1,2-DCA Natural Attenuation Evaluation in Groundwater: Insight by Dual Isotope 13C/37Cl and Molecular Analysis Approach
Abstract
1. Introduction
2. Materials and Methods
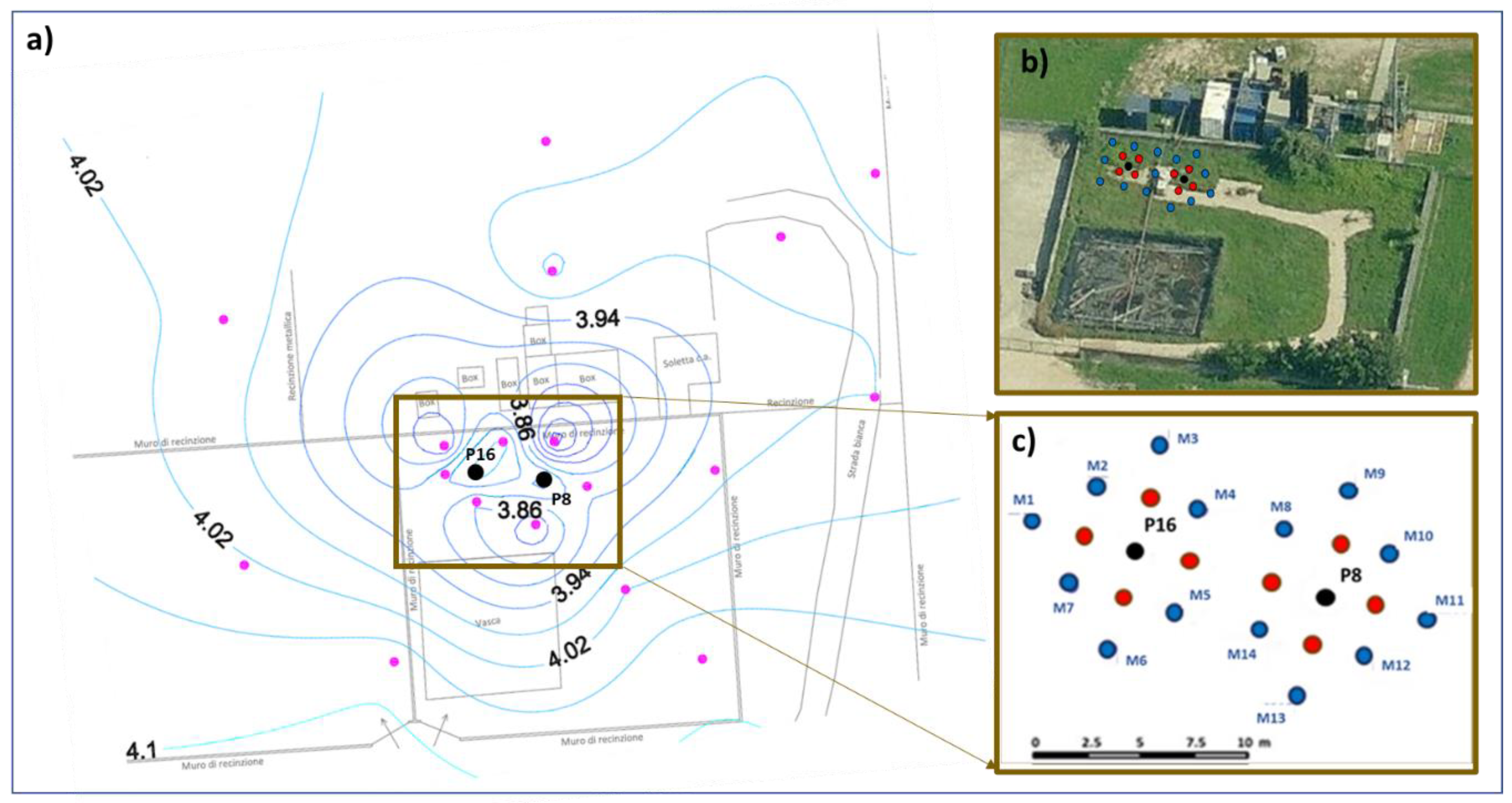
2.1. Site Description
2.2. Microcosm Experiments
2.3. 1,2-DCA Concentration, Isotope Analysis and Enrichment Factor Estimation
2.4. Characterization of Microbial Communities and Quantification of the Catabolic Gene of Geobacter-spp.
2.4.1. Microbial Population Analysis. DNA Extraction and Illumina-MiSeq® Sequencing
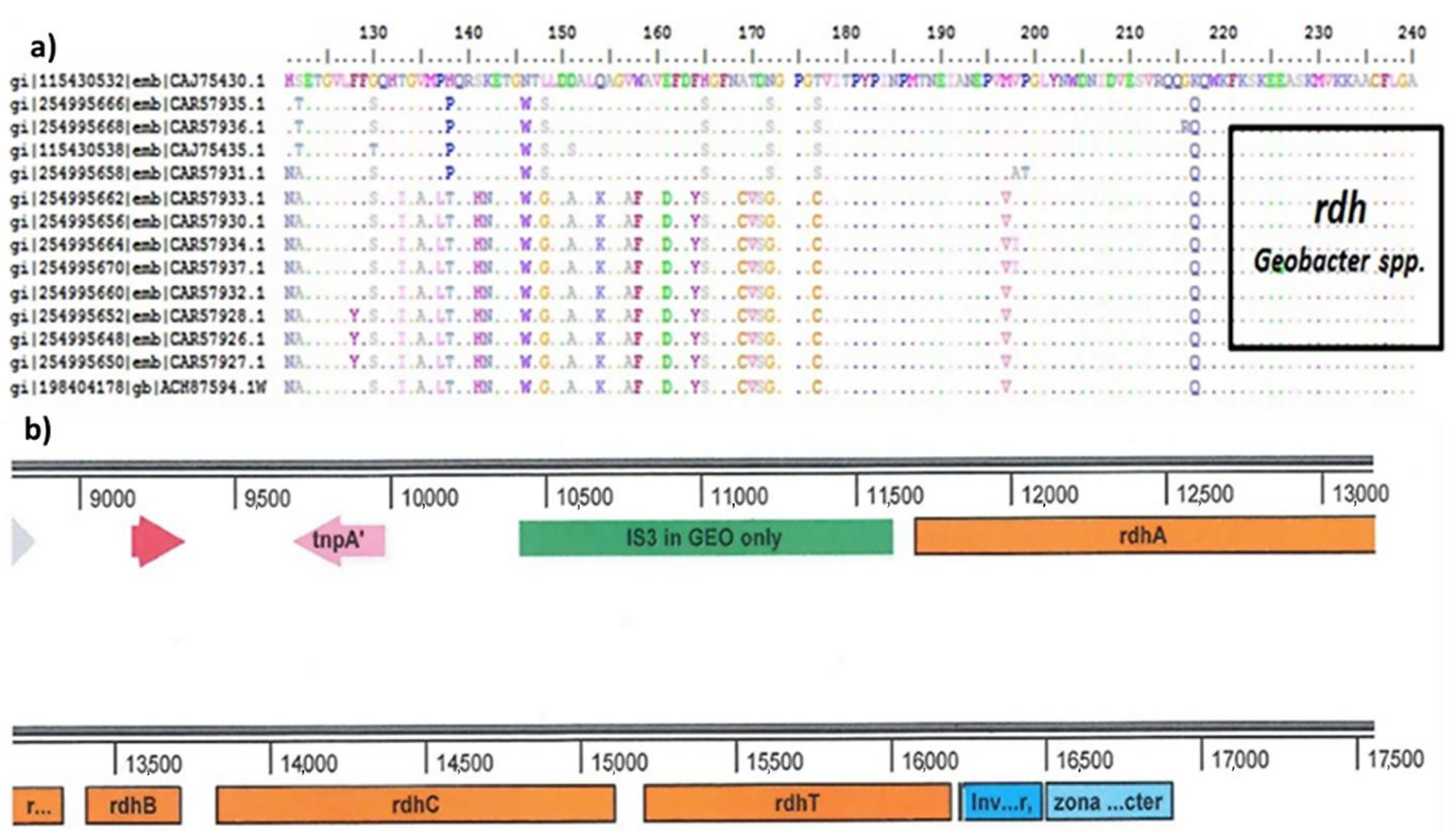
2.4.2. Quantitative Polymerase Chain Reaction (qPCR) for Dehalogenase (rdh) Gene Determination
3. Results and Discussion
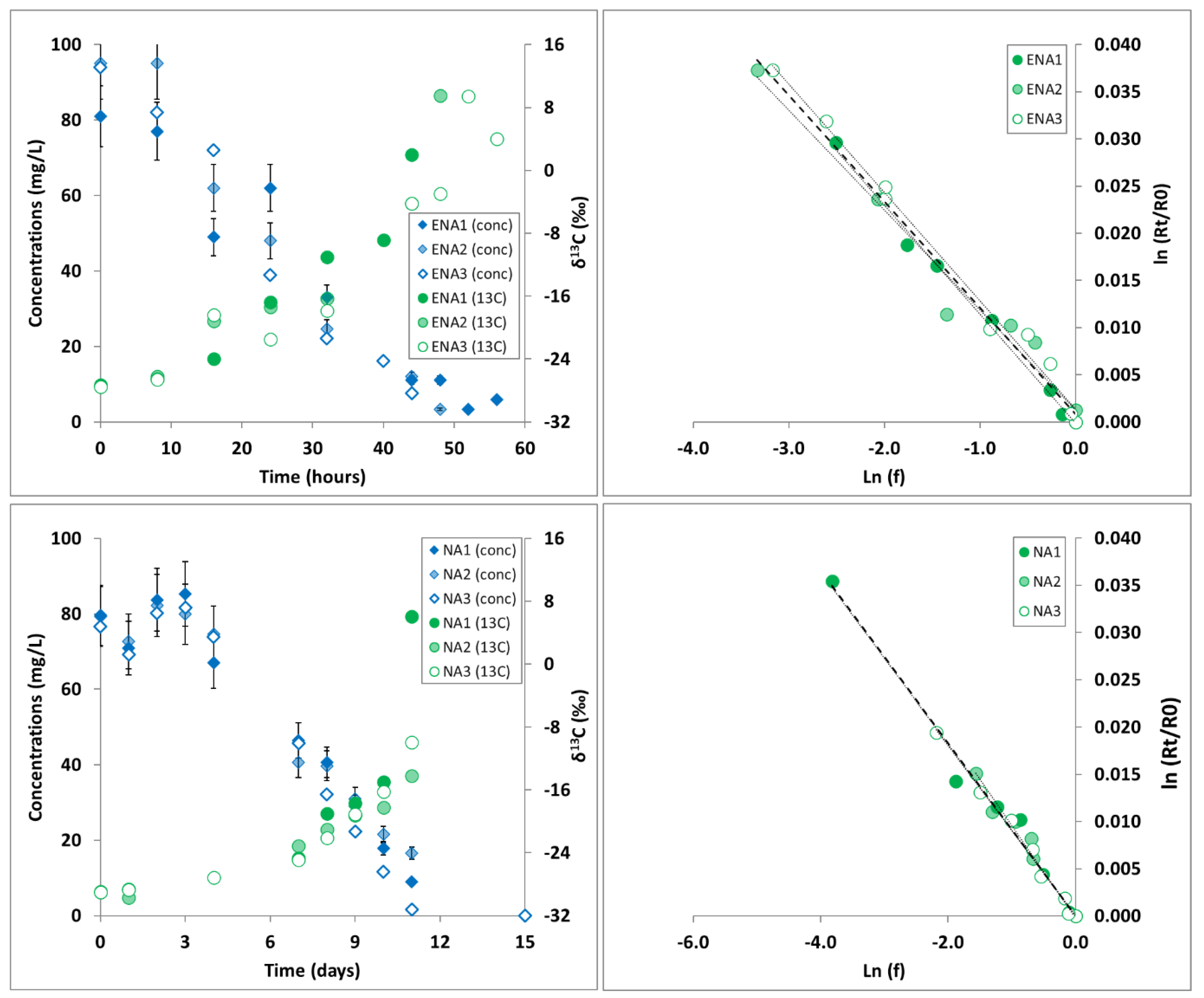
3.1. 1,2-DCA δ13C Fractionation in Microcosms
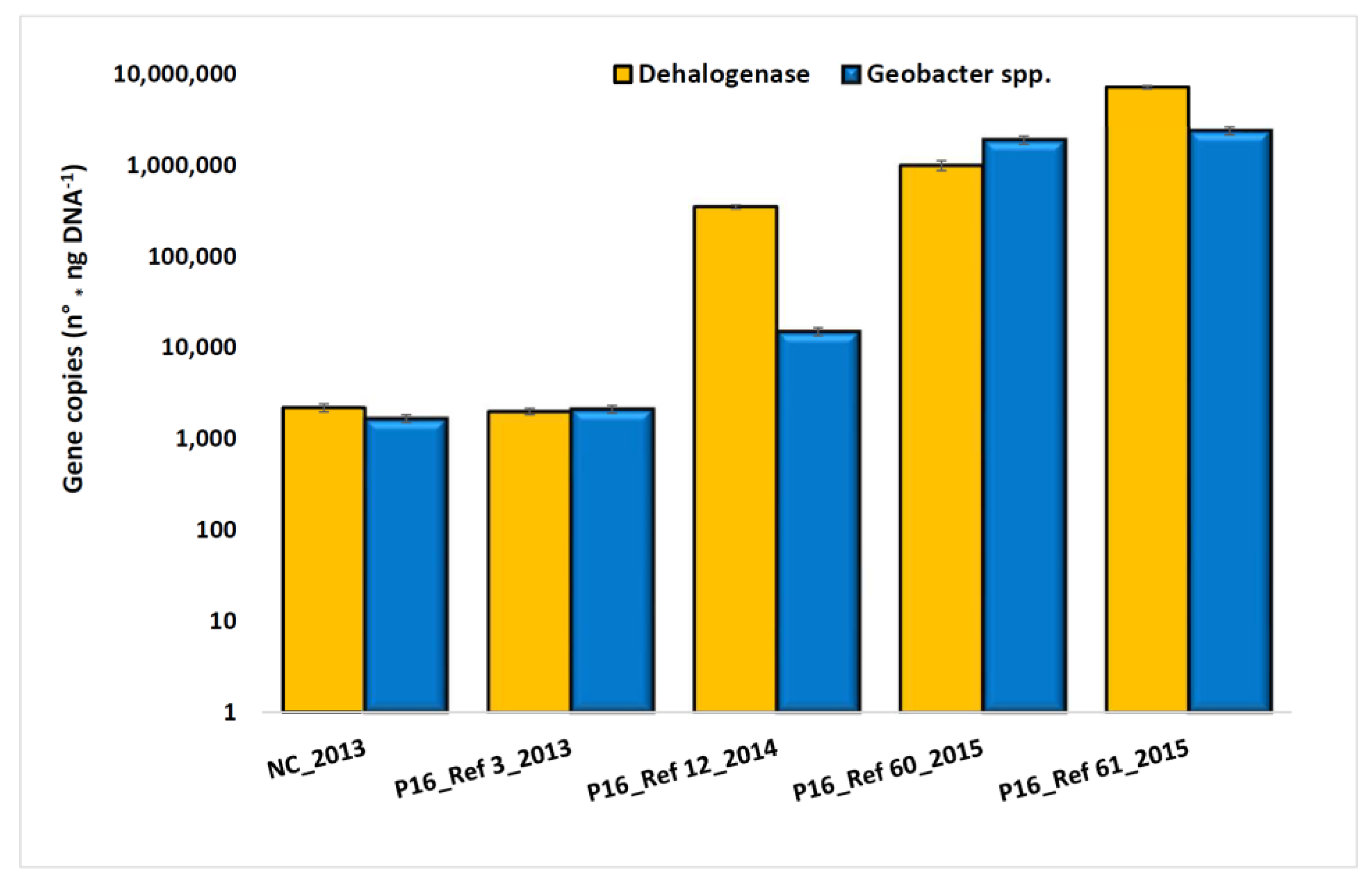
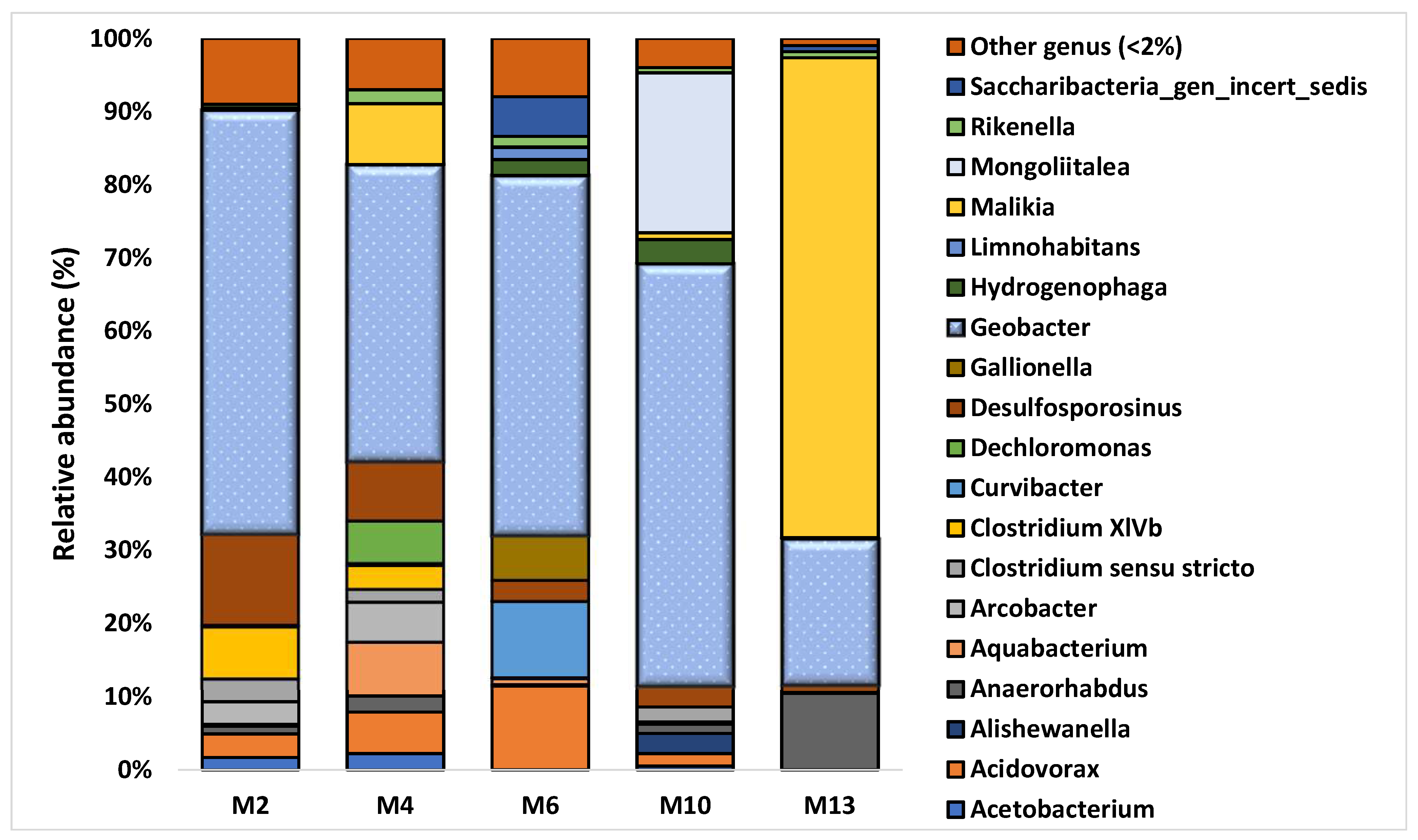
3.2. Characterization of Microbial Communities and Quantification of the Catabolic Gene and of Geobacter spp. from Microcosm Experiments
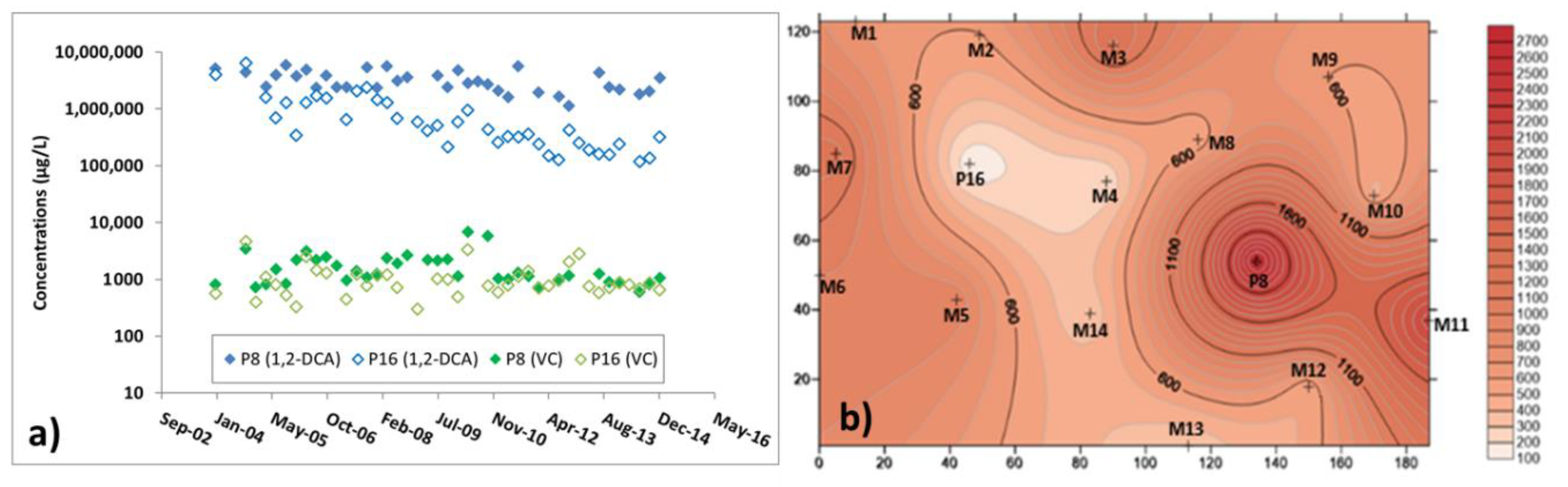
3.3. Microbiological and Isotopic Results from the Pilot Area
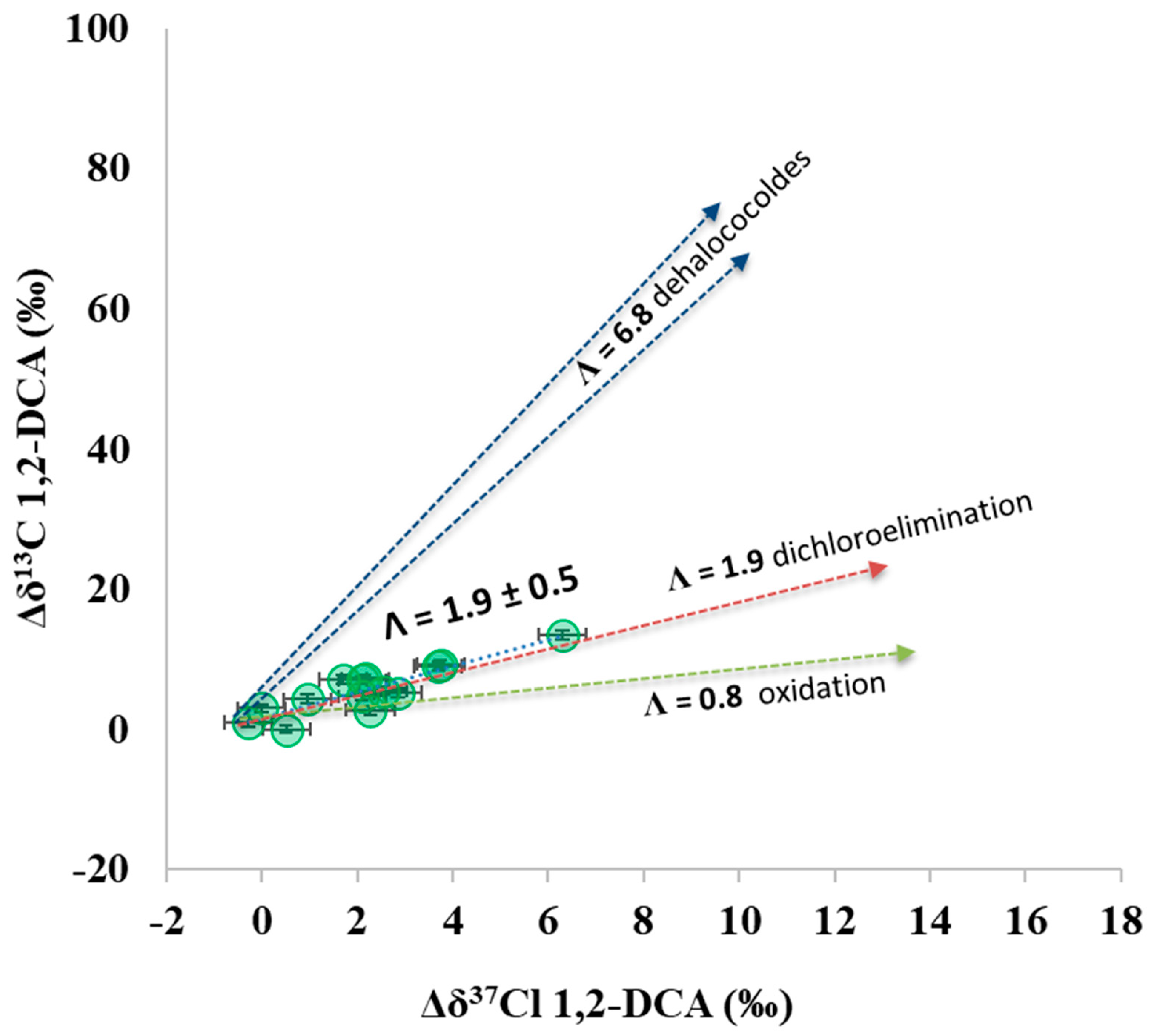
3.4. Additional Insights from the Dual 13C and 37Cl Isotopic Approach at the Pilot Area
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- United States Environmental Protection Agency. Proposed Designation of 1,2-Dichloroethane (CASRN 107-06-2) as a High-Priority Substance for Risk Evaluation; U.S. EPA.: Washington, DC, USA, 22 August 2019.
- Falta, R.W.; Bulsara, N.; Henderson, J.K.; Mayer, R.A. Leaded-Gasoline Additives Still Contaminate Groundwater. Environ. Sci. Technol. 2005, 39, 378A–384A. [Google Scholar] [CrossRef]
- Voudrias, E.A. Pump-and-Treat Remediation of Groundwater contaminated by Hazardous Waste: Can it really be achieved? Global Nest J. 2001, 3, 1–10. [Google Scholar]
- Chen, Q.; Fan, G.; Na, W.; Liu, J.; Cui, J.; Li, H. Past, Present, and Future of Groundwater Remediation Research: A Scien-tometric Analysis. Int. J. Environ. Res. Public Health 2019, 16, 3975. [Google Scholar] [CrossRef]
- De Wildeman, S.; Diekert, G.; Van Langenhove, H.; Verstraete, W. Stereoselective Microbial Dehalorespiration with Vicinal Dichlorinated Alkanes. Appl. Environ. Microbiol. 2003, 69, 5643–5647. [Google Scholar] [CrossRef]
- Duhamel, M.; Edwards, E.A. Microbial composition of chlorinated ethene-degrading cultures dominated by Dehalococ-coides. FEMS Microbiol. Ecol. 2006, 58, 538–549. [Google Scholar] [CrossRef] [PubMed]
- Duhamel, M.; Edwards, E.A. Growth and yields of dechlorinators, acetogens, and methanogens during reductive dechlo-rination of chlorinated ethenes and dihaloelimination of 1,2-dichloroethane. Environ. Sci. Technol. 2007, 41, 2303–2310. [Google Scholar] [CrossRef]
- Marzorati, M.; Balloi, A.; de Ferra, F.; Corallo, L.; Carpani, G.; Wittebolle, L.; Verstraete, W.; Daffonchio, D. Bacterial di-versity and reductive dehalogenase redundancy an a 1,2-dichloroethane-degradaing bacterial consortium enriched from a contaminated aquifer. Microb. Cell Fact. 2010, 9, 12. [Google Scholar] [CrossRef] [PubMed]
- Grostern, A.; Edwards, E.A. Characterization of a Dehalobacter Coculture That Dechlorinates 1,2-Dichloroethane to Eth-ene and Identification of the Putative Reductive Dehalogenase Gene. Appl. Environ. Microbiol. 2009, 75, 2684–2693. [Google Scholar] [CrossRef]
- Hirschorn, S.K.; Dinglasan, M.J.; Elsner, M.; Mancini, S.A.; Lacrampe-Couloume, G.; Edwards, E.A.; Lollar, B.S. Path-way dependent isotopic fractionation during aerobic biodegradation of 1,2-dichloroethane. Environmen. Sci. Technol. 2004, 38, 4775–4781. [Google Scholar] [CrossRef] [PubMed]
- Ansorge, W.J. Next-generation DNA sequencing techniques. New Biotechnol. 2009, 25, 195–203. [Google Scholar] [CrossRef] [PubMed]
- Marchesi, M.; Alberti, L.; Shouakar-Stash, O.; Pietrini, I.; de Ferra, F.; Carpani, G.; Aravena, R.; Franzetti, A.; Stella, T. 37Cl-compound specific isotope analysis and assessment of functional genes for monitoring monochlorobenzene (MCB) bio-degradation under aerobic conditions. Sci. Total Environ. 2018, 619–620, 784–793. [Google Scholar] [CrossRef]
- Hunkeler, D.; Aravena, R.; Butler, B.J. Monitoring Microbial Dechlorination of Tetrachloroethene (PCE) in Groundwater Using Compound-Specific Stable Carbon Isotope Ratios: Microcosm and Field Studies. Environ. Sci. Technol. 1999, 33, 2733–2738. [Google Scholar] [CrossRef]
- Meckenstock, R.U.; Morasch, B.; Griebler, C.; Richnow, H.H. Stable isotope fractionation analysis as a tool to monitor biodegradation in contaminated aquifers. J. Contam. Hydrol. 2004, 75, 215–255. [Google Scholar] [CrossRef] [PubMed]
- Hunkeler, D.; Meckenstock, R.U.; Sherwood Lollar, B.; Schmidt, T.C.; Wilson, J.T. CSIA EPA “A Guide for Assessing Bio-Degradation and Source Identification of Organic Groundwater Contaminants Using Compound Specific Isotope Analysis (CSIA)”; EPA 600/R-08/148; U.S. Environmental Protection Agency: Washington, DC, USA, 2008.
- Carter, J.F.; Barwick, V.J. Good Practice Guide for Isotope Mass Spectrometry; FIRMS: Bristol, UK, 2011; ISBN 978-0-948926-31-0. [Google Scholar]
- Alberti, L.; Marchesi, M.; Trefiletti, P.; Aravena, R. Compound-Specific Isotope Analysis (CSIA) Application for Source Apportionment and Natural Attenuation Assessment of Chlorinated Benzenes. Water 2017, 9, 872. [Google Scholar] [CrossRef]
- Hunkeler, D.; Aravena, R. Evidence of Substantial Carbon Isotope Fractionation among Substrate, Inorganic Carbon, and Biomass during Aerobic Mineralization of 1,2-Dichloroethane byXanthobacter autotrophicus. Appl. Environ. Microbiol. 2000, 66, 4870–4876. [Google Scholar] [CrossRef] [PubMed]
- Hunkeler, D.; Aravena, R.; Cox, E. Carbon Isotopes as a Tool to Evaluate the Origin and Fate of Vinyl Chloride: La-boratory Experiments and Modeling of Isotope Evolution. Environmen. Sci. Technol. 2002, 36, 3378–3384. [Google Scholar] [CrossRef] [PubMed]
- Hirschorn, S.K.; Dinglasan-Panlilio, M.J.; Edwards, E.A.; Lacrampe-Couloume, G.; Lollar, B.S. Isotope analysis as a natural reaction probe to determine mechanisms of biodegradation of 1,2-dichloroethane. Environ. Microbiol. 2007, 9, 1651–1657. [Google Scholar] [CrossRef]
- Schmidt, M.; Lege, S.; Nijenhuis, I. Comparison of 1,2-dichloroethane, dichloroethene and vinyl chloride carbon stable isotope fractionation during dechlorination by two Dehalococcoides strains. Water Res. 2014, 52, 146–154. [Google Scholar] [CrossRef]
- Palau, J.; Shouakar-Stash, O.; Mortan, S.H.; Yu, R.; Rosell, M.; Marco-Urrea, E.; Freedman, D.L.; Aravena, R.; Soler, A.; Hunkeler, D. Hydrogen Isotope Fractionation during the Biodegradation of 1,2-Dichloroethane: Potential for Pathway Identification Using a Multi-element (C, Cl, and H) Isotope Approach. Environ. Sci. Technol. 2017, 51, 10526–10535. [Google Scholar] [CrossRef]
- Palau, J.; Yu, R.; Mortan, S.H.; Shouakar-Stash, O.; Rosell, M.; Freedman, D.L.; Sbarbati, C.; Fiorenza, S.; Aravena, R.; Marco-Urrea, E.; et al. Distinct Dual C–Cl Isotope Fractionation Patterns during Anaerobic Biodegradation of 1,2-Dichloroethane: Potential To Characterize Microbial Degradation in the Field. Environ. Sci. Technol. 2017, 51, 2685–2694. [Google Scholar] [CrossRef]
- Marzorati, M.; De Ferra, F.; Van Raemdonck, H.; Borin, S.; Allifranchini, E.; Carpani, G.; Serbolisca, L.; Verstraete, W.; Boon, N.; Daffonchio, D. A Novel Reductive Dehalogenase, Identified in a Contaminated Groundwater Enrichment Culture and in Desulfitobacterium dichloroeliminans Strain DCA1, Is Linked to Dehalogenation of 1,2-Dichloroethane. Appl. Environ. Microbiol. 2007, 73, 2990–2999. [Google Scholar] [CrossRef]
- Marzorati, M.; Borin, S.; Brusetti, L.; Daffonchio, D.; Marsilli, C.; Carpani, G.; De Ferra, F. Response of 1,2-dichloroethane-adapted microbial communities to ex-situ biostimulation of polluted groundwater. Biogeochemistry 2005, 17, 41–56. [Google Scholar] [CrossRef]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef] [PubMed]
- Espinoza-Tofalos, A.; Daghio, M.; Palma, E.; Aulenta, F.; Franzetti, A. Structure and Functions of Hydrocarbon-Degrading Microbial Communities in Bioelectrochemical Systems. Water 2020, 12, 343. [Google Scholar] [CrossRef]
- Merlino, G.; Balloi, A.; Marzorati, M.; Mapelli, F.; Rizzi, A.; Lavazza, D.; de Ferra, F.; Carpani, G.; Daffonchio, D. Di-verse Reductive Dehalogenases Are Associated with Clostridiales-Enriched Microcosms Dechlorinating 1,2-Dichloroethane. BioMed Res. Int. 2015, 2015, 242856. [Google Scholar] [CrossRef] [PubMed]
- Morrill, P.L.; Sleep, B.E.; Seepersad, D.J.; McMaster, M.L.; Hood, E.D.; LeBron, C.; Major, D.W.; Edwards, E.A.; Sherwood Lollar, B. Variations in expression of carbon isotope fractionation of chlorinated ethenes during biologically en-hanced PCE dissolution close to a source zone. J. Contam. Hydrol. 2009, 110, 60–71. [Google Scholar] [CrossRef]
- Yuan, S.Y.; Chang, B.V. Anaerobic degradation of five polycyclic aromatic hydrocarbons from river sediment in Taiwan. J. Environ. Sci. Health B 2007, 42, 63–69. [Google Scholar] [CrossRef] [PubMed]
- Foght, J. Anaerobic Biodegradation of Aromatic Hydrocarbons: Pathways and Prospects. J. Mol. Microbiol. Biotechnol. 2008, 15, 93–120. [Google Scholar] [CrossRef]
- Xu, X.; Liu, W.; Tian, S.; Wang, W.; Qi, Q.; Jiang, P.; Gao, X.; Li, F.; Li, H.; Yu, H. Petroleum Hydrocarbon-Degrading Bacteria for the Remediation of Oil Pollution Under Aerobic Conditions: A Perspective Analysis. Front. Microbiol. 2018, 9, 2885. [Google Scholar] [CrossRef]
- Aburto, A.; Fahy, A.; Coulon, F.; Lethbridge, G.; Timmis, K.N.; Ball, A.S.; McGenity, T. Mixed aerobic and anaerobic microbial communities in benzene-contaminated groundwater. J. Appl. Microbiol. 2009, 106, 317–328. [Google Scholar] [CrossRef] [PubMed]
- Payne, K.A.P.; Quezada, C.P.; Fisher, K.; Dunstan, M.S.; Collins, F.A.; Sjuts, H.; Levy, C.; Hay, S.; Rigby, S.E.J.; Leys, D. Reductive dehalogenase structure suggests a mechanism for B12-dependent dehalogenation. Nat. Cell Biol. 2015, 517, 513–516. [Google Scholar] [CrossRef] [PubMed]

| M2 | M4 | M6 | M10 | M13 | |
|---|---|---|---|---|---|
| 1,2-DCA concentration (ppm) | 349 | 204 | 1000 | 460 | 255 |
| rdh gene copies (n° * ng DNA-1) | 5.63 × 105 | 6.81 × 105 | 1.42 × 106 | 1.07 × 106 | 1.78 × 105 |
| Relative abundance of Geobacter spp. (%) | 58 | 41 | 49 | 58 | 20 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carpani, G.; Marchesi, M.; Pietrini, I.; Alberti, L.; Zaninetta, L.M.; Shouakar-Stash, O.; de Ferra, F. 1,2-DCA Natural Attenuation Evaluation in Groundwater: Insight by Dual Isotope 13C/37Cl and Molecular Analysis Approach. Water 2021, 13, 728. https://doi.org/10.3390/w13050728
Carpani G, Marchesi M, Pietrini I, Alberti L, Zaninetta LM, Shouakar-Stash O, de Ferra F. 1,2-DCA Natural Attenuation Evaluation in Groundwater: Insight by Dual Isotope 13C/37Cl and Molecular Analysis Approach. Water. 2021; 13(5):728. https://doi.org/10.3390/w13050728
Chicago/Turabian StyleCarpani, Giovanna, Massimo Marchesi, Ilaria Pietrini, Luca Alberti, Luciano Massimo Zaninetta, Orfan Shouakar-Stash, and Francesca de Ferra. 2021. "1,2-DCA Natural Attenuation Evaluation in Groundwater: Insight by Dual Isotope 13C/37Cl and Molecular Analysis Approach" Water 13, no. 5: 728. https://doi.org/10.3390/w13050728
APA StyleCarpani, G., Marchesi, M., Pietrini, I., Alberti, L., Zaninetta, L. M., Shouakar-Stash, O., & de Ferra, F. (2021). 1,2-DCA Natural Attenuation Evaluation in Groundwater: Insight by Dual Isotope 13C/37Cl and Molecular Analysis Approach. Water, 13(5), 728. https://doi.org/10.3390/w13050728







