscape. Neoplasia 2021, 23, 1101–1109. [Google Scholar] [CrossRef]
- Baxter, E.J.; Scott, L.M.; Campbell, P.J.; East, C.; Fourouclas, N.; Swanton, S.; Vassiliou, G.S.; Bench, A.J.; Boyd, E.M.; Curtin, N.; et al. Acquired mutation of the tyrosine kinase JAK2 in human myeloproliferative disorders. Lancet 2005, 365, 1054–1061. [Google Scholar] [CrossRef]
- James, C.; Ugo, V.; Casadevall, N.; Constantinescu, S.N.; Vainchenker, W. A JAK2 mutation in myeloproliferative disorders: Pathogenesis and therapeutic and scientific prospects. Trends Mol. Med. 2005, 11, 546–554. [Google Scholar] [CrossRef]
- Pardanani, A.D.; Levine, R.L.; Lasho, T.; Pikman, Y.; Mesa, R.A.; Wadleigh, M.; Steensma, D.P.; Elliott, M.A.; Wolanskyj, A.P.; Hogan, W.J.; et al. MPL515 mutations in myeloproliferative and other myeloid disorders: A study of 1182 patients. Blood 2006, 108, 3472–3476. [Google Scholar] [CrossRef]
- Pikman, Y.; Lee, B.H.; Mercher, T.; Mcdowell, E.; Ebert, B.L.; Gozo, M.; Cuker, A.; Wernig, G.; Moore, S.; Galinsky, I.; et al. MPLW515L Is a Novel Somatic Activating Mutation in Myelofibrosis with Myeloid Metaplasia. PLoS Med. 2006, 3, e270. [Google Scholar] [CrossRef]
- Nangalia, J.; Massie, C.E.; Baxter, E.J.; Nice, F.L.; Gundem, G.; Wedge, D.C.; Avezov, E.; Li, J.; Kollmann, K.; Kent, D.G.; et al. Somatic CALR Mutations in Myeloproliferative Neoplasms with Nonmutated JAK2. N. Engl. J. Med. 2013, 369, 2391–2405. [Google Scholar] [CrossRef]
- Klampfl, T.; Gisslinger, H.; Harutyunyan, A.S.; Nivarthi, H.; Rumi, E.; Milosevic, J.D.; Them, N.C.C.; Berg, T.; Gisslinger, B.; Pietra, D.; et al. Somatic Mutations of Calreticulin in Myeloproliferative Neoplasms. N. Engl. J. Med. 2013, 369, 2379–2390. [Google Scholar] [CrossRef]
- Arber, D.A.; Orazi, A.; Hasserjian, R.; Thiele, J.; Borowitz, M.J.; Le Beau, M.M.; Bloomfield, C.D.; Cazzola, M.; Vardiman, J.W. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood 2016, 127, 2391–2405. [Google Scholar] [CrossRef]
- Mesa, R.A.; Verstovsek, S.; Cervantes, F.; Barosi, G.; Reilly, J.T.; Dupriez, B.; Levine, R.; Le Bousse-Kerdiles, M.C.; Wadleigh, M.; Campbell, P.J.; et al. Primary myelofibrosis (PMF), post polycythemia vera myelofibrosis (post-PV MF), post essential thrombocythemia myelofibrosis (post-ET MF), blast phase PMF (PMF-BP): Consensus on terminology by the international working group for myelofibrosis research and. Leuk. Res. 2007, 31, 737–740. [Google Scholar] [CrossRef]
- Jilg, S.; Reidel, V.; Müller-Thomas, C.; König, J.; Schauwecker, J.; Höckendorf, U.; Huberle, C.; Gorka, O.; Schmidt, B.; Burgkart, R.; et al. Blockade of BCL-2 proteins efficiently induces apoptosis in progenitor cells of high-risk myelodysplastic syndromes patients. Leukemia 2016, 30, 112–123. [Google Scholar] [CrossRef]
- Saenz, D.T.; Fiskus, W.; Manshouri, T.; Mill, C.P.; Qian, Y.; Raina, K.; Rajapakshe, K.; Coarfa, C.; Soldi, R.; Bose, P.; et al. Targeting nuclear β-catenin as therapy for post-myeloproliferative neoplasm secondary AML. Leukemia 2018, 33, 1373–1386. [Google Scholar] [CrossRef]
- Parker, J.E.; Mufti, G.J.; Rasool, F.; Mijovic, A.; Devereux, S.; Pagliuca, A. The role of apoptosis, proliferation, and the Bcl-2–related proteins in the myelodysplastic syndromes and acute myeloid leukemia secondary to MDS. Blood 2000, 96, 3932–3938. [Google Scholar] [CrossRef]
- Cogle, C.R. Incidence and Burden of the Myelodysplastic Syndromes. Curr. Hematol. Malig. Rep. 2015, 10, 272. [Google Scholar] [CrossRef]
- Vaht, K.; Göransson, M.; Carlson, K.; Isaksson, C.; Lenhoff, S.; Sandstedt, A.; Uggla, B.; Winiarski, J.; Ljungman, P.; Brune, M.; et al. Incidence and outcome of acquired aplastic anemia: Real-world data from patients diagnosed in Sweden from 2000–2011. Haematologica 2017, 102, 1683. [Google Scholar] [CrossRef]
- Prébet, T.; Gore, S.D.; Thépot, S.; Esterni, B.; Quesnel, B.; Beyne Rauzy, O.; Dreyfus, F.; Gardin, C.; Fenaux, P.; Vey, N. Outcome of acute myeloid leukaemia following myelodysplastic syndrome after azacitidine treatment failure. Br. J. Haematol. 2012, 157, 764. [Google Scholar] [CrossRef]
- Nilsson, C.; Linde, F.; Hulegårdh, E.; Garelius, H.; Lazarevic, V.; Antunovic, P.; Cammenga, J.; Deneberg, S.; Eriksson, A.; Jädersten, M.; et al. Characterization of therapy-related acute myeloid leukemia: Increasing incidence and prognostic implications. Haematologica 2023, 108, 1015. [Google Scholar] [CrossRef]
- Yanada, M.; Yamamoto, Y.; Iba, S.; Okamoto, A.; Inaguma, Y.; Tokuda, M.; Morishima, S.; Kanie, T.; Mizuta, S.; Akatsuka, Y.; et al. TP53 mutations in older adults with acute myeloid leukemia. Int. J. Hematol. 2016, 103, 429–435. [Google Scholar] [CrossRef]
- Michelis, F.V.; Atenafu, E.G.; Gupta, V.; Kim, D.D.; Kuruvilla, J.; Lipton, J.H.; Loach, D.; Seftel, M.D.; Uhm, J.; Alam, N.; et al. Comparable outcomes post allogeneic hematopoietic cell transplant for patients with de novo or secondary acute myeloid leukemia in first remission. Bone Marrow Transplant. 2015, 50, 907–913. [Google Scholar] [CrossRef]
- Rowe, J.M. The “7 + 3” regimen in acute myeloid leukemia. Haematologica 2022, 107, 3. [Google Scholar] [CrossRef]
- Hulegårdh, E.; Nilsson, C.; Lazarevic, V.; Garelius, H.; Antunovic, P.; Rangert Derolf, Å.; Möllgård, L.; Uggla, B.; Wennström, L.; Wahlin, A.; et al. Characterization and prognostic features of secondary acute myeloid leukemia in a population-based setting: A report from the Swedish Acute Leukemia Registry. Am. J. Hematol. 2015, 90, 208–214. [Google Scholar] [CrossRef] [PubMed]
- Østgård, L.S.G.; Medeiros, B.C.; Sengeløv, H.; Nørgaard, M.; Andersen, M.K.; Dufva, I.; Friis, L.S.; Kjeldsen, E.; Marcher, C.W.; Preiss, B.; et al. Epidemiology and clinical significance of secondary and therapy-related acute myeloid leukemia: A national population-based cohort study. J. Clin. Oncol. 2015, 33, 3641–3649. [Google Scholar] [CrossRef]
- Estey, E.; Hasserjian, R.P.; Döhner, H. Distinguishing AML from MDS: A fixed blast percentage may no longer be optimal. Blood 2022, 139, 323–332. [Google Scholar] [CrossRef]
- Bauer, M.; Vaxevanis, C.; Al-Ali, K.; Jaekel, N.; Le, C.; Naumann, H.; Schaffrath, J.; Rau, A.; Seliger, B.; Wickenhauser, C. Altered Spatial Composition of the Immune Cell Repertoire in Association to CD34 + Blasts in Myelodysplastic Syndromes and Secondary Acute Myeloid Leukemia. Cancers 2021, 13, 186. [Google Scholar] [CrossRef]
- Hasserjian, R.P.; Steensma, D.P.; Graubert, T.A.; Ebert, B.L. Clonal hematopoiesis and measurable residual disease assessment in acute myeloid leukemia. Blood 2020, 135, 1729–1738. [Google Scholar] [CrossRef]
- Sebert, M.; Phanie Gachet, S.; Leblanc, T.; Bluteau, D.; Gis Peffault De Latour, R.; Correspondence, J.S. Clonal hematopoiesis driven by chromosome 1q/MDM4 trisomy defines a canonical route toward leukemia in Fanconi anemia. Cell Stem Cell 2023, 30, 153–170.e9. [Google Scholar] [CrossRef]
- Malcovati, L.; Gallì, A.; Travaglino, E.; Ambaglio, I.; Rizzo, E.; Molteni, E.; Elena, C.; Ferretti, V.V.; Catricalà, S.; Bono, E.; et al. Clinical significance of somatic mutation in unexplained blood cytopenia. Blood 2017, 129, 3371–3378. [Google Scholar] [CrossRef] [PubMed]
- Weeks, L.D.; Niroula, A.; Neuberg, D.; Wong, W.; Lindsley, R.C.; Luskin, M.R.; Berliner, N.; Stone, R.M.; DeAngelo, D.J.; Soiffer, R.J.; et al. Prediction of Risk for Myeloid Malignancy in Clonal Hematopoiesis. NEJM Evid. 2023, 2, EVIDoa2200310. [Google Scholar] [CrossRef]
- Vucinic, V.; Ruhnke, L.; Sockel, K.; Rohnert, M.A.; Backhaus, D.; Brauer, D.; Franke, G.N.; Niederwieser, D.; Bornhauser, M.; Rollig, C.; et al. The diagnostic red blood cell distribution width as a prognostic factor in acute myeloid leukemia. Blood Adv. 2021, 5, 5584–5587. [Google Scholar] [CrossRef]
- Greenberg, P.L.; Tuechler, H.; Schanz, J.; Sanz, G.; Garcia-Manero, G.; Solé, F.; Bennett, J.M.; Bowen, D.; Fenaux, P.; Dreyfus, F.; et al. Revised international prognostic scoring system for myelodysplastic syndromes. Blood 2012, 120, 2454–2465. [Google Scholar] [CrossRef]
- Schanz, J.; Tüchler, H.; Solé, F.; Mallo, M.; Luño, E.; Cervera, J.; Granada, I.; Hildebrandt, B.; Slovak, M.L.; Ohyashiki, K.; et al. New comprehensive cytogenetic scoring system for primary myelodysplastic syndromes (MDS) and oligoblastic acute myeloid leukemia after MDS derived from an international database merge. J. Clin. Oncol. 2012, 30, 820–829. [Google Scholar] [CrossRef]
- O’Hagan Henderson, S.; Glaser, A.; Frietsch, J.J.; Hochhaus, A.; Hilgendorf, I. The incidental discovery of a constitutional trisomy 21 mosaicism in an adult female with myelodysplastic/myeloproliferative neoplasm. Ann. Hematol. 2022, 101, 919. [Google Scholar] [CrossRef]
- Begna, K.H.; Ali, W.; Naseema Gangat Elliott, M.A.; Al-Kali, A.; Litzow, M.R.; Christopher Hook, C.; Wolanskyj-Spinner, A.P.; Hogan, W.J.; Patnaik, M.M.; Pardanani, A.; et al. Mayo Clinic experience with 1123 adults with acute myeloid leukemia. Blood Cancer J. 2021, 113, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Gupta, V.; Kim, S.; Hu, Z.H.; Liu, Y.; Aljurf, M.; Bacher, U.; Beitinjaneh, A.; Cahn, J.Y.; Cerny, J.; Copelan, E.; et al. Comparison of outcomes of HCT in blast phase of BCR-ABL1− MPN with de novo AML and with AML following MDS. Blood Adv. 2020, 4, 4748–4757. [Google Scholar] [CrossRef]
- Ware, A.D. The complete blood count and white blood cell differential. In Contemporary Practice in Clinical Chemistry; Academic Press: Cambridge, MA, USA, 2020; pp. 429–444. [Google Scholar] [CrossRef]
- Boutault, R.; Peterlin, P.; Boubaya, M.; Sockel, K.; Chevallier, P.; Garnier, A.; Guillaume, T.; Le Bourgeois, A.; Debord, C.; Godon, C.; et al. A novel complete blood count-based score to screen for myelodysplastic syndrome in cytopenic patients. Br. J. Haematol. 2018, 183, 736–746. [Google Scholar] [CrossRef]
- Horvat, I.; Boban, A.; Zadro, R.; Antolic, M.R.; Serventi-Seiwerth, R.; Roncevic, P.; Radman, I.; Sertic, D.; Vodanovic, M.; Pulanic, D.; et al. Influence of Blood Count, Cardiovascular Risks, Inherited Thrombophilia, and JAK2 V617F Burden Allele on Type of Thrombosis in Patients with Philadelphia Chromosome Negative Myeloproliferative Neoplasms. Clin. Lymphoma Myeloma Leuk. 2019, 19, 53–63. [Google Scholar] [CrossRef]
- Wang, S.Y.; Cheng, W.Y.; Mao, Y.F.; Zhu, Y.M.; Liu, F.J.; Ma, T.T.; Shen, Y. Genetic alteration patterns and clinical outcomes of elderly and secondary acute myeloid leukemia. Hematol. Oncol. 2019, 37, 456–463. [Google Scholar] [CrossRef]
- Elessa, D.; Zhao, L.P.; de Oliveira, R.D.; Maslah, N.; Soret-Dulphy, J.; Verger, E.; Marcault, C.; Parquet, N.; Fenaux, P.; Adès, L.; et al. Clinical features and genomic landscape of myeloproliferative neoplasm (MPN) patients with autoimmune and inflammatory diseases (AID). Leukemia 2023, 378, 1741–1744. [Google Scholar] [CrossRef]
- Boddu, P.C.; Zeidan, A.M. Myeloid disorders after autoimmune disease. Best Pract. Res. Clin. Haematol. 2019, 32, 74. [Google Scholar] [CrossRef]
- Rodriguez-Meira, A.; Norfo, R.; Wen, S.; Chédeville, A.L.; Rahman, H.; O’Sullivan, J.; Wang, G.; Louka, E.; Kretzschmar, W.W.; Paterson, A.; et al. Single-cell multi-omics identifies chronic inflammation as a driver of TP53-mutant leukemic evolution. Nat. Genet. 2023, 55, 1531–1541. [Google Scholar] [CrossRef]
- Mendez Luque, L.F.; Blackmon, A.L.; Ramanathan, G.; Fleischman, A.G. Key Role of Inflammation in Myeloproliferative Neoplasms: Instigator of Disease Initiation, Progression. and Symptoms. Curr. Hematol. Malig. Rep. 2019, 14, 145–153. [Google Scholar] [CrossRef] [PubMed]
- Braun, T.; Carvalho, G.; Fabre, C.; Grosjean, J.; Fenaux, P.; Kroemer, G. Targeting NF-κB in hematologic malignancies. Cell Death Differ. 2006, 13, 748–758. [Google Scholar] [CrossRef] [PubMed]
- Karantanos, T.; Teodorescu, P.; Perkins, B.; Christodoulou, I.; Esteb, C.; Varadhan, R.; Helmenstine, E.; Rajkhowa, T.; Paun, B.C.; Bonifant, C.; et al. The role of the atypical chemokine receptor CCRL2 in myelodysplastic syndrome and secondary acute myeloid leukemia. Sci. Adv. 2022, 8, eabl8952. [Google Scholar] [CrossRef]
- Schinke, C.; Giricz, O.; Li, W.; Shastri, A.; Gordon, S.; Barreyro, L.; Bhagat, T.; Bhattacharyya, S.; Ramachandra, N.; Bartenstein, M.; et al. IL8-CXCR2 pathway inhibition as a therapeutic strategy against MDS and AML stem cells. Blood 2015, 125, 3144–3152. [Google Scholar] [CrossRef]
- Wang, X.; Teng, F.; Kong, L.; Yu, J. PD-L1 expression in human cancers and its association with clinical outcomes. Onco Targets Ther. 2016, 9, 5023. [Google Scholar] [CrossRef]
- Montes, P.; Bernal, M.; Campo, L.N.; González-Ramírez, A.R.; Jiménez, P.; Garrido, P.; Jurado, M.; Garrido, F.; Ruiz-Cabello, F.; Hernández, F. Tumor genetic alterations and features of the immune microenvironment drive myelodysplastic syndrome escape and progression. Cancer Immunol. Immunother. 2019, 68, 2015–2027. [Google Scholar] [CrossRef]
- Milosevic Feenstra, J.D.; Jäger, R.; Schischlik, F.; Ivanov, D.; Eisenwort, G.; Rumi, E.; Schuster, M.; Gisslinger, B.; Machherndl-Spandl, S.; Bettelheim, P.; et al. PD-L1 overexpression correlates with JAK2-V617F mutational burden and is associated with 9p uniparental disomy in myeloproliferative neoplasms. Am. J. Hematol. 2022, 97, 390–400. [Google Scholar] [CrossRef] [PubMed]
- Palumbo, G.A.; Parrinello, N.L.; Giallongo, C.; D’amico, E.; Zanghì, A.; Puglisi, F.; Conticello, C.; Chiarenza, A.; Tibullo, D.; Di Raimondo, F.; et al. Monocytic Myeloid Derived Suppressor Cells in Hematological Malignancies. Int. J. Mol. Sci. 2019, 20, 5459. [Google Scholar] [CrossRef]
- Vaxevanis, C.K.; Bauer, M.; Subbarayan, K.; Friedrich, M.; Massa, C.; Biehl, K.; Al-Ali, H.K.; Wickenhauser, C.; Seliger, B. Biglycan as a mediator of proinflammatory response and target for MDS and sAML therapy. Oncoimmunology 2023, 12, 2152998. [Google Scholar] [CrossRef] [PubMed]
- Rontauroli, S.; Carretta, C.; Parenti, S.; Bertesi, M.; Manfredini, R. Novel Molecular Insights into Leukemic Evolution of Myeloproliferative Neoplasms: A Single Cell Perspective. Int. J. Mol. Sci. 2022, 23, 15256. [Google Scholar] [CrossRef] [PubMed]
- Han, H.; Byun, J.M.; Shin, D.Y.; Yoon, S.S.; Koh, Y.; Hong, J.; Kim, I.; Lee, C.; Yoo, H.; Yun, H.; et al. Leukemic stem cell phenotype is associated with mutational profile in acute myeloid leukemia. Korean J. Intern. Med. 2021, 36, 401. [Google Scholar] [CrossRef]
- de Castro, F.A.; Mehdipour, P.; Chakravarthy, A.; Ettayebi, I.; Yau, H.L.; Medina, T.S.; Marhon, S.A.; de Almeida, F.C.; Bianco, T.M.; Arruda, A.G.F.; et al. Ratio of stemness to interferon signalling as a biomarker and therapeutic target of myeloproliferative neoplasm progression to acute myeloid leukaemia. Br. J. Haematol. 2023, 204, 206–220. [Google Scholar] [CrossRef]
- Guess, T.; Potts, C.R.; Bhat, P.; Cartailler, J.A.; Brooks, A.; Holt, C.; Yenamandra, A.; Wheeler, F.C.; Savona, M.R.; Cartailler, J.-P.; et al. Distinct Patterns of Clonal Evolution Drive Myelodysplastic Syndrome Progression to Secondary Acute Myeloid Leukemia. Blood Cancer Discov. 2022, 3, 316–329. [Google Scholar] [CrossRef]
- Miles, L.A.; Bowman, R.L.; Merlinsky, T.R.; Csete, I.S.; Ooi, A.T.; Durruthy-Durruthy, R.; Bowman, M.; Famulare, C.; Patel, M.A.; Mendez, P.; et al. Single cell mutation analysis of clonal evolution in myeloid malignancies. Nature 2020, 587, 477. [Google Scholar] [CrossRef]
- Luque Paz, D.; Kralovics, R.; Skoda, R.C. Genetic basis and molecular profiling in myeloproliferative neoplasms. Blood 2023, 141, 1909–1921. [Google Scholar] [CrossRef]
- Paz, D.L.; Jouanneau-Courville, R.; Riou, J.; Ianotto, J.C.; Boyer, F.; Chauveau, A.; Renard, M.; Chomel, J.C.; Cayssials, E.; Gallego-Hernanz, M.P.; et al. Leukemic evolution of polycythemia vera and essential thrombocythemia: Genomic profiles predict time to transformation. Blood Adv. 2020, 4, 4887–4897. [Google Scholar] [CrossRef]
- Makishima, H.; Yoshizato, T.; Yoshida, K.; Sekeres, M.A.; Radivoyevitch, T.; Suzuki, H.; Przychodzen, B.J.; Nagata, Y.; Meggendorfer, M.; Sanada, M.; et al. Dynamics of clonal evolution in myelodysplastic syndromes. Nat. Genet. 2016, 49, 204–212. [Google Scholar] [CrossRef]
- Taylor, J.; Lee, S.C.; Stanley Lee, C.C.; Sloan Kettering, M. Mutations in spliceosome genes and therapeutic opportunities in myeloid malignancies. Genes Chromosom. Cancer 2019, 58, 889–902. [Google Scholar] [CrossRef]
- Kar, S.A.; Jankowska, A.; Makishima, H.; Visconte, V.; Jerez, A.; Sugimoto, Y.; Muramatsu, H.; Traina, F.; Afable, M.; Guinta, K.; et al. Spliceosomal gene mutations are frequent events in the diverse mutational spectrum of chronic myelomonocytic leukemia but largely absent in juvenile myelomonocytic leukemia. Haematologica 2013, 98, 107. [Google Scholar] [CrossRef]
- Stegelmann, F.; Bullinger, L.; Schlenk, R.F.; Paschka, P.; Griesshammer, M.; Blersch, C.; Kuhn, S.; Schauer, S.; Döhner, H.; Döhner, K. DNMT3A mutations in myeloproliferative neoplasms. Leukemia 2011, 25, 1217–1219. [Google Scholar] [CrossRef] [PubMed]
- Brune, M.M.; Rau, A.; Overkamp, M.; Flaadt, T.; Bonzheim, I.; Schürch, C.M.; Federmann, B.; Dirnhofer, S.; Fend, F.; Tzankov, A. Molecular Progression of Myeloproliferative and Myelodysplastic/Myeloproliferative Neoplasms: A Study on Sequential Bone Marrow Biopsies. Cancers 2021, 13, 5605. [Google Scholar] [CrossRef] [PubMed]
- Benton, C.B.; Boddu, P.C.; DiNardo, C.D.; Bose, P.; Wang, F.; Assi, R.; Pemmaraju, N.; Devendra, K.C.; Pierce, S.; Patel, K.; et al. Janus kinase 2 variants associated with the transformation of myeloproliferative neoplasms into acute myeloid leukemia. Cancer 2019, 125, 1855–1866. [Google Scholar] [CrossRef] [PubMed]
- Stengel, A.; Baer, C.; Walter, W.; Meggendorfer, M.; Kern, W.; Haferlach, T.; Haferlach, C. Mutational patterns and their correlation to CHIP-related mutations and age in hematological malignancies. Blood Adv. 2021, 5, 4426–4434. [Google Scholar] [CrossRef] [PubMed]
- Harutyunyan, A.; Klampfl, T.; Cazzola, M.; Kralovics, R. p53 Lesions in Leukemic Transformation. N. Engl. J. Med. 2011, 364, 488–490. [Google Scholar] [CrossRef]
- Rampal, R.; Ahn, J.; Abdel-Wahaba, O.; Nahas, M.; Wang, K.; Lipson, D.; Otto, G.A.; Yelensky, R.; Hricik, T.; McKenney, A.S.; et al. Genomic and functional analysis of leukemic transformation of myeloproliferative neoplasms. Proc. Natl. Acad. Sci. USA 2014, 111, E5401–E5410. [Google Scholar] [CrossRef] [PubMed]
- Gaidzik, V.I.; Teleanu, V.; Papaemmanuil, E.; Weber, D.; Paschka, P.; Hahn, J.; Wallrabenstein, T.; Kolbinger, B.; Köhne, C.H.; Horst, H.A.; et al. RUNX1 mutations in acute myeloid leukemia are associated with distinct clinico-pathologic and genetic features. Leukemia 2016, 30, 2160–2168. [Google Scholar] [CrossRef]
- Jutzi, J.S.; Bogeska, R.; Nikoloski, G.; Schmid, C.A.; Seeger, T.S.; Stegelmann, F.; Schwemmers, S.; Gründer, A.; Peeken, J.C.; Gothwal, M.; et al. MPN patients harbor recurrent truncating mutations in transcription factor NF-E2. J. Exp. Med. 2013, 210, 1003–1019. [Google Scholar] [CrossRef] [PubMed]
- Goyal, H.; Chachoua, I.; Pecquet, C.; Vainchenker, W.; Constantinescu, S.N. A p53-JAK-STAT connection involved in myeloproliferative neoplasm pathogenesis and progression to secondary acute myeloid leukemia. Blood Rev. 2020, 42, 100712. [Google Scholar] [CrossRef]
- Zarka, J.; Short, N.J.; Kanagal-Shamanna, R.; Issa, G.C. Nucleophosmin 1 Mutations in Acute Myeloid Leukemia. Genes 2020, 11, 649. [Google Scholar] [CrossRef] [PubMed]
- Menssen, A.J.; Walter, M.J. Genetics of progression from MDS to secondary leukemia. Blood 2020, 136, 50–60. [Google Scholar] [CrossRef]
- Della Valle, V.; James, C.; Trannoy, S.; Jean-Pierre Le Couedic, B.S.; Robert, F.; Alberdi, A.; Lécluse, Y.; Plo, I.; Dreyfus, F.J.; Marzac, C.; et al. Mutation in TET2 in Myeloid Cancers. N. Engl. J. Med. 2009, 360, 2289–2301. [Google Scholar] [CrossRef]
- Abdel-Wahab, O.; Pardanani, A.; Rampal, R.; Lasho, T.L.; Levine, R.L.; Tefferi, A. DNMT3A mutational analysis in primary myelofibrosis, chronic myelomonocytic leukemia and advanced phases of myeloproliferative neoplasms. Leukemia 2011, 25, 1219–1220. [Google Scholar] [CrossRef]
- Shih, A.H.; Chung, S.S.; Dolezal, E.K.; Zhang, S.J.; Abdel-Wahab, O.I.; Park, C.Y.; Nimer, S.D.; Levine, R.L.; Klimek, V.M. Mutational analysis of therapy-related myelodysplastic syndromes and acute myelogenous leukemia. Haematologica 2013, 98, 908–912. [Google Scholar] [CrossRef]
- Guerra, V.A.; Yuanqing, Y.; Hsu, J.; Wang, F.; Alfayez, M.; Morita, K.; Song, X.; DiNardo, C.D.; Konopleva, M.Y.; Borthakur, G.M.; et al. Comprehensive Analysis of Genotype and Prior Exposures in Therapy-Related Myeloid Neoplasms (t-MNs). Blood 2019, 134, 458. [Google Scholar] [CrossRef]
- Lundberg, P.; Karow, A.; Nienhold, R.; Looser, R.; Hao-Shen, H.; Nissen, I.; Girsberger, S.; Lehmann, T.; Passweg, J.; Stern, M.; et al. Clonal evolution and clinical correlates of somatic mutations in myeloproliferative neoplasms. Blood 2014, 123, 2220–2228. [Google Scholar] [CrossRef]
- Soenen, V.; Preudhomme, C.; Roumier, C.; Daudignon, A.; Laï, J.L.; Fenaux, P. 17p Deletion in Acute Myeloid Leukemia and Myelodysplastic Syndrome. Analysis of Breakpoints and Deleted Segments by Fluorescence In Situ. Blood 1998, 91, 1008–1015. [Google Scholar] [CrossRef]
- Rücker, F.G.; Schlenk, R.F.; Bullinger, L.; Kayser, S.; Teleanu, V.; Kett, H.; Habdank, M.; Kugler, C.M.; Holzmann, K.; Gaidzik, V.I.; et al. TP53 alterations in acute myeloid leukemia with complex karyotype correlate with specific copy number alterations, monosomal karyotype, and dismal outcome. Blood 2012, 119, 2114–2121. [Google Scholar] [CrossRef]
- Bahaj, W.; Kewan, T.; Gurnari, C.; Durmaz, A.; Ponvilawan, B.; Pandit, I.; Kubota, Y.; Ogbue, O.D.; Zawit, M.; Madanat, Y.; et al. Novel scheme for defining the clinical implications of TP53 mutations in myeloid neoplasia. J. Hematol. Oncol. 2023, 16, 91. [Google Scholar] [CrossRef] [PubMed]
- Rahmé, R.; Braun, T.; Manfredi, J.J.; Fenaux, P. TP53 Alterations in Myelodysplastic Syndromes and Acute Myeloid Leukemia. Biomedicines 2023, 11, 1152. [Google Scholar] [CrossRef]
- Daver, N.G.; Maiti, A.; Kadia, T.M.; Vyas, P.; Majeti, R.; Wei, A.H.; Garcia-Manero, G.; Craddock, C.; Sallman, D.A.; Kantarjian, H.M. TP53-Mutated Myelodysplastic Syndrome and Acute Myeloid Leukemia: Biology, Current Therapy, and Future Directions. Cancer Discov. 2022, 12, 2516. [Google Scholar] [CrossRef]
- Arber, D.A.; Orazi, A.; Hasserjian, R.P.; Borowitz, M.J.; Calvo, K.R.; Kvasnicka, H.M.; Wang, S.A.; Bagg, A.; Barbui, T.; Branford, S.; et al. International Consensus Classification of Myeloid Neoplasms and Acute Leukemias: Integrating morphologic, clinical, and genomic data. Blood 2022, 140, 1200–1228. [Google Scholar] [CrossRef]
- Sebert, M.; Gachet, S.; Leblanc, T.; Rousseau, A.; Bluteau, O.; Rathana, K.I.M.; Benabelali, R.; de Fontbrune, F.S.; Maillard, L.; Fedronie, C.; et al. Clonal Hematopoiesis Driven By MDM4 Amplification Defines a Canonical Route Towards Secondary MDS/AML in Fanconi Anemia Patients. Blood 2021, 138, 860. [Google Scholar] [CrossRef]
- Pezeshki, A.; Podder, S.; Kamel, R.; Corey, S.J. Monosomy 7/del (7q) in inherited bone marrow failure syndromes: A systematic review. Pediatr. Blood Cancer 2017, 64, e26714. [Google Scholar] [CrossRef]
- Daniels, N.J.; Hershberger, C.E.; Gu, X.; Schueger, C.; DiPasquale, W.M.; Brick, J.; Saunthararajah, Y.; Maciejewski, J.P.; Padgett, R.A. Functional analyses of human LUC7-like proteins involved in splicing regulation and myeloid neoplasms. Cell Rep. 2021, 35, 108989. [Google Scholar] [CrossRef]
- Awada, H.; Durmaz, A.; Gurnari, C.; Kishtagari, A.; Meggendorfer, M.; Kerr, C.M.; Kuzmanovic, T.; Durrani, J.; Shreve, J.; Nagata, Y.; et al. Machine learning integrates genomic signatures for subclassification beyond primary and secondary acute myeloid leukemia. Blood 2021, 138, 1885–1895. [Google Scholar] [CrossRef]
- Guglielmelli, P.; Calabresi, L. The MPL mutation. Int. Rev. Cell Mol. Biol. 2021, 365, 163–178. [Google Scholar] [CrossRef]
- Chang, L.; Cui, Z.; Shi, D.; Chu, Y.; Wang, B.; Wan, Y.; Ma, Q.; Zhang, R.; Li, H.; Cheng, X.; et al. Polyclonal evolution of Fanconi anemia to MDS and AML revealed at single cell resolution. Exp. Hematol. Oncol. 2022, 11, 64. [Google Scholar] [CrossRef]
- Lee, C.; Kim, H.N.; Kwon, J.A.; Hwang, J.; Park, J.Y.; Shin, O.S.; Yoon, S.Y.; Yoon, J. Identification of a Complex Karyotype Signature with Clinical Implications in AML and MDS-EB Using Gene Expression Profiling. Cancers 2023, 15, 5289. [Google Scholar] [CrossRef]
- Tembrink, M.; Gerding, W.M.; Wieczorek, S.; Mika, T.; Schroers, R.; Nguyen, H.P.; Ben Vangala, D.; Nilius-Eliliwi, V. Novel NUP98::ASH1L Gene Fusion in Acute Myeloid Leukemia Detected by Optical Genome Mapping. Cancers 2023, 15, 2942. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhang, Y.; Wang, C.; Wang, X. TET (Ten-eleven translocation) family proteins: Structure, biological functions and applications. Signal Transduct. Target. Ther. 2023, 8, 297. [Google Scholar] [CrossRef]
- Ganster, C.; Müller-Thomas, C.; Haferlach, C.; Strupp, C.; Ogata, K.; Germing, U.; Hildebrandt, B.; Mallo, M.; Lübbert, M.; Müller, C.; et al. Comprehensive analysis of isolated der(1;7)(q10;p10) in a large international homogenous cohort of patients with myelodysplastic syndromes. Genes Chromosom. Cancer 2019, 58, 689–697. [Google Scholar] [CrossRef]
- Hebeda, K.; Boudova, L.; Beham-Schmid, C.; Orazi, A.; Kvasnicka, H.M.; Gianelli, U.; Tzankov, A. Progression, transformation, and unusual manifestations of myelodysplastic syndromes and myelodysplastic-myeloproliferative neoplasms: Lessons learned from the XIV European Bone Marrow Working Group Course 2019. Ann. Hematol. 2021, 100, 117–133. [Google Scholar] [CrossRef] [PubMed]
- Gurban, P.; Mambet, C.; Botezatu, A.; Necula, L.G.; Neagu, A.I.; Matei, L.; Pitica, I.M.; Nedeianu, S.; Chivu-Economescu, M.; Bleotu, C.; et al. Leukemic conversion involving RAS mutations of type 1 CALR-mutated primary myelofibrosis in a patient treated for HCV cirrhosis: A case report. Front. Oncol. 2023, 13, 1266996. [Google Scholar] [CrossRef]
- Wan, C.; Wen, J.; Liang, X.; Xie, Q.; Wu, W.; Wu, M.; Liu, Z. Identification of miR-320 family members as potential diagnostic and prognostic biomarkers in myelodysplastic syndromes. Sci. Rep. 2021, 11, 183. [Google Scholar] [CrossRef] [PubMed]
- Symeonidis, A.; Chatzilygeroudi, T.; Chondrou, V.; Sgourou, A. Contingent Synergistic Interactions between Non-Coding RNAs and DNA-Modifying Enzymes in Myelodysplastic Syndromes. Int. J. Mol. Sci. 2022, 23, 16069. [Google Scholar] [CrossRef] [PubMed]
- Lyu, C.; Liu, K.; Jiang, Y.; Wang, T.; Wang, Y.; Xu, R. Integrated analysis on mRNA microarray and microRNA microarray to screen immune-related biomarkers and pathways in myelodysplastic syndrome. Hematology 2021, 26, 417–431. [Google Scholar] [CrossRef] [PubMed]
- Thol, F.; Scherr, M.; Kirchner, A.; Shahswar, R.; Battmer, K.; Kade, S.; Chaturvedi, A.; Koenecke, C.; Stadler, M.; Platzbecker, U.; et al. Clinical and functional implications of microRNA mutations in a cohort of 935 patients with myelodysplastic syndromes and acute myeloid leukemia. Haematologica 2015, 100, e122. [Google Scholar] [CrossRef][Green Version]
- Kirimura, S.; Kurata, M.; Nakagawa, Y.; Onishi, I.; Abe-Suzuki, S.; Abe, S.; Yamamoto, K.; Kitagawa, M. Role of microRNA-29b in myelodysplastic syndromes during transformation to overt leukaemia. Pathology 2016, 48, 233–241. [Google Scholar] [CrossRef]
- Pavlovic, D.; Tosic, N.; Zukic, B.; Pravdic, Z.; Vukovic, N.S.; Pavlovic, S.; Gasic, V. Expression Profiles of Long Non-Coding RNA GAS5 and MicroRNA-222 in Younger AML Patients. Diagnostics 2022, 12, 86. [Google Scholar] [CrossRef]
- Qin, T.; Cheng, Y.; Wang, X. RNA-binding proteins as drivers of AML and novel therapeutic targets. Leuk. Lymphoma 2022, 63, 1045–1057. [Google Scholar] [CrossRef]
- Bauer, M.; Vaxevanis, C.; Heimer, N.; Al-Ali, H.K.; Jaekel, N.; Bachmann, M.; Wickenhauser, C.; Seliger, B. Expression, Regulation and Function of microRNA as Important Players in the Transition of MDS to Secondary AML and Their Cross Talk to RNA-Binding Proteins. Int. J. Mol. Sci. 2020, 21, 7140. [Google Scholar] [CrossRef]
- Jiang, Q.; Isquith, J.; Ladel, L.; Mark, A.; Holm, F.; Mason, C.; He, Y.; Mondala, P.; Oliver, I.; Pham, J.; et al. Inflammation-driven deaminase deregulation fuels human pre-leukemia stem cell evolution. Cell Rep. 2021, 34, 108670. [Google Scholar] [CrossRef] [PubMed]
- Maakaron, J.E.; Mims, A.S. Daunorubicin-cytarabine liposome (CPX-351) in the management of newly diagnosed secondary AML: A new twist on an old cocktail. Best. Pract. Res. Clin. Haematol. 2019, 32, 127–133. [Google Scholar] [CrossRef] [PubMed]
- Budziszewska, B.K.; Salomon-Perzyński, A.; Pruszczyk, K.; Barankiewicz, J.; Pluta, A.; Helbig, G.; Janowska, A.; Kuydowicz, M.; Bołkun, Ł.; Piszcz, J.; et al. Cladribine Combined with Low-Dose Cytarabine as Frontline Treatment for Unfit Elderly Acute Myeloid Leukemia Patients: Results from a Prospective Multicenter Study of Polish Adult Leukemia Group (PALG). Cancers 2021, 13, 4189. [Google Scholar] [CrossRef]
- Schieber, M.; Marinaccio, C.; Bolanos, L.C.; Haffey, W.D.; Greis, K.D.; Starczynowski, D.T.; Crispino, J.D. FBXO11 is a candidate tumor suppressor in the leukemic transformation of myelodysplastic syndrome. Blood Cancer J. 2020, 10, 98. [Google Scholar] [CrossRef]
- Roboz, G.J.; Mandrekar, S.J.; Desai, P.; Laumann, K.; Walker, A.R.; Wang, E.S.; Kolitz, J.E.; Powell, B.L.; Attar, E.C.; Stock, W.; et al. Randomized trial of 10 days of decitabine ± bortezomib in untreated older patients with AML: CALGB 11002 (Alliance). Blood Adv. 2018, 2, 3608–3617. [Google Scholar] [CrossRef]
- Malik, P.; Cashen, A.F. Decitabine in the treatment of acute myeloid leukemia in elderly patients. Cancer Manag. Res. 2014, 6, 53–61. [Google Scholar] [CrossRef]
- Zong, L.; Yin, M.; Kong, J.; Zhang, J.; Song, B.; Zhu, J.; Xue, S.; Wu, X.; Wu, D.; Bao, X.; et al. Development of a scoring system for predicting primary resistance to venetoclax plus hypomethylating agents (HMAs) in acute myeloid leukemia patients. Mol. Carcinog. 2023, 62, 1572–1584. [Google Scholar] [CrossRef]
- Haddad, F.; Daver, N. Targeting CD47/SIRPα in Acute Myeloid Leukemia and Myelodysplastic Syndrome: Preclinical and Clinical Developments of Magrolimab. J. Immunother. Precis. Oncol. 2021, 4, 67. [Google Scholar] [CrossRef]
- Maslah, N.; Salomao, N.; Drevon, L.; Verger, E.; Partouche, N.; Ly, P.; Aubin, P.; Naoui, N.; Schlageter, M.H.; Bally, C.; et al. Synergistic effects of PRIMA-1Met (APR-246) and 5-azacitidine in TP53-mutated myelodysplastic syndromes and acute myeloid leukemia. Haematologica 2020, 105, 1539. [Google Scholar] [CrossRef]
- Fiskus, W.; Mill, C.P.; Nabet, B.; Perera, D.; Birdwell, C.; Manshouri, T.; Lara, B.; Kadia, T.M.; DiNardo, C.; Takahashi, K.; et al. Superior efficacy of co-targeting GFI1/KDM1A and BRD4 against AML and post-MPN secondary AML cells. Blood Cancer J. 2021, 11, 98. [Google Scholar] [CrossRef]
- Tibes, R.; Bogenberger, J.M. Transcriptional Silencing of MCL-1 Through Cyclin-Dependent Kinase Inhibition in Acute Myeloid Leukemia. Front. Oncol. 2019, 9, 1205. [Google Scholar] [CrossRef]
- Fiskus, W.; Manshouri, T.; Birdwell, C.; Mill, C.P.; Masarova, L.; Bose, P.; Kadia, T.M.; Daver, N.; DiNardo, C.D.; Borthakur, G.; et al. Efficacy of CDK9 inhibition in therapy of post-myeloproliferative neoplasm (MPN) secondary (s) AML cells. Blood Cancer J. 2022, 12, 23. [Google Scholar] [CrossRef]
- Crews, L.A.; Ma, W.; Ladel, L.; Pham, J.; Balaian, L.; Steel, S.K.; Mondala, P.K.; Diep, R.H.; Wu, C.N.; Mason, C.N.; et al. Reversal of malignant ADAR1 splice isoform switching with Rebecsinib. Cell Stem Cell 2023, 30, 250–263.e6. [Google Scholar] [CrossRef]
- Qing, Y.; Su, R.; Chen, J. RNA modifications in hematopoietic malignancies: A new research frontier. Blood 2021, 138, 637–648. [Google Scholar] [CrossRef]
- Eisenberg, E.; Levanon, E.Y. A-to-I RNA editing—Immune protector and transcriptome diversifier. Nat. Rev. Genet. 2018, 19, 473–490. [Google Scholar] [CrossRef]
- Schmaelter, A.K.; Labopin, M.; Socié, G.; Itälä-Remes, M.; Blaise, D.; Yakoub-Agha, I.; Forcade, E.; Cornelissen, J.; Ganser, A.; Beelen, D.; et al. Inferior outcome of allogeneic stem cell transplantation for secondary acute myeloid leukemia in first complete remission as compared to de novo acute myeloid leukemia. Blood Cancer J. 2020, 10, 26. [Google Scholar] [CrossRef]
- Nilsson, C.; Hulegårdh, E.; Garelius, H.; Möllgård, L.; Brune, M.; Wahlin, A.; Lenhoff, S.; Frödin, U.; Remberger, M.; Höglund, M.; et al. Secondary Acute Myeloid Leukemia and the Role of Allogeneic Stem Cell Transplantation in a Population-Based Setting. Biol. Blood Marrow Transplant. 2019, 25, 1770–1778. [Google Scholar] [CrossRef]
- Guolo, F.; Fianchi, L.; Minetto, P.; Clavio, M.; Gottardi, M.; Galimberti, S.; Rizzuto, G.; Rondoni, M.; Bertani, G.; Dargenio, M.; et al. CPX-351 treatment in secondary acute myeloblastic leukemia is effective and improves the feasibility of allogeneic stem cell transplantation: Results of the Italian compassionate use program. Blood Cancer J. 2020, 10, 96. [Google Scholar] [CrossRef]
- Limongello, R.; Marra, A.; Mancusi, A.; Bonato, S.; Hoxha, E.; Ruggeri, L.; Hui, S.; Velardi, A.; Pierini, A. Novel Immune Cell-Based Therapies to Eradicate High-Risk Acute Myeloid Leukemia. Front. Immunol. 2021, 12, 695051. [Google Scholar] [CrossRef]
- Zhang, H.; Gan, W.T.; Hao, W.G.; Wang, P.F.; Li, Z.Y.; Chang, L.J. Successful Anti-CLL1 CAR T-Cell Therapy in Secondary Acute Myeloid Leukemia. Front. Oncol. 2020, 10, 685. [Google Scholar] [CrossRef]
- Haubner, S.; Perna, F.; Köhnke, T.; Schmidt, C.; Berman, S.; Augsberger, C.; Schnorfeil, F.M.; Krupka, C.; Lichtenegger, F.S.; Liu, X.; et al. Coexpression profile of leukemic stem cell markers for combinatorial targeted therapy in AML. Leukemia 2018, 33, 64–74. [Google Scholar] [CrossRef]
- Sugita, M.; Galetto, R.; Zong, H.; Ewing-Crystal, N.; Trujillo-Alonso, V.; Mencia-Trinchant, N.; Yip, W.; Filipe, S.; Lebuhotel, C.; Gouble, A.; et al. Allogeneic TCRαβ deficient CAR T-cells targeting CD123 in acute myeloid leukemia. Nat. Commun. 2022, 13, 2227. [Google Scholar] [CrossRef]
- Bester, A.C.; Lee, J.D.; Chavez, A.; Lee, Y.R.; Nachmani, D.; Vora, S.; Victor, J.; Sauvageau, M.; Monteleone, E.; Rinn, J.L.; et al. An Integrated Genome-wide CRISPRa Approach to Functionalize lncRNAs in Drug Resistance. Cell 2018, 173, 649–664.e20. [Google Scholar] [CrossRef]
- Labanieh, L.; Majzner, R.G.; Mackall, C.L. Programming CAR-T cells to kill cancer. Nat. Biomed. Eng. 2018, 2, 377–391. [Google Scholar] [CrossRef]
- Marofi, F.; Rahman, H.S.; Al-Obaidi, Z.M.J.; Jalil, A.T.; Abdelbasset, W.K.; Suksatan, W.; Dorofeev, A.E.; Shomali, N.; Chartrand, M.S.; Pathak, Y.; et al. Novel CAR T therapy is a ray of hope in the treatment of seriously ill AML patients. Stem Cell Res. Ther. 2021, 12, 465. [Google Scholar] [CrossRef]
- Oliai, C.; Schiller, G. How to address second and therapy-related acute myelogenous leukaemia. Br. J. Haematol. 2020, 188, 116–128. [Google Scholar] [CrossRef]
- Baumeister, S.H.; Murad, J.; Werner, L.; Daley, H.; Trebeden-Negre, H.; Gicobi, J.K.; Schmucker, A.; Reder, J.; Sentman, C.L.; Gilham, D.E.; et al. Phase i trial of autologous CAR T cells targeting NKG2D ligands in patients with AML/MDS and multiple myeloma. Cancer Immunol Res. 2019, 7, 100–112. [Google Scholar] [CrossRef]
- Bertulfo, K.; Debackere, K.; Miller, H.; Najac, R.; Ma, C.; Ferrando, A.A.; Palomero, T. Therapeutic Targeting of JAK/STAT Signaling and Histone Acetyltransferase Activity in Post-Myeloproliferative Neoplasm Secondary AML. In Proceedings of the 65th Annual ASH Meeting & E Position, San Diego, CA, USA, 6–15 December 2023; ASH: Washington, DC, USA, 2023; p. 2792. [Google Scholar]
- Rahmé, R.; Vidal, V.; Hueso, T.; Le Meur, L.; Rigal, M.; Ivanoff, S.; Brechignac, S.; De Latour, R.P.; Gardin, C.; Braun, T. Treatment of Adverse-Risk and Refractory/Relapsed Acute Myeloid Leukemia (AML) Patients with FLAG-IDA ± Venetoclax and CLAG-M: A Monocentric Experience. In Proceedings of the 65th Annual ASH Meeting & Exposition, San Diego, CA, USA, 6–15 December 2023; ASH: Washington, DC, USA, 2023; p. 1520. [Google Scholar]
| Clinical Overview | Healthy | MDS | MPN | sAML | De Novo AML |
|---|---|---|---|---|---|
| Age (y) | 18–65 | 53–98 | 18–92 | 21–77 | 18–59 |
| White Blood Cells (109/L) | 4–11 | 1.1–17.9 | 7.2–14.7 | 0.8–144.1 | 0.77–419.9 |
| Platelets (109/L) | 150–450 | 8–505 | 376–720 | 3–752 | 30–171 |
| Hemoglobin (g/L) | 120–175 | 47–149 | 109–173 | 34–143 | 2–1726 |
| Clinical record | Free of treatment and transfusion | Both therapy free and treatment * | 3 + 7 regimen Hypomethylation or palliative treatment | 3 + 7 regimen | |
| Reference | [62] | [63] | [64] | [65] | [65] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Auerbach, S.; Puka, B.; Golla, U.; Chachoua, I. Recent Advances towards the Understanding of Secondary Acute Myeloid Leukemia Progression. Life 2024, 14, 309. https://doi.org/10.3390/life14030309
Auerbach S, Puka B, Golla U, Chachoua I. Recent Advances towards the Understanding of Secondary Acute Myeloid Leukemia Progression. Life. 2024; 14(3):309. https://doi.org/10.3390/life14030309
Chicago/Turabian StyleAuerbach, Scott, Beana Puka, Upendarrao Golla, and Ilyas Chachoua. 2024. "Recent Advances towards the Understanding of Secondary Acute Myeloid Leukemia Progression" Life 14, no. 3: 309. https://doi.org/10.3390/life14030309
APA StyleAuerbach, S., Puka, B., Golla, U., & Chachoua, I. (2024). Recent Advances towards the Understanding of Secondary Acute Myeloid Leukemia Progression. Life, 14(3), 309. https://doi.org/10.3390/life14030309







