Peptides for Infectious Diseases: From Probe Design to Diagnostic Microarrays
Abstract
1. Introduction
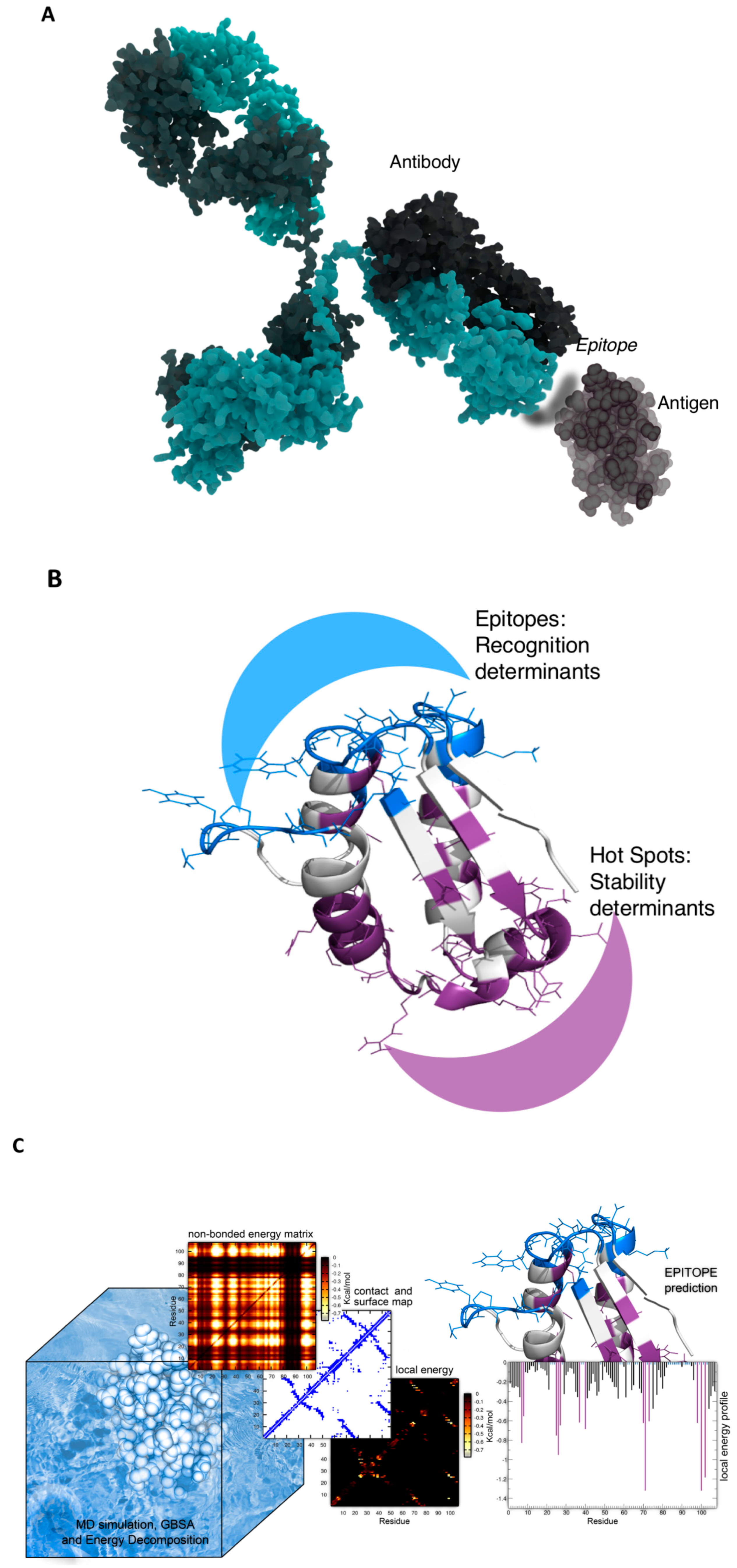
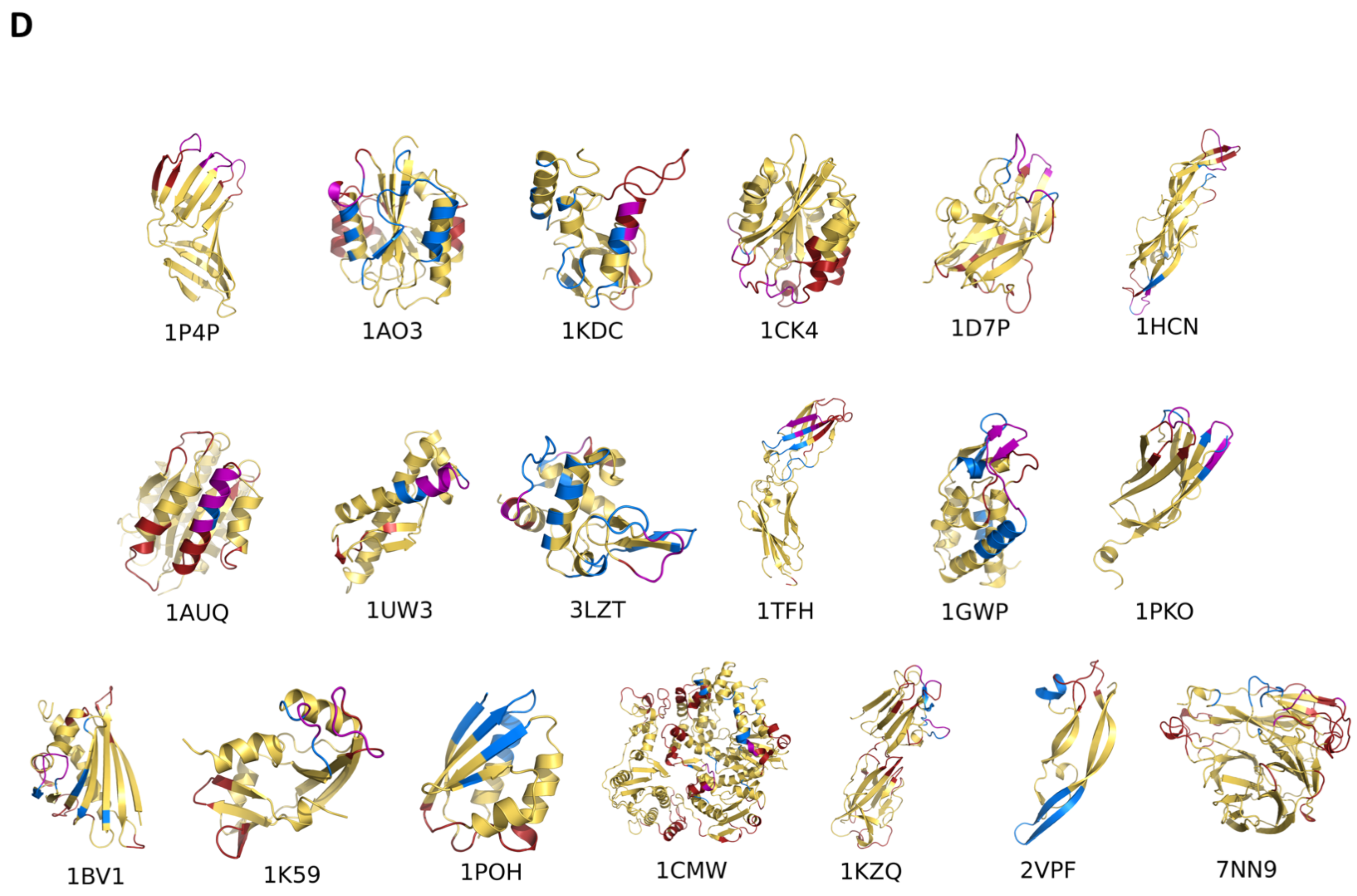
2. Epitope Prediction
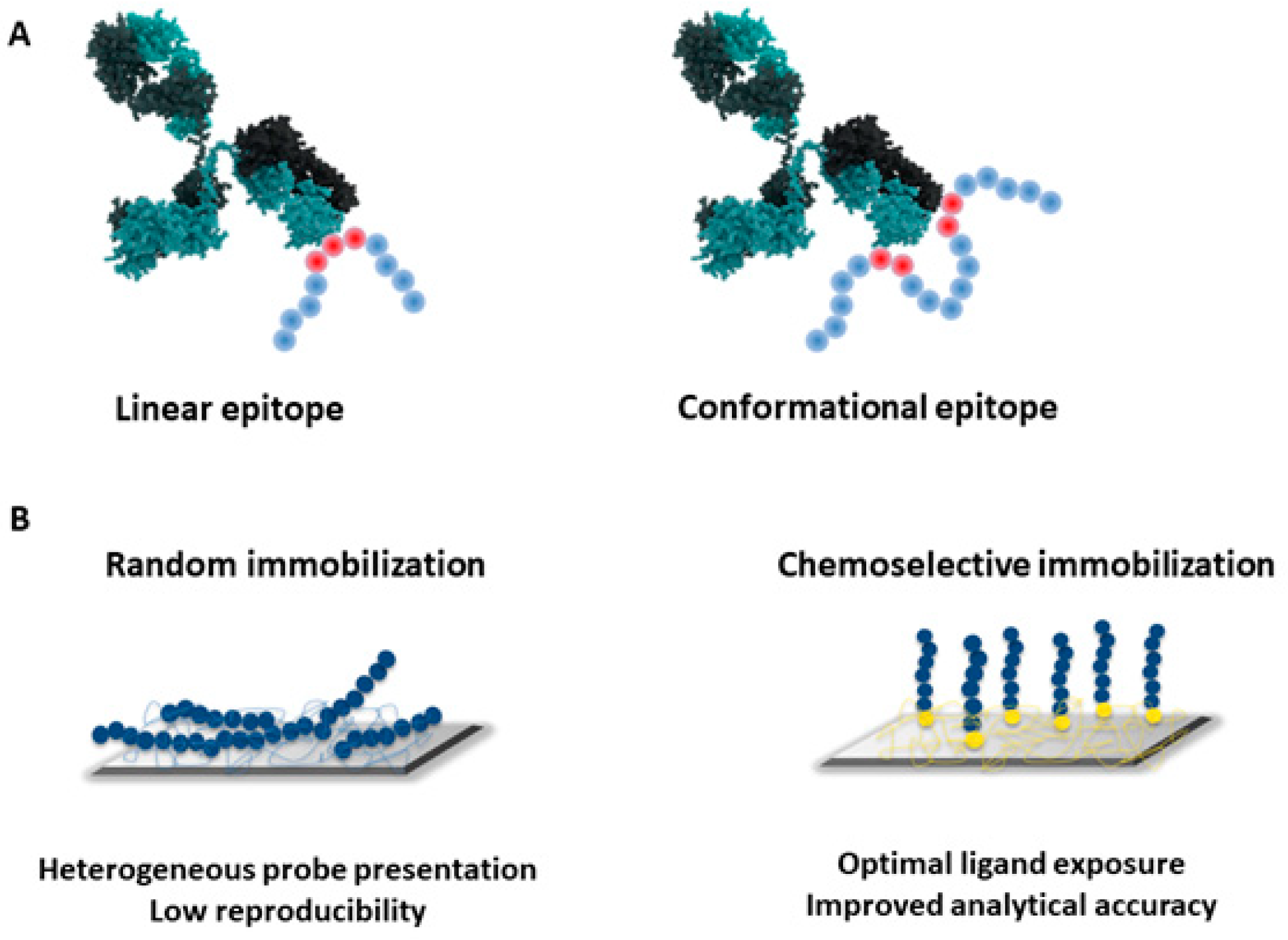
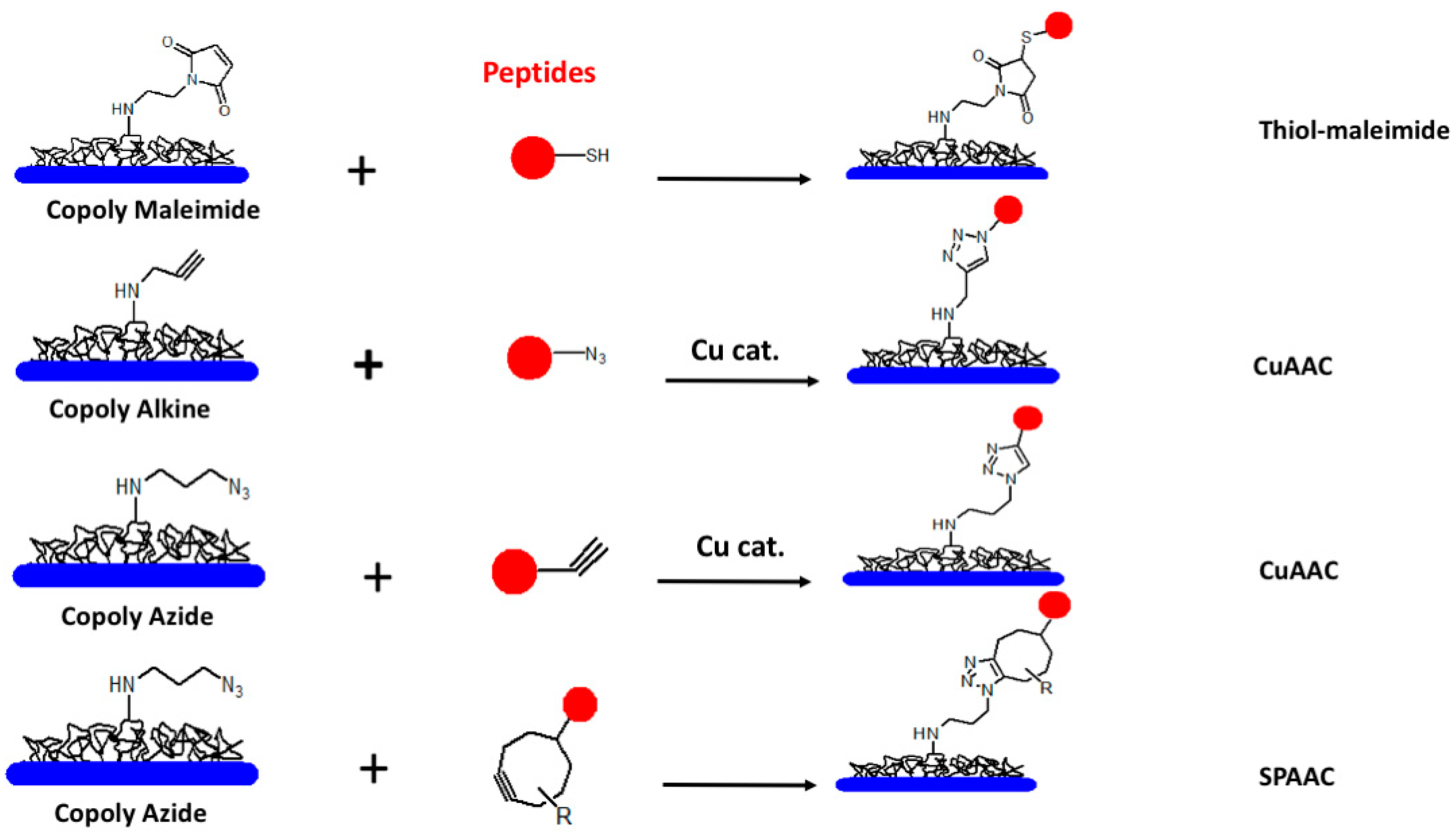
3. The Importance of Probe Presentation: Conformation and Surface Chemistry Matters
4. Conclusions and Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Meng, X.; Wei, J.; Wang, Y.; Zhang, H.; Wang, Z. The role of peptide microarrays in biomedical research. Anal. Methods 2018, 10, 4614–4624. [Google Scholar] [CrossRef]
- Grötzinger, C. Peptide microarrays for medical applications in autoimmunity, infection, and cancer. Meth. Mol. Biol. 2016. [Google Scholar] [CrossRef]
- Li, T.; Liu, D.; Wang, Z. Screening Kinase Inhibitors with a Microarray-Based Fluorescent and Resonance Light Scattering Assay. Anal. Chem. 2010, 82, 3067–3072. [Google Scholar] [CrossRef] [PubMed]
- Carmona, S.J.; Nielsen, M.; Schafer-Nielsen, C.; Mucci, J.; Altcheh, J.; Balouz, V.; Tekiel, V.; Frasch, A.C.; Campetella, O.; Buscaglia, C.A.; et al. Towards High-throughput Immunomics for Infectious Diseases: Use of Next-generation Peptide Microarrays for Rapid Discovery and Mapping of Antigenic Determinants. Mol. Cell. Proteom. 2015, 14, 1871–1884. [Google Scholar] [CrossRef] [PubMed]
- Hecker, M.; Lorenz, P.; Steinbeck, F.; Hong, L.; Riemekasten, G.; Li, Y.; Zettl, U.K.; Thiesen, H.J. Computational analysis of high-density peptide microarray data with application from systemic sclerosis to multiple sclerosis. Autoimmun. Rev. 2012, 11, 180–190. [Google Scholar] [CrossRef] [PubMed]
- Emmadi, R.; Boonyaratanakornkit, J.B.; Selvarangan, R.; Shyamala, V.; Zimmer, B.L.; Williams, L.; Bryant, B.; Schutzbank, T.; Schoonmaker, M.M.; Amos Wilson, J.A.; et al. Molecular methods and platforms for infectious diseases testing a review of FDA-approved and cleared assays. J. Mol. Diagn. 2011, 13, 583–604. [Google Scholar] [CrossRef] [PubMed]
- Tenover, F.C. The role for rapid molecular diagnostic tests for infectious diseases in precision medicine. Expert Rev. Precis. Med. Drug Dev. 2018, 3, 69–77. [Google Scholar] [CrossRef]
- Tettelin, H.; Saunders, N.J.; Heidelberg, J.; Jeffries, A.C.; Nelson, K.E.; Eisen, J.A.; Ketchum, K.A.; Hood, D.W.; Peden, J.F.; Dodson, R.J.; et al. Complete genome sequence of Neisseria meningitidis serogroup B strain MC58. Science 2000, 287, 1809–1815. [Google Scholar] [CrossRef] [PubMed]
- Rappuoli, R. Reverse vaccinology, a genome-based approach to vaccine development. Vaccine 2001, 19, 2688–2691. [Google Scholar] [CrossRef]
- De Gregorio, E.; Rappuoli, R. From empiricism to rational design: A personal perspective of the evolution of vaccine development. Nat. Rev. Immunol. 2014, 14, 505–514. [Google Scholar] [CrossRef]
- Rappuoli, R.; Bottomley, M.J.; D’Oro, U.; Finco, O.; De Gregorio, E. Reverse vaccinology 2.0: Human immunology instructs vaccine antigen design. J. Exp. Med. 2016, 213, 469–481. [Google Scholar] [CrossRef]
- Toseland, C.P.; Clayton, D.J.; McSparron, H.; Hemsley, S.L.; Blythe, M.J.; Paine, K.; Doytchinova, I.A.; Guan, P.; Hattotuwagama, C.K.; Flower, D.R. AntiJen: A quantitative immunology database integrating functional, thermodynamic, kinetic, biophysical, and cellular data. Immunome Res. 2005, 1, 4. [Google Scholar] [CrossRef][Green Version]
- Lefranc, M.P.; Giudicelli, V.; Ginestoux, C.; Jabado-Michaloud, J.; Folch, G.; Bellahcene, F.; Wu, Y.; Gemrot, E.; Brochet, X.; Lane, J.; et al. IMGT, the international ImMunoGeneTics information system. Nucleic Acids Res. 2009, 37, D1006–D1012. [Google Scholar] [CrossRef]
- Ponomarenko, J.; Bui, H.H.; Li, W.; Fusseder, N.; Bourne, P.E.; Sette, A.; Peters, B. ElliPro: A new structure-based tool for the prediction of antibody epitopes. BMC Bioinform. 2008, 9, 514. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Wu, D.; Xu, T.; Wang, X.; Xu, X.; Tao, L.; Li, Y.X.; Cao, Z.W. SEPPA: A computational server for spatial epitope prediction of protein antigens. Nucleic Acids Res. 2009, 37, W612–W616. [Google Scholar] [CrossRef]
- Zhang, G.L.; Khan, A.M.; Srinivasan, K.N.; August, J.T.; Brusic, V. MULTIPRED: A computational system for prediction of promiscuous HLA binding peptides. Nucleic Acids Res. 2005, 33, W172–W179. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Chen, Y.; Wong, H.-S.; Zhou, S.; Mamitsuka, H.; Zhu, S. TEPITOPEpan: Extending TEPITOPE for Peptide Binding Prediction Covering over 700 HLA-DR Molecules. PLoS ONE 2012, 7, e30483. [Google Scholar] [CrossRef]
- Singh, H.; Raghava, G.P.S. ProPred: Prediction of HLA-DR binding sites. Bioinformatics 2001, 17, 1236–1237. [Google Scholar] [CrossRef]
- Ansari, H.R.; Raghava, G.P.S. Identification of conformational B-cell Epitopes in an antigen from its primary sequence. Immunome Res. 2010, 6. [Google Scholar] [CrossRef]
- Dalkas, G.A.; Rooman, M. SEPIa, a knowledge-driven algorithm for predicting conformational B-cell epitopes from the amino acid sequence. BMC Bioinform. 2017, 18, 95. [Google Scholar] [CrossRef]
- Fiorucci, S.; Zacharias, M. Prediction of protein-protein interaction sites using electrostatic desolvation profiles. Biophys. J. 2010, 98, 1921–1930. [Google Scholar] [CrossRef]
- Scarabelli, G.; Morra, G.; Colombo, G. Predicting interaction sited from the energetics of isolated proteins: A new approach to epitope mapping. Biophys. J. 2010, 98, 1966–1975. [Google Scholar] [CrossRef]
- Soriani, M.; Petit, P.; Grifantini, R.; Petracca, R.; Gancitano, G.; Frigimelica, E.; Nardelli, F.; Garcia, C.; Spinelli, S.; Scarabelli, G.; et al. Exploiting antigenic diversity for vaccine design: The chlamydia ArtJ paradigm. J. Biol. Chem. 2010, 285, 30126–30138. [Google Scholar] [CrossRef]
- Tiana, G.; Simona, F.; De Mori, G.M.S.; Broglia, R.A.; Colombo, G. Understanding the determinants of stability and folding of small globular proteins from their energetics. Protein Sci. 2004, 13, 113–124. [Google Scholar] [CrossRef]
- Morra, G.; Colombo, G. Relationship between energy distribution and fold stability: Insights from molecular dynamics simulations of native and mutant proteins. Proteins Struct. Funct. Bioinf. 2008, 72, 660–672. [Google Scholar] [CrossRef]
- Genoni, A.; Morra, G.; Colombo, G. Identification of Domains in Protein Structures from the Analysis of Intramolecular Interactions. J. Phys. Chem. B 2012, 116, 3331–3343. [Google Scholar] [CrossRef]
- Cheng, A.C.; Currie, B.J. Melioidosis: Epidemiology, pathophysiology, and management. Clin. Microbiol. Rev. 2005, 18, 383–416. [Google Scholar] [CrossRef]
- Gourlay, L.J.; Lassaux, P.; Thomas, R.J.; Peri, C.; Conchillo-Sole, O.; Nithichanon, A.; Ferrer-Navarro, M.; Vila, J.; Daura, X.; Lertmemongkolchai, G.; et al. Flagellar subunits as targets for structure-based epitope discovery approaches and melioidosis vaccine development. FEBS J. 2015, 282, 338. [Google Scholar]
- Gourlay, L.J.; Peri, C.; Ferrer-Navarro, M.; Conchillo-Sole, O.; Gori, A.; Rinchai, D.; Thomas, R.J.; Champion, O.L.; Michell, S.L.; Kewcharoenwong, C.; et al. Exploiting the Burkholderia pseudomallei Acute Phase Antigen BPSL2765 for Structure-Based Epitope Discovery/Design in Structural Vaccinology. Chem. Biol. 2013, 20, 1147–1156. [Google Scholar] [CrossRef]
- Lassaux, P.; Peri, C.; Ferrer-Navarro, M.; Gourlay, L.; Gori, A.; Conchillo-Solé, O.; Rinchai, D.; Lertmemongkolchai, G.; Longhi, R.; Daura, X.; et al. A structure-based strategy for epitope discovery in Burkholderia pseudomallei OppA antigen. Structure 2013, 21, 1–9. [Google Scholar] [CrossRef]
- Nithichanon, A.; Rinchai, D.; Gori, A.; Lassaux, P.; Peri, C.; Conchillio-Sole, O.; Ferrer-Navarro, M.; Gourlay, L.J.; Nardini, M.; Vila, J.; et al. Sequence- and Structure-Based Immunoreactive Epitope Discovery for Burkholderia pseudomallei Flagellin. PLoS Negl. Trop. Dis. 2015, 9. [Google Scholar] [CrossRef]
- Gori, A.; Peri, C.; Quilici, G.; Nithichanon, A.; Gaudesi, D.; Longhi, R.; Gourlay, L.; Bolognesi, M.; Lertmemongkolchai, G.; Musco, G.; et al. Flexible vs Rigid Epitope Conformations for Diagnostic- and Vaccine-Oriented Applications: Novel Insights from the Burkholderia pseudomallei BPSL2765 Pa13 Epitope. ACS Infect. Dis. 2016, 2, 221–230. [Google Scholar] [CrossRef]
- Peri, C.; Gagni, P.; Combi, F.; Gori, A.; Chiari, M.; Longhi, R.; Cretich, M.; Colombo, G. Rational epitope design for protein targeting. ACS Chem. Biol. 2013, 8, 397–404. [Google Scholar] [CrossRef]
- Peri, C.; Gori, A.; Gagni, P.; Sola, L.; Girelli, D.; Sottotetti, S.; Cariani, L.; Chiari, M.; Cretich, M.; Colombo, G. Evolving serodiagnostics by rationally designed peptide arrays: The Burkholderia paradigm in Cystic Fibrosis. Sci. Rep. 2016, 6, 32873. [Google Scholar] [CrossRef]
- Sola, L.; Gagni, P.; D’Annessa, I.; Capelli, R.; Bertino, C.; Romanato, A.; Damin, F.; Bergamaschi, G.; Marchisio, E.; Cuzzocrea, A.; et al. Enhancing Antibody Serodiagnosis Using a Controlled Peptide Coimmobilization Strategy. ACS Infect. Dis. 2018, 4, 998–1006. [Google Scholar] [CrossRef]
- Sweredoski, M.J.; Baldi, P. COBEpro: A novel system for predicting continuous B-cell epitopes. Protein Eng. Des. Sel. 2009, 22, 113–120. [Google Scholar] [CrossRef]
- Sweredoski, M.J.; Baldi, P. PEPITO: Improved discontinuous B-cell epitope prediction using multiple distance thresholds and half sphere exposure. Bioinformatics 2008, 24, 1459–1460. [Google Scholar] [CrossRef]
- Sela-Culang, I.; Benhnia, M.R.-E.-I.; Matho, M.H.; Kaever, T.; Maybeno, M.; Schlossman, A.; Nimrod, G.; Li, S.; Xiang, Y.; Zajonc, D.; et al. Using a Combined Computational-Experimental Approach to Predict Antibody-Specific B Cell Epitopes. Structure 2014, 22, 646–657. [Google Scholar] [CrossRef]
- Pandit, D.; Tuske, S.J.; Coales, S.J.; Liu, A.; Lee, J.E.; Morrow, J.A.; Nemeth, J.F.; Hamuro, Y. Mapping of discontinuous conformational epitopes by amide hydrogen/deuterium exchange mass spectrometry and computational docking. J. Mol. Recognit. 2012, 25, 114–124. [Google Scholar] [CrossRef]
- Jemmerson, R.; Paterson, Y. Mapping epitopes on a protein antigen by the proteolysis of antigen-antibody complexes. Science 1986, 232, 1001–1004. [Google Scholar] [CrossRef]
- Suckau, D.; Köhl, J.; Karwath, G.; Schneider, K.; Casaretto, M.; Bitter-Suermann, D.; Przybylski, M. Molecular epitope identification by limited proteolysis of an immobilized antigen-antibody complex and mass spectrometric peptide mapping. Proc. Natl. Acad. Sci. USA 1990, 87, 9848–9852. [Google Scholar] [CrossRef]
- Urosev, D.; Ferrer-Navarro, M.; Pastorello, I.; Cartocci, E.; Costenaro, L.; Zhulenkovs, D.; Maréchal, J.-D.; Leonchiks, A.; Reverter, D.; Serino, L.; et al. Crystal structure of c5321: A protective antigen present in uropathogenic Escherichia coli strains displaying an SLR fold. BMC Struct. Biol. 2013, 13, 19. [Google Scholar] [CrossRef]
- Nuccitelli, A.; Rinaudo, C.D.; Brogioni, B.; Cozzi, R.; Ferrer-Navarro, M.; Yero, D.; Telford, J.L.; Grandi, G.; Daura, X.; Zacharias, M.; et al. Understanding the molecular determinants driving the immunological specificity of the protective pilus 2a backbone protein of group B streptococcus. PLoS Comput. Biol. 2013, 9, e1003115. [Google Scholar] [CrossRef]
- Baerga-Ortiz, A.; Hughes, C.A.; Mandell, J.G.; Komives, E.A. Epitope mapping of a monoclonal antibody against human thrombin by H/D-exchange mass spectrometry reveals selection of a diverse sequence in a highly conserved protein. Protein Sci. 2002, 11, 1300–1308. [Google Scholar] [CrossRef]
- Coales, S.J.; Tuske, S.J.; Tomasso, J.C.; Hamuro, Y. Epitope mapping by amide hydrogen/deuterium exchange coupled with immobilization of antibody, on-line proteolysis, liquid chromatography and mass spectrometry. Rapid Commun. Mass Spectrom. 2009, 23, 639–647. [Google Scholar] [CrossRef]
- Doran, T.M.; Sarkar, M.; Kodadek, T. Chemical Tools To Monitor and Manipulate Adaptive Immune Responses. J. Am. Chem. Soc. 2016, 138, 6076–6094. [Google Scholar] [CrossRef]
- Szymczak, L.C.; Kuo, H.-Y.; Mrksich, M. Peptide Arrays: Development and Application. Anal. Chem. 2018, 90, 266–282. [Google Scholar] [CrossRef]
- Timmerman, P.; Puijk, W.C.; Boshuizen, R.S.; Van Dijken, P.; Slootstra, J.W.; Beurskens, F.J.; Parren, P.W.H.I.; Huber, A.; Bachmann, M.F.; Meloen, R.H. Functional Reconstruction of Structurally Complex Epitopes using CLIPS TM Technology. Open Vac. J. 2009, 2, 56–67. [Google Scholar] [CrossRef]
- Timmerman, P.; Puijk, W.C.; Meloen, R.H. Functional reconstruction and synthetic mimicry of a conformational epitope using CLIPS technology. J. Mol. Recognit. 2007, 20, 283–299. [Google Scholar] [CrossRef]
- Platt, G.W.; Damin, F.; Swann, M.J.; Metton, I.; Skorski, G.; Cretich, M.; Chiari, M. Allergen immobilisation and signal amplification by quantum dots for use in a biosensor assay of IgE in serum. Biosens. Bioelectron. 2014, 52, 82–88. [Google Scholar] [CrossRef]
- Chiari, M.; Cretich, M.; Damin, F.; Di Carlo, G.; Oldani, C. Advanced polymers for molecular recognition and sensing at the interface. J. Chromatogr. B 2008, 866, 89–103. [Google Scholar] [CrossRef]
- Cretich, M.; Pirri, G.; Damin, F.; Solinas, I.; Chiari, M. A new polymeric coating for protein microarrays. Anal. Biochem. 2004, 332, 67–74. [Google Scholar] [CrossRef]
- Sola, L.; Damin, F.; Gagni, P.; Consonni, R.; Chiari, M. Synthesis of Clickable Coating Polymers by Postpolymerization Modification: Applications in Microarray Technology. Langmuir 2016, 32, 10284–10295. [Google Scholar] [CrossRef] [PubMed]
- Gori, A.; Longhi, R. Chemoselective Strategies to Peptide and Protein Bioprobes Immobilization on Microarray Surfaces. Methods Mol. Biol. 2016, 1352, 145–156. [Google Scholar]
- Gori, A.; Sola, L.; Gagni, P.; Bruni, G.; Liprino, M.; Peri, C.; Colombo, G.; Cretich, M.; Chiari, M. Screening Complex Biological Samples with Peptide Microarrays: The Favorable Impact of Probe Orientation via Chemoselective Immobilization Strategies on Clickable Polymeric Coatings. Bioconjugate Chem. 2016, 27, 2669–2677. [Google Scholar] [CrossRef]
- Gori, A.; Cretich, M.; Vanna, R.; Sola, L.; Gagni, P.; Bruni, G.; Liprino, M.; Gramatica, F.; Burastero, S.; Chiari, M. Multiple epitope presentation and surface density control enabled by chemoselective immobilization lead to enhanced performance in IgE-binding fingerprinting on peptide microarrays. Anal. Chim. Acta 2017, 983, 189–197. [Google Scholar] [CrossRef]
- Gagni, P.; Romanato, A.; Bergamaschi, G.; Bettotti, P.; Vanna, R.; Piotto, C.; Morasso, C.F.; Chiari, M.; Cretich, M.; Gori, A. A self-assembling peptide hydrogel for ultrarapid 3D bioassays. Nanoscale Adv. 2019. [Google Scholar] [CrossRef]
- Mammen, M.; Choi, S.-K.; Whitesides, G.M. Polyvalent Interactions in Biological Systems: Implications for Design and Use of Multivalent Ligands and Inhibitors. Angew. Chem. Int. Ed. 1998, 37, 2754–2794. [Google Scholar] [CrossRef]
- Vetro, M.; Costa, B.; Donvito, G.; Arrighetti, N.; Cipolla, L.; Perego, P.; Compostella, F.; Ronchetti, F.; Colombo, D. Anionic glycolipids related to glucuronosyldiacylglycerol inhibit protein kinase Akt. Org. Biomol. Chem. 2015, 13, 1091–1099. [Google Scholar] [CrossRef]
- Compostella, F.; Pitirollo, O.; Silvestri, A.; Polito, L. Glyco gold nanoparticles: Synthesis and applications. Beilstein J. Org. Chem. 2017, 13, 1008–1021. [Google Scholar] [CrossRef]
- Legnani, L.; Compostella, F.; Sansone, F.; Toma, L. Cone Calix 4 arenes with Orientable Glycosylthioureido Groups at the Upper Rim: An In-Depth Analysis of Their Symmetry Properties. J. Org. Chem. 2015, 80, 7412–7418. [Google Scholar] [CrossRef]
- Morelli, L.; Fallarini, S.; Lombardi, G.; Colombo, C.; Lay, L.; Compostella, F. Synthesis and Biological Evaluation of a Trisaccharide Repeating Unit Derivative of Streptococcus pneumoniae 19A Capsular Polysaccharide. Bioorg. Med. Chem. 2018, 26, 5682–5690. [Google Scholar] [CrossRef]
- De Mori, G.M.S.; Micheletti, C.; Colombo, G. All-atom folding simulations of the villin headpiece from stochastically selected coarse-grained structures. J. Phys. Chem. B 2004, 108, 12267–12270. [Google Scholar] [CrossRef]
- Ferraro, M.; D’Annessa, I.; Moroni, E.; Morra, G.; Paladino, A.; Rinaldi, S.; Compostella, F.; Colombo, G. Allosteric Modulators of HSP90 and HSP70: Dynamics Meets Function through Structure-Based Drug Design. J. Med. Chem. 2019, 62, 60–87. [Google Scholar] [CrossRef]
- Sherwood, P.; Brooks, B.R.; Sansom, M.S.P. Multiscale methods for macromolecular simulations. Curr. Opin. Struct. Biol. 2008, 18, 630–640. [Google Scholar] [CrossRef]
- Aliper, A.; Plis, S.; Artemov, A.; Ulloa, A.; Mamoshina, P.; Zhavoronkov, A. Deep Learning Applications for Predicting Pharmacological Properties of Drugs and Drug Repurposing Using Transcriptomic Data. Mol. Pharm. 2016, 13, 2524–2530. [Google Scholar] [CrossRef]
- Schneider, G. From Theory to Bench Experiment by Computer Assisted Drug Design. Chimia 2012, 66, 120–124. [Google Scholar] [CrossRef]
- Schneider, G. Virtual Screening: An endless staircase? Nat. Rev. Drug. Disc. 2010, 9, 273–276. [Google Scholar] [CrossRef]
- Gawehn, E.; Hiss, J.A.; Schneider, G. Deep Learning in Drug Discovery. Mol. Inform. 2016, 35, 3–14. [Google Scholar] [CrossRef]
- Lima, A.N.; Philot, E.A.; Trossini, G.H.; Scott, L.P.; Maltarollo, V.G.; Honorio, K.M. Use of machine learning approaches for novel drug discovery. Expert Opin. Drug Discov. 2016, 11, 225–239. [Google Scholar] [CrossRef]
- Li, Z.; Kermode, J.R.; De Vita, A. Molecular Dynamics with On-the-Fly Machine Learning of Quantum-Mechanical Forces. Phys. Rev. Lett. 2015, 114, 096405. [Google Scholar] [CrossRef]
- Mamoshina, P.; Vieira, A.; Putin, E.; Zhavoronkov, A. Applications of deep learning in biomedicine. Mol. Pharm. 2016, 13, 1445–1454. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cretich, M.; Gori, A.; D’Annessa, I.; Chiari, M.; Colombo, G. Peptides for Infectious Diseases: From Probe Design to Diagnostic Microarrays. Antibodies 2019, 8, 23. https://doi.org/10.3390/antib8010023
Cretich M, Gori A, D’Annessa I, Chiari M, Colombo G. Peptides for Infectious Diseases: From Probe Design to Diagnostic Microarrays. Antibodies. 2019; 8(1):23. https://doi.org/10.3390/antib8010023
Chicago/Turabian StyleCretich, Marina, Alessandro Gori, Ilda D’Annessa, Marcella Chiari, and Giorgio Colombo. 2019. "Peptides for Infectious Diseases: From Probe Design to Diagnostic Microarrays" Antibodies 8, no. 1: 23. https://doi.org/10.3390/antib8010023
APA StyleCretich, M., Gori, A., D’Annessa, I., Chiari, M., & Colombo, G. (2019). Peptides for Infectious Diseases: From Probe Design to Diagnostic Microarrays. Antibodies, 8(1), 23. https://doi.org/10.3390/antib8010023





