1. Introduction
The human pathogenic
Cryptococcus (HPC) is a group of globally distributed basidiomycete yeasts. These yeasts are opportunistic pathogens to humans and other mammals. They are commonly found in soil, avian excretion and rotting tree barks [
1,
2,
3,
4]. HPC consists of two species complexes, the
Cryptococcus neoformans species complex (CNSC) and the
Cryptococcus gatti species complex (CGSC). Globally, most human infections are caused by strains of CNSC. CNSC tends to infect immunocompromised hosts and is a leading cause of death in HIV patients [
5,
6]. Infections can lead to systemic cryptococcosis, with the most common and detrimental form being cryptococcal meningitis. In 2014, there were ~223,100 recorded cases of cryptococcal meningitis resulting in ~180,100 deaths worldwide [
5,
6]. With cases on the rise over the past five decades due to increasing populations of immunocompromised hosts [
6,
7], it is important that we improve our understanding of the global distribution and genetic diversity of
C. neoformans.CNSC is a highly heterogeneous group of organisms, with divergent lineages showing >10% nucleotide sequence divergence [
1]. Over the last 50 years, a variety of molecular markers have been used to identify strains of CNSC [
8]. These markers have revealed divergent lineages within CNSC. The current emerging consensus separates CNSC into two species,
C. neoformans (serotype A) and
C. deneoformans (serotype D).
C. neoformans (Serotype A) is further divided into three major molecular types, VNI, VNB and VNII, while
C. deneoformans (serotype D) corresponds to the molecular type VNIV. In addition to these four major molecular types, VNB was further divided into two subtypes, VNBI and VNBII, and diploid/aneuploid hybrids have been observed in nature and are referred to as VNIII or serotype AD hybrids [
1,
8,
9,
10].
To help standardize the genotyping system and make it easy to share information among labs, in 2007, the International Society for Human and Animal Mycology (ISHAM) established a committee to set up a multi-locus sequence typing (MLST) method. The recommended system was published in 2009 and included partial DNA sequences at the following seven loci:
CAP59, GPD1, LAC1, PLB1, SOD1, URA5 and
IGS1 [
11]. Subsequently, an online international fungal multi-locus serotyping database (IFMLST) was established for storing and comparing the MLST data. This data repository is comprised of allelic profiles for each recorded sequence type (ST) and the nucleotide sequence for each determined allele type (AT) at each of the seven loci [
12]. Newly genotyped strains can be compared to the database to determine their AT and ST profiles. The MLST system is a great tool for the consistent and efficient comparison of strain genotypes across labs. However, little analysis has been conducted utilizing the publicly available dataset.
Considering the high global burden of infections by CNSC, it is important to understand the global population genetic variation of this species complex. In this paper, we investigate the global genotype distribution and population structure by analyzing the 4200 CNSC isolates with MLST data published in 41 studies. We aim to estimate the overall geographical pattern of genetic variation and determine if recombination plays a role in shaping the diversity observed within individual species and individual molecular types of CNSC. We hypothesize that the genetic variations within CNSC are geographically structured, and recombination plays an important role during the evolution of this species complex.
3. Results
As of January 2022, there were 657 total sequence types (STs) deposited into the Cryptococcus MLST database for CNSC, with associated DNA sequence data for all seven loci. Of the 657 STs, the geographical location information was documented for 296 (45%), while the remaining 361 (55%) STs had unknown geographic information. Our population genetic analyses focused on the 296 STs. The 296 STs represented 4200 CNSC isolates, as extracted from 41 published reports. The metadata for all isolates were retrieved from these published reports. Below, we summarize the retrieved data on the 4200 isolates and present the results of our analyses.
3.1. Geographical and Ecological Distributions
The geographic distribution of the non-redundant 4200 CNSC isolates is presented in
Table 1. These isolates were from 31 countries and five continents, with the majority being found in Asia (61.9%), followed by Africa (12.6%), Europe (14.3%), South America (10.9%) and North America (0.3%). At the country level, the highest number of isolates in this dataset came from China (1216 isolates; 28.95%), while the lowest came from the Congo, the Dominican Republic and the Democratic Republic of Congo (with one isolate each). In between these two extremes, the second largest national population of CNSC in the retrieved dataset was from Thailand (524 isolates; 12.27%), followed by India (380; 9.05%), Brazil (318 isolates; 7.57%), South Africa (268 isolates; 6.38%), Uganda (241 isolates; 5.74%), France (226; 5.38%), Italy (151 isolates; 3.60%), Germany (145 isolates, 3.45%), Vietnam (136 isolates; 3.24%) and Japan (119 isolates; 2.83%). The remaining 20 countries each had <100 isolates analyzed, and, together, they contributed 476 isolates to the analyzed dataset. The geographic associations of the 296 STs are presented in
Supplementary Table S1. Among the 296 STs, ST5 was the most abundant; it was found across 18 countries on four continents (
Supplementary Table S1).
Of the 657 STs in the MLST database, only 284 had ecological niche information (
Supplementary Table S2;
Table 2). These 284 STs represented a total of 4064 isolates, while the remaining 373 STs had no ecological niche/source data. Here, we broadly categorized the isolates into three ecological sources: clinical, environmental and veterinary. The majority of the 4200 isolates were collected from clinical sources (3370 isolates; 80.24%), followed by environmental (648 isolates; 15.43%), and veterinary (46 isolates; 1.10%) sources, leaving 3.24% of isolates with unknown source information. The ecological distributions of the individual STs are shown in
Supplementary Table S2. A total of 14 STs were found in all three niches; 3 STs were found from both clinical and veterinary sources only; 32 STs were found in both clinical and environmental sources only; and no ST was shared between only environmental and veterinary sources. The remaining 235 STs with ecological niche information were each found in only one of the three ecological niches (
Table 2).
Table 2 summarizes the geographic and ecological distributions of the 296 STs in the published MLST literature for CNSC. Geographically, among the 296 STs, 15 STs (representing a total of 2675 isolates) were found in all four continents, 9 STs (representing a total of 656 isolates) were found in three of the four continents, 28 STs (representing a total of 312 isolates) were found in two of the four continents and 244 (representing 557 isolates) were found in only one of the continents (
Table 2). Among the 244 STs, 176 were each represented by only one isolate in the database. Ecologically, among the 296 STs, 284 STs, including 4064 isolates, had ecological niche data. Of these 284 STs, 14 (representing 2550 isolates) were found in all three ecological niches, 35 STs (representing 1034 isolates) were found in two of the three ecological niches and 235 STs (representing 480 isolates) were found in one niche only (
Table 2). The detailed geographic and ecological distributions for each of the 296 STs are shown in
Supplementary Tables S1 and S2. At the country level, 78 (26%) sequence types were reported from two or more countries each (
Supplementary Table S3).
3.2. DNA Sequence Variation
The allelic profiles of each ST, including the allele type (AT) number at each of the seven MLST loci (
CAP59, GPD1, LAC1, IGS1, PLB1, SOD1, URA5), were retrieved for all 657 STs in the database. Summaries of the allele types across the total sample, the serotype A sample and the serotype D sample are shown in
Table 3. The differences in length of bp per allele type for each gene range from 0 difference for
CAP59 to 45bp difference for
IGS1. Among the seven loci, in the total sample,
PLB1 had the fewest number of alleles (44) while
IGS1 had the most (93). A largely similar pattern was observed for the serotype A and serotype D samples, where
IGS1 had the highest allele number in both. However,
GPD1 had the lowest allele number in the serotype D sample. The range of occurrence of each allele type in each of the samples is shown in
Supplementary Table S3.
3.3. Phylogenetic Analysis
The
C. neoformans species complex is commonly grouped into five broad molecular types: VNI, VNII, VNIII, VNB and VNIV. The strains of VNI, VNII and VNB belonged to
C. neoformans (serotype A); the strains of VNIV belonged to
C. deneoformans (serotype D); and the strains of VNIII (serotype AD) represented hybrids of serotypes A and D. For some VNB strains, they were further classified into VNBI and VNBII [
8,
10]. The molecular type designations were mostly based on the restriction enzyme digest pattern of the
URA5 sequence, amplified fragment length polymorphisms or PCR fingerprinting [
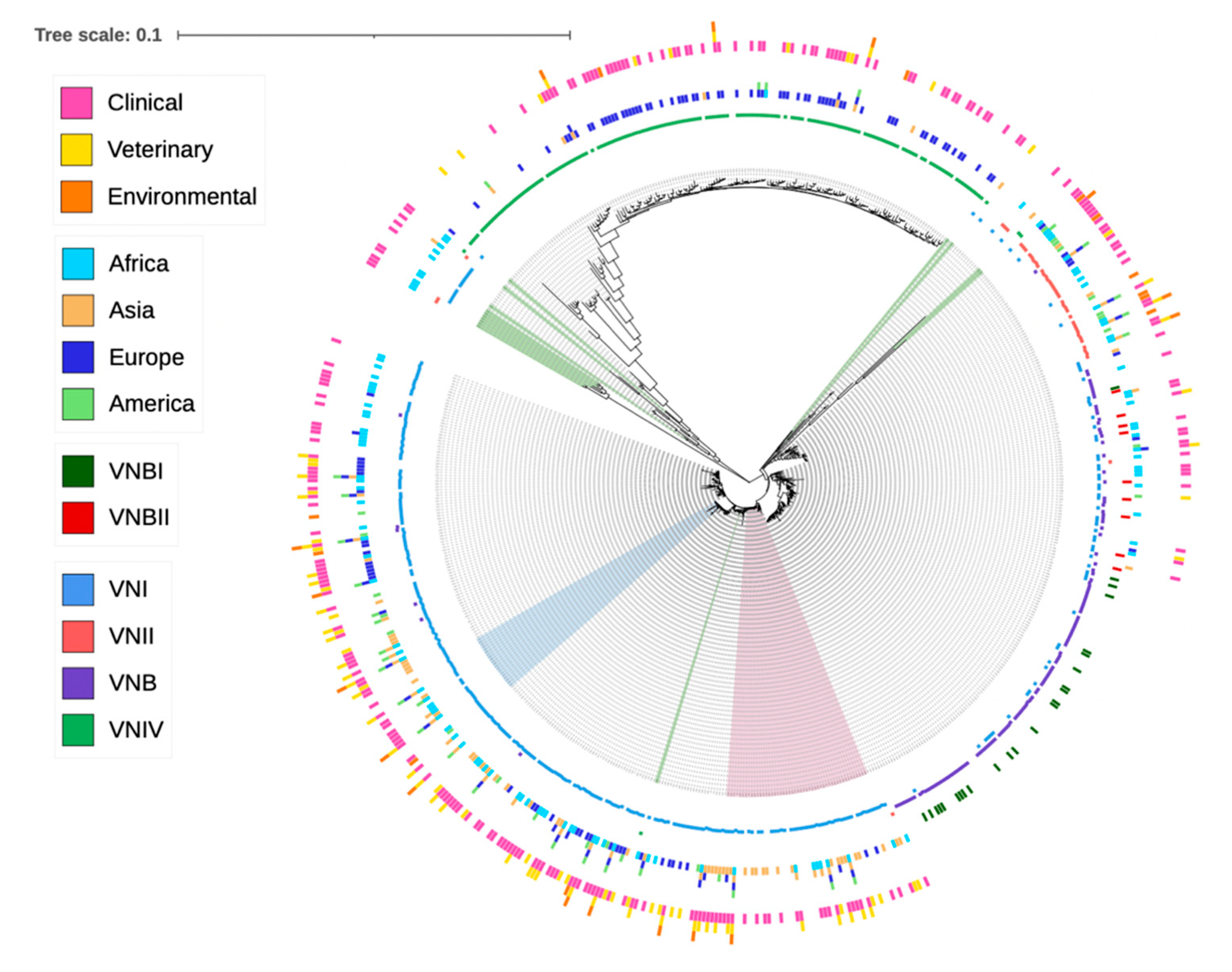
8]. Analyses of the concatenated sequences at the seven MLST loci showed a largely consistent clustering of STs into their original molecular type designations, with VNIV being the most distant from VNI, VNII and VNB (
Figure 1). Similarly, except for one ST (ST434) that was originally assigned to VNBII but was clustered more closely with VNBI strains, all other STs originally assigned to VNBI and VNBII were clearly separated into two groups (
Figure 1). However, there were several other notable inconsistencies. Specifically, six STs originally assigned to VNIV (ST521, ST254, ST266, ST355, ST489 and ST538) showed a closer relationship with the VNI clade. In contrast, 14 STs originally assigned to VNI (ST210, ST224, ST225, ST249, ST259, ST263, ST326, ST345, ST353, ST354, ST358, ST365, ST366 and ST651) and three STs originally assigned to VNII (ST221, ST222 and ST363) showed intermediate phylogenetic placing between the major serotypes A and D genotypes. Interestingly, 15 of the above 23 STs contained alleles with mixed clustering patterns, where some of the alleles belonged to the serotype A cluster, while others belonged to the serotype D allele cluster (
Table 4;
Supplementary Figures S1–S7). In addition, multiple STs originally assigned to molecular types VNI, VNII and VNB showed ambiguous placements within serotype A, often showing large distances from the three main molecular types (
Figure 1). In contrast, 51 STs with previously undefined molecular type assignments were grouped into various species/molecular types (
Figure 1). Phylogenic trees showing relationships among allele sequences for each of the seven genes can be seen in
Supplementary Figures S1–S7.
3.4. AMOVA
Because of the highly skewed population sizes among countries, with seven countries each having fewer than five isolates represented, our AMOVA was conducted separately at the continental and country levels, instead of through a two-level hierarchical analysis. At the country level, only those with more than five isolates represented are included. The overall objective of our AMOVA was to assess how much geographic separations contributed to the total genetic variation. Below, we briefly summarize the results.
At the continental level analyses, in the none-clone-corrected sample, genetic variations within continents contributed 72%, 78% and 84% of the total observed genetic variations in the total CNSC population, the serotype A population and the serotype D population, respectively. The remaining 28%, 22% and 16% were attributed among continents. The within-continent and among-continent contributions for each of the three taxonomic populations were statistically significant at the
p < 0.001 level (
Table 5). In the three clone-corrected samples, genetic variations within continents contributed 96%, 98% and 96% of the total observed genetic variations in the entire CNSC population, the serotype A population and the serotype D population, respectively. These percentages were significantly greater than those without clone corrections. The remaining 4%, 2% and 4% were attributed among continents. Despite the smaller percentages of contributions, the among-continent contributions for two of the three population types were statistically significant at
p < 0.001, while the serotype D population was significant at
p = 0.03 (
Table 5). The pairwise comparisons between continents for the three taxonomic samples are shown in
Table 6.
At the country level, in the non-clone-corrected sample analyses, genetic variations within countries contributed 39%, 41% and 74% of the total observed genetic variations in the total CNSC sample, the serotype A sample and the serotype D sample, respectively. The remaining 61%, 59% and 26% were attributed among countries. The within-country and among-country contributions for each of the three taxonomic sample types were statistically significant at the
p < 0.001 level (
Table 7). In the three clone-corrected samples, genetic variations within countries contributed 83%, 79% and 99% of the total observed genetic variations in the total CNSC sample, the serotype A sample and the serotype D sample, respectively. Similar to those observed at the continental level, these percentages by within-countries in the clone-corrected samples were significantly greater than those without clone corrections. The remaining 17%, 21% and 1% were attributed among countries. Despite the smaller percentages of contributions, except for the serotype D sample, the remaining two among-country contributions for the three sample types were statistically significant at the
p < 0.001 level (
Table 7). The pairwise comparisons among countries for the three taxonomic samples are shown in
Supplementary Tables S4–S6.
3.5. Recombination & Linkage Disequilibrium
We investigated the potential signatures of recombination among different samples of CNSC using two common indicators: phylogenetic incompatibility and linkage equilibrium. Here, aside from the three large taxonomic samples (the total CNSC, serotype A and serotype D), we also separately analyzed the three major molecular types (VNI, VNII and VNB) within serotype A, two subclades (VNBI and VNBII) within VNB as well as the two genotype clusters closely related to the two most dominant STs (ST5 and ST93) in the global sample.
In the phylogenetic incompatibility test, we found that none of the 10 samples showed 100% phylogenetic compatibility (
Table 8). Specifically, 4 (the total CNSC, serotype A, serotype D and clade VNI) of the 10 analyzed datasets showed no phylogenetic compatibility among the seven loci, which was consistent with the evidence of recombination among all 21 pairs of loci within each of the four samples. For VNII-, VNBI- and ST5-associated genotype groups, 16 of the 21 pairwise loci combinations were phylogenetically incompatible, with only five pairs (23.8%) being phylogenetically compatible. For the VNB and VNBII datasets, 19 of the 21 pairwise loci were phylogenetically incompatible. For the ST93-associated genotype group, 5 of the 21 pairs showed phylogenetic incompatibility, which was also consistent with the evidence for recombination in this sample.
Linkage disequilibrium analyses revealed that in nine of the ten samples, the null hypothesis of random recombination was rejected (
Table 8). The only exception was the ST93-associated genotype group, where the null hypothesis of random recombination was not rejected, likely due to the small sample size and the lack of statistical power to reject the null hypothesis. However, variable numbers of pairs of loci within each of the ten samples showed no significant deviation from those expected under the random recombination hypothesis (
Supplementary Table S7). For example, in the VNII sample, 4 of the 21 loci pairs had observed genotype frequencies not significantly different from random recombination, with all 4 involving the
IGS1 locus. In the VNB sample, 9 of the 21 loci pairs had observed genotype frequencies not significantly different from random recombination. Interestingly, while no evidence for linkage equilibrium across all 21 loci pairs was observed in the ST93-associated genotype cluster, there was abundant evidence for linkage equilibrium between pairs of loci within the ST5-associated genotype cluster (
Table 9). The complete allelic profiles for these clusters are shown in
Supplementary Tables S8 and S9.
4. Discussion
This study analyzed the genetic structure of geographic populations of the
C. neoformans complex based on published multilocus sequence data. We focused on 296 STs with available geographical data. Our analyses included a robust global population of CNSC, with 4200 isolates originating across 31 countries and four continents (South and North America combined as one continent). Of the 296 sequence types with geographical data, 244 (82%) were sampled from a single continent, with 24 of these 244 STs found across multiple countries within a continent. The remaining 18% STs were distributed across multiple continents, with 28 STs (9%) found in two continents, 9 STs (3%) among three continents and 15 STs (5%) across all four continents. The broad distributions of multiple sequence types across multiple continents and countries are consistent with the recent and frequent gene flow in CNSC. Contemporary factors such as wind, animal and human migrations and other anthropogenic activities could all have facilitated the dispersals of genes and genotypes, causing wide distributions of certain genotypes [
8,
59,
60,
61].
However, our population genetic analyses revealed statistically significant differentiations among continental and national populations of CNSC. At the whole CNSC level, the observed genetic differentiations were contributed by differences in the distributions of the four molecular types and by the localized clonal expansion of specific sequence types. Indeed, evidence for the clonal expansion of specific genotypes was found for all molecular types. For example, the most abundantly collected ST in the analyzed data was ST5, represented by 1332 isolates. The second most abundant was ST93, represented by 460 isolates. Even though both were found in all four continents, ST5 and ST93 were mainly found in Asia (1211 of the 1332 isolates) and the Americas (224 of the 460 isolates), respectively. Among the serotype D (VNIV) isolates, ST160 was the most recorded sequence type, representing 78 isolates across three continents. Localized clonal expansion can significantly skew the allele and genotype frequencies and contribute to observed genetic differences among geographic populations [
62]. Thus, we analyzed clone-corrected samples where only one representative from each country was included for analyses. Using the clone-corrected samples, the amount of contribution due to geographic separation to the total genetic variance reduced by 70–93% at the continental level and by 60–93% at the country level for the total CNSC, the serotype A and the serotype D samples, respectively. This result is consistent with the presence of indigenous genetic variations within most national and continental populations, likely due to historical differentiations.
Interestingly, out of the 529 isolates representing 109 STs collected within Africa, only 2 isolates belonged to serotype D (VNIV). Our meta-analysis was consistent with an earlier study that analyzed the molecular types of 505 isolates from Africa and found that none of them belonged to molecular type VNIV [
61]. However, these results do not mean that VNIV is unimportant in Africa. For example, one study that analyzed 252 isolates from South Africa identified 5 cases of molecular type VNIV [
36]. However, those isolates were not genotyped using MLST. At present, Africa accounts for the greatest global burden of HPC infection [
63,
64] and contains the most genetically diverse population of serotype A [
36], including the relatively frequent distributions of both mating types. The high genetic diversity of serotype A strains in Africa has led to the “Out of Africa” hypothesis for the origin and spread of serotype A [
65]. Interestingly, four STs from Africa that were originally identified as belonging to VNI were clustered to the basal clade of VNIV (
Figure 1). These STs likely represent some of the ancestor genotypes of VNIV or recent hybrids between the VNI and VNIV strains.
Among the 72 serotype D (VNIV) STs, 65 (90%) were represented in Europe. In comparison, only 23% of serotype A STs were found in Europe. The results are consistent with multiple studies reporting a relatively high prevalence of serotype D within Europe [
66]. The relatively broad distributions of both serotype A and serotype D strains in Europe are likely the main contributors to the frequent observations of serotype AD hybrids within Europe [
66].
Although there is an abundant
C. neoformans population in North America [
67,
68], only 12 isolates have been analyzed using the ISHAM MLST scheme. Such a lack of MLST data from North America was not due to the lack of samples for analyses. Indeed, between 1992 and 1994, the US CDC conducted a large-scale surveillance of the agents of cryptococcosis [
69]. Those isolates were genotyped using random amplified polymorphic DNA and/or multilocus enzyme electrophoresis, the commonly used molecular markers at that time, revealing abundant genetic variations, including at least three independent hybridizing events between serotypes A and D [
8,
59,
69,
70]. However, those strains, as well as many strains isolated afterwards from North America, have not been genotyped using the ISHAM MLST scheme, which was published in 2009. It would be very interesting to analyze the North American population of
C. neoformans and to compare them with those from other parts of the world. In contrast to the lack of MLST data from North America, there is a large representation of
C. neoformans from China, making that population of CNSC one of the best for understanding fine-scale spatial and temporal structures of CNSC.
In this study, we performed a phylogenetic analysis of all STs based on their concatenated DNA sequences. Overall, the phylogenetic results were consistent with the molecular type designations for most isolates based on PCR fingerprinting, AFLP and/or PCR-RFLP of the URA5 gene fragment. However, we found that the placements of a small number of STs in the phylogenetic trees were inconsistent with their original molecular type designations (
Figure 1). Such inconsistencies were found among molecular types within serotype A as well as between serotype A and serotype D. Among these inconsistently placed STs, 15 contained mixtures of alleles from serotype A and serotype D (
Table 4). These STs were likely recombinants derived from the hybridization between serotypes A and D strains. After hybridization, either meiotic or mitotic recombination could have led to a loss of heterozygosity (LOH) to generate the haploid recombinant genotypes observed here. Indeed, LOH in serotype AD hybrids has been observed through both meiosis and mitosis, with environmental stress facilitating LOH [
71,
72]. The observed recombination is also consistent with the results from nuclear and mitochondrial genome phylogenetic comparisons, where the mitochondrial genome-based phylogeny showed several differences with that based on nuclear genome-based phylogeny within CNSC [
73]. Furthermore, some of these STs had distinct alleles and showed significant divergence with the main serotype A and the main serotype D genotypes and clades, a result suggesting the existence of distinct novel lineages within CNSC (
Figure 1,
Supplementary Figures S1–S7). Finally, our phylogenetic analyses successfully placed 51 previously undefined STs into the phylogenetic framework and revealed their possible origins. The presence of these hybrids, as well as many intermediate STs with ambiguous molecular type assignments in the phylogeny, supports the continued use of CNSC for this group of fungal pathogens [
9].
CNSC has been shown to reproduce predominantly asexually in nature, but evidence for recombination has been reported for both the serotype A and serotype D populations [
28,
74,
75,
76]. Sexual reproduction can accelerate adaptation to diverse environments and remove deleterious mutations more effectively than asexual reproduction [
77]. Across all four major molecular types (VNI, VNII, VNB and VNIV) as well as two subtypes of VNB (VNBI and VNBII), we found clear evidence of recombination within each. Significantly, evidence for recombination was also found in two presumed “clonal ST clusters” within VNI. The observed results suggest that, even in geographic populations of CNSC dominated by a few STs, recombination is still possible. Such recombination could be achieved through same-sex mating or opposite-sex mating, as suggested previously when regional populations of CNSC were found to contain evidence of recombination, despite only one mating type (MATα) being found in those analyzed samples [
74,
75].
In conclusion, analyses of the published MLST data for CNSC allowed us to quantify the genetic diversity within and among geographic populations of this important human pathogenic yeast. Our analyses revealed evidence for historical geographic differentiations of CNSC, both at the whole CNSC level as well as within the populations of serotypes A and D. Not surprisingly, evidence for the clonal expansion of many STs was found at both the local population level as well as across countries and continents, suggesting the potentially important roles of recent anthropogenic activities in the dispersals of alleles and genotypes of CNSC. Importantly, we found evidence for recombination within all molecular types, including at least two presumed “clonal ST clusters” within VNI. The results indicate the diverse methods of CNSC reproduction in nature. While a large number of isolates were analyzed in this study for population genetic patterns, about 55% of the 657 STs in the ISHAM MLST database were not included for geographic structure analyses due to the lack of geographic location information associated with these 361 STs. In the future, authors should be required to submit the metadata associated with each isolate and each sequence type that they publish in their MLST study. Additional information on these STs, as well as more MLST data, especially from under-reported regions such as North America, will provide a more comprehensive understanding of the global population structure of CNSC and help develop more realistic models of global cryptococcal threat predictions and management strategies against cryptococcosis [
78].