Plant Recognition Using Morphological Feature Extraction and Transfer Learning over SVM and AdaBoost
Abstract
1. Introduction
- Proposing a robust, precise, and fast plant-species recognition model;
- Making use of morphology-based features from leaf images with low dimensionality;
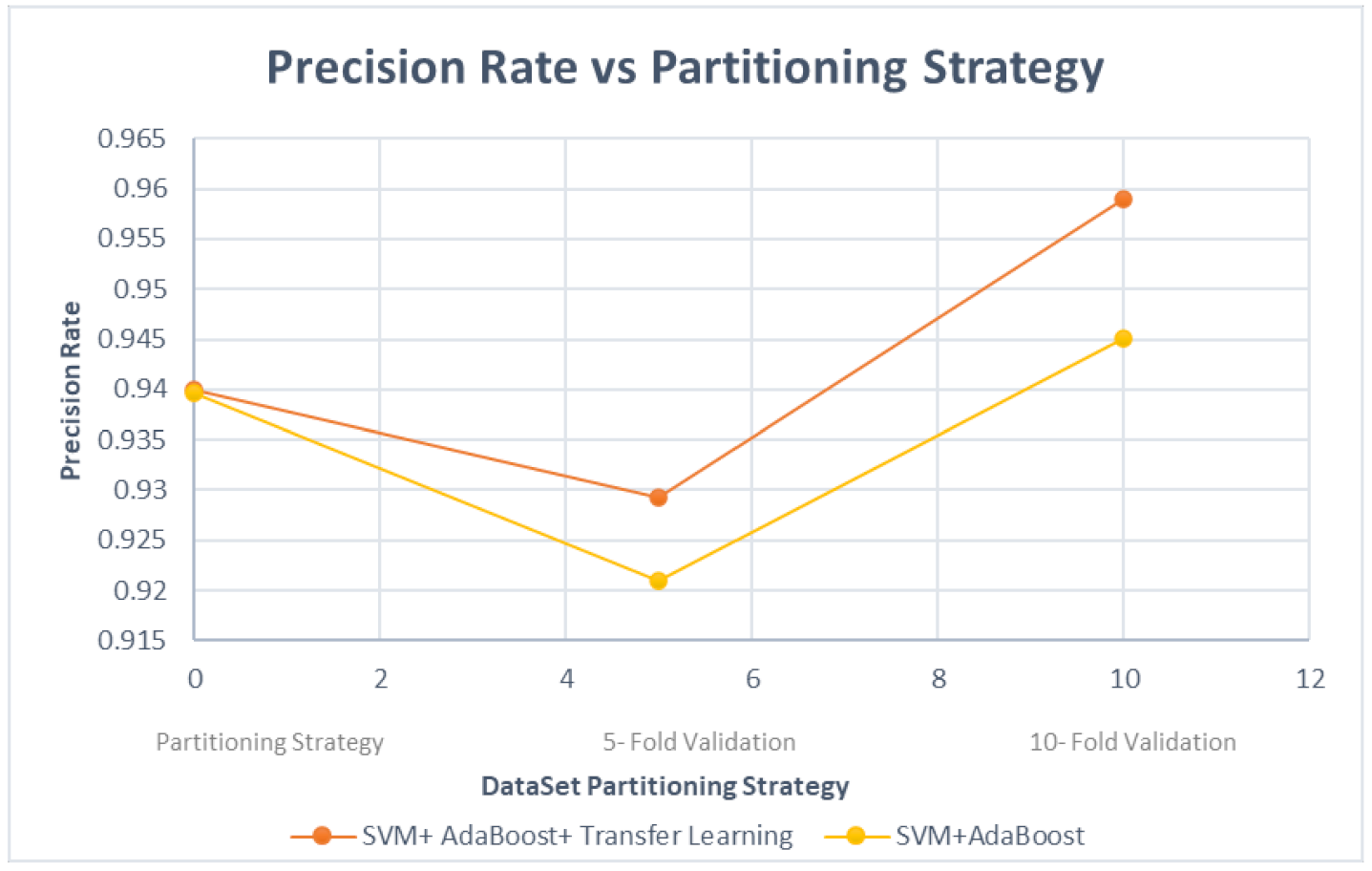
- Using features by transfer learning from low-dimensional ConvNet architecture;
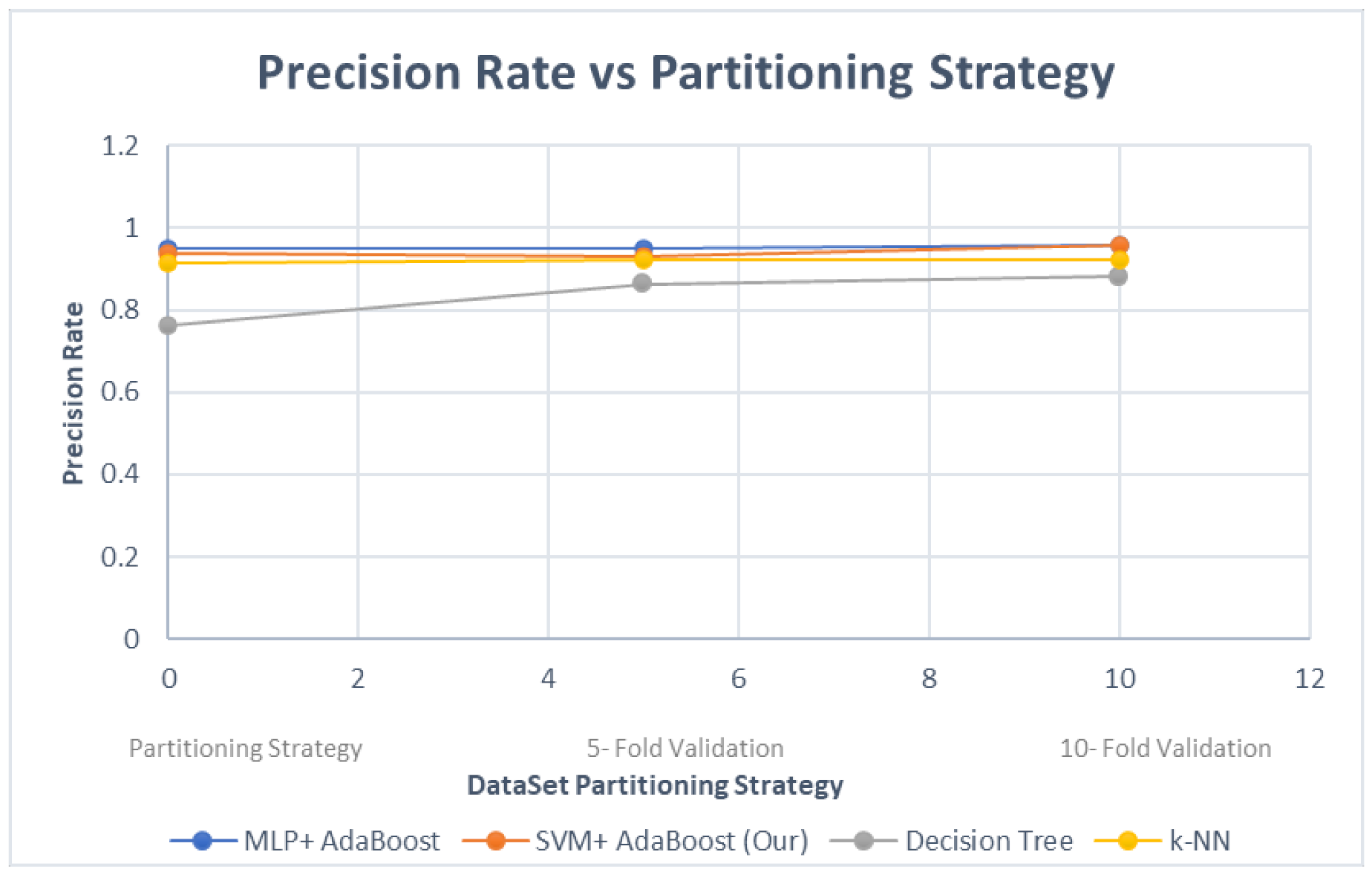
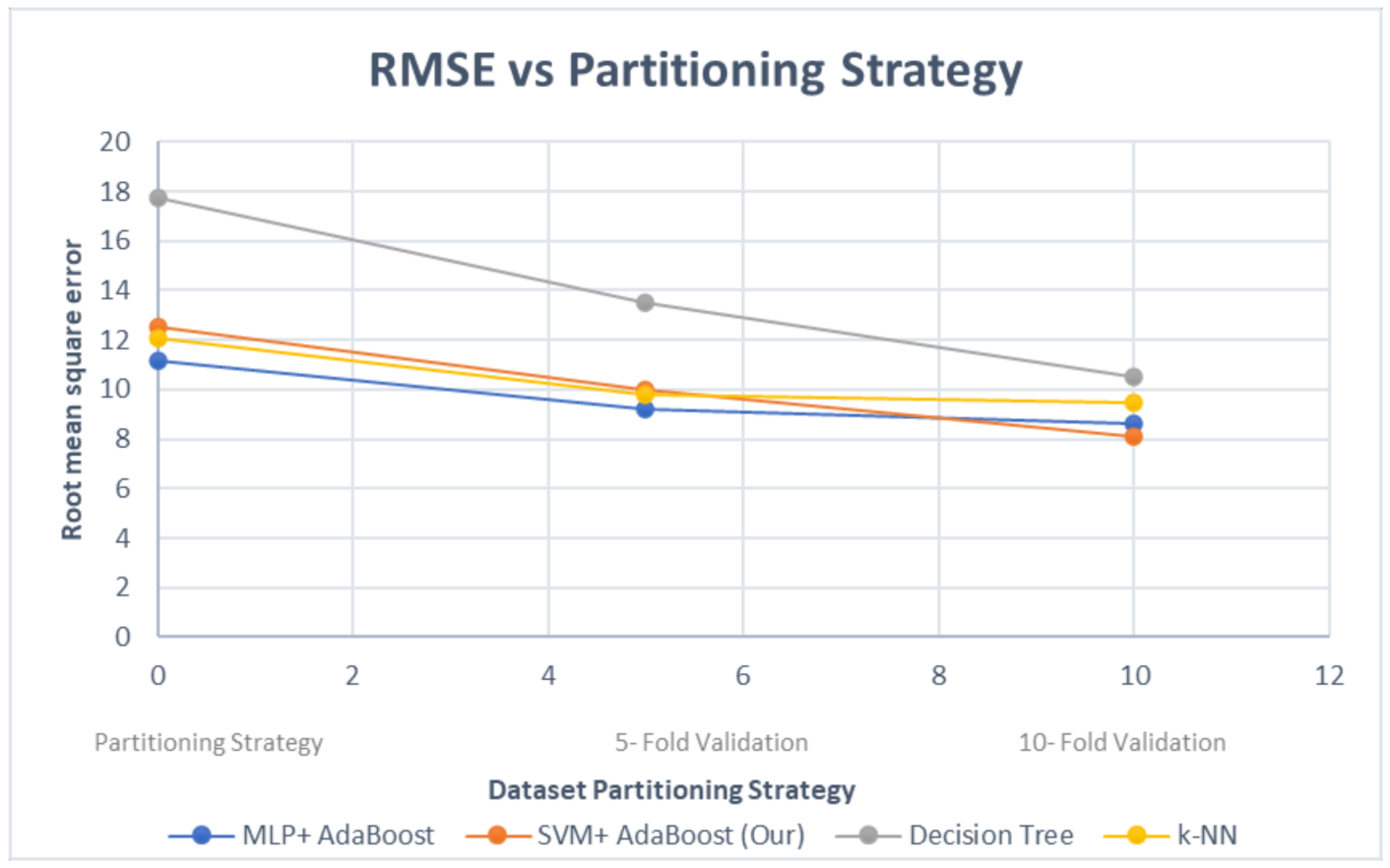
- Evaluation of different classifiers using controlled comparison;
- Enhancing the classification results via adaptive boosting.
1.1. Literature Review
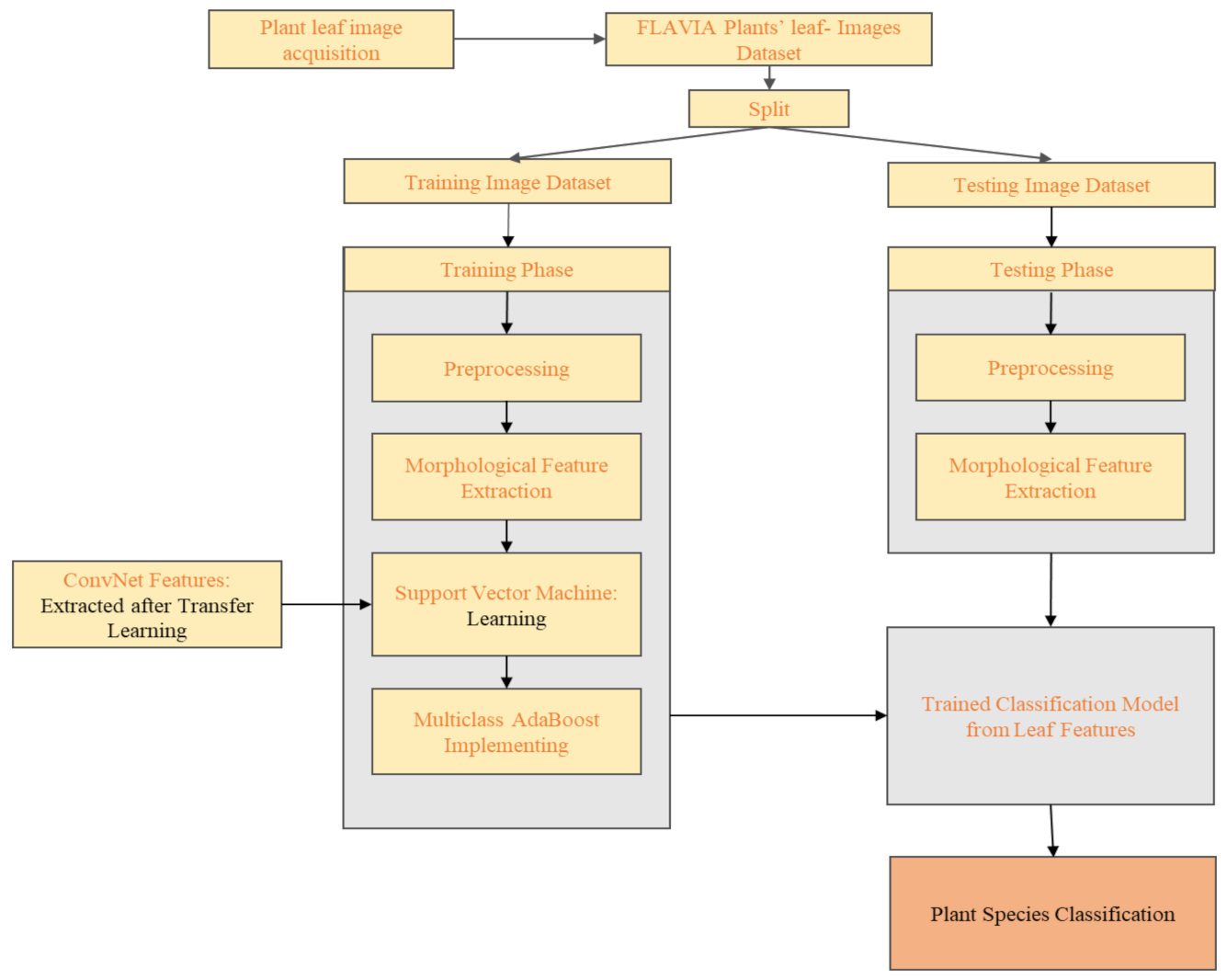
2. Proposed Methodology
2.1. Support Vector Machine (SVM)
2.2. Adaptive Boosting (AdaBoost)
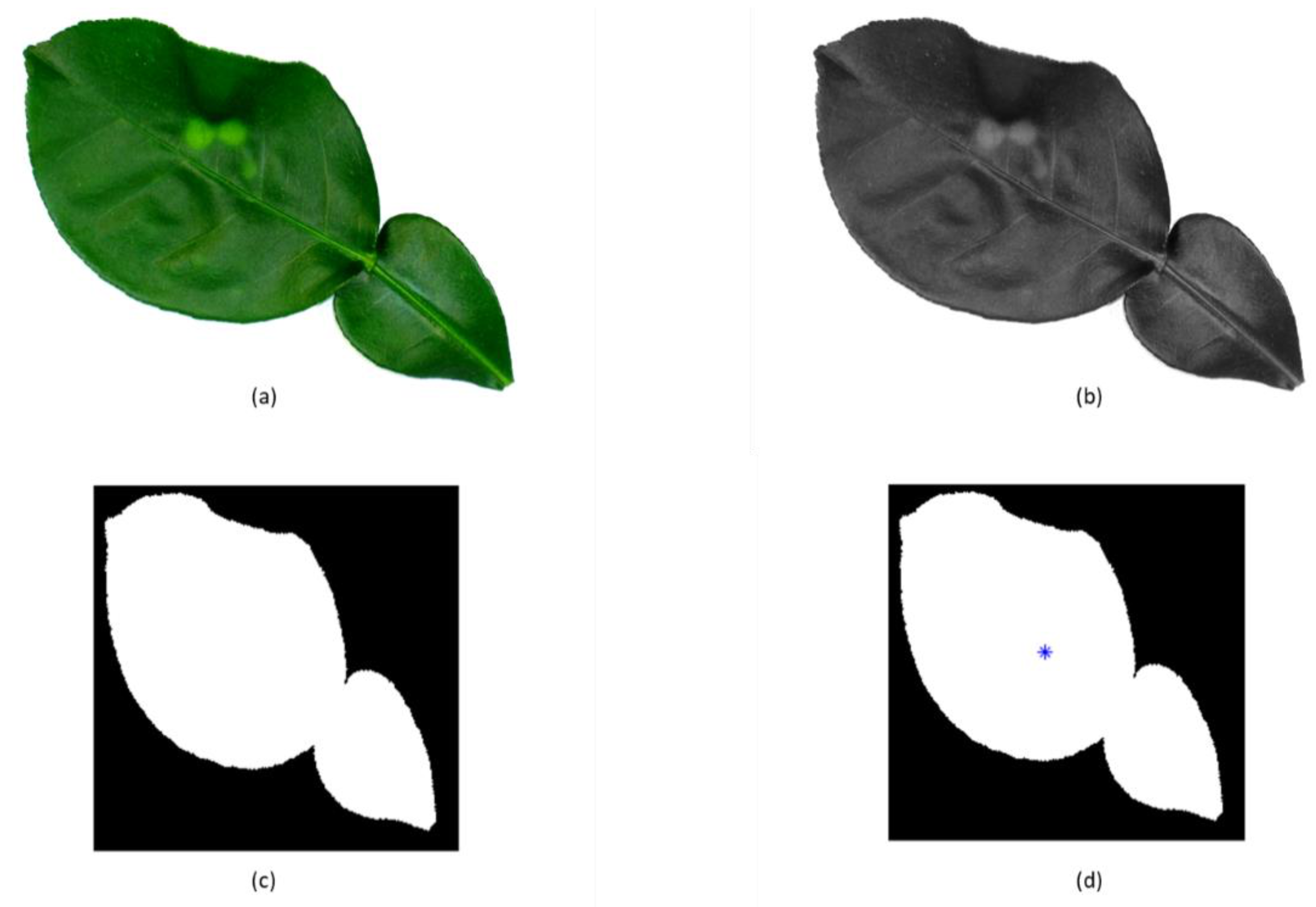
2.3. Feature Extraction and Selection
2.4. Model Highlights
3. Data and Training
4. Evaluation and Results
5. Credibility of the Methodology
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sun, R.; Zhang, M.; Yang, K.; Liu, J. Data enhancement for plant disease classification using generated lesions. Appl. Sci. 2020, 10, 466. [Google Scholar]
- Yang, K.; Zhong, W.; Li, F. Leaf segmentation and classification with a complicated background using deep learning. Agronomy 2020, 10, 1721. [Google Scholar] [CrossRef]
- Zhang, Y.; Cui, J.; Wang, Z.; Kang, J.; Min, Y. Leaf Image Recognition Based on Bag of Features. Appl. Sci. 2020, 10, 5177. [Google Scholar] [CrossRef]
- Azlah, M.A.; Chua, L.S.; Rahmad, F.R.; Abdullah, F.I.; Wan Alwi, S.R. Review on techniques for plant leaf classification and recognition. Computers 2019, 8, 77. [Google Scholar] [CrossRef]
- Kumar, M.; Gupta, S.; Gao, X.Z.; Singh, A. Plant Species Recognition Using Morphological Features and Adaptive Boosting Methodology. IEEE Access 2019, 7, 163912–163918. [Google Scholar] [CrossRef]
- Jeon, W.S.; Rhee, S.Y. Plant leaf recognition using a convolution neural network. Int. J. Fuzzy Log. Intell. Syst. 2017, 17, 26–34. [Google Scholar]
- Pankaja, K.; Suma, V. Leaf Recognition and Classification Using GLCM and Hierarchical Centroid Based Technique. In Proceedings of the 2018 International Conference on Inventive Research in Computing Applications (ICIRCA), Coimbatore, India, 11–12 July 2018; IEEE: Piscataway, NJ, USA; pp. 1190–1194. [Google Scholar]
- Kumar, S. Leaf Color, Area and Edge features-based approach for Identification of Indian Medicinal Plants. Int. J. Comput. Sci. Eng. 2012, 3, 436–442. [Google Scholar]
- Kadir, A.; Nugroho, L.E.; Susanto, A.; Santosa, P.I. Performance improvement of leaf identification system using principal component analysis. Int. J. Adv. Sci. Technol. 2012, 44, 113–124. [Google Scholar]
- Priya, C.A.; Balasaravanan, T.; Thanamani, A.S. An efficient leaf recognition algorithm for plant classification using support vector machine. In Proceedings of the InInternational conference on pattern recognition, informatics and medical engineering (PRIME-2012), Salem, India, 21–23 March 2012; IEEE: Piscataway, NJ, USA; pp. 428–432. [Google Scholar]
- Anami, B.S.; Nandyal, S.S.; Govardhan, A. A combined color, texture and edge features-based approach for identification and classification of indian medicinal plants. Int. J. Comput. Appl. 2010, 6, 45–51. [Google Scholar] [CrossRef]
- Sambhaji, E.S.; Andore, D.B. Leaf recognition algorithm using neural network-based image processing. Asian J. Eng. Technol. Innov. 2014, 2, 10–16. [Google Scholar]
- Sun, Y.; Liu, Y.; Wang, G.; Zhang, H. Deep learning for plant identification in natural environment. Comput. Intell. Neurosci. 2017, 22, 2017. [Google Scholar] [CrossRef] [PubMed]
- Cortes, C.; Vapnik, V. Support-vector networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- Duan, K.B.; Keerthi, S.S. Which is the best multiclass SVM method? An empirical study. In International Workshop on Multiple Classifier Systems, Proceedings of the Multiple Classifier Systems; Springer: Berlin/Heidelberg, Germany, 2005; pp. 278–285. [Google Scholar]
- Hastie, T.; Tibshirani, R. Classification by pairwise coupling. In Advances in Neural Information Processing Systems; MIT Press: Cambridge, MA, USA, 1998; pp. 507–513. [Google Scholar]
- Freund, Y.; Schapire, R.E. A decision-theoretic generalization of on-line learning and an application to boosting. J. Comput. Syst. Sci. 1997, 55, 119–139. [Google Scholar] [CrossRef]
- Boosting Algorithms: AdaBoost, Gradient Boosting and XGBoost. 5 May 2018. Available online: hackernoon.com (accessed on 4 January 2020).
- Hastie, T.; Rosset, S.; Zhu, J.; Zou, H. Multi-class adaboost. Stat. Its Interface 2009, 2, 349–360. [Google Scholar] [CrossRef]
- Kim, T.H.; Park, D.C.; Woo, D.M.; Jeong, T.; Min, S.Y. Multi-class classifier-based adaboost algorithm. In International Conference on Intelligent Science and Intelligent Data Engineering, Proceedings of the Intelligent Science and Intelligent Data Engineering; Springer: Berlin/Heidelberg, Germany, 2011; pp. 122–127. [Google Scholar]
- CS231n: Convolutional Neural Networks for Visual Recognition. Available online: http://cs231n.github.io/transfer-learning/ (accessed on 10 January 2021).
- Yosinski, J.; Clune, J.; Bengio, Y.; Lipson, H. How transferable are features in deep neural networks? arXiv 2014, arXiv:1411.1792. [Google Scholar]
- Plant Leaf Recognition. Available online: http://cs229.stanford.edu/proj2016/report/LiuHuang-PlantLeafRecognition-report.pdf (accessed on 5 January 2021).
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Boston, MA, USA, 7–12 June 2015. [Google Scholar]
- Li, X.; Wang, L.; Sung, E. AdaBoost with SVM-based component classifiers. Eng. Appl. Artif. Intell. 2008, 21, 785–795. [Google Scholar] [CrossRef]
- Govindaraj, D. Can Boosting with SVM as Week Learners Help? arXiv 2016, preprint. arXiv:1604.05242. [Google Scholar]
- García, E.; Lozano, F. Boosting Support Vector Machines. MLDM Posters. 2007, pp. 153–167. Available online: https://www.sciencedirect.com/science/article/abs/pii/S0957417420301457 (accessed on 10 January 2020).
- Flavia (at a Glance). Available online: http://flavia.sourceforge.net/ (accessed on 2 January 2021).
- Sangeetha, R.; Kalpana, D.B. Identifying efficient kernel function in multiclass support vector machines. Int. J. Comput. Appl. 2011, 28, 18–23. [Google Scholar] [CrossRef]
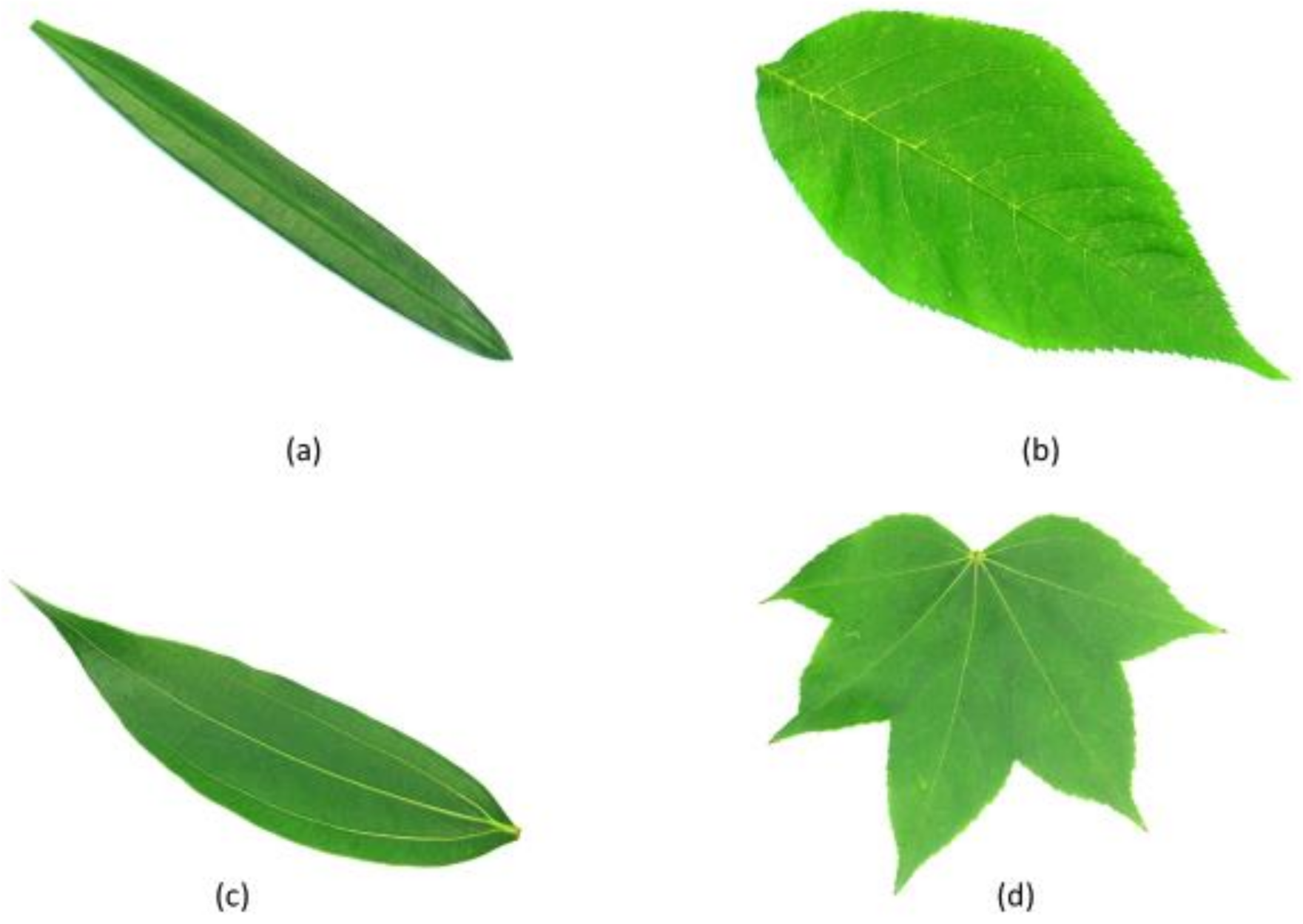
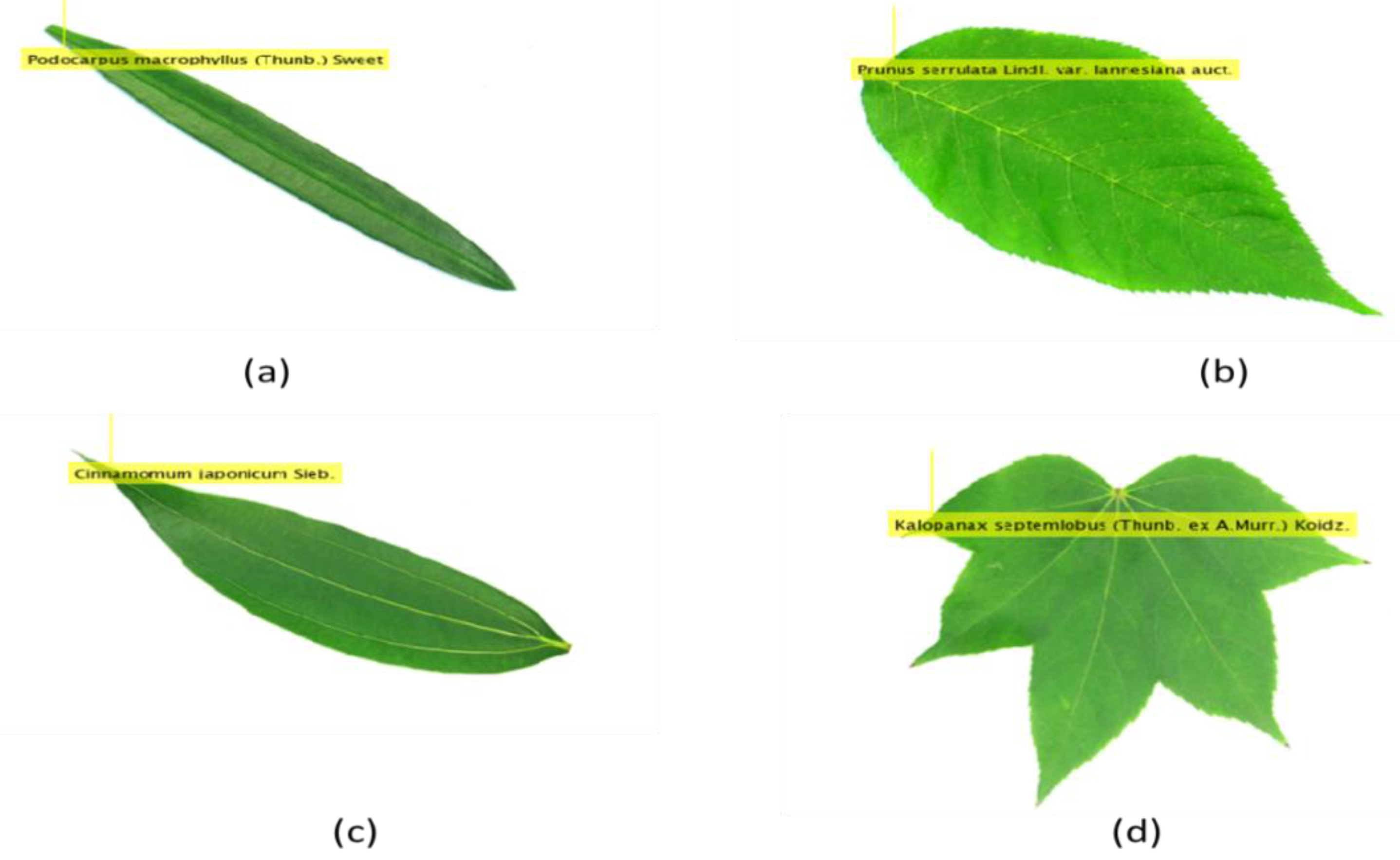






| Feature | Explanation | Formula |
|---|---|---|
| Solidity | Area fraction of the region compared to its convex hull, or the extent of the pixels in the convex hull that is additionally in the area. | |
| Centroid | Center of mass of a region or area. | |
| Perimeter | Path length of the shape externally. | |
| Major axis length | Line connecting one end, called a base point, to the tip of the leaf. | |
| Minor axis length | Line drawn perpendicular to the major axis. | |
| Orientation | Angle between the x-axis and the major axis of the ellipse. | |
| ConvNet via transfer learning | Features extracted from ResNet50 via transfer learning |
| Element | Value/Detail |
|---|---|
| Fundamental algorithm | Support vector machine (polynomial kernel) |
| Secondary algorithm | Multiclass adaptive boosting |
| Dataset | FLAVIA (32 classes distributed among 1907 instances) |
| Features | Morphological and from transfer learning (ResNet50) |
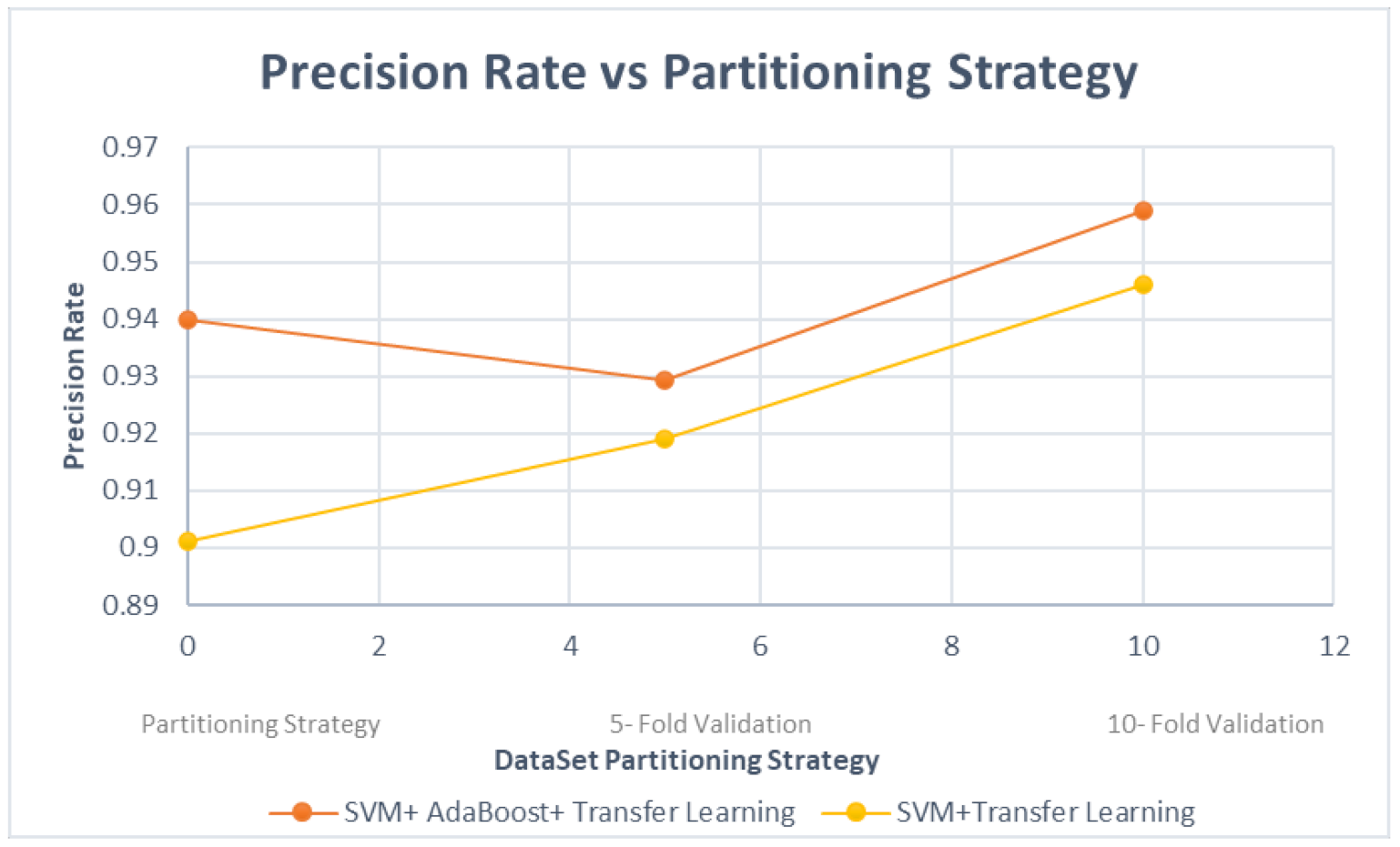
| Partitioning strategy | 80:20 split, five-fold CV and 10-fold CV (best: 10-fold CV) |
| Fundamental model | Support vector machine and adaptive boosting |
| Serial | Research Study | Accuracy (%) |
|---|---|---|
| 1 | CNN-based [6] | 94.00 |
| 2 | CNN-based [13] | 91.78 |
| 3 | Bag of features-based [3] | 94.22 |
| 4 | Our study (hybrid) | 94.72 |
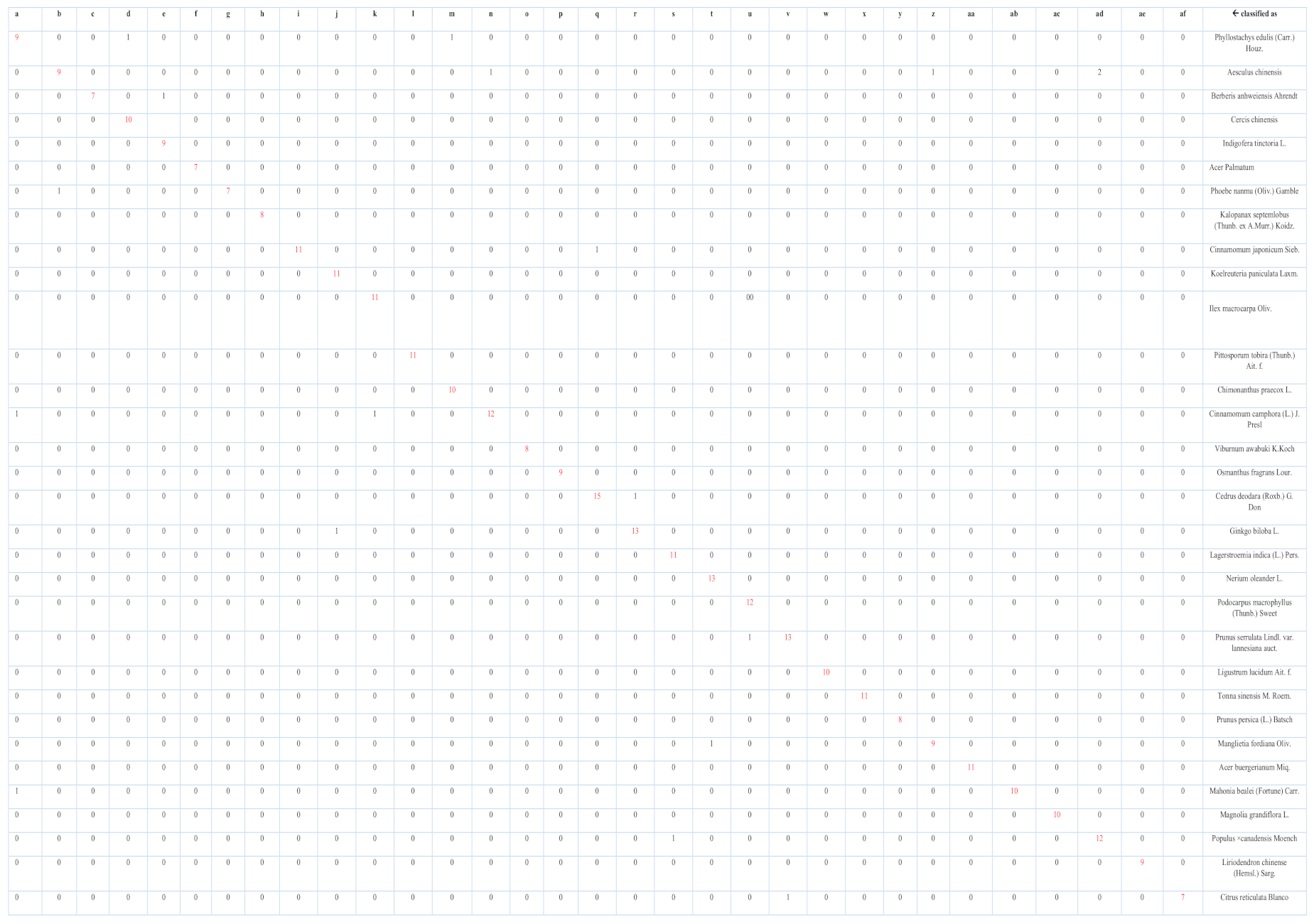
| Serial | Class | Precision | Class | Precision |
|---|---|---|---|---|
| 1 | Phyllostachys edulis (Carr.) Houz. | 1.0000 | Cedrus deodara (Roxb.) G. Don | 0.93750 |
| 2 | Aesculus chinensis | 0.9000 | Ginkgo biloba L. | 0.92860 |
| 3 | Berberis anhweiensis Ahrendt | 1.0000 | Lagerstroemia indica (L.) Pers. | 0.91660 |
| 4 | Cercis chinensis | 0.9090 | Nerium oleander L. | 0.92860 |
| 5 | Indigofera tinctoria L. | 0.9000 | Podocarpus macrophyllus (Thunb.) Sweet | 0.92300 |
| 6 | Acer palmatum | 1.0000 | Prunus s. Lindl var. l. auct. | 0.92857 |
| 7 | Phoebe nanmu (Oliv.) Gamble | 1.0000 | Ligustrum lucidum Ait. f. | 1.00000 |
| 8 | Kalopanax s. Koidz. | 1.0000 | Tonna sinensis M. Roem. | 1.00000 |
| 9 | Cinnamomum japonicum Sieb. | 1.0000 | Prunus persica (L.) Batsch | 1.000000 |
| 10 | Koelreuteria paniculata Laxm. | 0.9166 | Manglietia fordiana Oliv. | 0.90000 |
| 11 | Ilex macrocarpa Oliv. | 0.9166 | Acer buergerianum Miq. | 1.00000 |
| 12 | Pittosporum tobira Ait. f. | 1.0000 | Mahonia bealei (Fortune) Carr. | 0.90900 |
| 13 | Chimonanthus praecox L. | 0.9090 | Magnolia grandiflora L. | 1.00000 |
| 14 | Cinnamomum camphora (L.) J. Presl | 0.9231 | Populus× canadensis Moench | 0.92307 |
| 15 | Viburnum awabuki K. Koch | 1.0000 | Liriodendron chinense Sarg. | 1.00000 |
| 16 | Osmanthus fragrans Lour. | 1.0000 | Citrus reticulata Blanco | 1.00000 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mahajan, S.; Raina, A.; Gao, X.-Z.; Kant Pandit, A. Plant Recognition Using Morphological Feature Extraction and Transfer Learning over SVM and AdaBoost. Symmetry 2021, 13, 356. https://doi.org/10.3390/sym13020356
Mahajan S, Raina A, Gao X-Z, Kant Pandit A. Plant Recognition Using Morphological Feature Extraction and Transfer Learning over SVM and AdaBoost. Symmetry. 2021; 13(2):356. https://doi.org/10.3390/sym13020356
Chicago/Turabian StyleMahajan, Shubham, Akshay Raina, Xiao-Zhi Gao, and Amit Kant Pandit. 2021. "Plant Recognition Using Morphological Feature Extraction and Transfer Learning over SVM and AdaBoost" Symmetry 13, no. 2: 356. https://doi.org/10.3390/sym13020356
APA StyleMahajan, S., Raina, A., Gao, X.-Z., & Kant Pandit, A. (2021). Plant Recognition Using Morphological Feature Extraction and Transfer Learning over SVM and AdaBoost. Symmetry, 13(2), 356. https://doi.org/10.3390/sym13020356






