Zinc Metalloproteins in Epigenetics and Their Crosstalk
Abstract
1. Introduction
2. Molecular Bases of Epigenetic Modifications
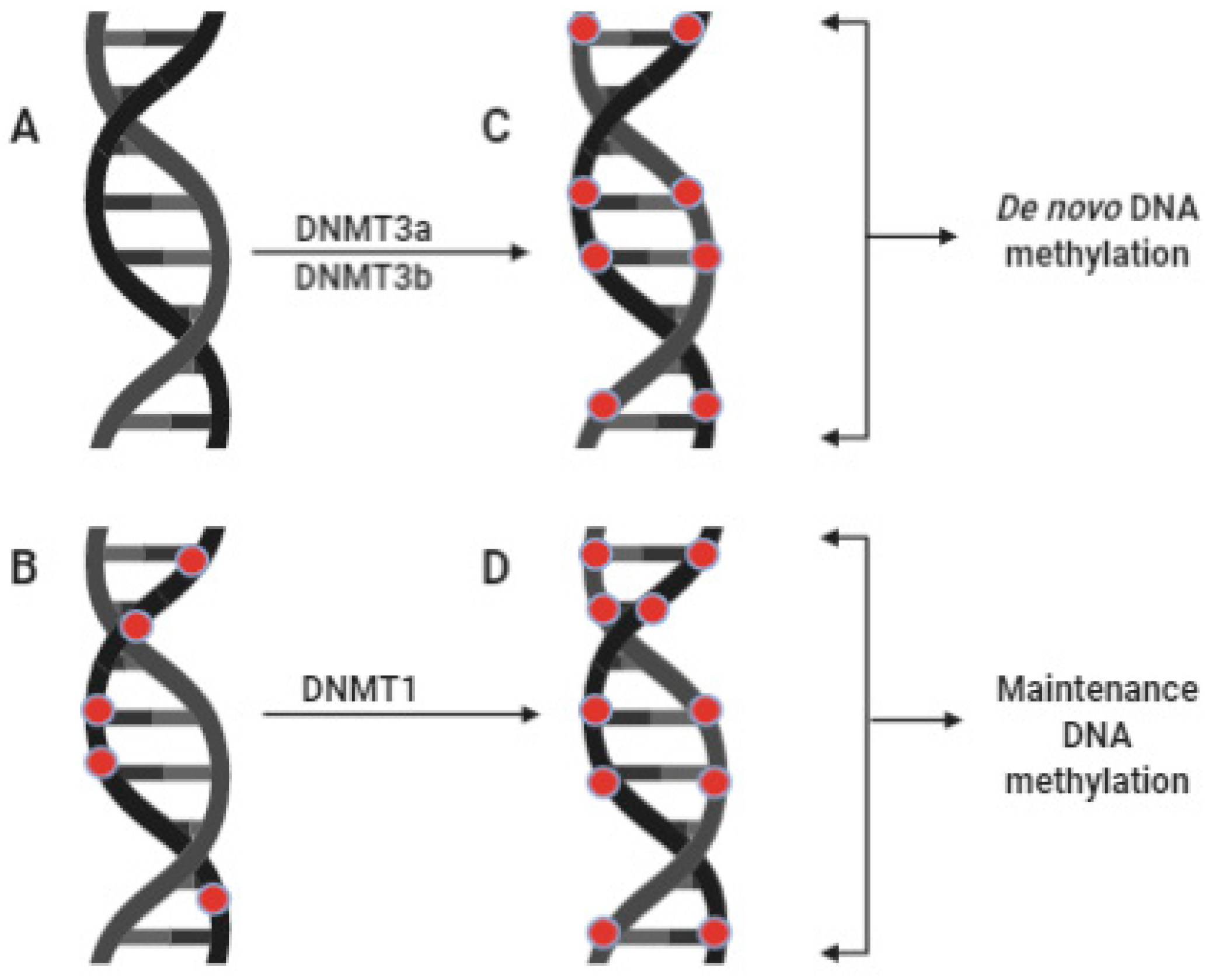
2.1. DNA Methylation
2.2. Histone Post-Translational Modifications and Chromatin Remodeling
3. Molecular Bases of Epigenome Regulation Associated with Zinc Metalloproteins
3.1. Role of Zinc in DNA Methylation
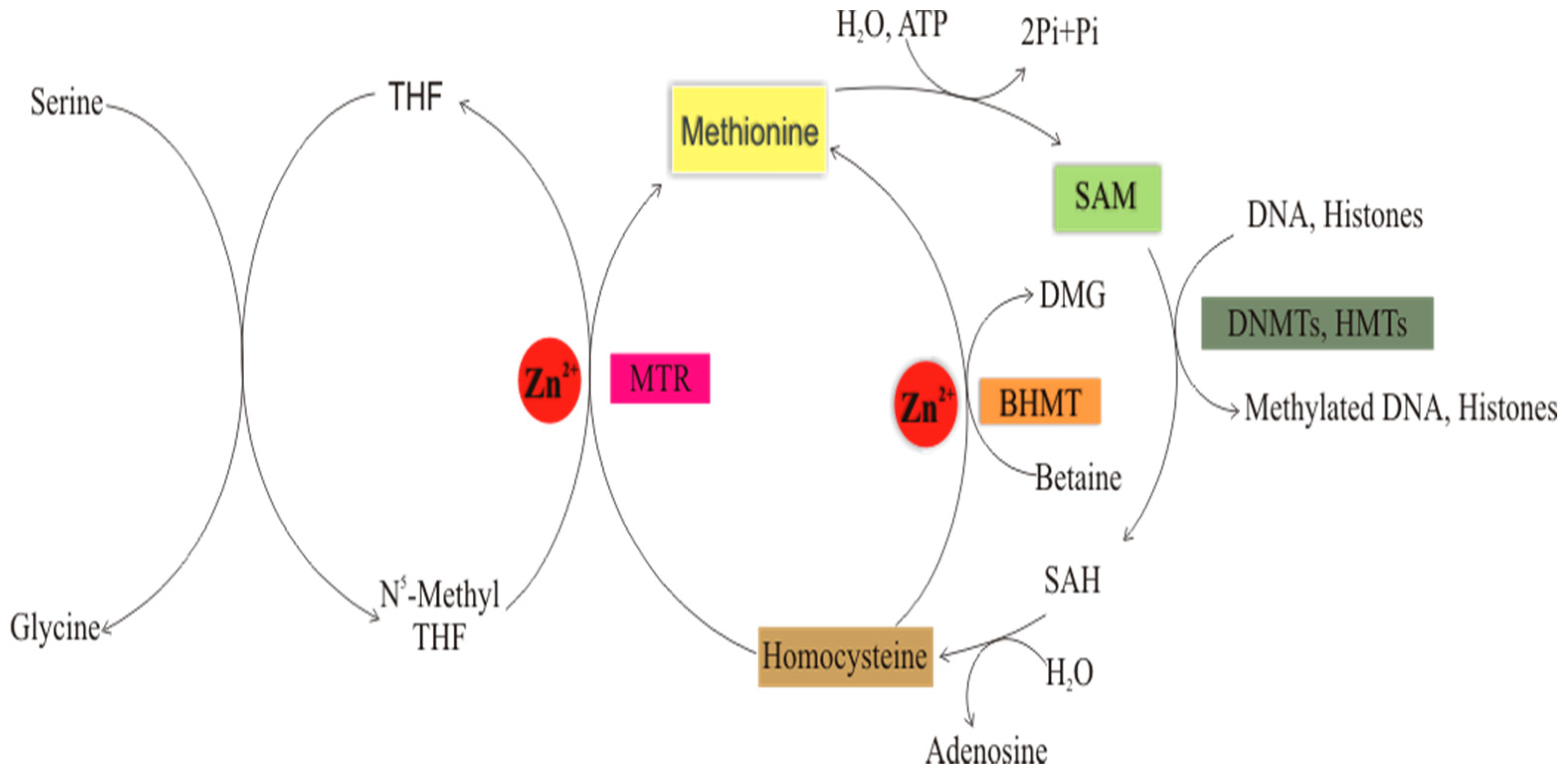
Role of Zinc in Folate-Mediated One-Carbon Metabolism
3.2. Role of Zinc in Histone Methylation
3.3. Role of Zinc in Histone Demethylation
3.4. Role of Zinc in Histone Acetylation
3.5. Role of Zinc in Histone Deacetylation
3.6. Role of Zinc in Histone Ubiquitination
3.7. Role of Zinc in Histone Deubiquitination
3.8. Zinc Finger Proteins That Read Epigenetic Modifications
3.8.1. DNA Methylation Readers and Their Binding Mechanisms
3.8.2. Histone Modifications Readers and Their Binding Mechanisms
3.9. Role of Zinc in Epigenome Editing
Role of Zinc in DNA Methylation Editing
4. Summary of Zinc Metalloproteins in Epigenetics and Their Crosstalk
5. Conclusions and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chasapis, C.T.; Loutsidou, A.C.; Spiliopoulou, C.A.; Stefanidou, M.E. Zinc and human health: An update. Arch. Toxicol. 2012, 86, 521–534. [Google Scholar] [CrossRef]
- Hambidge, K.M.; Krebs, N.F. Zinc Deficiency: A Special Challenge. J. Nutr. 2007, 137, 1101–1105. [Google Scholar] [CrossRef] [PubMed]
- Andreini, C.; Banci, L.; Bertini, I.; Rosato, A. Counting the Zinc-Proteins Encoded in the Human Genome. J. Proteome Res. 2006, 5, 196–201. [Google Scholar] [CrossRef]
- Brito, S.; Lee, M.-G.; Bin, B.-H.; Lee, J.-S. Zinc and Its Transporters in Epigenetics. Mol. Cells 2020, 43, 323–330. [Google Scholar] [PubMed]
- Chasapis, C.T.; Ntoupa, P.-S.A.; Spiliopoulou, C.A.; Stefanidou, M.E. Recent aspects of the effects of zinc on human health. Arch. Toxicol. 2020, 94, 1443–1460. [Google Scholar] [CrossRef]
- Roohani, N.; Hurrell, R.; Kelishadi, R.; Schulin, R. Zinc and its importance for human health: An integrative review. J. Res. Med. Sci. 2013, 18, 144–157. [Google Scholar] [CrossRef]
- Foster, M.; Samman, S. Zinc and Regulation of Inflammatory Cytokines: Implications for Cardiometabolic Disease. Nutrients 2012, 4, 676–694. [Google Scholar] [CrossRef] [PubMed]
- Blanquart, C.; Linot, C.; Cartron, P.-F.; Tomaselli, D.; Mai, A.; Bertrand, P. Epigenetic Metalloenzymes. Curr. Med. Chem. 2019, 26, 2748–2785. [Google Scholar] [CrossRef]
- Porter, N.J.; Christianson, D.W. Structure, mechanism, and inhibition of the zinc-dependent histone deacetylases. Curr. Opin. Struct. Biol. 2019, 59, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Hodges, A.J.; Hudson, N.O.; Buck-Koehntop, B.A. Cys2His2 Zinc Finger Methyl-CpG Binding Proteins: Getting a Handle on Methylated DNA. J. Mol. Biol. 2020, 432, 1640–1660. [Google Scholar] [CrossRef] [PubMed]
- Matthews, J.M.; Bhati, M.; Lehtomaki, E.; Mansfield, R.E.; Cubeddu, L.; Mackay, J.P. It Takes Two to Tango: The Structure and Function of LIM, RING, PHD and MYND Domains. Curr. Pharm. Des. 2009, 15, 3681–3696. [Google Scholar] [CrossRef] [PubMed]
- Wessels, I.; Maywald, M.; Rink, L. Zinc as a Gatekeeper of Immune Function. Nutrients 2017, 9, 1286. [Google Scholar] [CrossRef] [PubMed]
- Coneyworth, L.J.; Jackson, K.A.; Tyson, J.; Bosomworth, H.J.; Van Der Hagen, E.; Hann, G.M.; Ogo, O.A.; Swann, D.C.; Mathers, J.C.; Valentine, R.A.; et al. Identification of the human zinc transcriptional regulatory element (ZTRE): A palindromic protein-binding DNA se-quence responsible for zinc-induced transcriptional repression. J. Biol. Chem. 2012, 287, 36567–36581. [Google Scholar] [CrossRef]
- Bin, B.-H.; Seo, J.; Kim, S.T. Function, Structure, and Transport Aspects of ZIP and ZnT Zinc Transporters in Immune Cells. J. Immunol. Res. 2018, 2018, 9365747. [Google Scholar] [CrossRef] [PubMed]
- Myers, S.A.; Nield, A.; Myers, M. Zinc Transporters, Mechanisms of Action and Therapeutic Utility: Implications for Type 2 Diabetes Mellitus. J. Nutr. Metab. 2012, 2012, 1–13. [Google Scholar] [CrossRef]
- Gu, H.F.; Zhang, X. Zinc Deficiency and Epigenetics. In Handbook of Famine, Starvation, and Nutrient Deprivation: From Biology to Policy; Preedy, V., Patel, V.B., Eds.; Springer: New York City, NY, USA, 2017; pp. 1–18. [Google Scholar] [CrossRef]
- Lappalainen, T.; Greally, J.M. Associating cellular epigenetic models with human phenotypes. Nat. Rev. Genet. 2017, 18, 441–451. [Google Scholar] [CrossRef]
- Greally, J.M. A user’s guide to the ambiguous word ’epigenetics’. Nat. Rev. Mol. Cell Biol. 2018, 19, 207–208. [Google Scholar] [CrossRef] [PubMed]
- Felsenfeld, G. A Brief History of Epigenetics. Cold Spring Harb. Perspect. Biol. 2014, 6, a018200. [Google Scholar] [CrossRef]
- Kungulovski, G.; Jeltsch, A. Epigenome Editing: State of the Art, Concepts, and Perspectives. Trends Genet. 2016, 32, 101–113. [Google Scholar] [CrossRef]
- Van Otterdijk, S.D.; Michels, K.B. Transgenerational epigenetic inheritance in mammals: How good is the evidence? FASEB J. 2016, 30, 2457–2465. [Google Scholar] [CrossRef] [PubMed]
- Kanherkar, R.R.; Bhatia-Dey, N.; Csoka, A.B. Epigenetics across the human lifespan. Front. Cell Dev. Biol. 2014, 2, 49. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Qu, J.; Liu, G.-H.; Belmonte, J.C.I. The ageing epigenome and its rejuvenation. Nat. Rev. Mol. Cell Biol. 2020, 21, 137–150. [Google Scholar] [CrossRef] [PubMed]
- Moore, L.D.; Le, T.; Fan, G. DNA Methylation and Its Basic Function. Neuropsychopharmacology 2013, 38, 23–38. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, M.A.; Greif, P.A.; Diamond, J.; Sharaf, O.; Maxwell, P.; Montironi, R.; Young, R.A.; Hamilton, P.W. Epigenetic events, remodelling enzymes and their relationship to chromatin organization in prostatic intraepithelial neoplasia and prostatic adenocarcinoma. BJU Int. 2007, 99, 908–915. [Google Scholar] [CrossRef]
- Korkmaz, A.; Manchester, L.; Topal, T.; Ma, S.; Tan, D.; Reiter, R. Epigenetic mechanisms in human physiology and diseases. J. Exp. Integr. Med. 2011, 1, 139–147. [Google Scholar] [CrossRef]
- Lin, C.-C.; Chen, Y.-P.; Yang, W.-Z.; Shen, J.C.K.; Yuan, H.S. Structural insights into CpG-specific DNA methylation by human DNA methyltransferase 3B. Nucleic Acids Res. 2020, 48, 3949–3961. [Google Scholar] [CrossRef]
- Maret, W. Zinc Biochemistry: From a Single Zinc Enzyme to a Key Element of Life. Adv. Nutr. 2013, 4, 82–91. [Google Scholar] [CrossRef]
- An, J.; Rao, A.; Ko, M. TET family dioxygenases and DNA demethylation in stem cells and cancers. Exp. Mol. Med. 2017, 49, e323. [Google Scholar] [CrossRef] [PubMed]
- Gowher, H.; Jeltsch, A. Mammalian DNA methyltransferases: New discoveries and open questions. Biochem. Soc. Trans. 2018, 46, 1191–1202. [Google Scholar] [CrossRef]
- Ren, W.; Gao, L.; Song, J. Structural Basis of DNMT1 and DNMT3A-Mediated DNA Methylation. Genes 2018, 9, 620. [Google Scholar] [CrossRef]
- Venkatesh, S.; Workman, J.L. Histone exchange, chromatin structure and the regulation of transcription. Nat. Rev. Mol. Cell Biol. 2015, 16, 178–189. [Google Scholar] [CrossRef]
- Ebrahimi, V.; Soleimanian, A.; Ebrahimi, T.; Azargun, R.; Yazdani, P.; Eyvazi, S.; Tarhriz, V. Epigenetic modifications in gastric cancer: Focus on DNA methylation. Gene 2020, 742, 144577. [Google Scholar] [CrossRef]
- Furukawa, A.; Wakamori, M.; Arimura, Y.; Ohtomo, H.; Tsunaka, Y.; Kurumizaka, H.; Umehara, T.; Nishimura, Y. Acetylated histone H4 tail enhances histone H3 tail acetylation by altering their mutual dynamics in the nucleosome. Proc. Natl. Acad. Sci. USA 2020, 117, 19661–19663. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Dun, Y.; Fu, H.; Wang, L.; Pan, X.; Yang, X.; Fang, H. Design, synthesis, and preliminary bioactivity evaluation of N-benzylpyrimidin-2-amine derivatives as novel histone deacetylase inhibitor. Chem. Biol. Drug Des. 2017, 90, 936–942. [Google Scholar] [CrossRef] [PubMed]
- Moosavi, A.; Ardekani, A.M. Role of Epigenetics in Biology and Human Diseases. Iran. Biomed. J. 2016, 20, 246–258. [Google Scholar]
- Hayashi-Takanaka, Y.; Kina, Y.; Nakamura, F.; Becking, L.; Nakao, Y.; Nagase, T.; Nozaki, N.; Kimura, H. Histone mod-ification dynamics as revealed by a multicolor immunofluorescence-based single-cell analysis. J. Cell Sci. 2020, 133. [Google Scholar] [CrossRef]
- Obri, A.; Serra, D.; Herrero, L.; Mera, P. The role of epigenetics in the development of obesity. Biochem. Pharmacol. 2020, 177, 113973. [Google Scholar] [CrossRef]
- Blum, R. Stepping inside the realm of epigenetic modifiers. Biomol. Concepts 2015, 6, 119–136. [Google Scholar] [CrossRef]
- Sánchez, R.; Zhou, M.-M. The PHD finger: A versatile epigenome reader. Trends Biochem. Sci. 2011, 36, 364–372. [Google Scholar] [CrossRef]
- Marsh, D.J.; Dickson, K.A. Writing Histone Monoubiquitination in Human Malignancy—The Role of RING Finger E3 Ubiquitin Ligases. Genes 2019, 10, 67. [Google Scholar] [CrossRef] [PubMed]
- Jeltsch, A.; Jurkowska, R.Z. Allosteric control of mammalian DNA methyltransferases—A new regulatory paradigm. Nucleic Acids Res. 2016, 44, 8556–8575. [Google Scholar] [CrossRef] [PubMed]
- Bronner, C.; Krifa, M.; Mousli, M. Increasing role of UHRF1 in the reading and inheritance of the epigenetic code as well as in tumorogenesis. Biochem. Pharmacol. 2013, 86, 1643–1649. [Google Scholar] [CrossRef] [PubMed]
- Rothbart, S.B.; Krajewski, K.; Nady, N.; Tempel, W.; Xue, S.; Badeaux, A.I.; Barsyte-Lovejoy, D.; Martinez, J.Y.; Bedford, M.T.; Fuchs, S.M.; et al. Association of UHRF1 with methylated H3K9 directs the maintenance of DNA methylation. Nat. Struct. Mol. Biol. 2012, 19, 1155–1160. [Google Scholar] [CrossRef] [PubMed]
- Gu, B.; Lee, M.G. Histone H3 lysine 4 methyltransferases and demethylases in self-renewal and differentiation of stem cells. Cell Biosci. 2013, 3, 39. [Google Scholar] [CrossRef]
- Thakore, P.I.; Black, J.B.; Hilton, I.B.; Gersbach, C.A. Editing the epigenome: Technologies for programmable transcription and epigenetic modulation. Nat. Methods 2016, 13, 127–137. [Google Scholar] [CrossRef]
- Laity, J.H.; Lee, B.M.; Wright, P.E. Zinc finger proteins: New insights into structural and functional diversity. Curr. Opin. Struc. Biol. 2011, 11, 39–46. [Google Scholar] [CrossRef]
- Cui, D.; Xu, X. DNA Methyltransferases, DNA Methylation, and Age-Associated Cognitive Function. Int. J. Mol. Sci. 2018, 19, 1315. [Google Scholar] [CrossRef]
- Pradhan, M.; Estève, P.-O.; Chin, H.G.; Samaranayke, M.; Kim, G.-D.; Pradhan, S. CXXC Domain of Human DNMT1 Is Essential for Enzymatic Activity. Biochem. 2008, 47, 10000–10009. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.-M.; Liu, S.; Lin, K.; Luo, Y.; Perry, J.J.; Wang, Y.; Song, J. Crystal Structure of Human DNA Methyltransferase 1. J. Mol. Biol. 2015, 427, 2520–2531. [Google Scholar] [CrossRef]
- Dhe-Paganon, S.; Syeda, F.; Park, L. DNA methyl transferase 1: Regulatory mechanisms and implications in health and disease. Int. J. Biochem. Mol. Boil. 2011, 2, 58–66. [Google Scholar]
- Niinuma, T.; Kitajima, H.; Kai, M.; Yamamoto, E.; Yorozu, A.; Ishiguro, K.; Sasaki, H.; Sudo, G.; Toyota, M.; Hatahira, T.; et al. UHRF1 depletion and HDAC inhibition reactivate epigenetically silenced genes in colorectal cancer cells. Clin. Epigenet. 2019, 11, 1–13. [Google Scholar] [CrossRef]
- Argentaro, A.; Yang, J.-C.; Chapman, L.; Kowalczyk, M.S.; Gibbons, R.J.; Higgs, D.R.; Neuhaus, D.; Rhodes, D. Structural consequences of disease-causing mutations in the ATRX-DNMT3-DNMT3L (ADD) domain of the chromatin-associated protein ATRX. Proc. Natl. Acad. Sci. USA 2007, 104, 11939–11944. [Google Scholar] [CrossRef] [PubMed]
- Chédin, F.; Lieber, M.R.; Hsieh, C.-L. The DNA methyltransferase-like protein DNMT3L stimulates de novo methylation by Dnmt3a. Proc. Natl. Acad. Sci. USA 2002, 99, 16916–16921. [Google Scholar] [CrossRef]
- Bourc’His, D.; Xu, G.-L.; Lin, C.-S.; Bollman, B.; Bestor, T.H. DNMT3L and the Establishment of Maternal Genomic Imprints. Science 2001, 294, 2536–2539. [Google Scholar] [CrossRef]
- Suetake, I.; Shinozaki, F.; Miyagawa, J.; Takeshima, H.; Tajima, S. DNMT3L Stimulates the DNA Methylation Activity of DNMT3a and DNMT3b through a Direct Interaction. J. Biol. Chem. 2004, 279, 27816–27823. [Google Scholar] [CrossRef]
- Vlachogiannis, G.; Niederhuth, C.E.; Tuna, S.; Stathopoulou, A.; Viiri, K.; de Rooij, D.G.; Jenner, R.G.; Schmitz, R.J.; Ooi, S.K. The Dnmt3L ADD Domain Controls Cytosine Methylation Establishment during Spermatogenesis. Cell Rep. 2015, 10, 944–956. [Google Scholar] [CrossRef]
- Jin, J.; Guo, Y.; Dong, X.; Liu, J.; He, Y. Methylation-associated silencing of miR-193b improves the radiotherapy sensitivity of esophageal cancer cells by targeting cyclin D1 in areas with zinc deficiency. Radiother. Oncol. 2020, 150, 104–113. [Google Scholar] [CrossRef]
- Mahmood, N.; Rabbani, S.A. DNA Methylation Readers and Cancer: Mechanistic and Therapeutic Applications. Front. Oncol. 2019, 9, 489. [Google Scholar] [CrossRef]
- Hu, Y.-D.; Pang, W.; He, C.-C.; Lu, H.; Liu, W.; Wang, Z.-Y.; Liu, Y.-Q.; Huang, C.-Y.; Jiang, Y.-G. The cognitive impairment induced by zinc deficiency in rats aged 0~2 months related to BDNF DNA methylation changes in the hippocampus. Nutr. Neurosci. 2016, 20, 519–525. [Google Scholar] [CrossRef] [PubMed]
- Dandi, Ε.; Kalamari, A.; Touloumi, O.; Lagoudaki, R.; Nousiopoulou, E.; Simeonidou, C.; Spandou, E.; Tata, D.A. Beneficial effects of environmental enrichment on behavior, stress reactivity and synaptophysin/BDNF expression in hippocampus following early life stress. Int. J. Dev. Neurosci. 2018, 67, 19–32. [Google Scholar] [CrossRef] [PubMed]
- Miranda, M.; Morici, J.F.; Zanoni, M.B.; Bekinschtein, P. Brain-Derived Neurotrophic Factor: A Key Molecule for Memory in the Healthy and the Pathological Brain. Front. Cell. Neurosci. 2019, 13, 363. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.-G.; Wang, Y.-H.; Zhang, H.; Wang, Z.-Y.; Liu, Y.-Q. Effects of early-life zinc deficiency on learning and memory in offspring and the changes in DNA methylation patterns. Nutr. Neurosci. 2020, 1–10. [Google Scholar] [CrossRef]
- Asai, A.; Konno, M.; Koseki, J.; Taniguchi, M.; Vecchione, A.; Ishii, H. One-carbon metabolism for cancer diagnostic and therapeutic approaches. Cancer Lett. 2020, 470, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Clare, C.E.; Brassington, A.H.; Kwong, W.Y.; Sinclair, K.D. One-Carbon Metabolism: Linking Nutritional Biochemistry to Epigenetic Programming of Long-Term Development. Annu. Rev. Anim. Biosci. 2019, 7, 263–287. [Google Scholar] [CrossRef]
- Choi, S.-W.; Friso, S. Epigenetics: A New Bridge between Nutrition and Health. Adv. Nutr. 2010, 1, 8–16. [Google Scholar] [CrossRef]
- Koutmos, M.; Pejchal, R.; Bomer, T.M.; Matthews, R.G.; Smith, J.L.; Ludwig, M.L. Metal active site elasticity linked to activation of homocysteine in methionine synthases. Proc. Natl. Acad. Sci. USA 2008, 105, 3286–3291. [Google Scholar] [CrossRef] [PubMed]
- Jing, M.; Rech, L.; Wu, Y.; Goltz, D.; Taylor, C.G.; House, J.D. Effects of zinc deficiency and zinc supplementation on homocysteine levels and related enzyme expression in rats. J. Trace Elem. Med. Biol. 2015, 30, 77–82. [Google Scholar] [CrossRef] [PubMed]
- Danchin, A.; Sekowska, A.; You, C. One-carbon metabolism, folate, zinc and translation. Microb. Biotechnol. 2020, 13, 899–925. [Google Scholar] [CrossRef]
- Castro, C.; Millian, N.S.; Garrow, T.A. Liver betaine-homocysteine S-methyltransferase activity undergoes a redox switch at the active site zinc. Arch. Biochem. Biophys. 2008, 472, 26–33. [Google Scholar] [CrossRef]
- Wu, L.; Zhou, X.; Linli, H.; He, J.; Huang, L.; Ouyang, Z.; He, L.; Wei, T.; He, Q. Improved Sp1 and Betaine Homocysteine-S-Methyltransferase Expression and Homocysteine Clearance Are Involved in the Effects of Zinc on Oxidative Stress in High-Fat-Diet-Pretreated Mice. Biol. Trace Elem. Res. 2018, 184, 436–441. [Google Scholar] [CrossRef] [PubMed]
- Sawan, C.; Herceg, Z. Histone modifications and cancer. Adv. Genet. 2010, 70, 57–85. [Google Scholar] [CrossRef] [PubMed]
- Wood, A.; Shilatifard, A. Posttranslational Modifications of Histones by Methylation. Adv. Protein Chem. 2004, 67, 201–222. [Google Scholar] [CrossRef]
- Qian, C.; Zhou, M.-M. SET domain protein lysine methyltransferases: Structure, specificity and catalysis. Cell. Mol. Life Sci. CMLS 2006, 63, 2755–2763. [Google Scholar] [CrossRef]
- Elgin, S.C.; Reuter, G. Position-Effect Variegation, Heterochromatin Formation, and Gene Silencing in Drosophila. Cold Spring Harb. Perspect. Biol. 2013, 5, a017780. [Google Scholar] [CrossRef] [PubMed]
- Min, J.; Zhang, X.; Cheng, X.; Grewal, S.I.; Xu, R.-M. Structure of the SET domain histone lysine methyltransferase Clr4. Nat. Struct. Biol. 2002, 9, 828–832. [Google Scholar] [CrossRef]
- Zhang, X.; Tamaru, H.; Khan, S.I.; Horton, J.R.; Keefe, L.J.; Selker, E.U.; Cheng, X. Structure of the Neurospora SET Domain Protein DIM-5, a Histone H3 Lysine Methyltransferase. Cell 2002, 111, 117–127. [Google Scholar] [CrossRef]
- Ruesch, C.E.; Ramakrishnan, M.; Park, J.; Li, N.; Chong, H.S.; Zaman, R.; Joska, T.M.; Belden, W.J. The Histone H3 Lysine 9 Methyltransferase DIM-5 Modifies Chromatin atfrequencyand Represses Light-Activated Gene Expression. G3 Genes Genomes Genet. 2015, 5, 93–101. [Google Scholar] [CrossRef]
- Dillon, S.C.; Zhang, X.; Trievel, R.C.; Cheng, X. The SET-domain protein superfamily: Protein lysine methyltransferases. Genome Biol. 2005, 6, 227. [Google Scholar] [CrossRef]
- Li, B.; Zheng, Y.; Yang, L. The Oncogenic Potential of SUV39H2: A Comprehensive and Perspective View. J. Cancer 2019, 10, 721–729. [Google Scholar] [CrossRef] [PubMed]
- Hublitz, P.; Albert, M.; Hfmpeters, A. Mechanisms of transcriptional repression by histone lysine methylation. Int. J. Dev. Biol. 2009, 53, 335–354. [Google Scholar] [CrossRef] [PubMed]
- Hosseini, A.; Minucci, S. A comprehensive review of lysine-specific demethylase 1 and its roles in cancer. Epigenomics 2017, 9, 1123–1142. [Google Scholar] [CrossRef]
- Yan, M.; Yang, X.; Wang, H.; Shao, Q. The critical role of histone lysine demethylase KDM2B in cancer. Am. J. Transl. Res. 2018, 10, 2222–2233. [Google Scholar]
- Karytinos, A.; Forneris, F.; Profumo, A.; Ciossani, G.; Battaglioli, E.; Binda, C.; Mattevi, A. A Novel Mammalian Flavin-dependent Histone Demethylase. J. Biol. Chem. 2009, 284, 17775–17782. [Google Scholar] [CrossRef] [PubMed]
- Burg, J.M.; Link, J.E.; Morgan, B.S.; Heller, F.J.; Hargrove, A.E.; McCafferty, D.G. KDM1 class flavin-dependent protein lysine demethylases. Biopolymers 2015, 104, 213–246. [Google Scholar] [CrossRef]
- Zhang, Q.; Qi, S.; Xu, M.; Yu, L.; Tao, Y.; Deng, Z.; Wu, W.; Li, J.; Chen, Z.; Wong, J. Structure-function analysis reveals a novel mechanism for regulation of histone demethylase LSD2/AOF1/KDM1b. Cell Res. 2012, 23, 225–241. [Google Scholar] [CrossRef] [PubMed]
- Isshiki, Y.; Nakajima-Takagi, Y.; Oshima, M.; Aoyama, K.; Rizk, M.; Kurosawa, S.; Saraya, A.; Kondo, T.; Sakaida, E.; Nakaseko, C.; et al. KDM2B in polycomb repressive complex 1.1 functions as a tumor suppressor in the initiation of T-cell leukemogenesis. Blood Adv. 2019, 3, 2537–2549. [Google Scholar] [CrossRef]
- Chrun, E.S.; Modolo, F.; Daniel, F.I. Histone modifications: A review about the presence of this epigenetic phenomenon in carcinogenesis. Pathol. Res. Pract. 2017, 213, 1329–1339. [Google Scholar] [CrossRef]
- Calcagno, D.Q.; Wisnieski, F.; Mota, E.R.D.S.; De Sousa, S.B.M.; Da Silva, J.M.C.; Leal, M.F.; Gigek, C.O.; Santos, L.C.; Rasmussen, L.T.; Assumpção, P.P.; et al. Role of histone acetylation in gastric cancer: Implications of dietetic compounds and clinical perspectives. Epigenomics 2019, 11, 349–362. [Google Scholar] [CrossRef]
- Zhou, Z.; Lin, Z.; Pang, X.; Tariq, M.A.; Ao, X.; Li, P.; Wang, J. Epigenetic regulation of long non-coding RNAs in gastric cancer. Oncotarget 2017, 9, 19443–19458. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.; Marmorstein, R. Histone acetyltransferases: Rising ancient counterparts to protein kinases. Biopolymers 2012, 99, 98–111. [Google Scholar] [CrossRef]
- Yang, X.-J.; Ullah, M.F. MOZ and MORF, two large MYSTic HATs in normal and cancer stem cells. Oncogene 2007, 26, 5408–5419. [Google Scholar] [CrossRef]
- Katsumoto, T.; Yoshida, N.; Kitabayashi, I. Roles of the histone acetyltransferase monocytic leukemia zinc finger protein in normal and malignant hematopoiesis. Cancer Sci. 2008, 99, 1523–1527. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.-J. MOZ and MORF acetyltransferases: Molecular interaction, animal development and human disease. Biochim. Biophys. Acta (BBA) Mol. Cell Res. 2015, 1853, 1818–1826. [Google Scholar] [CrossRef]
- Seto, E.; Yoshida, M. Erasers of Histone Acetylation: The Histone Deacetylase Enzymes. Cold Spring Harb. Perspect. Biol. 2014, 6, a018713. [Google Scholar] [CrossRef]
- D’Mello, S.R. Histone deacetylases 1, 2 and 3 in nervous system development. Curr. Opin. Pharmacol. 2020, 50, 74–81. [Google Scholar] [CrossRef] [PubMed]
- Asfaha, Y.; Schrenk, C.; Avelar, L.A.A.; Hamacher, A.; Pflieger, M.; Kassack, M.U.; Kurz, T. Recent advances in class IIa histone deacetylases research. Bioorg. Med. Chem. 2019, 27, 115087. [Google Scholar] [CrossRef]
- Athanasopoulos, D.; Karagiannis, G.; Tsolaki, M. Recent Findings in Alzheimer Disease and Nutrition Focusing on Epigenetics. Adv. Nutr. 2016, 7, 917–927. [Google Scholar] [CrossRef] [PubMed]
- Michan, S.; Sinclair, D. Sirtuins in mammals: Insights into their biological function. Biochem. J. 2007, 404, 1–13. [Google Scholar] [CrossRef]
- Lombardi, P.M.; Cole, K.E.; Dowling, D.P.; Christianson, D.W. Structure, mechanism, and inhibition of histone deacetylases and related metalloenzymes. Curr. Opin. Struct. Biol. 2011, 21, 735–743. [Google Scholar] [CrossRef] [PubMed]
- Lindskog, S. Structure and mechanism of carbonic anhydrase. Pharmacol. Ther. 1997, 74, 1–20. [Google Scholar] [CrossRef]
- Liu, H.; Zhang, F.; Wang, K.; Tang, X.; Wu, R. Conformational dynamics and allosteric effect modulated by the unique zinc-binding motif in class IIa HDACs. Phys. Chem. Chem. Phys. 2019, 21, 12173–12183. [Google Scholar] [CrossRef]
- Yahaya, T. Role of Epigenetics in the Pathogenesis and Management of Type 2 Diabetes Mellitus. Transl. Univ. Toledo J. Med. Sci. 2019, 6, 20–28. [Google Scholar] [CrossRef]
- Yamato, E. High dose of histone deacetylase inhibitors affects insulin secretory mechanism of pancreatic beta cell line. Endocr. Regul. 2018, 52, 21–26. [Google Scholar] [CrossRef] [PubMed]
- Christensen, D.P.; Dahllöf, M.; Lundh, M.; Rasmussen, D.N.; Nielsen, M.D.; Billestrup, N.; Grunnet, L.G.; Mandrup-Poulsen, T. Histone Deacetylase (HDAC) Inhibition as a Novel Treatment for Diabetes Mellitus. Mol. Med. 2011, 17, 378–390. [Google Scholar] [CrossRef]
- Chen, J.; Li, D.; Li, W.; Yin, J.; Zhang, Y.; Yuan, Z.; Gao, C.; Liu, F.; Jiang, Y. Design, synthesis and anticancer evaluation of acridine hydroxamic acid derivatives as dual Topo and HDAC inhibitors. Bioorg. Med. Chem. 2018, 26, 3958–3966. [Google Scholar] [CrossRef]
- Yuan, Z.; Sun, Q.; Li, D.; Miao, S.; Chen, S.; Song, L.; Gao, C.; Chen, Y.; Tan, C.; Jiang, Y. Design, synthesis and anticancer potential of NSC-319745 hydroxamic acid derivatives as DNMT and HDAC inhibitors. Eur. J. Med. Chem. 2017, 134, 281–292. [Google Scholar] [CrossRef] [PubMed]
- Heaton, S.M.; Borg, N.A.; Dixit, V.M. Ubiquitin in the activation and attenuation of innate antiviral immunity. J. Exp. Med. 2016, 213, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Hammond-Martel, I.; Yu, H.; Affar, E.B. Roles of ubiquitin signaling in transcription regulation. Cell. Signal. 2012, 24, 410–421. [Google Scholar] [CrossRef]
- Callis, J. The Ubiquitination Machinery of the Ubiquitin System. Arab. Book 2014, 12, e0174. [Google Scholar] [CrossRef] [PubMed]
- Borden, K.L.; Freemont, P.S. The RING finger domain: A recent example of a sequence—Structure family. Curr. Opin. Struct. Biol. 1996, 6, 395–401. [Google Scholar] [CrossRef]
- Uckelmann, M.; Sixma, T.K. Histone ubiquitination in the DNA damage response. DNA Repair 2017, 56, 92–101. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Zee, B.M.; Wang, Y.; Garcia, B.A.; Dou, Y. The RING Finger Protein MSL2 in the MOF Complex Is an E3 Ubiquitin Ligase for H2B K34 and Is Involved in Crosstalk with H3 K4 and K79 Methylation. Mol. Cell 2011, 43, 132–144. [Google Scholar] [CrossRef] [PubMed]
- Lang, G.; Bonnet, J.; Umlauf, D.; Karmodiya, K.; Koffler, J.; Stierle, M.; Devys, D.; Tora, L. The Tightly Controlled Deubiquitination Activity of the Human SAGA Complex Differentially Modifies Distinct Gene Regulatory Elements. Mol. Cell. Biol. 2011, 31, 3734–3744. [Google Scholar] [CrossRef]
- Morgan, M.T.; Haj-Yahya, M.; Ringel, A.E.; Bandi, P.; Brik, A.; Wolberger, C. Structural basis for histone H2B deubiquitination by the SAGA DUB module. Science 2016, 351, 725–728. [Google Scholar] [CrossRef]
- Koehler, C.; Bonnet, J.; Stierle, M.; Romier, C.; Devys, D.; Kieffer, B. DNA Binding by Sgf11 Protein Affects Histone H2B Deubiquitination by Spt-Ada-Gcn5-Acetyltransferase (SAGA). J. Biol. Chem. 2014, 289, 8989–8999. [Google Scholar] [CrossRef]
- Hudson, N.O.; Buck-Koehntop, B.A. Zinc Finger Readers of Methylated DNA. Molecules 2018, 23, 2555. [Google Scholar] [CrossRef]
- Cassandri, M.; Smirnov, A.; Novelli, F.; Pitolli, C.; Agostini, M.; Malewicz, M.; Melino, G.; Raschellà, G. Zinc-finger proteins in health and disease. Cell Death Discov. 2017, 3, 17071. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhang, X.; Blumenthal, R.M.; Cheng, X. A common mode of recognition for methylated CpG. Trends Biochem. Sci. 2013, 38, 177–183. [Google Scholar] [CrossRef]
- Kribelbauer, J.F.; Lu, X.-J.; Rohs, R.; Mann, R.S.; Bussemaker, H.J. Toward a Mechanistic Understanding of DNA Methylation Readout by Transcription Factors. J. Mol. Biol. 2020, 432, 1801–1815. [Google Scholar] [CrossRef]
- Shimbo, T.; Wade, P.A. Proteins That Read DNA Methylation. Adv. Exp. Med. Biol. 2016, 945, 303–320. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; Xiong, J.; Wang, M.; Yang, N.; Wong, J.; Zhu, B.; Xu, R.-M. Structural Basis for Hydroxymethylcytosine Recognition by the SRA Domain of UHRF2. Mol. Cell 2014, 54, 879–886. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.; Hon, G.C.; Szulwach, K.E.; Song, C.-X.; Zhang, L.; Kim, A.; Li, X.; Dai, Q.; Shen, Y.; Park, B.; et al. Base-Resolution Analysis of 5-Hydroxymethylcytosine in the Mammalian Genome. Cell 2012, 149, 1368–1380. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; D’Alessio, A.C.; Ito, S.; Wang, Z.; Cui, K.; Zhao, K.; Sun, Y.E.; Zhang, Y. Genome-wide analysis of 5-hydroxymethylcytosine distribution reveals its dual function in transcriptional regulation in mouse embryonic stem cells. Genes Dev. 2011, 25, 679–684. [Google Scholar] [CrossRef] [PubMed]
- Yun, M.; Wu, J.; Workman, J.L.; Li, B. Readers of histone modifications. Cell Res. 2011, 21, 564–578. [Google Scholar] [CrossRef] [PubMed]
- Yap, K.L.; Zhou, M.-M. Structure and Mechanisms of Lysine Methylation Recognition by the Chromodomain in Gene Transcription. Biochemistry 2011, 50, 1966–1980. [Google Scholar] [CrossRef]
- Li, H.; Ilin, S.; Wang, W.; Duncan, E.M.; Wysocka, J.; Allis, C.D.; Patel, D.J. Molecular basis for site-specific read-out of histone H3K4me3 by the BPTF PHD finger of NURF. Nat. Cell Biol. 2006, 442, 91–95. [Google Scholar] [CrossRef]
- Jurkowski, T.P.; Ravichandran, M.; Stepper, P. Synthetic epigenetics—Towards intelligent control of epigenetic states and cell identity. Clin. Epigenet. 2015, 7, 18. [Google Scholar] [CrossRef]
- Cui, C.; Gan, Y.; Gu, L.; Wilson, J.; Liu, Z.; Zhang, B.; Deng, D. P16-specific DNA methylation by engineered zinc finger methyltransferase inactivates gene transcription and promotes cancer metastasis. Genome Biol. 2015, 16, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Meng, H.-X.; Miao, S.-S.; Chen, K.; Li, H.-N.; Yao, G.; Geng, J.; Wang, H.; Shi, Q.-T.; He, J.; Mao, X.; et al. Association of p16 as Prognostic Factors for Oropharyngeal Cancer: Evaluation of p16 in 1470 Patients for a 16 Year Study in Northeast China. BioMed Res. Int. 2018, 2018, 1–8. [Google Scholar] [CrossRef]
- Chen, H.; Kazemier, H.G.; De Groote, M.L.; Ruiters, M.H.J.; Xu, G.-L.; Rots, M.G. Induced DNA demethylation by targeting Ten-Eleven Translocation 2 to the human ICAM-1 promoter. Nucleic Acids Res. 2014, 42, 1563–1574. [Google Scholar] [CrossRef] [PubMed]



| ZBD | Function | Example(s) | References |
|---|---|---|---|
| ADD | Allosteric control of the catalytic domain | DNMT3a/3b/3L | [31,41,57] |
| BTB/POZ | Readers of fully methylated DNA (double methylated CpG islands) | Kaiso (ZBTB33), ZBTB4, ZBTB38. | [10,117] |
| CXXC | Substrate recognition and self-regulation | DNMT1. | [49,50] |
| JmjC | H3K36me2 demethylation | KDM2B | [82] |
| KRAB | Reads fully methylated DNA (double methylated CpG islands) | ZFP57 | [10] |
| MYST | HAT activity (H3K9/K14 acetylation) | MOZ, MORF | [94] |
| PHD | Reads a broad range of histone marks and has E3-ubiquitin ligase activity. | UHRF1, UHRF2, BPTF | [11,40] |
| RING | E3-ubiquitin ligase | RNF168, PRC1, RNF20-RNF40 complex | [41] |
| SET | H3K4 methyltransferase activity | SET1A, SET1B | [45,76] |
| SRA | Reads hemimethylated DNA. | UHRF1, UHRF2. | [121,122] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yusuf, A.P.; Abubakar, M.B.; Malami, I.; Ibrahim, K.G.; Abubakar, B.; Bello, M.B.; Qusty, N.; Elazab, S.T.; Imam, M.U.; Alexiou, A.; et al. Zinc Metalloproteins in Epigenetics and Their Crosstalk. Life 2021, 11, 186. https://doi.org/10.3390/life11030186
Yusuf AP, Abubakar MB, Malami I, Ibrahim KG, Abubakar B, Bello MB, Qusty N, Elazab ST, Imam MU, Alexiou A, et al. Zinc Metalloproteins in Epigenetics and Their Crosstalk. Life. 2021; 11(3):186. https://doi.org/10.3390/life11030186
Chicago/Turabian StyleYusuf, Abdurrahman Pharmacy, Murtala Bello Abubakar, Ibrahim Malami, Kasimu Ghandi Ibrahim, Bilyaminu Abubakar, Muhammad Bashir Bello, Naeem Qusty, Sara T. Elazab, Mustapha Umar Imam, Athanasios Alexiou, and et al. 2021. "Zinc Metalloproteins in Epigenetics and Their Crosstalk" Life 11, no. 3: 186. https://doi.org/10.3390/life11030186
APA StyleYusuf, A. P., Abubakar, M. B., Malami, I., Ibrahim, K. G., Abubakar, B., Bello, M. B., Qusty, N., Elazab, S. T., Imam, M. U., Alexiou, A., & Batiha, G. E.-S. (2021). Zinc Metalloproteins in Epigenetics and Their Crosstalk. Life, 11(3), 186. https://doi.org/10.3390/life11030186









