Unwinding BRAHMA Functions in Plants
Abstract
1. Introduction
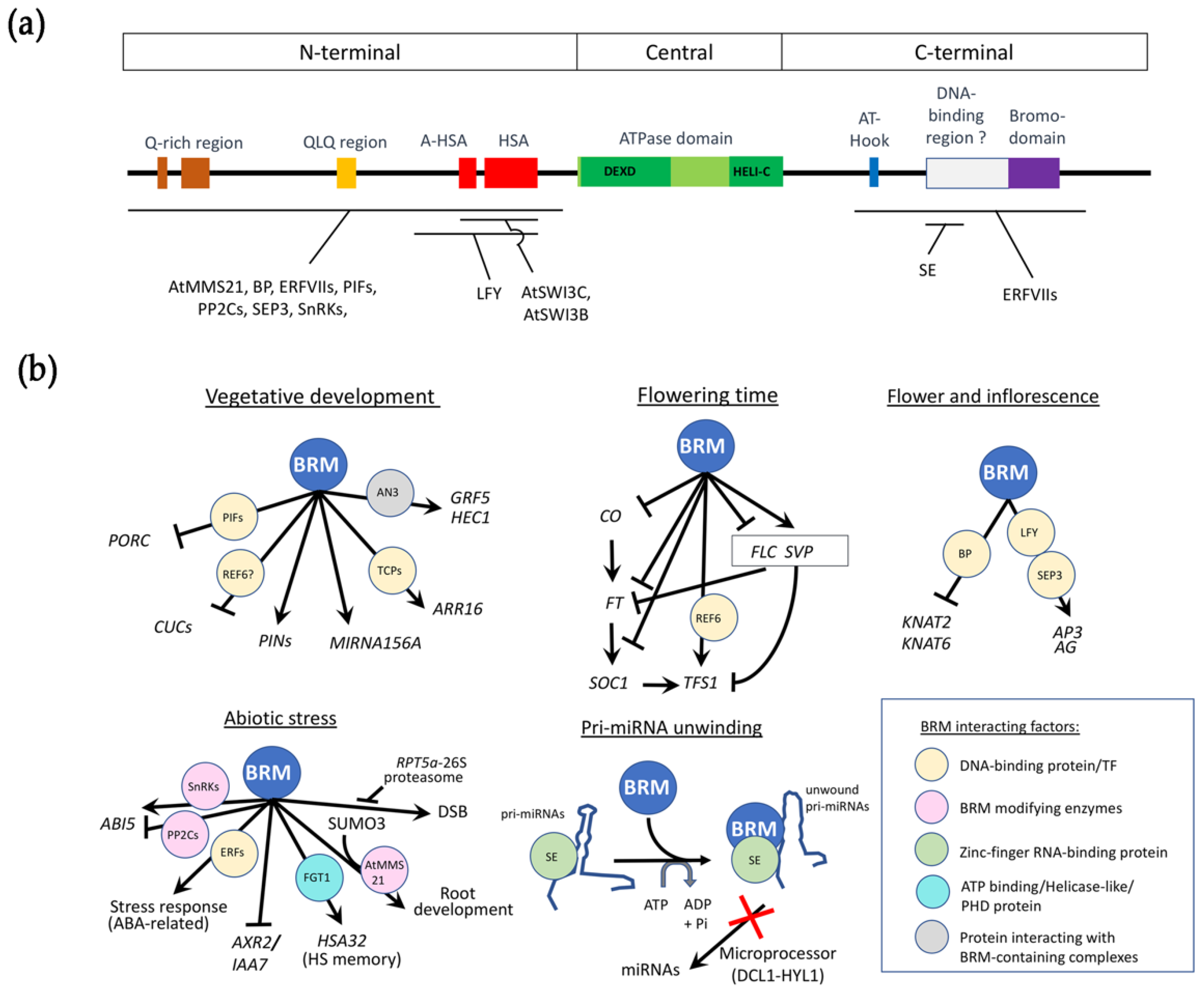
2. BRM Functional Domains Are Conserved in Plants
3. Genome-Wide Functions of BRM in Plants
4. BRM Is Required throughout Plant Development
4.1. BRM Is Involved in Vegetative Development
4.2. BRM Controls Flowering Time
4.3. BRM Shapes Inflorescence and Flower Development
4.4. BRM Has Roles in Abiotic Stress and Hormone Pathways
5. BRM Plays a Chromatin-Independent Role in pri-miRNA Processing
6. Discussion
7. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Luger, K.; Mäder, A.W.; Richmond, R.K.; Sargent, D.F.; Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 1997, 389, 251–260. [Google Scholar] [CrossRef] [PubMed]
- Tamkun, J.W.; Deuring, R.; Scott, M.P.; Kissinger, M.; Pattatucci, A.M.; Kaufman, T.C.; Kennison, J.A. brahma: A regulator of Drosophila homeotic genes structurally related to the yeast transcriptional activator SNF2SWI2. Cell 1992, 68, 561–572. [Google Scholar] [CrossRef]
- Narlikar, G.J.; Fan, H.-Y.; Kingston, R.E. Cooperation between Complexes that Regulate Chromatin Structure and Transcription. Cell 2002, 108, 475–487. [Google Scholar] [CrossRef]
- Phelan, M.L.; Narlikar, G.J.; Kingston, R.E. Reconstitution of a Core Chromatin Remodeling Complex from SWI/SNF Subunits. Mol. Cell 1999, 3, 247–253. [Google Scholar] [CrossRef]
- Sarnowska, E.; Gratkowska, D.M.; Sacharowski, S.P.; Cwiek, P.; Tohge, T.; Fernie, A.R.; Siedlecki, J.A.; Koncz, C.; Sarnowski, T.J. The Role of SWI/SNF Chromatin Remodeling Complexes in Hormone Crosstalk. Trends Plant Sci. 2016, 21, 594–608. [Google Scholar] [CrossRef] [PubMed]
- Bezhani, S.; Winter, C.; Hershman, S.; Wagner, J.D.; Kennedy, J.F.; Kwon, C.S.; Pfluger, J.; Su, Y.; Wagner, D. Unique, Shared, and Redundant Roles for the Arabidopsis SWI/SNF Chromatin Remodeling ATPases BRAHMA and SPLAYED. Plant Cell 2007, 19, 403–416. [Google Scholar] [CrossRef]
- Wagner, D.; Meyerowitz, E.M. SPLAYED, a novel SWI/SNF ATPase homolog, controls reproductive development in Arabidopsis. Curr. Biol. 2002, 12, 85–94. [Google Scholar] [CrossRef]
- Farrona, S.; Hurtado, L.; Bowman, J.L.; Reyes, J.C. The Arabidopsis thaliana SNF2 homolog AtBRM controls shoot development and flowering. Development 2004, 131, 4965–4975. [Google Scholar] [CrossRef]
- Hurtado, L.; Farrona, S.; C, R.J. The putative SWI/SNF complex subunit BRAHMA activates flower homeotic genes in Arabidopsis thaliana. Plant Mol. Biol. 2006, 62, 291–304. [Google Scholar] [CrossRef]
- Kwon, C.S.; Hibara, K.; Pfluger, J.; Bezhani, S.; Metha, H.; Aida, M.; Tasaka, M.; Wagner, D. A role for chromatin remodeling in regulation of CUC gene expression in the Arabidopsis cotyledon boundary. Development 2006, 133, 3223–3230. [Google Scholar] [CrossRef]
- Han, S.-K.; Wu, M.-F.; Cui, S.; Wagner, D. Roles and activities of chromatin remodeling ATPases in plants. Plant J. 2015, 83, 62–77. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.M.-F.; Sang, Y.Y.; Bezhani, S.; Yamaguchi, N.; Han, S.-K.S.-K.; Li, Z.; Su, Y.; Slewinski, T.L.; Wagner, D. SWI2/SNF2 chromatin remodeling ATPases overcome polycomb repression and control floral organ identity with the LEAFY and SEPALLATA3 transcription factors. Proc. Natl. Acad. Sci. USA 2012, 109, 3576–3581. [Google Scholar] [CrossRef]
- Trotter, K.W.; Fan, H.-Y.; Ivey, M.L.; Kingston, R.E.; Archer, T.K. The HSA domain of BRG1 mediates critical interactions required for glucocorticoid receptor-dependent transcriptional activation in vivo. Mol. Cell. Biol. 2008, 28, 1413–1426. [Google Scholar] [CrossRef] [PubMed]
- Farrona, S.; Hurtado, L.; Reyes, J.C. A Nucleosome Interaction Module Is Required for Normal Function of Arabidopsis thaliana BRAHMA. J. Mol. Biol. 2007, 373, 240–250. [Google Scholar] [CrossRef] [PubMed]
- Zaware, N.; Zhou, M.-M. Bromodomain biology and drug discovery. Nat. Struct. Mol. Biol. 2019, 26, 870–879. [Google Scholar] [CrossRef]
- Papillon, J.P.N.; Nakajima, K.; Adair, C.D.; Hempel, J.; Jouk, A.O.; Karki, R.G.; Mathieu, S.; Mo, H.; Ntaganda, R.; Smith, T.; et al. Discovery of Orally Active Inhibitors of Brahma Homolog (BRM)/SMARCA2 ATPase Activity for the Treatment of Brahma Related Gene 1 (BRG1)/SMARCA4-Mutant Cancers. J. Med. Chem. 2018, 61, 10155–10172. [Google Scholar] [CrossRef] [PubMed]
- Shen, W.; Xu, C.; Huang, W.; Zhang, J.; Carlson, J.E.; Tu, X.; Wu, J.; Shi, Y. Solution structure of human Brg1 bromodomain and its specific binding to acetylated histone tails. Biochemistry 2007, 46, 2100–2110. [Google Scholar] [CrossRef]
- Archacki, R.; Sarnowski, T.J.; Halibart-Puzio, J.; Brzeska, K.; Buszewicz, D.; Prymakowska-Bosak, M.; Koncz, C.; Jerzmanowski, A. Genetic analysis of functional redundancy of BRM ATPase and ATSWI3C subunits of Arabidopsis SWI/SNF chromatin remodelling complexes. Planta 2009, 229, 1281–1292. [Google Scholar] [CrossRef]
- Gebuhr, T.C.; Bultman, S.J.; Magnuson, T. Pc-G/trx-G and the SWI/SNF Connection : Developmental Gene Regulation Through Chromatin Remodeling. Genesis 2000, 26, 189–197. [Google Scholar] [CrossRef]
- Schuettengruber, B.; Bourbon, H.M.; Di Croce, L.; Cavalli, G. Genome Regulation by Polycomb and Trithorax: 70 Years and Counting. Cell 2017, 171, 34–57. [Google Scholar] [CrossRef]
- Xu, Y.; Guo, C.; Zhou, B.; Li, C.; Wang, H.; Zheng, B.; Ding, H.; Zhu, Z.; Peragine, A.; Cui, Y.; et al. Regulation of Vegetative Phase Change by SWI2/SNF2 Chromatin Remodeling ATPase BRAHMA. Plant Physiol. 2016, 172, 2416–2428. [Google Scholar] [CrossRef]
- Li, C.; Chen, C.; Gao, L.; Yang, S.; Nguyen1, V.; Shi, X.; Siminovitch, K.; Kohalmi, S.E.; Huang, S.; Wu, K.; et al. The Arabidopsis SWI2/SNF2 Chromatin Remodeler BRAHMA Regulates Polycomb Function during Vegetative Development and Directly Activates the Flowering Repressor SVP. PLoS Genet. 2015, 11, e1004944. [Google Scholar] [CrossRef]
- Li, C.; Gu, L.; Gao, L.; Chen, C.; Wei, C.-Q.; Qiu, Q.; Chien, C.-W.; Wang, S.; Jiang, L.; Ai, L.-F.; et al. Concerted genomic targeting of H3K27 demethylase REF6 and chromatin-remodeling ATPase BRM in Arabidopsis. Nat. Genet. 2016, 48, 687–693. [Google Scholar] [CrossRef]
- Lu, F.; Cui, X.; Zhang, S.; Jenuwein, T.; Cao, X. Arabidopsis REF6 is a histone H3 lysine 27 demethylase. Nat. Genet. 2011, 43, 715–719. [Google Scholar] [CrossRef] [PubMed]
- Torres, E.S.; Deal, R.B. The histone variant H2A.Z and chromatin remodeler BRAHMA act coordinately and antagonistically to regulate transcription and nucleosome dynamics in Arabidopsis. Plant J. 2019, 99, 144–162. [Google Scholar] [CrossRef] [PubMed]
- Archacki, R.; Yatusevich, R.; Buszewicz, D.; Krzyczmonik, K.; Patryn, J.; Iwanicka-nowicka, R.; Biecek, P.; Wilczynski, B.; Koblowska, M.; Jerzmanowski, A.; et al. Arabidopsis SWI/SNF chromatin remodeling complex binds both promoters and terminators to regulate gene expression. Nucleic Acids Res. 2017, 45, 3116–3129. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, T.; Tsujimoto-inui, Y.; Sotta, N.; Hirakawa, T.; Matsunaga, T.M.; Fukao, Y.; Matsunaga, S.; Fujiwara, T. Proteasomal degradation of BRAHMA promotes Boron tolerance in Arabidopsis. Nat. Commun. 2018, 9, 5285. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.; Lu, F.; Qiu, Q.; Zhou, B.; Gu, L.; Zhang, S.; Kang, Y.; Cui, X.; Ma, X.; Yao, Q.; et al. REF6 recognizes a specific DNA sequence to demethylate H3K27me3 and regulate organ boundary formation in Arabidopsis. Nat. Genet. 2016, 48, 694–699. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Leichty, A.R.; Hu, T.; Poethig, R.S. H2A.Z promotes the transcription of MIR156A and MIR156C in Arabidopsis by facilitating the deposition of H3K4me3. Development 2018, 145, dev152868. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Li, Y.; Zhang, X.; Zha, P.; Lin, R. The SWI2/SNF2 Chromatin-Remodeling ATPase BRAHMA Regulates Chlorophyll Biosynthesis in Arabidopsis. Mol. Plant 2017, 10, 155–167. [Google Scholar] [CrossRef]
- Jégu, T.; Veluchamy, A.; Ramirez-prado, J.S.; Rizzi-paillet, C.; Perez, M.; Lhomme, A.; Latrasse, D.; Coleno, E.; Vicaire, S.; Legras, S.; et al. The Arabidopsis SWI/SNF protein BAF60 mediates seedling growth control by modulating DNA accessibility. Genome Biol. 2017, 18, 114. [Google Scholar] [CrossRef]
- Tang, X.; Hou, A.; Babu, M.; Nguyen, V.; Hurtado, L.; Lu, Q.; Reyes, J.C.; Wang, A.; Keller, W.A.; Harada, J.J.; et al. The Arabidopsis BRAHMA chromatin-remodeling ATPase is involved in repression of seed maturation genes in leaves. Plant Physiol. 2008, 147, 1143–1157. [Google Scholar] [CrossRef] [PubMed]
- Vercruyssen, L.; Verkest, A.; Gonzalez, N.; Heyndrickx, K.S.; Eeckhout, D.; Han, S.; Archacki, R.; Van Leene, J.; Andriankaja, M.; De Bodt, S.; et al. ANGUSTIFOLIA3 Binds to SWI/SNF Chromatin Remodeling Complexes to Regulate Transcription during Arabidopsis Leaf Development. Plant Cell 2014, 26, 210–229. [Google Scholar] [CrossRef] [PubMed]
- Farrona, S.; Coupland, G.; Turck, F. The impact of chromatin regulation on the floral transition. Semin. Cell Dev. Biol. 2008, 19, 560–573. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Farrona, S.; Hurtado, L.; March-Diaz, R.; Schmitz, R.J.; Florencio, F.J.; Turck, F.; Amasino, R.M.; Reyes, J.C. Brahma Is Required for Proper Expression of the Floral Repressor FLC in Arabidopsis. PLoS ONE 2011, 6, e17997. [Google Scholar] [CrossRef] [PubMed]
- Richter, R.; Kinoshita, A.; Vincent, C.; Martinez-gallegos, R.; Gao, H.; Van Driel, A.D.; Hyun, Y.; Mateos, J.L.; Coupland, G. Floral regulators FLC and SOC1 directly regulate expression of the B3-type transcription factor TARGET OF FLC AND SVP 1 at the Arabidopsis shoot apex via antagonistic chromatin modifications. PLoS Genet. 2019, 15, e1008065. [Google Scholar] [CrossRef]
- Deal, R.B.; Topp, C.N.; McKinney, E.C.; Meagher, R.B. Repression of Flowering in Arabidopsis Requires Activation of FLOWERING LOCUS C Expression by the Histone Variant H2A.Z. Plant Cell 2007, 19, 74–83. [Google Scholar] [CrossRef]
- Kumar, S.V.; Wigge, P.A. H2A.Z-Containing Nucleosomes Mediate the Thermosensory Response in Arabidopsis. Cell 2010, 140, 136–147. [Google Scholar] [CrossRef] [PubMed]
- Martin-Trillo, M.; Lázaro, A.; Poethig, R.S.; Gómez-mena, C.; Piñeiro, M.A.; Martinez-zapater, J.M.; Jarillo, J.A. EARLY IN SHORT DAYS 1 (ESD1) encodes ACTIN-RELATED PROTEIN 6 (AtARP6), a putative component of chromatin remodelling complexes that positively regulates FLC accumulation in Arabidopsis. Development 2006, 1252, 1241–1252. [Google Scholar] [CrossRef]
- Levin, J.Z.; Fletcher, J.C.; Chen, X.; Meyerowitz, E.M. A Genetic Screen for Modifiers of UFO Meristem\rActivity Identifies Three Novel FUSED FLORAL\rORGANS Genes Required for Early Flower\rDevelopment in Arabidopsis. Genetics 1998, 149, 579–595. [Google Scholar]
- Sayou, C.; Monniaux, M.; Nanao, M.H.; Moyroud, E.; Brockington, S.F.; Thévenon, E.; Chahtane, H.; Warthmann, N.; Melkonian, M.; Zhang, Y.; et al. A promiscuous intermediate underlies the evolution of LEAFY DNA binding specificity. Science 2014, 343, 645–648. [Google Scholar] [CrossRef]
- Muiño, J.M.; de Bruijn, S.; Pajoro, A.; Geuten, K.; Vingron, M.; Angenent, G.C.; Kaufmann, K. Evolution of DNA-Binding Sites of a Floral Master Regulatory Transcription Factor. Mol. Biol. Evol. 2016, 33, 185–200. [Google Scholar] [CrossRef] [PubMed]
- Smaczniak, C.; Immink, R.G.H.; Muiño, J.M.; Blanvillain, R.; Busscher, M. Characterization of MADS-domain transcription factor complexes in Arabidopsis flower development. Proc. Natl. Acad. Sci. USA 2012, 109, 1560–1565. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Yang, S.; Chen, C.; Li, C.; Shan, W. Arabidopsis BREVIPEDICELLUS Interacts with the SWI2/SNF2 Chromatin Remodeling ATPase BRAHMA to Regulate KNAT2 and KNAT6 Expression in Control of Inflorescence Architecture. PLoS Genet. 2015, 11, e1005125. [Google Scholar] [CrossRef] [PubMed]
- Smith, H.M.S.; Boschke, I.; Hake, S. Selective interaction of plant homeodomain proteins mediates high DNA-binding affinity. Proc. Natl. Acad. Sci. USA 2002, 99, 9579–9584. [Google Scholar] [CrossRef]
- Han, S.-K.; Sang, Y.; Rodrigues, A.; F2010, B.; Wu, M.-F.; Rodriguez, P.L.; Wagner, D. The SWI2/SNF2 Chromatin Remodeling ATPase BRAHMA Represses Abscisic Acid Responses in the Absence of the Stress Stimulus in Arabidopsis. Plant Cell 2012, 24, 4892–4906. [Google Scholar] [CrossRef]
- Peirats-Llobet, M.; Han, S.K.; Gonzalez-Guzman, M.; Jeong, C.W.; Rodriguez, L.; Belda-Palazon, B.; Wagner, D.; Rodriguez, P.L. A Direct Link between Abscisic Acid Sensing and the Chromatin-Remodeling ATPase BRAHMA via Core ABA Signaling Pathway Components. Mol. Plant 2016, 9, 136–147. [Google Scholar] [CrossRef]
- Archacki, R.; Buszewicz, D.; Sarnowski, T.J.; Sarnowska, E.; Rolicka, A.T. BRAHMA ATPase of the SWI/SNF Chromatin Remodeling Complex Acts as a Positive Regulator of Gibberellin- Mediated Responses in Arabidopsis. PLoS ONE 2013, 8, e58588. [Google Scholar] [CrossRef]
- Efroni, I.; Han, S.; Kim, H.J.; Wu, M.; Steiner, E.; Birnbaum, K.D.; Hong, J.C.; Eshed, Y.; Wagner, D. Regulation of Leaf Maturation by Chromatin-Mediated Modulation of Cytokinin Responses. Dev. Cell 2013, 24, 438–445. [Google Scholar] [CrossRef]
- Yang, S.; Li, C.; Zhao, L.; Gao, S.; Lu, J.; Zhao, M.; Chen, C.-Y.; Liu, X.; Luo, M.; Cui, Y.; et al. The Arabidopsis SWI2/SNF2 Chromatin Remodeling ATPase BRAHMA Targets Directly to PINs and Is Required for Root Stem Cell Niche Maintenance. Plant Cell 2015, 27, 1670–1680. [Google Scholar] [CrossRef]
- Nguyen, H.N.; Jung, C.; Cheong, J. Chromatin remodeling for the transcription of type 2C protein phosphatase genes in response to salt stress. Plant Physiol. Biochem. 2019, 141, 325–331. [Google Scholar] [CrossRef] [PubMed]
- Vain, T.; Raggi, S.; Ferro, N.; Barange, D.K.; Kieffer, M.; Ma, Q.; Doyle, S.M.; Thelander, M.; Pařízková, B.; Novák, O.; et al. Selective auxin agonists induce specific AUX/IAA protein degradation to modulate plant development. Proc. Natl. Acad. Sci. USA 2019, 116, 6463–6472. [Google Scholar] [CrossRef] [PubMed]
- Vicente, J.; Mendiondo, G.M.; Movahedi, M.; Charng, Y.; Gray, J.E.; Holdsworth, M.J.; Vicente, J.; Mendiondo, G.M.; Movahedi, M.; Peirats-llobet, M.; et al. The Cys-Arg/N-End Rule Pathway Is a General Sensor of Abiotic Stress in Flowering Plants Report The Cys-Arg/N-End Rule Pathway Is a General Sensor of Abiotic Stress in Flowering Plants. Curr. Biol. 2017, 27, 3183–3190. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Lai, J.; Wang, F.; Yang, S.; He, Z.; Jiang, J.; Li, Q.; Wu, Q.; Liu, Y.; Yu, M.; et al. A SUMO Ligase AtMMS21 Regulates the Stability of the Chromatin Remodeler BRAHMA in Root Development. Plant Physiol. 2017, 173, 1574–1582. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Qi, Y.; Liu, M.; Yang, C. SUMO E3 Ligase AtMMS21 Regulates Drought Tolerance in Arabidopsis thaliana. J. Integr. Plant Biol. 2013, 55, 83–95. [Google Scholar] [CrossRef]
- Liu, Y.; Lai, J.; Yu, M.; Wang, F.; Zhang, J.; Jiang, J.; Hu, H.; Wu, Q.; Lu, G.; Xu, P.; et al. The Arabidopsis SUMO E3 Ligase AtMMS21 Dissociates the E2Fa/DPa Complex in Cell Cycle Regulation. Plant Cell 2016, 28, 2225–2237. [Google Scholar] [CrossRef]
- Yuan, D.; Lai, J.; Xu, P.; Zhang, S.; Zhang, J.; Li, C.; Wang, Y.; Du, J.; Liu, Y.; Yang, C. AtMMS21 regulates DNA damage response and homologous recombination repair in Arabidopsis. DNA Repair (Amst.) 2014, 21, 140–147. [Google Scholar] [CrossRef]
- Brzezinka, K.; Altmann, S.; Czesnick, H.; Nicolas, P.; Gorka, M.; Benke, E.; Kabelitz, T.; Jähne, F.; Graf, A.; Kappel, C.; et al. Arabidopsis FORGETTER1 mediates stress-induced chromatin memory through nucleosome remodeling. Elife 2016, 5, 1–23. [Google Scholar] [CrossRef]
- Wang, Z.; Sun, D.; Li, Y.; Yu, B.; Zhao, B.; Li, P.; Zhang, X. SWI2/SNF2 ATPase CHR2 remodels pri-miRNAs via Serrate to impede miRNA production. Nature 2018, 557, 516–521. [Google Scholar] [CrossRef]
- Vachon, G.; Engelhorn, J.; Carles, C.C. Interactions between transcription factors and chromatin regulators in the control of flower development. J. Exp. Bot. 2018, 69, 2461–2471. [Google Scholar] [CrossRef]
- Hou, X.; Zhou, J.; Liu, C.; Liu, L.; Shen, L.; Yu, H. Nuclear factor Y-mediated H3K27me3 demethylation of the SOC1 locus orchestrates flowering responses of Arabidopsis. Nat. Commun. 2014, 5, 4601. [Google Scholar] [CrossRef] [PubMed]
- To, T.; Kim, J.-M. Epigenetic regulation of gene responsiveness in Arabidopsis. Front. Plant Sci. 2014, 4. [Google Scholar] [CrossRef] [PubMed]
- Jégu, T.; Latrasse, D.; Delarue, M.; Hirt, H.; Domenichini, S.; Ariel, F.; Crespi, M.; Bergounioux, C.; Raynaud, C.; Benhamed, M. The BAF60 Subunit of the SWI/SNF Chromatin-Remodeling Complex Directly Controls the Formation of a Gene Loop at FLOWERING LOCUS C in Arabidopsis. Plant Cell 2014, 26, 538–551. [Google Scholar] [CrossRef] [PubMed]
- Xue, L.; Wang, P.; Wang, L.; Renzi, E.; Radivojac, P.; Tang, H.; Arnold, R.; Zhu, J.-K.; Tao, W.A. Quantitative Measurement of Phosphoproteome Response to Osmotic Stress in Arabidopsis Based on Library-Assisted eXtracted Ion Chromatogram (LAXIC). Mol. Cell. Proteom. 2013, 12, 2354–2369. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Gannon, L.; Hassall, K.L.; Deery, M.J.; Gibbs, D.J.; Holdsworth, M.J.; Van Der Hoorn, R.A.L.; Lilley, K.S.; Theodoulou, F.L. N-terminomics reveals control of Arabidopsis seed storage proteins and proteases by the Arg/N-end rule pathway. New Phytol. 2017, 218, 1106–1126. [Google Scholar] [CrossRef] [PubMed]
- Côté, J.; Quinn, J.; Workman, J.L.; Peterson, C.L. Stimulation of GAL4 derivative binding to nucleosomal DNA by the yeast SWI/SNF complex. Science (80-) 1994, 265, 53–60. [Google Scholar] [CrossRef]
- Batsché, E.; Yaniv, M.; Muchardt, C. The human SWI/SNF subunit Brm is a regulator of alternative splicing. Nat. Struct. Mol. Biol. 2006, 13, 22–29. [Google Scholar] [CrossRef]
- Zhu, Y.; Rowley, M.J.; Böhmdorfer, G.; Wierzbicki, A.T. A SWI/SNF Chromatin-Remodeling Complex Acts in Noncoding RNA-Mediated Transcriptional Silencing. Mol. Cell 2013, 49, 298–309. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Thouly, C.; Le Masson, M.; Lai, X.; Carles, C.C.; Vachon, G. Unwinding BRAHMA Functions in Plants. Genes 2020, 11, 90. https://doi.org/10.3390/genes11010090
Thouly C, Le Masson M, Lai X, Carles CC, Vachon G. Unwinding BRAHMA Functions in Plants. Genes. 2020; 11(1):90. https://doi.org/10.3390/genes11010090
Chicago/Turabian StyleThouly, Caroline, Marie Le Masson, Xuelei Lai, Cristel C. Carles, and Gilles Vachon. 2020. "Unwinding BRAHMA Functions in Plants" Genes 11, no. 1: 90. https://doi.org/10.3390/genes11010090
APA StyleThouly, C., Le Masson, M., Lai, X., Carles, C. C., & Vachon, G. (2020). Unwinding BRAHMA Functions in Plants. Genes, 11(1), 90. https://doi.org/10.3390/genes11010090





