Bacterial Aspartyl-tRNA Synthetase Has Glutamyl-tRNA Synthetase Activity
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Overexpression and Purification of aaRSs
2.3. In Vivo Transcription and Purification of tRNAs
2.4. Initial Rate Aminoacylation Assays
2.5. Extended Aminoacylation Assays (90 Min)
2.6. Acid Gel Electrophoresis and Northern Blot Analysis
3. Results
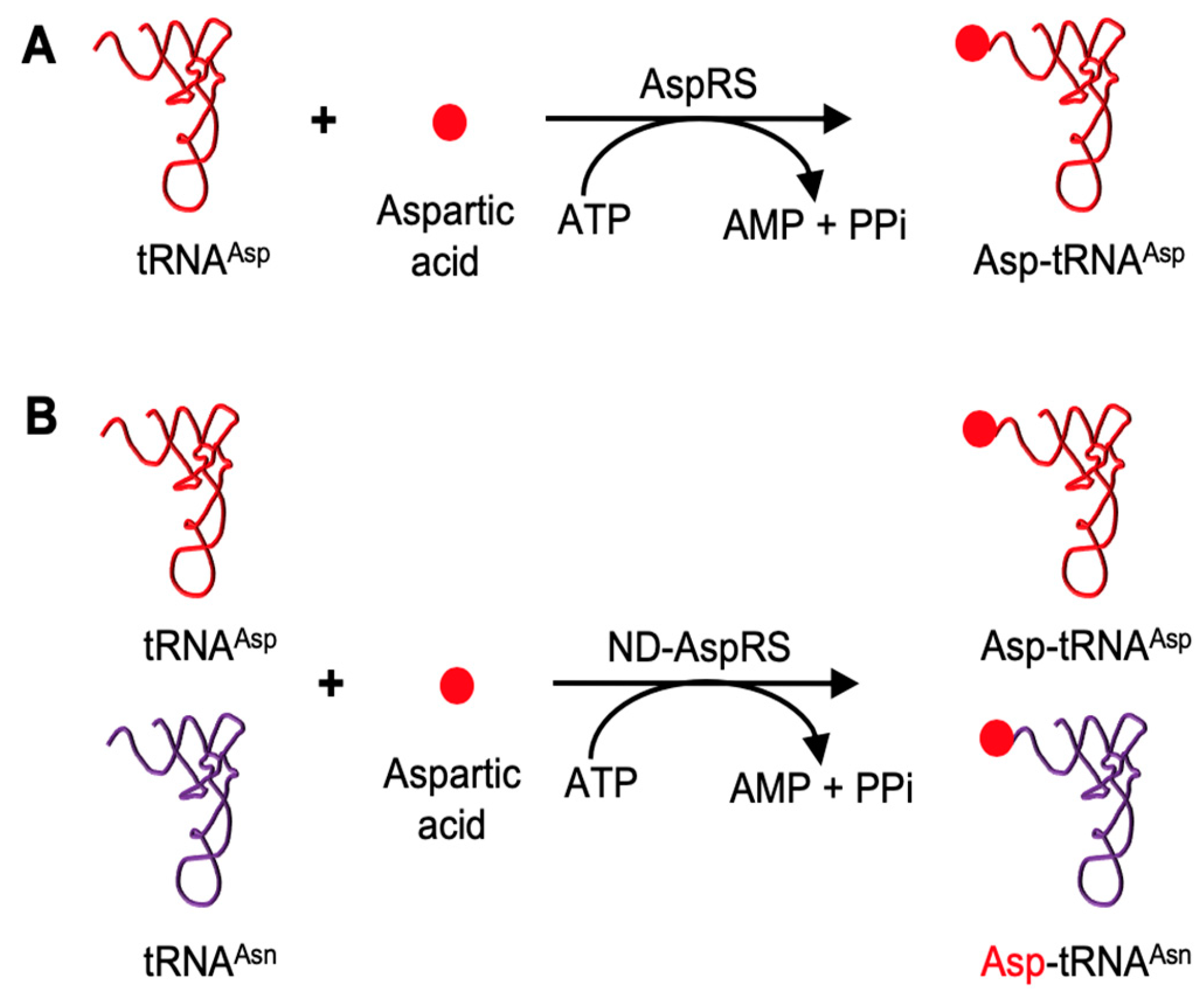
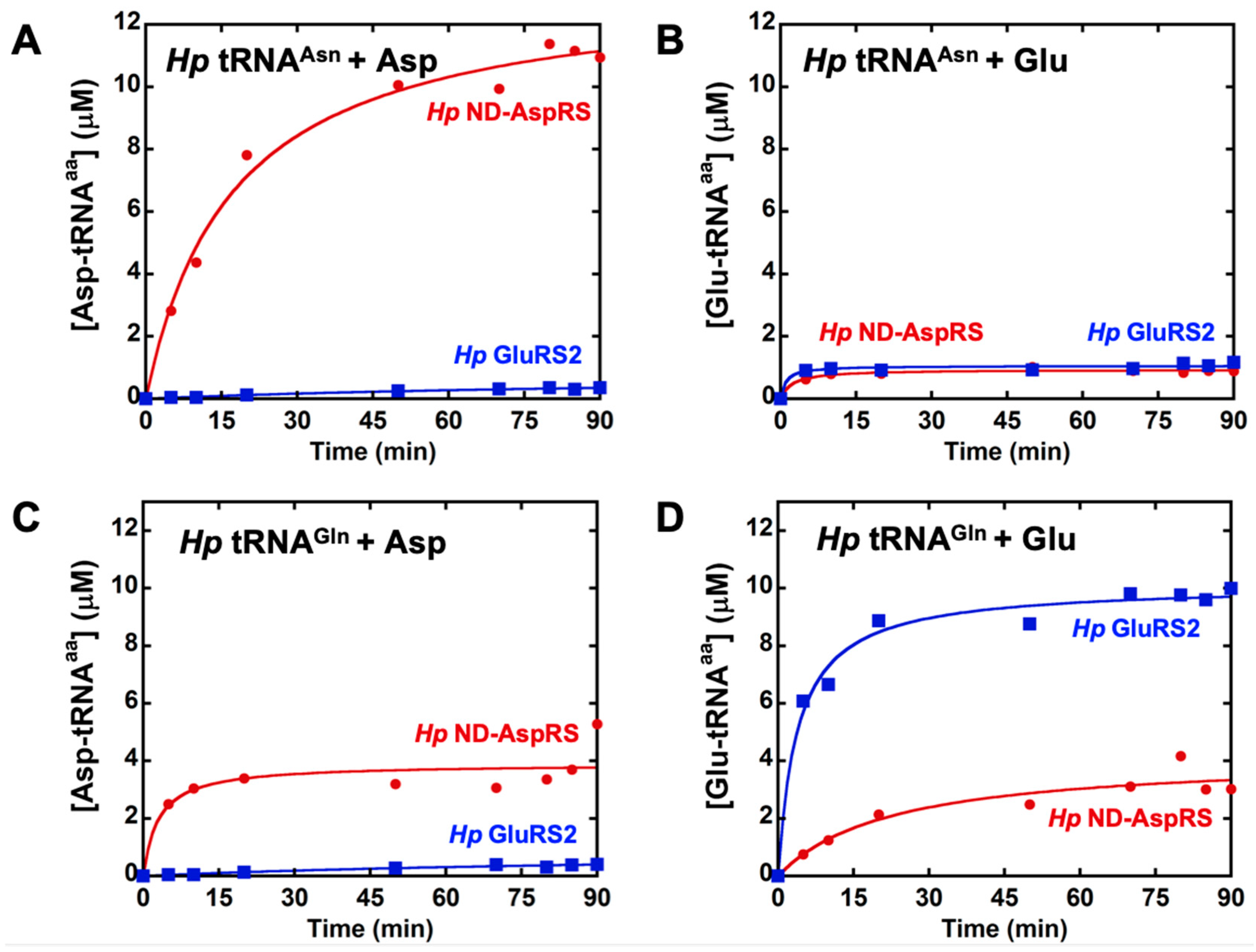
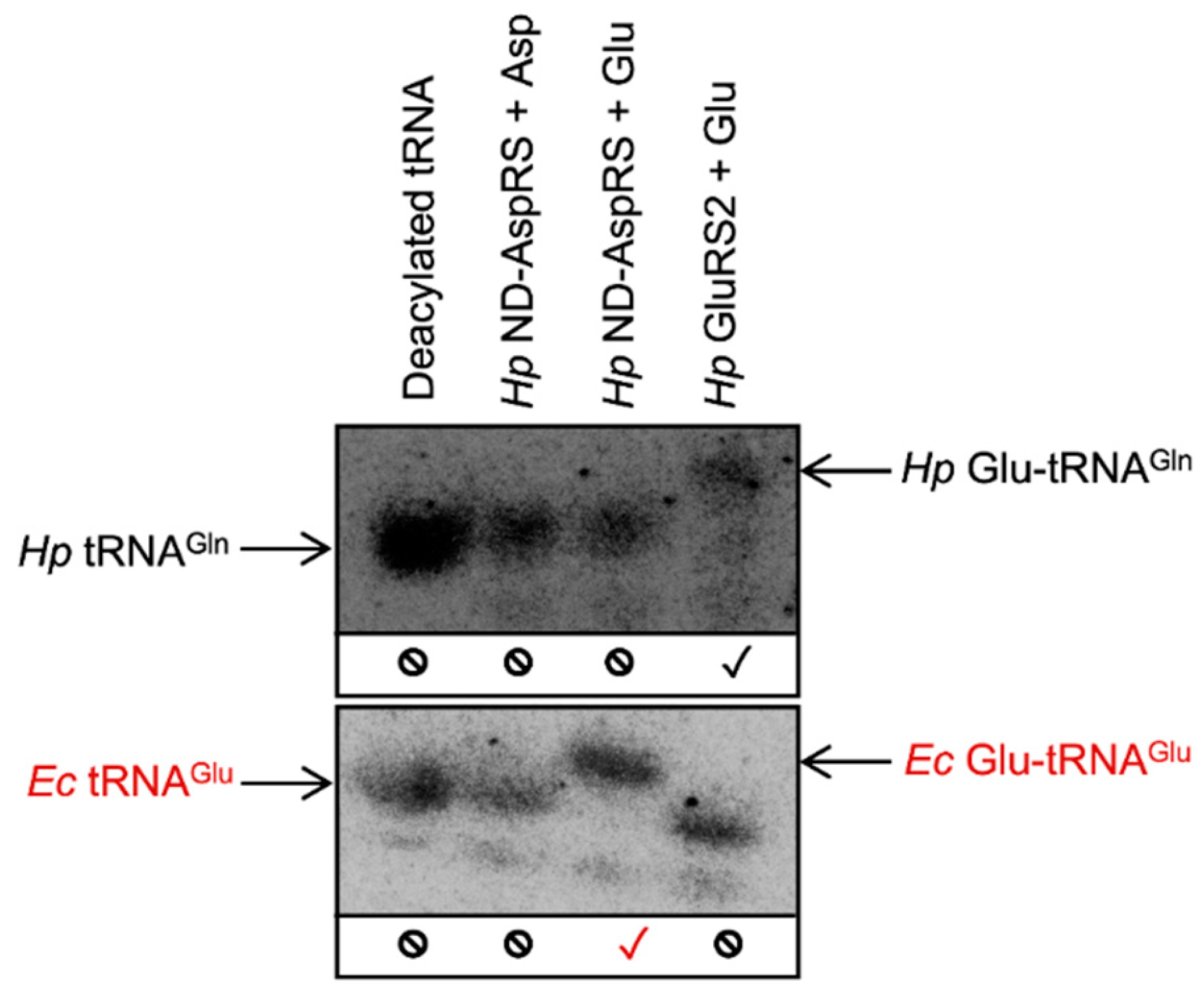
3.1. H. pylori ND-AspRS Appears to Aminoacylate H. pylori tRNAGln with Aspartate and Glutamate
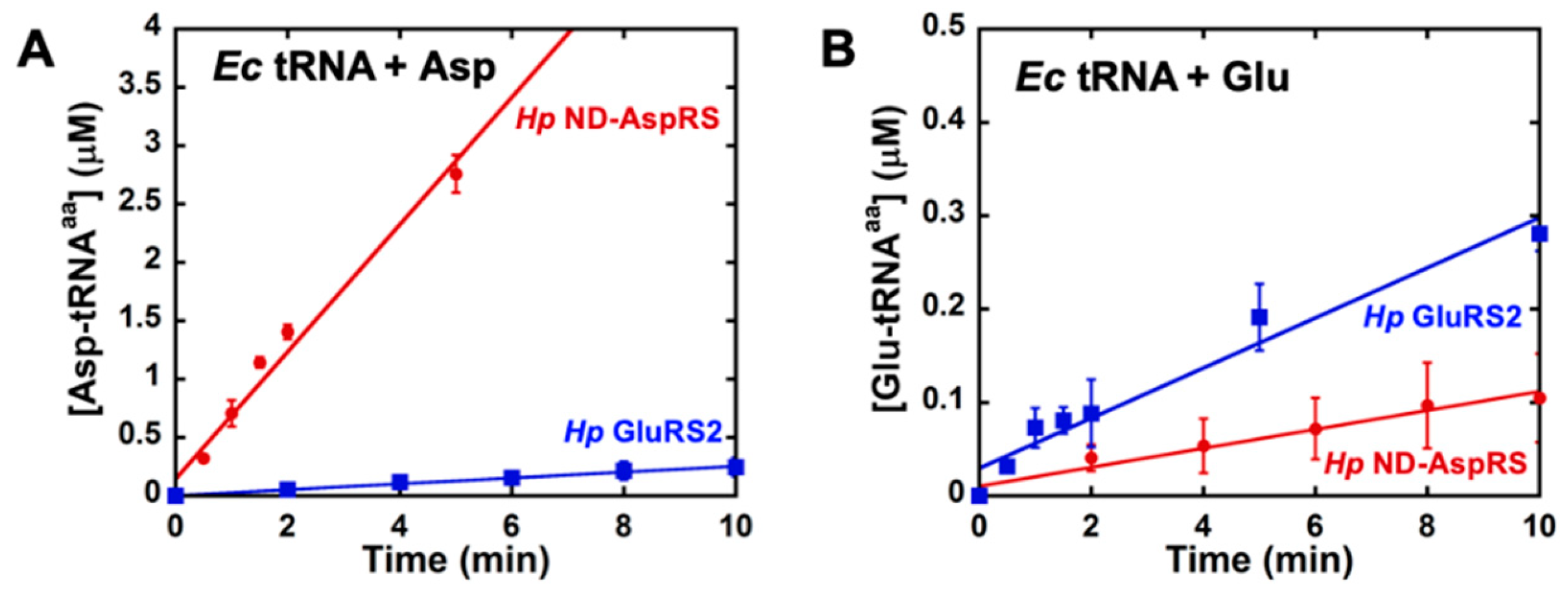
3.2. H. pylori ND-AspRS Aminoacylates E. coli tRNAGlu with Glutamate to Produce Glu-tRNAGlu
3.3. Glu-tRNAGlu Production Is Common among Bacterial Non-Discriminating and Discriminating AspRSs
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ribas de Pouplana, L.; Schimmel, P. A view into the origin of life: aminoacyl-tRNA synthetases. Cell. Mol. Life Sci. 2000, 57, 865–870. [Google Scholar] [CrossRef] [PubMed]
- Ibba, M.; Soll, D. Aminoacyl-tRNA synthesis. Annu. Rev. Biochem. 2000, 69, 617–650. [Google Scholar] [CrossRef]
- Giege, R.; Springer, M. Aminoacyl-tRNA synthetases in the bacterial world. EcoSal Plus 2016, 7. [Google Scholar] [CrossRef]
- Schimmel, P.; Giege, R.; Moras, D.; Yokoyama, S. An operational RNA code for amino acids and possible relationship to genetic code. Proc. Natl. Acad. Sci. USA 1993, 90, 8763–8768. [Google Scholar] [CrossRef]
- Schimmel, P.; Ribas de Pouplana, L. Transfer RNA: From minihelix to genetic code. Cell 1995, 81, 983–986. [Google Scholar] [CrossRef]
- Kubyshkin, V.; Acevedo-Rocha, C.G.; Budisa, N. On universal coding events in protein biogenesis. Biosystems 2018, 164, 16–25. [Google Scholar] [CrossRef] [PubMed]
- Mohler, K.; Ibba, M. Translational fidelity and mistranslation in the cellular response to stress. Nat. Microbiol. 2017, 2, 17117. [Google Scholar] [CrossRef]
- Ribas de Pouplana, L.; Santos, M.A.; Zhu, J.H.; Farabaugh, P.J.; Javid, B. Protein mistranslation: Friend or foe? Trends Biochem. Sci. 2014, 39, 355–362. [Google Scholar] [CrossRef]
- Reynolds, N.M.; Lazazzera, B.A.; Ibba, M. Cellular mechanisms that control mistranslation. Nat. Rev. Microbiol. 2010, 8, 849–856. [Google Scholar] [CrossRef]
- Wiltrout, E.; Goodenbour, J.M.; Frechin, M.; Pan, T. Misacylation of tRNA with methionine in Saccharomyces cerevisiae. Nucleic Acids Res. 2012, 40, 10494–10506. [Google Scholar] [CrossRef]
- Schwartz, M.H.; Pan, T. Determining the fidelity of tRNA aminoacylation via microarrays. Methods 2017, 113, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Javid, B.; Sorrentino, F.; Toosky, M.; Zheng, W.; Pinkham, J.T.; Jain, N.; Pan, M.; Deighan, P.; Rubin, E.J. Mycobacterial mistranslation is necessary and sufficient for rifampicin phenotypic resistance. Proc. Natl. Acad. Sci. USA 2014, 111, 1132–1137. [Google Scholar] [CrossRef] [PubMed]
- Woese, C.R.; Olsen, G.J.; Ibba, M.; Soll, D. Aminoacyl-tRNA synthetases, the genetic code, and the evolutionary process. MMBR 2000, 64, 202–236. [Google Scholar] [CrossRef] [PubMed]
- Curnow, A.W.; Ibba, M.; Soll, D. tRNA-dependent asparagine formation. Nature 1996, 382, 589–590. [Google Scholar] [CrossRef] [PubMed]
- Becker, H.D.; Kern, D. Thermus thermophilus: A link in evolution of the tRNA-dependent amino acid amidation pathways. Proc. Natl. Acad. Sci. USA 1998, 95, 12832–12837. [Google Scholar] [CrossRef]
- Ibba, M.; Soll, D. Aminoacyl-tRNAs: Setting the limits of the genetic code. Genes Dev. 2004, 18, 731–738. [Google Scholar] [CrossRef] [PubMed]
- Schon, A.; Kannangara, C.G.; Gough, S.; Soll, D. Protein biosynthesis in organelles requires misaminoacylation of tRNA. Nature 1988, 331, 187–190. [Google Scholar] [CrossRef]
- Curnow, A.W.; Hong, K.; Yuan, R.; Kim, S.; Martins, O.; Winkler, W.; Henkin, T.M.; Soll, D. Glu-tRNAGln amidotransferase: A novel heterotrimeric enzyme required for correct decoding of glutamine codons during translation. Proc. Natl. Acad. Sci. USA 1997, 94, 11819–11826. [Google Scholar] [CrossRef]
- Rathnayake, U.M.; Wood, W.N.; Hendrickson, T.L. Indirect tRNA aminoacylation during accurate translation and phenotypic mistranslation. Curr. Opin. Chem. Biol. 2017, 41, 114–122. [Google Scholar] [CrossRef]
- Tumbula, D.L.; Becker, H.D.; Chang, W.Z.; Soll, D. Domain-specific recruitment of amide amino acids for protein synthesis. Nature 2000, 407, 106–110. [Google Scholar] [CrossRef] [PubMed]
- Akochy, P.M.; Bernard, D.; Roy, P.H.; Lapointe, J. Direct glutaminyl-tRNA biosynthesis and indirect asparaginyl-tRNA biosynthesis in Pseudomonas aeruginosa PAO1. J. Bacteriol. 2004, 186, 767–776. [Google Scholar] [CrossRef] [PubMed]
- Chuawong, P.; Hendrickson, T.L. The nondiscriminating aspartyl-tRNA synthetase from Helicobacter pylori: Anticodon-binding domain mutations that impact tRNA specificity and heterologous toxicity. Biochemistry 2006, 45, 8079–8087. [Google Scholar] [CrossRef]
- Mladenova, S.R.; Stein, K.R.; Bartlett, L.; Sheppard, K. Relaxed tRNA specificity of the Staphylococcus aureus aspartyl-tRNA synthetase enables RNA-dependent asparagine biosynthesis. FEBS Lett. 2014, 588, 1808–1812. [Google Scholar] [CrossRef] [PubMed]
- Becker, H.D.; Reinbolt, J.; Kreutzer, R.; Giege, R.; Kern, D. Existence of two distinct aspartyl-tRNA synthetases in Thermus thermophilus. Structural and biochemical properties of the two enzymes. Biochemistry 1997, 36, 8785–8797. [Google Scholar] [CrossRef]
- Min, B.; Pelaschier, J.T.; Graham, D.E.; Tumbula-Hansen, D.; Soll, D. Transfer RNA-dependent amino acid biosynthesis: an essential route to asparagine formation. Proc. Natl. Acad. Sci. USA 2002, 99, 2678–2683. [Google Scholar] [CrossRef]
- Skouloubris, S.; Ribas de Pouplana, L.; De Reuse, H.; Hendrickson, T.L. A noncognate aminoacyl-tRNA synthetase that may resolve a missing link in protein evolution. Proc. Natl. Acad. Sci. USA 2003, 100, 11297–11302. [Google Scholar] [CrossRef]
- Shimizu, Y.; Inoue, A.; Tomari, Y.; Suzuki, T.; Yokogawa, T.; Nishikawa, K.; Ueda, T. Cell-free translation reconstituted with purified components. Nat. Biotechnol. 2001, 19, 751–755. [Google Scholar] [CrossRef]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.C.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol. 1999, 112, 531–552. [Google Scholar]
- Zhao, L.; Rathnayake, U.M.; Dewage, S.W.; Wood, W.N.; Veltri, A.J.; Cisneros, G.A.; Hendrickson, T.L. Characterization of tunnel mutants reveals a catalytic step in ammonia delivery by an aminoacyl-tRNA amidotransferase. FEBS Lett. 2016, 590, 3122–3132. [Google Scholar] [CrossRef]
- Varshney, U.; Lee, C.P.; RajBhandary, U.L. Direct analysis of aminoacylation levels of tRNAs in vivo. Application to studying recognition of Escherichia coli initiator tRNA mutants by glutaminyl-tRNA synthetase. J. Biol. Chem. 1991, 266, 24712–24718. [Google Scholar]
- Salazar, J.C.; Ahel, I.; Orellana, O.; Tumbula-Hansen, D.; Krieger, R.; Daniels, L.; Soll, D. Coevolution of an aminoacyl-tRNA synthetase with its tRNA substrates. Proc. Natl. Acad. Sci. USA 2003, 100, 13863–13868. [Google Scholar] [CrossRef]
- Kasai, H.; Oashi, Z.; Harada, F.; Nishimura, S.; Oppenheimer, N.J.; Crain, P.F.; Liehr, J.G.; von Minden, D.L.; McCloskey, J.A. Structure of the modified nucleoside Q isolated from Escherichia coli transfer ribonucleic acid. 7-(4,5-cis-Dihydroxy-1-cyclopenten-3-ylaminomethyl)-7-deazaguanosine. Biochemistry 1975, 14, 4198–4208. [Google Scholar] [CrossRef]
- Sylvers, L.A.; Rogers, K.C.; Shimizu, M.; Ohtsuka, E.; Soll, D. A 2-thiouridine derivative in tRNAGlu is a positive determinant for aminoacylation by Escherichia coli glutamyl-tRNA synthetase. Biochemistry 1993, 32, 3836–3841. [Google Scholar] [CrossRef]
- Giege, R.; Sissler, M.; Florentz, C. Universal rules and idiosyncratic features in tRNA identity. Nucleic Acids Res. 1998, 26, 5017–5035. [Google Scholar] [CrossRef]
- Hasegawa, T.; Himeno, H.; Ishikura, H.; Shimizu, M. Discriminator base of tRNAAsp is involved in amino acid acceptor activity. Biochem. Biophys. Res. Commun. 1989, 163, 1534–1538. [Google Scholar] [CrossRef]
- Nameki, N.; Tamura, K.; Himeno, H.; Asahara, H.; Hasegawa, T.; Shimizu, M. Escherichia coli tRNAAsp recognition mechanism differing from that of the yeast system. Biochem. Biophys. Res. Commun. 1992, 189, 856–862. [Google Scholar] [CrossRef]
- Gregory, S.T.; Dahlberg, A.E. Effects of mutations at position 36 of tRNAGlu on missense and nonsense suppression in Escherichia coli. FEBS Lett. 1995, 361, 25–28. [Google Scholar] [CrossRef]
- Sekine, S.; Nureki, O.; Sakamoto, K.; Niimi, T.; Tateno, M.; Go, M.; Kohno, T.; Brisson, A.; Lapointe, J.; Yokoyama, S. Major identity determinants in the “augmented D helix” of tRNAGlu from Escherichia coli. J. Mol. Biol. 1996, 256, 685–700. [Google Scholar] [CrossRef] [PubMed]
- Sekine, S.; Nureki, O.; Tateno, M.; Yokoyama, S. The identity determinants required for the discrimination between tRNAGlu and tRNAAsp by glutamyl-tRNA synthetase from Escherichia coli. Eur. J. Biochem. 1999, 261, 354–360. [Google Scholar] [CrossRef]
- Rodin, S.N.; Ohno, S. Four primordial modes of tRNA-synthetase recognition, determined by the (G,C) operational code. Proc. Natl. Acad. Sci. USA 1997, 94, 5183–5188. [Google Scholar] [CrossRef]
- Ribas de Pouplana, L.; Schimmel, P. Two classes of tRNA synthetases suggested by sterically compatible dockings on tRNA acceptor stem. Cell 2001, 104, 191–193. [Google Scholar] [CrossRef]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rathnayake, U.M.; Hendrickson, T.L. Bacterial Aspartyl-tRNA Synthetase Has Glutamyl-tRNA Synthetase Activity. Genes 2019, 10, 262. https://doi.org/10.3390/genes10040262
Rathnayake UM, Hendrickson TL. Bacterial Aspartyl-tRNA Synthetase Has Glutamyl-tRNA Synthetase Activity. Genes. 2019; 10(4):262. https://doi.org/10.3390/genes10040262
Chicago/Turabian StyleRathnayake, Udumbara M., and Tamara L. Hendrickson. 2019. "Bacterial Aspartyl-tRNA Synthetase Has Glutamyl-tRNA Synthetase Activity" Genes 10, no. 4: 262. https://doi.org/10.3390/genes10040262
APA StyleRathnayake, U. M., & Hendrickson, T. L. (2019). Bacterial Aspartyl-tRNA Synthetase Has Glutamyl-tRNA Synthetase Activity. Genes, 10(4), 262. https://doi.org/10.3390/genes10040262




