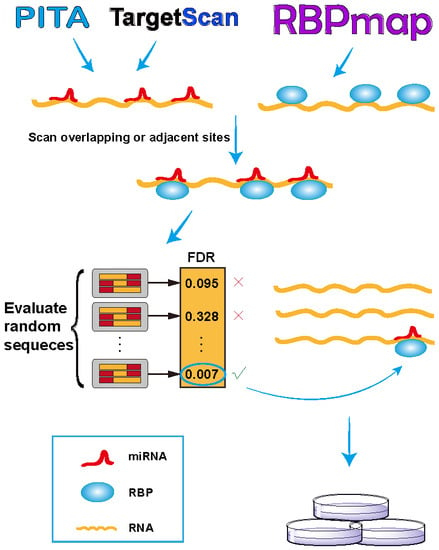
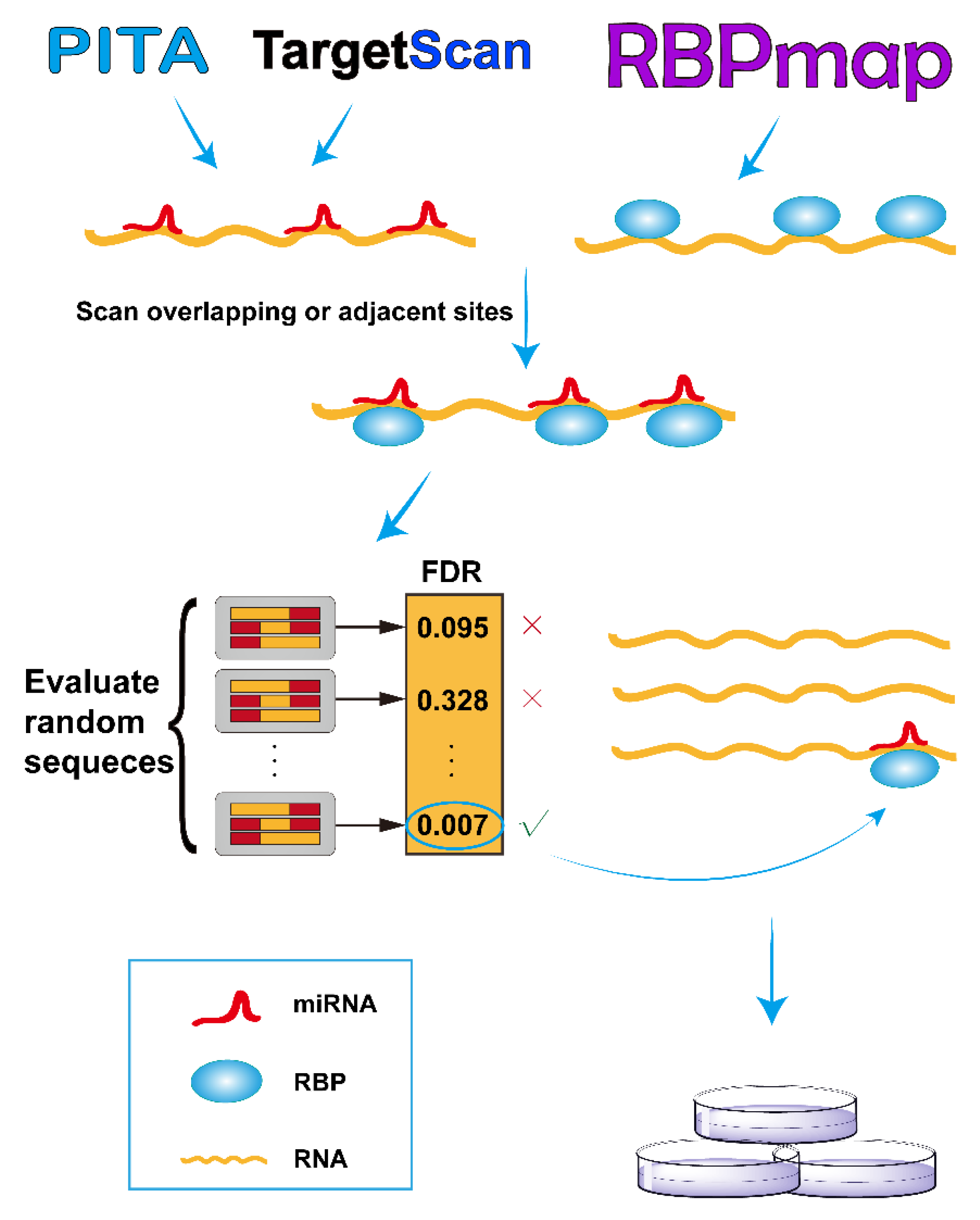
RBPvsMIR: A Computational Pipeline to Identify Competing miRNAs and RNA-Binding Protein Pairs Regulating the Shared Transcripts
Abstract
1. Introduction
2. Materials and Methods
2.1. RBPvsMIR Prediction Pipeline
2.2. Construction of RBPvsMIR Web Server
2.3. Experimental Confirmation
3. Results
3.1. RBPvsMIR Pipeline and Web Server
3.2. Predicting and Confirming the Competing RNA-Binding Protein and microRNA Pairs on Long Non-coding RNA MALAT1
3.3. A Computational Landscape of Competing microRNA and RNA-Binding Protein Pairs on Human Functional Long Non-coding RNAs
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Liao, X.; Lu, N.; Liu, W.; Wong, C.W. Chromatin-modifying drugs induce miRNA-153 expression to suppress Irs-2 in glioblastoma cell lines. Int. J. Cancer 2011, 129, 2527–2531. [Google Scholar] [CrossRef] [PubMed]
- Harada, M.; Luo, X.; Murohara, T.; Yang, B.; Dobrev, D.; Nattel, S. MicroRNA regulation and cardiac calcium signaling: Role in cardiac disease and therapeutic potential. Circ. Res. 2014, 114, 689–705. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, N.; Corcoran, D.L.; Nusbaum, J.D.; Reid, D.W.; Georgiev, S.; Hafner, M.; Ascano, M., Jr.; Tuschl, T.; Ohler, U.; Keene, J.D. Integrative regulatory mapping indicates that the RNA-binding protein HuR couples pre-mRNA processing and mRNA stability. Mol. Cell 2011, 43, 327–339. [Google Scholar] [CrossRef] [PubMed]
- Licatalosi, D.D.; Darnell, R.B. RNA processing and its regulation: Global insights into biological networks. Nat.Rev.Genet. 2010, 11, 75–87. [Google Scholar] [CrossRef] [PubMed]
- Tominaga, K.; Srikantan, S.; Lee, E.K.; Subaran, S.S.; Martindale, J.L.; Abdelmohsen, K.; Gorospe, M. Competitive regulation of nucleolin expression by HuR and miR-494. Mol.Cell. Biol. 2011, 31, 4219–4231. [Google Scholar] [CrossRef] [PubMed]
- Sosanya, N.M.; Huang, P.P.; Cacheaux, L.P.; Chen, C.J.; Nguyen, K.; Perrone-Bizzozero, N.I.; Raab-Graham, K.F. Degradation of high affinity HuD targets releases Kv1.1 mRNA from miR-129 repression by mTORC1. J.Cell Biol. 2013, 202, 53–69. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Abdelmohsen, K.; Yang, X.; De, S.; Grammatikakis, I.; Noh, J.H.; Gorospe, M. LncRNA OIP5-AS1/cyrano sponges RNA-binding protein HuR. Nucl. Acids Res. 2016, 44, 2378–2392. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Ouyang, M.; Rao, J.N.; Zou, T.; Xiao, L.; Chung, H.K.; Wu, J.; Donahue, J.M.; Gorospe, M.; Wang, J.Y. Competition between RNA-binding proteins CELF1 and HuR modulates MYC translation and intestinal epithelium renewal. Mol. Biol.Cell 2015, 26, 1797–1810. [Google Scholar] [CrossRef] [PubMed]
- Kawahara, H.; Imai, T.; Imataka, H.; Tsujimoto, M.; Matsumoto, K.; Okano, H. Neural RNA-binding protein Musashi1 inhibits translation initiation by competing with eIF4G for PABP. J. Cell Biol. 2008, 181, 639–653. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Xu, J.; Yang, D.; Tan, X.; Wang, H. Computational approaches for microRNA studies: A review. Mamm. Genome 2010, 21, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, V.; Bell, G.W.; Nam, J.W.; Bartel, D.P. Predicting effective microRNA target sites in mammalian mRNAs. Elife 2015, 4. [Google Scholar] [CrossRef] [PubMed]
- Kertesz, M.; Iovino, N.; Unnerstall, U.; Gaul, U.; Segal, E. The role of site accessibility in microRNA target recognition. Nat.Genet. 2007, 39, 1278–1284. [Google Scholar] [CrossRef] [PubMed]
- Ray, D.; Kazan, H.; Chan, E.T.; Pena Castillo, L.; Chaudhry, S.; Talukder, S.; Blencowe, B.J.; Morris, Q.; Hughes, T.R. Rapid and systematic analysis of the RNA recognition specificities of RNA-binding proteins. Nat. Biotechnol. 2009, 27, 667–670. [Google Scholar] [CrossRef] [PubMed]
- Paz, I.; Kosti, I.; Ares, M., Jr.; Cline, M.; Mandel-Gutfreund, Y. RBPmap: A web server for mapping binding sites of RNA-binding proteins. Nucl. Acids Res. 2014, 42, W361–W367. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R.Stat. Soc. B 1995, 57, 289–300. [Google Scholar]
- Quek, X.C.; Thomson, D.W.; Maag, J.L.; Bartonicek, N.; Signal, B.; Clark, M.B.; Gloss, B.S.; Dinger, M.E. lncRNAdb v2.0: Expanding the reference database for functional long noncoding RNAs. Nucl. Acids Res. 2015, 43, D168–D173. [Google Scholar] [CrossRef] [PubMed]
- Edmond, V.; Moysan, E.; Khochbin, S.; Matthias, P.; Brambilla, C.; Brambilla, E.; Gazzeri, S.; Eymin, B. Acetylation and phosphorylation of SRSF2 control cell fate decision in response to cisplatin. EMBO J. 2011, 30, 510–523. [Google Scholar] [CrossRef] [PubMed]
- Komeno, Y.; Huang, Y.J.; Qiu, J.; Lin, L.; Xu, Y.; Zhou, Y.; Chen, L.; Monterroza, D.D.; Li, H.; DeKelver, R.C.; et al. SRSF2 is essential for hematopoiesis and its myelodysplastic syndrome-related mutations dysregulate alternative pre-mRNA splicing. Mol.Cell Biol. 2015, 35, 3071–3082. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, V.; Ellis, J.D.; Shen, Z.; Song, D.Y.; Pan, Q.; Watt, A.T.; Freier, S.M.; Bennett, C.F.; Sharma, A.; Bubulya, P.A.; et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol. Cell 2010, 39, 925–938. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Li, M.; Wang, Z.; Han, S.; Tang, X.; Ge, Y.; Zhou, L.; Zhou, C.; Yuan, Q.; Yang, M. Silencing of long noncoding RNA MALAT1 by miR-101 and miR-217 inhibits proliferation, migration and invasion of esophageal squamous cell carcinoma cells. J. Biol. Chem. 2015, 290, 3925–3935. [Google Scholar] [CrossRef] [PubMed]
- Shen, W.J.; Zhang, F.; Zhao, X.; Xu, J. LncRNAs and esophageal squamous cell carcinoma—implications for pathogenesis and drug development. J. Cancer 2016, 7, 1258–1264. [Google Scholar] [CrossRef] [PubMed]
- Ji, Q.; Zhang, L.; Liu, X.; Zhou, L.; Wang, W.; Han, Z.; Sui, H.; Tang, Y.; Wang, Y.; Liu, N.; et al. Long non-coding RNA MALAT1 promotes tumour growth and metastasis in colorectal cancer through binding to SFPQ and releasing oncogene PTBP2 from SFPQ/PTBP2 complex. Br.J. Cancer 2014, 111, 736–748. [Google Scholar] [CrossRef] [PubMed]
- Blin, K.; Dieterich, C.; Wurmus, R.; Rajewsky, N.; Landthaler, M.; Akalin, A. DoRiNA 2.0—Upgrading the doRiNA database of RNA interactions in post-transcriptional regulation. Nucl. Acids Res. 2015, 43, D160–D167. [Google Scholar] [CrossRef] [PubMed]
- Schirle, N.T.; Sheu-Gruttadauria, J.; MacRae, I.J. Structural basis for microRNA targeting. Science 2014, 346, 608–613. [Google Scholar] [CrossRef] [PubMed]



© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, X.; Chen, D.; Cai, Y.; Zhang, F.; Xu, J. RBPvsMIR: A Computational Pipeline to Identify Competing miRNAs and RNA-Binding Protein Pairs Regulating the Shared Transcripts. Genes 2018, 9, 426. https://doi.org/10.3390/genes9090426
Zhao X, Chen D, Cai Y, Zhang F, Xu J. RBPvsMIR: A Computational Pipeline to Identify Competing miRNAs and RNA-Binding Protein Pairs Regulating the Shared Transcripts. Genes. 2018; 9(9):426. https://doi.org/10.3390/genes9090426
Chicago/Turabian StyleZhao, Xing, Danze Chen, Yujie Cai, Fan Zhang, and Jianzhen Xu. 2018. "RBPvsMIR: A Computational Pipeline to Identify Competing miRNAs and RNA-Binding Protein Pairs Regulating the Shared Transcripts" Genes 9, no. 9: 426. https://doi.org/10.3390/genes9090426
APA StyleZhao, X., Chen, D., Cai, Y., Zhang, F., & Xu, J. (2018). RBPvsMIR: A Computational Pipeline to Identify Competing miRNAs and RNA-Binding Protein Pairs Regulating the Shared Transcripts. Genes, 9(9), 426. https://doi.org/10.3390/genes9090426