Genome-Wide Identification and Analysis of Class III Peroxidases in Allotetraploid Cotton (Gossypium hirsutum L.) and their Responses to PK Deficiency
Abstract
1. Introduction
2. Materials and Methods
2.1. Sequence Retrieval for POD Proteins in Cotton
2.2. Phylogenetic Analysis
2.3. Gene Structure Analysis
2.4. Analysis of Chromosomal Location and Gene Duplication
2.5. Cotton Culture and Expression Analysis of POD Genes under PK Deficiency
2.6. Localization of POD Proteins
3. Results
3.1. Identification of POD Genes
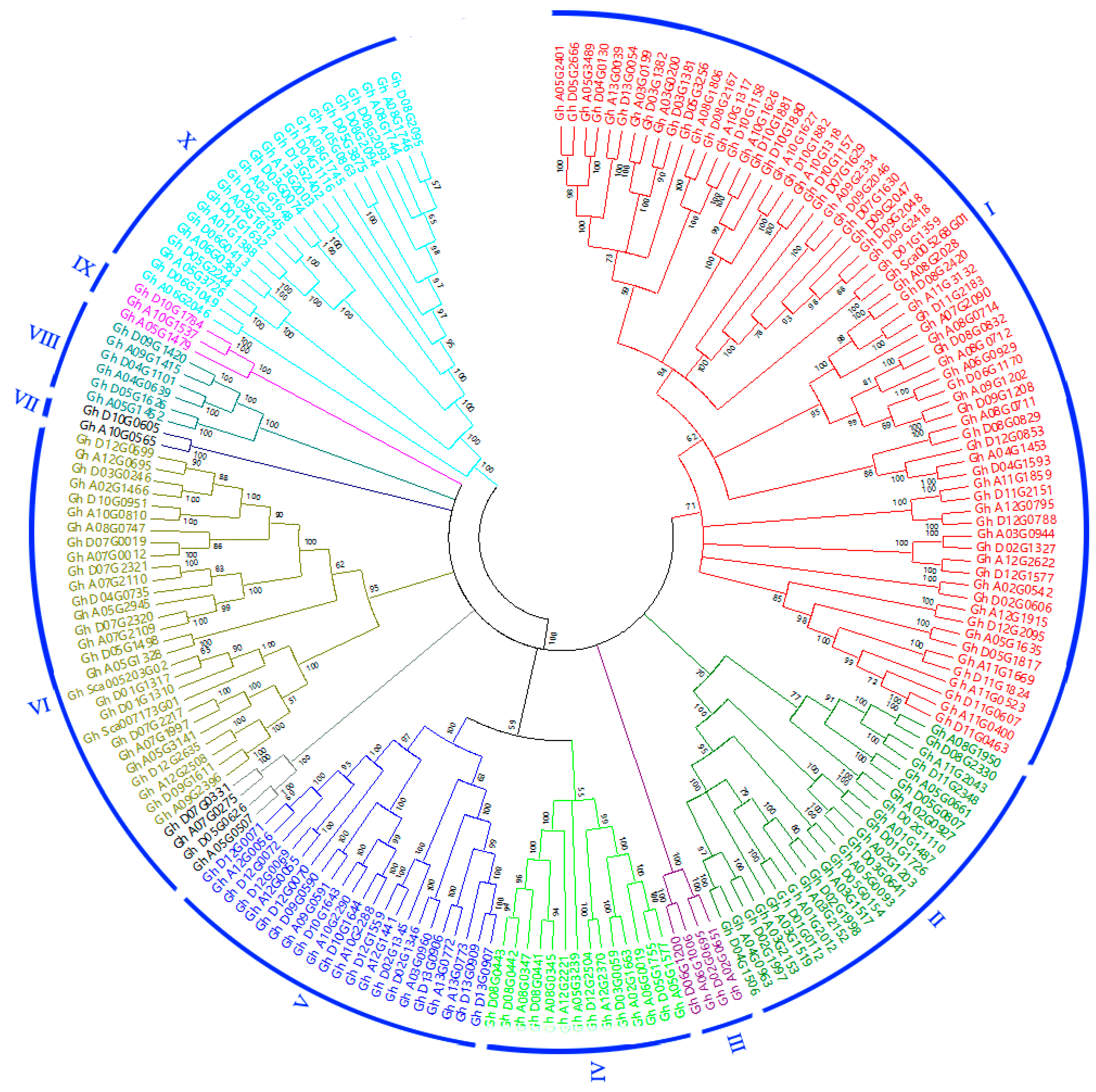
3.2. Phylogenetic Analysis
3.3. Gene Structures
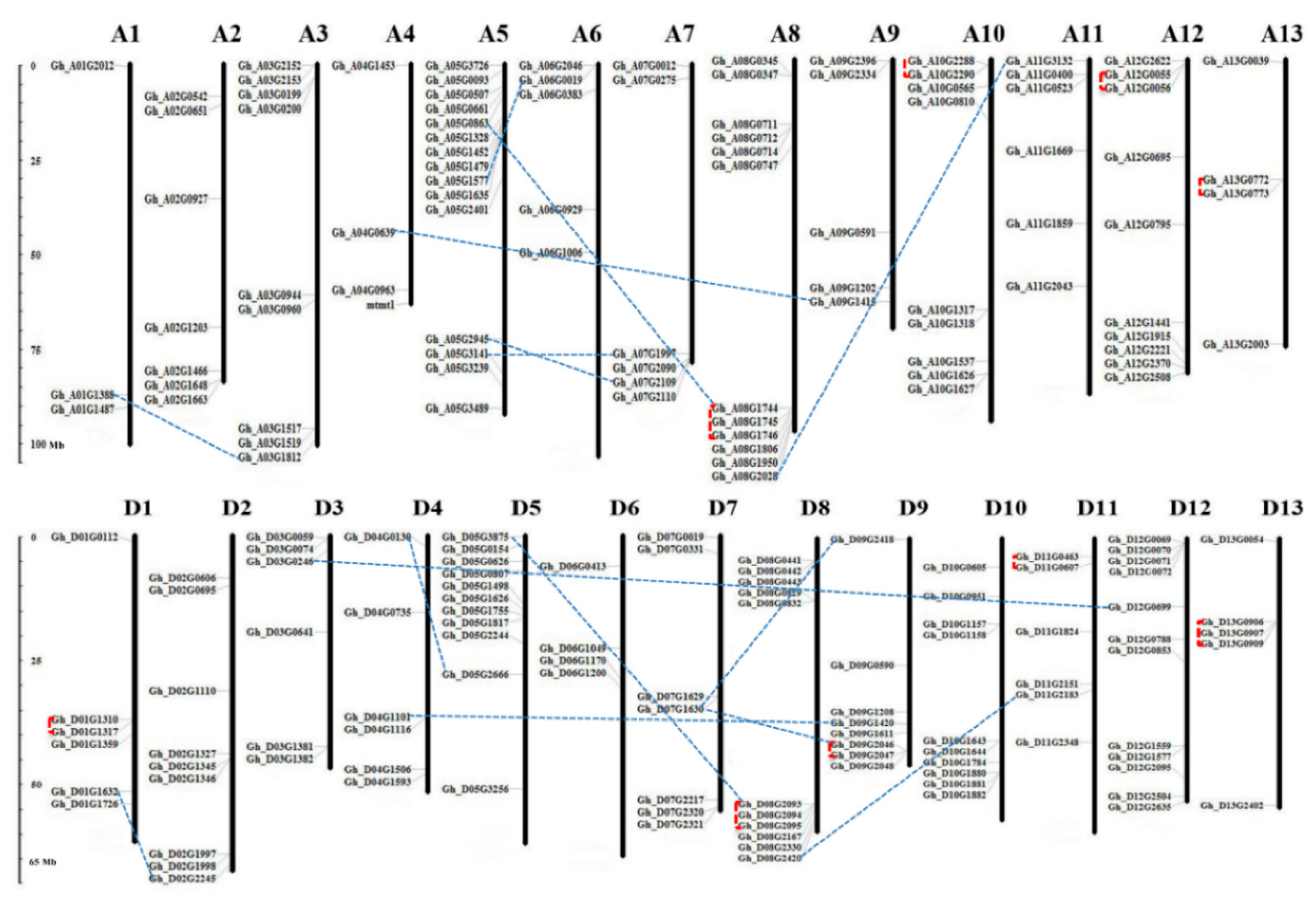
3.4. Chromosomal Location and Gene Duplication
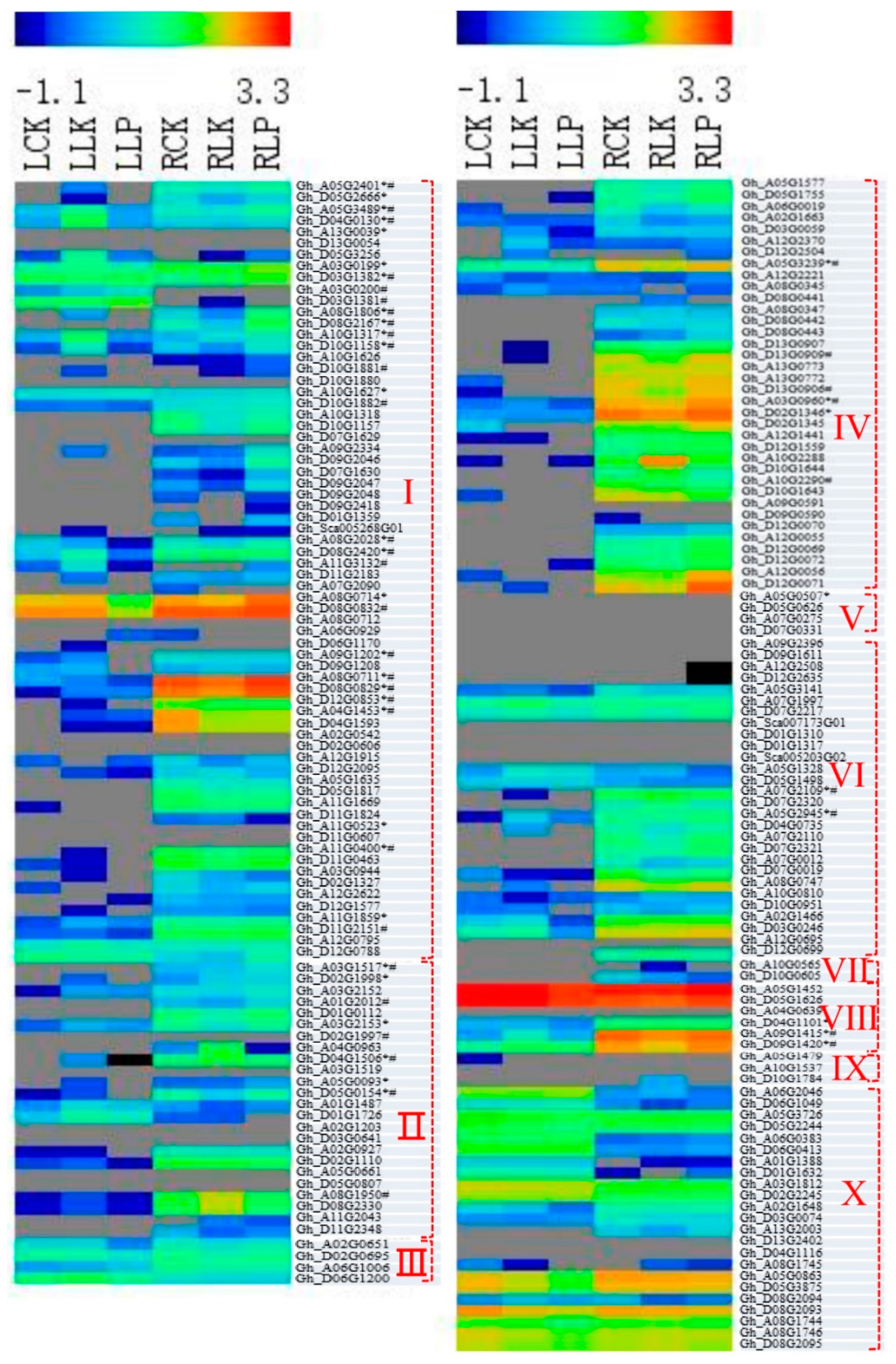
3.5. Responses of POD Genes to PK Deficiency in Cotton Roots and Leaves
3.6. Secret Traits of Cotton POD
4. Discussion
4.1. Identification of Cotton POD Genes and Their Expansion
4.2. Expression Profiles of GhPOD Genes
Author Contributions
Funding
Conflicts of Interest
References
- Passardi, F.; Theiler, G.; Zamocky, M.; Cosio, C.; Rouhier, N.; Teixera, F.; Margis-Pinheiro, M.; Ioannidis, V.; Penel, C.; Falquet, L.; et al. PeroxiBase: The peroxidase database. Phytochemistry 2007, 68, 1605–1611. [Google Scholar] [CrossRef] [PubMed]
- Moural, T.W.; Lewis, K.M.; Barnaba, C.; Zhu, F.; Palmer, N.A.; Sarath, G.; Scully, E.D.; Jones, J.P.; Sattler, S.E.; Kang, C. Characterization of class III peroxidases from switchgrass. Plant Physiol. 2017, 173, 417–433. [Google Scholar] [CrossRef] [PubMed]
- Ren, L.L.; Liu, Y.J.; Liu, H.J.; Qian, T.T.; Qi, L.W.; Wang, X.R.; Zeng, Q.Y. Subcellular relocalization and positive selection play key roles in the retention of duplicate genes of Populus class III peroxidase family. Plant Cell 2014, 26, 2404–2419. [Google Scholar] [CrossRef] [PubMed]
- Passardi, F.; Longet, D.; Penel, C.; Dunand, C. The class III peroxidase multigenic family in rice and its evolution in land plants. Phytochemistry 2004, 65, 1879–1893. [Google Scholar] [CrossRef] [PubMed]
- Tognolli, M.; Penel, C.; Greppin, H.; Simon, P. Analysis and expression of the class III peroxidase large gene family in Arabidopsis thaliana. Gene 2002, 288, 129–138. [Google Scholar] [CrossRef]
- Mei, W.; Qin, Y.; Song, W.; Li, J.; Zhu, Y. Cotton GhPOX1 encoding plant class III peroxidase may be responsible for the high level of reactive oxygen species production that is related to cotton fiber elongation. J. Genet. Genom. 2009, 36, 141–150. [Google Scholar] [CrossRef]
- Martinez-Rubio, R.; Acebes, J.L.; Encina, A.; Karkonen, A. Class III peroxidases in cellulose deficient cultured maize cells during cell wall remodeling. Physiol. Plant. 2018, 164, 45–55. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Xin, W.; Wang, S.; Zhang, X.; Dai, H.; Sun, R.; Frazier, T.; Zhang, B.; Wang, Q. Xylem sap in cotton contains proteins that contribute to environmental stress response and cell wall development. Funct. Integr. Genom. 2015, 15, 17–26. [Google Scholar] [CrossRef] [PubMed]
- Delannoy, E.; Jalloul l, A.; Assigbetse, K.; Marmey, P.; Geiger, J.P.; Lherminier, J.; Daniel, J.F.; Martinez, C.; Nicole, M. Activity of class III peroxidases in the defense of cotton to bacterial blight. Mol. Plant Microbe Interact. 2003, 16, 1030–1038. [Google Scholar] [CrossRef]
- Racz, A.; Hideg, E.; Czegeny, G. Selective responses of class III plant peroxidase isoforms to environmentally relevant UV-B doses. J. Plant Physiol. 2018, 221, 101–106. [Google Scholar] [CrossRef]
- Passardi, F.; Cosio, C.; Penel, C.; Dunand, C. Peroxidases have more functions than a Swiss army knife. Plant Cell Rep. 2005, 24, 255–265. [Google Scholar] [CrossRef] [PubMed]
- Almagro, L.; Gomez Ros, L.V.; Belchi-Navarro, S.; Bru, R.; Ros Barcelo, A.; Pedreno, M.A. Class III peroxidases in plant defence reactions. J. Exp. Bot. 2009, 60, 377–390. [Google Scholar] [CrossRef] [PubMed]
- Marjamaa, K.; Hilden, K.; Kukkola, E.; Lehtonen, M.; Holkeri, H.; Haapaniemi, P.; Koutaniemi, S.; Teeri, T.H.; Fagerstedt, K.; Lundell, T. Cloning, characterization and localization of three novel class III peroxidases in lignifying xylem of Norway spruce (Picea abies). Plant Mol. Biol. 2006, 61, 719–732. [Google Scholar] [CrossRef] [PubMed]
- Francoz, E.; Ranocha, P.; Nguyen-Kim, H.; Jamet, E.; Burlat, V.; Dunand, C. Roles of cell wall peroxidases in plant development. Phytochemistry 2015, 112, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Chen, S.; Alvarez, S.; Asirvatham, V.S.; Schachtman, D.P.; Wu, Y.; Sharp, R.E. Cell wall proteome in the maize primary root elongation zone. I. Extraction and identification of water-soluble and lightly ionically bound proteins. Plant Physiol. 2006, 140, 311–325. [Google Scholar] [CrossRef]
- Girma, K.; Teal, R.K.; Freeman, K.W.; Boman, R.K.; Raun, W.R. Cotton lint yield and quality as affected by applications of N, P, and K fertilizers. J. Cotton Sci. 2007, 11, 12–19. [Google Scholar]
- Dong, H.; Kong, X.; Li, W.; Tang, W.; Zhang, D. Effects of plant density and nitrogen and potassium fertilization on cotton yield and uptake of major nutrients in two fields with varying fertility. Field Crops Res. 2010, 119, 106–113. [Google Scholar] [CrossRef]
- Chen, D.; Ding, Y.; Guo, W.; Zhang, T. Molecular cloning and characterization of a flower-specific class III peroxidase gene in G. hirsutum. Mol. Biol. Rep. 2009, 36, 461–469. [Google Scholar] [CrossRef]
- Delannoy, E.; Marmey, P.; Jalloul, A.; Etienne, H.; Nicole, M. Molecular analysis of class III peroxidases from cotton. J. Cotton Sci. 2006, 10, 53–60. [Google Scholar]
- Zhang, T.; Hu, Y.; Jiang, W.; Fang, L.; Guan, X.; Chen, J.; Zhang, J.; Saski, C.A.; Scheffler, B.E.; Stelly, D.M.; et al. Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat. Biotechnol. 2015, 33, 531–537. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Han, Y.; Meng, D.; Li, D.; Jin, Q.; Lin, Y.; Cai, Y. Structural, evolutionary, and functional analysis of the class III peroxidase gene family in Chinese Pear (Pyrus bretschneideri). Front. Plant Sci. 2016, 7, 1874. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, Q.; Zhao, Y.; Han, G.; Zhu, S. Systematic analysis of maize class III peroxidase gene family reveals a conserved subfamily involved in abiotic stress response. Gene 2015, 566, 95–108. [Google Scholar] [CrossRef]
- Moore, R.C.; Purugganan, M.D. The early stages of duplicate gene evolution. Proc. Natl. Acad. Sci. USA 2003, 100, 15682–15687. [Google Scholar] [CrossRef] [PubMed]
- Matsui, T.; Tabayashi, A.; Iwano, M.; Shinmyo, A.; Kato, K.; Nakayama, H. Activity of the C-terminal-dependent vacuolar sorting signal of horseradish peroxidase C1a is enhanced by its secondary structure. Plant Cell Physiol. 2011, 52, 413–420. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Gong, Q.; Wang, L.; Jin, Y.; Xi, J.; Li, Z.; Qin, W.; Yang, Z.; Lu, L.; Chen, Q.; et al. Genome-wide study of YABBY genes in upland cotton and their expression patterns under different stresses. Front. Genet. 2018, 9, 33. [Google Scholar] [CrossRef]
- Ma, J.; Wang, Q.; Sun, R.; Xie, F.; Jones, D.C.; Zhang, B. Genome-wide identification and expression analysis of TCP transcription factors in Gossypium raimondii. Sci. Rep. 2014, 4, 6645. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.; Zhao, T.; Li, J.; Liu, B.; Fang, L.; Hu, Y.; Zhang, T. Identification and characterization of the GhHsp20 gene family in Gossypium hirsutum. Sci. Rep. 2016, 6, 32517. [Google Scholar] [CrossRef]
- Li, F.; Fan, K.; Ma, F.; Yue, E.; Bibi, N.; Wang, M.; Shen, H.; Hasan, M.M.; Wang, X. Genomic identification and comparative expansion analysis of the non-specific lipid transfer protein gene family in Gossypium. Sci. Rep. 2016, 6, 38948. [Google Scholar] [CrossRef]
- Rogozin, I.B.; Wolf, Y.I.; Sorokin, A.V.; Mirkin, B.G.; Koonin, E.V. Remarkable interkingdom conservation of intron positions and massive, lineage-specific intron loss and gain in eukaryotic evolution. Curr. Biol. 2003, 13, 1512–1517. [Google Scholar] [CrossRef]
- Robertson, H.M. The large srh family of chemoreceptor genes in Caenorhabditis nematodes reveals processes of genome evolution involving large duplications and deletions and intron gains and losses. Genome Res. 2000, 10, 192–203. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Trotman, C.N. Introns-early: Slipping lately? Trends Genet. 1998, 14, 132–134. [Google Scholar] [CrossRef]
- Valerio, L.; De Meyer, M.; Penel, C.; Dunand, C. Expression analysis of the Arabidopsis peroxidase multigenic family. Phytochemistry 2004, 65, 1331–1342. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Chao, M.; Wang, S.; Bu, J.; Tang, J.; Li, F.; Wang, Q.; Zhang, B. Proteome quantification of cotton xylem sap suggests the mechanisms of potassium-deficiency-induced changes in plant resistance to environmental stresses. Sci. Rep. 2016, 6, 21060. [Google Scholar] [CrossRef] [PubMed]
- Shigeto, J.; Tsutsumi, Y. Diverse functions and reactions of class III peroxidases. New Phytol. 2016, 209, 1395–1402. [Google Scholar] [CrossRef]


| Protein Name/ID | Chr Location | Gene/CDS Size (bp) | PL (aa)/MW (Kda)/PI | SignalP/SeretomeP/TargetP |
|---|---|---|---|---|
| GhPOD01/Gh_A01G1388 | chrA01:87059677-87066826 | 7150/867 | 288/31.86/5.64 | N/S/- |
| GhPOD02/Gh_A01G1487 | chrA01:90340312-90341927 | 1616/972 | 323/34.9/5.69 | S/N/S |
| GhPOD03/Gh_A01G2012 | chrA01:175229-177095 | 1867/975 | 324/35.23/9.57 | S/N/- |
| GhPOD04/Gh_A02G0542 | chrA02:8131024-8132267 | 1244/1002 | 333/36.13/4.12 | S/N/S |
| GhPOD05/Gh_A02G0651 | chrA02:10404884-10407370 | 2487/1029 | 342/37.15/5.16 | S/N/S |
| GhPOD06/Gh_A02G0927 | chrA02:35288344-35290345 | 2002/1020 | 339/37.96/8.3 | S/N/S |
| GhPOD07/Gh_A02G1203 | chrA02:69325920-69327130 | 1211/984 | 327/35.33/8.2 | N/N/S |
| GhPOD08/Gh_A02G1466 | chrA02:80770130-80772880 | 2751/993 | 330/36.02/9.44 | S/N/S |
| GhPOD09/Gh_A02G1648 | chrA02:82884707-82887518 | 2812/909 | 302/33.98/9.11 | N/S/- |
| GhPOD10/Gh_A02G1663 | chrA02:82988074-82989301 | 1228/954 | 317/34.25/9.27 | S/N/S |
| GhPOD11/Gh_A03G0199 | chrA03:3093785-3095982 | 2198/969 | 322/34.57/9.93 | S/N/S |
| GhPOD12/Gh_A03G0200 | chrA03:3102427-3103937 | 1511/744 | 247/26.58/5.73 | N/S/C |
| GhPOD13/Gh_A03G0944 | chrA03:60522384-60523564 | 1181/1008 | 335/36.76/5.19 | S/N/S |
| GhPOD14/Gh_A03G0960 | chrA03:61923097-61977652 | 54556/1971 | 656/71.66/8.68 | S/N/S |
| GhPOD15/Gh_A03G1517 | chrA03:95777005-95778170 | 1166/984 | 327/35.68/9.58 | S/N/S |
| GhPOD16/Gh_A03G1519 | chrA03:95814422-95816062 | 1641/723 | 240/25.77/6.99 | S/N/S |
| GhPOD17/Gh_A03G1812 | chrA03:99201877-99205979 | 4103/867 | 288/31.94/7.2 | N/N/- |
| GhPOD18/Gh_A03G2152 | chrA03:16442-17606 | 1165/984 | 327/35.9/8.94 | S/N/S |
| GhPOD19/Gh_A03G2153 | chrA03:28781-29958 | 1178/987 | 328/35.15/7.68 | S/N/S |
| GhPOD20/Gh_A04G0639 | chrA04:44156365-44158523 | 2159/990 | 329/37.28/7.27 | S/N/S |
| GhPOD21/Gh_A04G0963 | chrA04:59248781-59249939 | 1159/996 | 331/36/8.26 | S/N/S |
| GhPOD22/Gh_A04G1453 | chrA04:7944-10075 | 2132/1020 | 339/37.62/7.73 | N/S/S |
| GhPOD23/Gh_A05G0093 | chrA05:1157788-1159020 | 1233/963 | 320/34.93/7.34 | S/N/S |
| GhPOD24/Gh_A05G0507 | chrA05:5454946-5456283 | 1338/999 | 332/36.14/5.05 | N/N/S |
| GhPOD25/Gh_A05G0661 | chrA05:6938388-6950337 | 11950/1014 | 337/36.17/6.73 | S/N/S |
| GhPOD26/Gh_A05G0863 | chrA05:8607073-8608956 | 1884/753 | 250/27.59/6.35 | N/S/- |
| GhPOD27/Gh_A05G1328 | chrA05:13602610-13604358 | 1749/1053 | 350/38.23/9.51 | S/N/S |
| GhPOD28/Gh_A05G1452 | chrA05:15020810-15022360 | 1551/999 | 332/37.66/8.54 | S/N/S |
| GhPOD29/Gh_A05G1479 | chrA05:15196748-15197863 | 1116/1011 | 336/36.82/8.25 | N/S/- |
| GhPOD30/Gh_A05G1577 | chrA05:16171654-16172991 | 1338/951 | 316/34.19/9.5 | S/N/S |
| GhPOD31/Gh_A05G1635 | chrA05:16892245-16893617 | 1373/1050 | 349/37.73/7.7 | S/N/S |
| GhPOD32/Gh_A05G2401 | chrA05:29807925-29809249 | 1325/972 | 323/34.96/9.47 | S/N/S |
| GhPOD33/Gh_A05G2945 | chrA05:72277688-72279245 | 1558/972 | 323/35.2/4.67 | S/N/S |
| GhPOD34/Gh_A05G3141 | chrA05:81059455-81060420 | 966/966 | 321/35.48/8.86 | S/N/S |
| GhPOD35/Gh_A05G3239 | chrA05:84740299-84741875 | 1577/972 | 323/35.83/5.06 | N/S/S |
| GhPOD36/Gh_A05G3489 | chrA05:90428029-90429659 | 1631/1014 | 337/36.06/9.57 | S/N/S |
| GhPOD37/Gh_A05G3726 | chrA05:122462-125928 | 3467/1182 | 393/42.73/9.35 | N/S/C |
| GhPOD38/Gh_A06G0019 | chrA06:84546-85807 | 1262/954 | 317/34.23/9.28 | S/N/S |
| GhPOD39/Gh_A06G0383 | chrA06:6376778-6380906 | 4129/1167 | 388/42.25/7.68 | N/S/C |
| GhPOD40/Gh_A06G0929 | chrA06:38044180-38045572 | 1393/1038 | 345/37.15/4.4 | S/N/S |
| GhPOD41/Gh_A06G1006 | chrA06:49554553-49556908 | 2356/987 | 328/35.73/8.27 | S/N/S |
| GhPOD42/Gh_A06G2046 | chrA06:45925-48981 | 3057/960 | 319/34.42/8.88 | N/S/C |
| GhPOD43/Gh_A07G0012 | chrA07:136360-137613 | 1254/996 | 331/36.45/9.58 | S/N/S |
| GhPOD44/Gh_A07G0275 | chrA07:3399593-3400898 | 1306/996 | 331/35.82/4.82 | S/N/- |
| GhPOD45/Gh_A07G1997 | chrA07:76170501-76172204 | 1704/1026 | 341/37.65/8.37 | S/N/S |
| GhPOD46/Gh_A07G2090 | chrA07:77469444-77471552 | 2109/1101 | 366/40.24/6.74 | N/S/- |
| GhPOD47/Gh_A07G2109 | chrA07:77657753-77658858 | 1106/966 | 321/34.93/4.26 | S/N/S |
| GhPOD48/Gh_A07G2110 | chrA07:77659654-77660804 | 1151/975 | 324/35.43/7.68 | S/N/S |
| GhPOD49/Gh_A08G0345 | chrA08:4189416-4190503 | 1088/1002 | 333/35.9/7.9 | S/N/S |
| GhPOD50/Gh_A08G0347 | chrA08:4289654-4290385 | 732/732 | 243/25.94/7.89 | N/S/- |
| GhPOD51/Gh_A08G0711 | chrA08:17779406-17781156 | 1751/999 | 332/35.79/5.71 | N/S/S |
| GhPOD52/Gh_A08G0712 | chrA08:18361761-18367521 | 5761/1056 | 351/38.2/4.88 | S/N/S |
| GhPOD53/Gh_A08G0714 | chrA08:18413356-18418531 | 5176/1032 | 343/36.85/4.51 | S/N/S |
| GhPOD54/Gh_A08G0747 | chrA08:24039157-24042529 | 3373/990 | 329/36.31/10.43 | S/N/S |
| GhPOD55/Gh_A08G1744 | chrA08:97409160-97411219 | 2060/753 | 250/27.54/5.6 | N/N/- |
| GhPOD56/Gh_A08G1745 | chrA08:97440700-97442620 | 1921/726 | 241/26.72/7.74 | N/N/- |
| GhPOD57/Gh_A08G1746 | chrA08:97442829-97446411 | 3583/867 | 288/32.07/6.91 | N/N/- |
| GhPOD58/Gh_A08G1806 | chrA08:98615798-98617217 | 1420/960 | 319/34.42/5.68 | S/N/S |
| GhPOD59/Gh_A08G1950 | chrA08:100557586-100558616 | 1031/951 | 316/34.48/9.77 | S/N/S |
| GhPOD60/Gh_A08G2028 | chrA08:101461027-101462441 | 1415/969 | 322/34.96/7.75 | S/N/S |
| GhPOD61/Gh_A09G0591 | chrA09:48054359-48058181 | 3823/777 | 258/28.28/8.82 | N/S/- |
| GhPOD62/Gh_A09G1202 | chrA09:63774095-63775448 | 1354/1020 | 339/36.01/5.24 | S/N/S |
| GhPOD63/Gh_A09G1415 | chrA09:67412511-67414343 | 1833/990 | 329/37.12/6.58 | S/N/S |
| GhPOD64/Gh_A09G2334 | chrA09:93316-94482 | 1167/951 | 316/34.58/10.02 | S/N/S |
| GhPOD65/Gh_A09G2396 | chrA09:35011-35983 | 973/906 | 301/33.44/8.46 | S/N/S |
| GhPOD66/Gh_A10G0565 | chrA10:7031511-7032708 | 1198/1041 | 346/38.29/6.03 | S/N/S |
| GhPOD67/Gh_A10G0810 | chrA10:16610264-16611518 | 1255/993 | 330/35.91/9.88 | S/N/M |
| GhPOD68/Gh_A10G1317 | chrA10:69770818-69772754 | 1937/966 | 321/33.76/8.33 | S/N/S |
| GhPOD69/Gh_A10G1318 | chrA10:69892772-69893920 | 1149/954 | 317/34.64/9.07 | S/N/S |
| GhPOD70/Gh_A10G1537 | chrA10:84415649-84417960 | 2312/972 | 323/35.59/8.64 | S/N/S |
| GhPOD71/Gh_A10G1626 | chrA10:87538433-87544412 | 5980/909 | 302/33.3/10.36 | N/S/- |
| GhPOD72/Gh_A10G1627 | chrA10:87544899-87546095 | 1197/987 | 328/35.3/9.64 | S/N/S |
| GhPOD73/Gh_A10G2288 | chrA10:1167442-1169647 | 2206/984 | 327/35.78/10.03 | S/N/S |
| GhPOD74/Gh_A10G2290 | chrA10:1189614-1190983 | 1370/993 | 330/36.03/8.94 | S/N/S |
| GhPOD75/Gh_A11G0400 | chrA11:3718938-3720282 | 1345/996 | 331/36.51/9.37 | S/N/S |
| GhPOD76/Gh_A11G0523 | chrA11:4917637-4918870 | 1234/927 | 308/34.89/9.73 | N/S/S |
| GhPOD77/Gh_A11G1669 | chrA11:25291980-25293232 | 1253/1005 | 334/36.93/8.46 | S/N/S |
| GhPOD78/Gh_A11G1859 | chrA11:45617267-45622682 | 5416/930 | 309/34.02/4.72 | N/N/- |
| GhPOD79/Gh_A11G2043 | chrA11:63260806-63261288 | 483/483 | 160/17.1/4.89 | N/S/- |
| GhPOD80/Gh_A11G3132 | chrA11:6967-8208 | 1242/966 | 321/34.97/8 | S/N/S |
| GhPOD81/Gh_A12G0055 | chrA12:735887-756659 | 20773/972 | 323/35.31/8.26 | S/N/S |
| GhPOD82/Gh_A12G0056 | chrA12:760206-761426 | 1221/972 | 323/35.56/8.99 | S/N/S |
| GhPOD83/Gh_A12G0695 | chrA12:26988350-26989090 | 741/561 | 186/20.48/6.66 | N/S/- |
| GhPOD84/Gh_A12G0795 | chrA12:45772636-45776292 | 3657/1068 | 355/39.81/5.12 | S/N/S |
| GhPOD85/Gh_A12G1441 | chrA12:73250331-73252001 | 1671/1023 | 340/37.17/9.36 | N/N/M |
| GhPOD86/Gh_A12G1915 | chrA12:81887117-81888429 | 1313/1014 | 337/37.91/6 | S/N/S |
| GhPOD87/Gh_A12G2221 | chrA12:84939580-84946625 | 7046/3291 | 1096/122.66/7.51 | N/N/- |
| GhPOD88/Gh_A12G2370 | chrA12:86207288-86208988 | 1701/972 | 323/35.63/8.88 | S/N/S |
| GhPOD89/Gh_A12G2508 | chrA12:87451770-87452774 | 1005/1005 | 334/37.17/7.73 | S/N/S |
| GhPOD90/Gh_A12G2622 | chrA12:49766-51061 | 1296/1011 | 336/37.08/5.05 | S/N/S |
| GhPOD91/Gh_A13G0039 | chrA13:393916-396005 | 2090/978 | 325/35.42/9.01 | S/N/S |
| GhPOD92/Gh_A13G0772 | chrA13:33291935-33293456 | 1522/975 | 324/34.9/6.65 | S/N/S |
| GhPOD93/Gh_A13G0773 | chrA13:33310901-33312163 | 1263/987 | 328/35.42/7.36 | S/N/S |
| GhPOD94/Gh_A13G2003 | chrA13:79484281-79487307 | 3027/762 | 253/28/5.19 | N/N/- |
| GhPOD95/Gh_D01G0112 | chrD01:855518-857368 | 1851/975 | 324/35.25/9.56 | S/N/- |
| GhPOD96/Gh_D01G1310 | chrD01:36863811-36864806 | 996/996 | 331/36.9/8.19 | S/N/S |
| GhPOD97/Gh_D01G1317 | chrD01:37193921-37194913 | 993/993 | 330/36.56/7.96 | S/N/S |
| GhPOD98/Gh_D01G1359 | chrD01:40012715-40014005 | 1291/969 | 322/34.7/7.93 | S/N/S |
| GhPOD99/Gh_D01G1632 | chrD01:51431707-51437024 | 5318/867 | 288/31.78/5.81 | N/S/- |
| GhPOD100/Gh_D01G1726 | chrD01:53949584-53951128 | 1545/972 | 323/34.9/5.4 | S/N/S |
| GhPOD101/Gh_D02G0606 | chrD02:8255229-8256474 | 1246/1014 | 337/36.66/4.13 | S/N/S |
| GhPOD102/Gh_D02G0695 | chrD02:9863175-9865761 | 2587/1116 | 371/40.62/7.7 | N/S/M |
| GhPOD103/Gh_D02G1110 | chrD02:31109209-31111160 | 1952/1020 | 339/37.91/8.45 | S/N/S |
| GhPOD104/Gh_D02G1327 | chrD02:43855329-43856508 | 1180/1008 | 335/36.76/5.38 | S/N/S |
| GhPOD105/Gh_D02G1345 | chrD02:44875923-44877373 | 1451/990 | 329/36.19/9.18 | S/N/S |
| GhPOD106/Gh_D02G1346 | chrD02:44991381-44992620 | 1240/984 | 327/35.44/7.72 | S/N/S |
| GhPOD107/Gh_D02G1997 | chrD02:64059836-64065386 | 5551/987 | 328/35.24/7.94 | S/N/S |
| GhPOD108/Gh_D02G1998 | chrD02:64081876-64083040 | 1165/984 | 327/35.7/9.37 | S/N/S |
| GhPOD109/Gh_D02G2245 | chrD02:66344682-66348942 | 4261/867 | 288/31.96/7.2 | N/N/- |
| GhPOD110/Gh_D03G0059 | chrD03:390493-391719 | 1227/954 | 317/34.27/9.27 | S/N/S |
| GhPOD111/Gh_D03G0074 | chrD03:531888-534705 | 2818/912 | 303/33.96/8.22 | N/S/- |
| GhPOD112/Gh_D03G0246 | chrD03:2618435-2620271 | 1837/993 | 330/35.99/9.44 | S/N/S |
| GhPOD113/Gh_D03G0641 | chrD03:19207034-19208004 | 971/699 | 232/25.15/8.87 | N/N/S |
| GhPOD114/Gh_D03G1381 | chrD03:42417094-42420536 | 3443/1158 | 385/42.12/7.65 | N/S/- |
| GhPOD115/Gh_D03G1382 | chrD03:42426148-42428382 | 2235/969 | 322/34.55/9.72 | S/N/S |
| GhPOD116/Gh_D04G0130 | chrD04:1796590-1798244 | 1655/1014 | 337/36.04/9.22 | S/N/S |
| GhPOD117/Gh_D04G0735 | chrD04:15156949-15158096 | 1148/972 | 323/35.41/8.64 | S/N/S |
| GhPOD118/Gh_D04G1101 | chrD04:36441991-36444139 | 2149/987 | 328/37.19/6.73 | S/N/S |
| GhPOD119/Gh_D04G1116 | chrD04:36784560-36786018 | 1459/723 | 240/26.55/6.51 | N/N/- |
| GhPOD120/Gh_D04G1506 | chrD04:46932159-46969978 | 37820/2016 | 671/72.76/7.71 | S/N/S |
| GhPOD121/Gh_D04G1593 | chrD04:48023771-48025856 | 2086/1020 | 339/37.4/7.06 | N/N/S |
| GhPOD122/Gh_D05G0154 | chrD05:1545607-1546840 | 1234/978 | 325/35.03/7.99 | S/N/S |
| GhPOD123/Gh_D05G0626 | chrD05:5026931-5028271 | 1341/1002 | 333/36.15/5.05 | N/N/S |
| GhPOD124/Gh_D05G0807 | chrD05:6771535-6772831 | 1297/1014 | 337/36.1/6.73 | S/N/S |
| GhPOD125/Gh_D05G1498 | chrD05:13463061-13464814 | 1754/1059 | 352/38.37/9.62 | N/S/C |
| GhPOD126/Gh_D05G1626 | chrD05:14640100-14641647 | 1548/999 | 332/37.65/8.54 | S/N/S |
| GhPOD127/Gh_D05G1755 | chrD05:15837493-15838825 | 1333/951 | 316/34.22/9.47 | S/N/S |
| GhPOD128/Gh_D05G1817 | chrD05:16541131-16542504 | 1374/1050 | 349/37.76/7.7 | S/N/S |
| GhPOD129/Gh_D05G2244 | chrD05:21543390-21547747 | 4358/1218 | 405/43.82/9.15 | N/S/C |
| GhPOD130/Gh_D05G2666 | chrD05:27862573-27863898 | 1326/972 | 323/34.85/8.92 | S/N/S |
| GhPOD131/Gh_D05G3256 | chrD05:51004089-51005627 | 1539/951 | 316/33.69/8.5 | S/N/S |
| GhPOD132/Gh_D05G3875 | chrD05:76982-78848 | 1867/753 | 250/27.57/6.04 | N/S/- |
| GhPOD133/Gh_D06G0413 | chrD06:5889650-5894261 | 4612/1353 | 450/49.69/7.3 | N/S/C |
| GhPOD134/Gh_D06G1049 | chrD06:22550424-22553640 | 3217/1008 | 335/36.1/9.95 | N/S/C |
| GhPOD135/Gh_D06G1170 | chrD06:28537753-28539152 | 1400/1038 | 345/37.22/4.46 | S/N/S |
| GhPOD136/Gh_D06G1200 | chrD06:30337737-30340827 | 3091/1050 | 349/37.92/8.39 | S/N/S |
| GhPOD137/Gh_D07G0019 | chrD07:200078-201330 | 1253/996 | 331/36.42/9.11 | S/N/S |
| GhPOD138/Gh_D07G0331 | chrD07:3515650-3516955 | 1306/996 | 331/35.86/4.66 | S/N/S |
| GhPOD139/Gh_D07G1629 | chrD07:32319582-32320538 | 957/792 | 263/28.65/9.4 | N/S/- |
| GhPOD140/Gh_D07G1630 | chrD07:32393320-32394431 | 1112/939 | 312/34.24/9.89 | S/N/S |
| GhPOD141/Gh_D07G2217 | chrD07:53125200-53126909 | 1710/1026 | 341/37.7/8.37 | S/N/S |
| GhPOD142/Gh_D07G2320 | chrD07:54647068-54648167 | 1100/966 | 321/35.08/4.35 | S/N/S |
| GhPOD143/Gh_D07G2321 | chrD07:54648867-54650018 | 1152/975 | 324/35.39/7.24 | S/N/S |
| GhPOD144/Gh_D08G0441 | chrD08:4640562-4641585 | 1024/858 | 285/30.9/8.72 | N/S/C |
| GhPOD145/Gh_D08G0442 | chrD08:4670537-4671627 | 1091/1005 | 334/35.7/8.55 | S/N/S |
| GhPOD146/Gh_D08G0443 | chrD08:4687890-4688979 | 1090/1005 | 334/35.66/8.55 | S/N/S |
| GhPOD147/Gh_D08G0829 | chrD08:13789103-13790820 | 1718/999 | 332/35.8/6.5 | S/N/S |
| GhPOD148/Gh_D08G0832 | chrD08:14037040-14038366 | 1327/1065 | 354/38.22/4.6 | S/N/S |
| GhPOD149/Gh_D08G2093 | chrD08:59783046-59784995 | 1950/741 | 246/27.13/6.04 | N/S/- |
| GhPOD150/Gh_D08G2094 | chrD08:59804611-59806683 | 2073/738 | 245/27.03/5.9 | N/N/- |
| GhPOD151/Gh_D08G2095 | chrD08:59806893-59810058 | 3166/879 | 292/32.56/6.95 | N/N/- |
| GhPOD152/Gh_D08G2167 | chrD08:61090294-61091717 | 1424/960 | 319/34.38/5.68 | S/N/S |
| GhPOD153/Gh_D08G2330 | chrD08:63032857-63033881 | 1025/945 | 314/34.22/9.44 | S/N/S |
| GhPOD154/Gh_D08G2420 | chrD08:64145436-64146846 | 1411/969 | 322/34.92/7.75 | S/N/S |
| GhPOD155/Gh_D09G0590 | chrD09:28473937-28477562 | 3626/990 | 329/36.6/9.74 | S/N/S |
| GhPOD156/Gh_D09G1208 | chrD09:39009612-39010932 | 1321/1020 | 339/35.98/4.95 | S/N/S |
| GhPOD157/Gh_D09G1420 | chrD09:41767263-41769101 | 1839/987 | 328/36.92/6.28 | S/N/S |
| GhPOD158/Gh_D09G1611 | chrD09:43695508-43696494 | 987/987 | 328/36.13/6.76 | S/N/S |
| GhPOD159/Gh_D09G2046 | chrD09:47806422-47807579 | 1158/951 | 316/34.5/9.58 | S/N/S |
| GhPOD160/Gh_D09G2047 | chrD09:47809072-47810196 | 1125/885 | 294/31.91/9.35 | S/N/S |
| GhPOD161/Gh_D09G2048 | chrD09:47810407-47821212 | 10806/2019 | 672/73.64/9.25 | S/N/S |
| GhPOD162/Gh_D09G2418 | chrD09:1692-2815 | 1124/1026 | 341/37.69/9.36 | S/N/S |
| GhPOD163/Gh_D10G0605 | chrD10:6317691-6318891 | 1201/1041 | 346/38.32/6.12 | S/N/S |
| GhPOD164/Gh_D10G0951 | chrD10:12771913-12773186 | 1274/993 | 330/36.01/9.99 | S/N/M |
| GhPOD165/Gh_D10G1157 | chrD10:19241093-19242236 | 1144/954 | 317/34.55/9.06 | S/N/S |
| GhPOD166/Gh_D10G1158 | chrD10:19299752-19300772 | 1021/705 | 234/24.46/8.21 | N/S/- |
| GhPOD167/Gh_D10G1643 | chrD10:45423530-45424891 | 1362/993 | 330/36.1/8.98 | S/N/S |
| GhPOD168/Gh_D10G1644 | chrD10:45445937-45448111 | 2175/984 | 327/35.72/10.04 | S/N/S |
| GhPOD169/Gh_D10G1784 | chrD10:50389073-50391342 | 2270/972 | 323/35.64/8.8 | S/N/S |
| GhPOD170/Gh_D10G1880 | chrD10:52703553-52704762 | 1210/1041 | 346/37.99/8.78 | S/N/S |
| GhPOD171/Gh_D10G1881 | chrD10:52813459-52814911 | 1453/972 | 323/35.59/10.39 | S/N/S |
| GhPOD172/Gh_D10G1882 | chrD10:52815399-52816603 | 1205/987 | 328/35.25/9.64 | S/N/S |
| GhPOD173/Gh_D11G0463 | chrD11:3963558-3964899 | 1342/996 | 331/36.57/9.22 | S/N/S |
| GhPOD174/Gh_D11G0607 | chrD11:5302020-5303236 | 1217/978 | 325/36.13/8.95 | N/S/S |
| GhPOD175/Gh_D11G1824 | chrD11:20830683-20831933 | 1251/969 | 322/35.55/8.46 | S/N/S |
| GhPOD176/Gh_D11G2151 | chrD11:32727567-32730117 | 2551/972 | 323/35.51/4.66 | S/N/S |
| GhPOD177/Gh_D11G2183 | chrD11:34071064-34072304 | 1241/966 | 321/34.9/7.74 | S/N/S |
| GhPOD178/Gh_D11G2348 | chrD11:45744028-45745072 | 1045/957 | 318/35.11/10.5 | S/N/M |
| GhPOD179/Gh_D12G0069 | chrD12:815867-853649 | 37783/1764 | 587/64.17/7.56 | N/S/- |
| GhPOD180/Gh_D12G0070 | chrD12:860573-861841 | 1269/972 | 323/35.07/6.5 | S/N/S |
| GhPOD181/Gh_D12G0071 | chrD12:866590-867867 | 1278/1029 | 342/37.73/8.99 | N/S/S |
| GhPOD182/Gh_D12G0072 | chrD12:876401-877438 | 1038/789 | 262/28.85/8.06 | N/S/- |
| GhPOD183/Gh_D12G0699 | chrD12:15282699-15283971 | 1273/993 | 330/35.78/8.49 | S/N/S |
| GhPOD184/Gh_D12G0788 | chrD12:22644947-22646101 | 1155/1068 | 355/39.65/4.9 | S/N/S |
| GhPOD185/Gh_D12G0853 | chrD12:27599646-27601009 | 1364/1050 | 349/38.71/6.76 | S/N/S |
| GhPOD186/Gh_D12G1559 | chrD12:46570546-46572226 | 1681/1023 | 340/37.12/9.36 | N/S/M |
| GhPOD187/Gh_D12G1577 | chrD12:46856738-46864694 | 7957/1803 | 600/68.11/6.77 | N/S/- |
| GhPOD188/Gh_D12G2095 | chrD12:53969820-53971114 | 1295/1002 | 333/37.49/5.48 | S/N/S |
| GhPOD189/Gh_D12G2504 | chrD12:57927732-57929447 | 1716/972 | 323/35.48/8.72 | S/N/S |
| GhPOD190/Gh_D12G2635 | chrD12:59085174-59086178 | 1005/1005 | 334/37.2/7.99 | S/N/S |
| GhPOD191/Gh_D13G0054 | chrD13:451962-454035 | 2074/978 | 325/35.63/9.66 | S/N/S |
| GhPOD192/Gh_D13G0906 | chrD13:18641774-18643302 | 1529/978 | 325/34.95/6.77 | S/N/S |
| GhPOD193/Gh_D13G0907 | chrD13:18659873-18661135 | 1263/987 | 328/35.47/7.41 | S/N/S |
| GhPOD194/Gh_D13G0909 | chrD13:18677172-18678434 | 1263/987 | 328/35.5/7.41 | S/N/S |
| GhPOD195/Gh_D13G2402 | chrD13:60076810-60079762 | 2953/750 | 249/27.31/5.17 | N/N/- |
| GhPOD196/Gh_Sca005203G02 | chrSca:12082-12672 | 591/591 | 196/22.12/8.79 | N/S/- |
| GhPOD197/Gh_Sca005268G01 | chrSca:3248-4662 | 1415/960 | 319/34.49/7.93 | S/N/S |
| GhPOD198/Gh_Sca007173G01 | chrSca:8026-8850 | 825/825 | 275/30.21/6.74 | S/N/S |
| Paralogous Pairs | Ks | Ka | Ka/Ks | Duplicate Type |
|---|---|---|---|---|
| Gh_A01G1388-Gh_A03G1812 | 0.446 | 0.065 | 0.15 | Segmental |
| Gh_D03G0246-Gh_D12G0699 | 1.037 | 0.099 | 0.10 | Segmental |
| Gh_D04G0130-Gh_D05G2666 | 0.573 | 0.086 | 0.15 | Segmental |
| Gh_D04G1101-Gh_D09G1420 | 0.452 | 0.095 | 0.21 | Segmental |
| Gh_D05G3875-Gh_D08G2093 | 0.373 | 0.03 | 0.08 | Segmental |
| Gh_D07G1630-Gh_D09G2046 | 0.298 | 0.063 | 0.21 | Segmental |
| Gh_D07G1630-Gh_D09G2418 | 0.205 | 0.046 | 0.22 | Segmental |
| Gh_D08G2420-Gh_D11G2183 | 0.479 | 0.066 | 0.14 | Segmental |
| Gh_A04G0639-Gh_A09G1415 | 0.486 | 0.095 | 0.20 | Segmental |
| Gh_A05G0863-Gh_A08G1744 | 0.148 | 0.074 | 0.50 | Segmental |
| Gh_A05G1577-Gh_A06G0019 | 0.508 | 0.07 | 0.14 | Segmental |
| Gh_A05G2945-Gh_A07G2109 | 0.79 | 0.096 | 0.12 | Segmental |
| Gh_A05G3141-Gh_A07G1997 | 0.721 | 0.079 | 0.11 | Segmental |
| Gh_A08G2028-Gh_A11G3132 | 0.483 | 0.073 | 0.15 | Segmental |
| Gh_D01G1632-Gh_D02G2245 | 0.452 | 0.063 | 0.14 | Segmental |
| Gh_D08G2093-Gh_D08G2094 | 0.087 | 0.011 | 0.13 | Tandem |
| Gh_D08G2094-Gh_D08G2095 | 0.087 | 0.017 | 0.20 | Tandem |
| Gh_D09G2046-Gh_D09G2047 | 0.355 | 0.046 | 0.13 | Tandem |
| Gh_D11G0463-Gh_D11G0607 | 0.129 | 0.039 | 0.30 | Tandem |
| Gh_D13G0906-Gh_D13G0907 | 0.205 | 0.07 | 0.34 | Tandem |
| Gh_D13G0907-Gh_D13G0909 | 0.009 | 0.001 | 0.11 | Tandem |
| Gh_A08G1744-Gh_A08G1745 | 0.039 | 0.039 | 1.00 | Tandem |
| Gh_A08G1745-Gh_A08G1746 | 0.033 | 0.033 | 1.00 | Tandem |
| Gh_A10G2288-Gh_A10G2290 | 0.306 | 0.093 | 0.30 | Tandem |
| Gh_A12G0055-Gh_A12G0056 | 0.289 | 0.061 | 0.21 | Tandem |
| Gh_A13G0772-Gh_A13G0773 | 0.205 | 0.076 | 0.37 | Tandem |
| Gh_D01G1310-Gh_D01G1317 | 0.018 | 0.008 | 0.44 | Tandem |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Duan, P.; Wang, G.; Chao, M.; Zhang, Z.; Zhang, B. Genome-Wide Identification and Analysis of Class III Peroxidases in Allotetraploid Cotton (Gossypium hirsutum L.) and their Responses to PK Deficiency. Genes 2019, 10, 473. https://doi.org/10.3390/genes10060473
Duan P, Wang G, Chao M, Zhang Z, Zhang B. Genome-Wide Identification and Analysis of Class III Peroxidases in Allotetraploid Cotton (Gossypium hirsutum L.) and their Responses to PK Deficiency. Genes. 2019; 10(6):473. https://doi.org/10.3390/genes10060473
Chicago/Turabian StyleDuan, Pengfei, Guo Wang, Maoni Chao, Zhiyong Zhang, and Baohong Zhang. 2019. "Genome-Wide Identification and Analysis of Class III Peroxidases in Allotetraploid Cotton (Gossypium hirsutum L.) and their Responses to PK Deficiency" Genes 10, no. 6: 473. https://doi.org/10.3390/genes10060473
APA StyleDuan, P., Wang, G., Chao, M., Zhang, Z., & Zhang, B. (2019). Genome-Wide Identification and Analysis of Class III Peroxidases in Allotetraploid Cotton (Gossypium hirsutum L.) and their Responses to PK Deficiency. Genes, 10(6), 473. https://doi.org/10.3390/genes10060473






