The Hearing Impairment Ontology: A Tool for Unifying Hearing Impairment Knowledge to Enhance Collaborative Research
Abstract
:1. Introduction
2. Materials and Methods
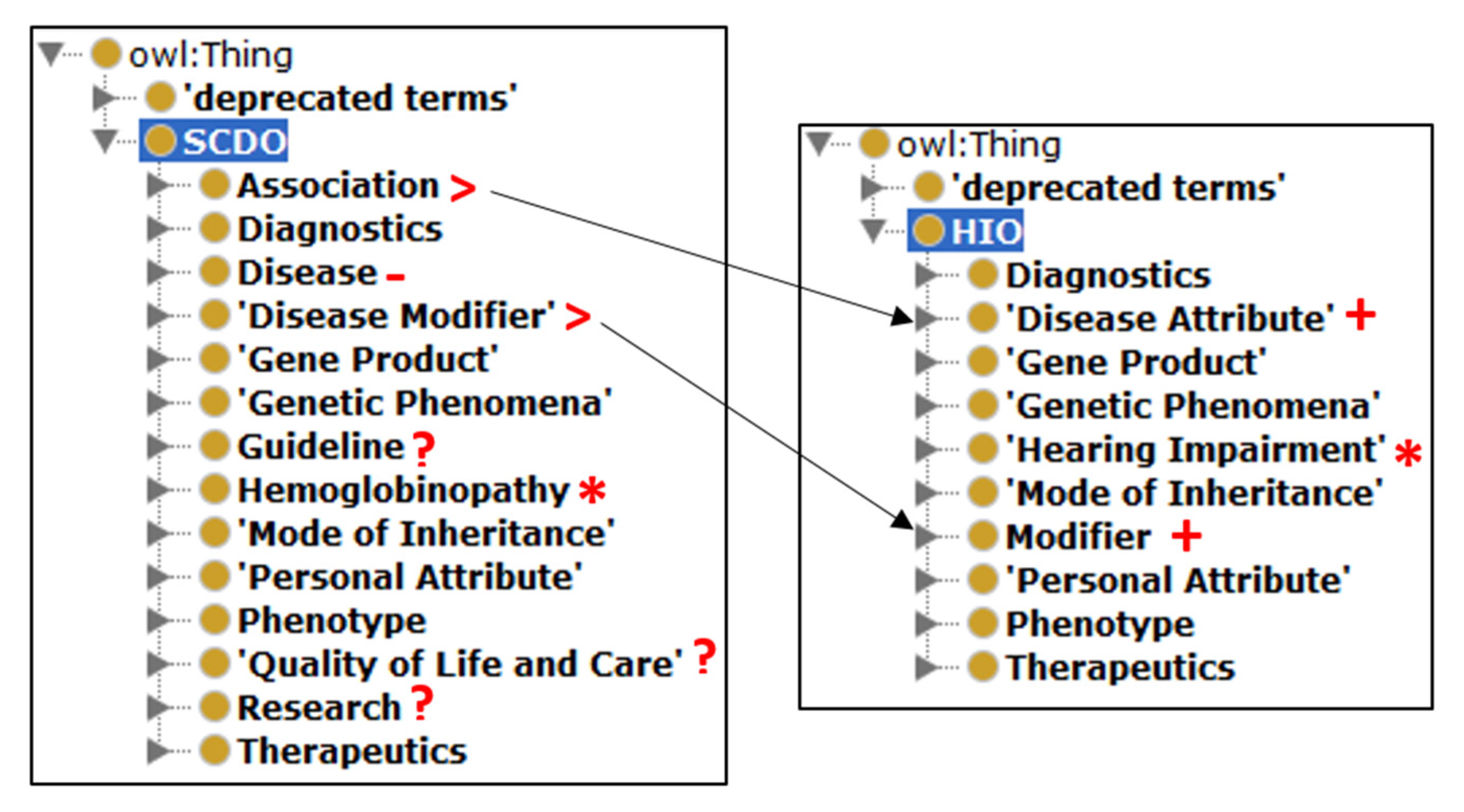
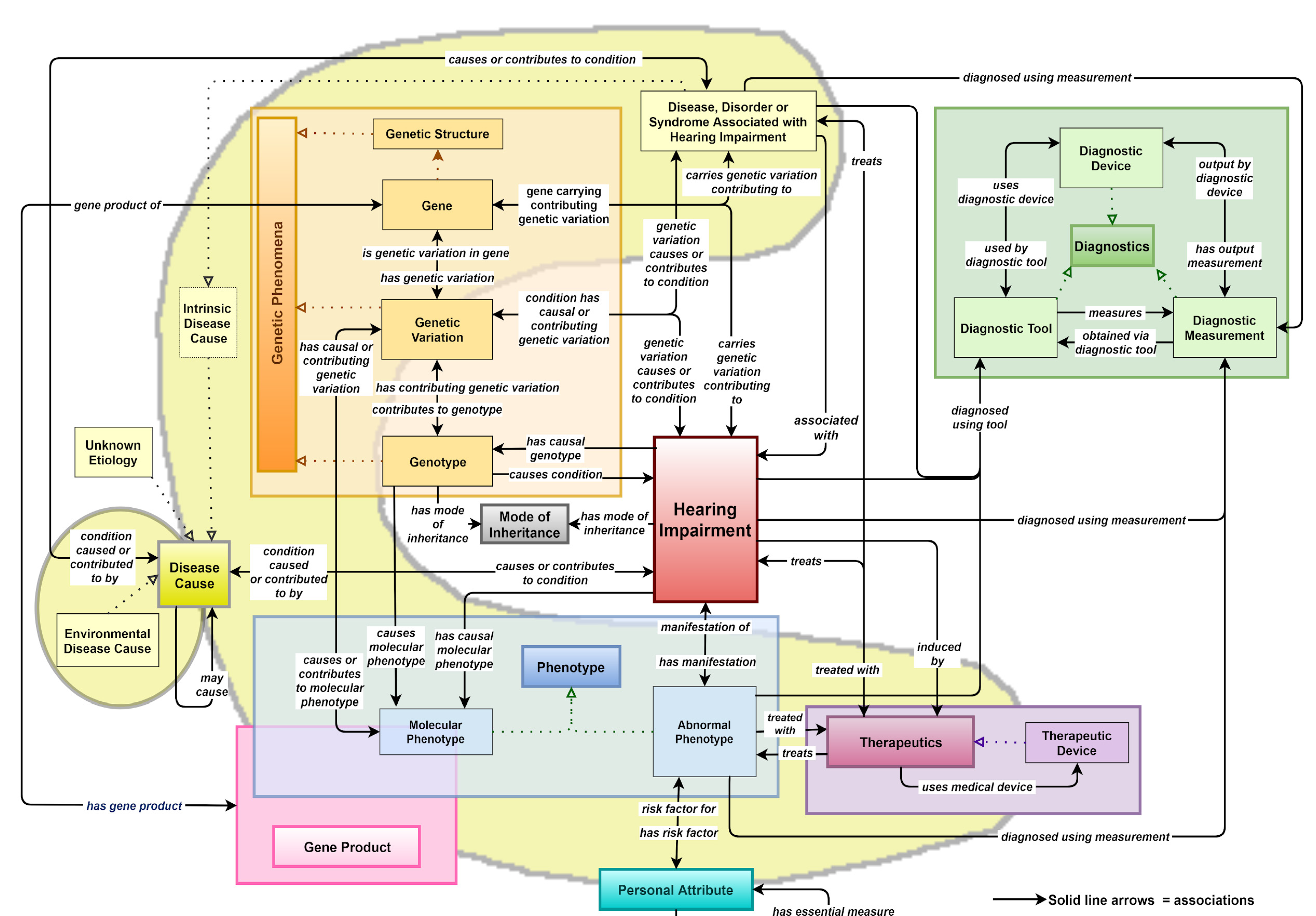
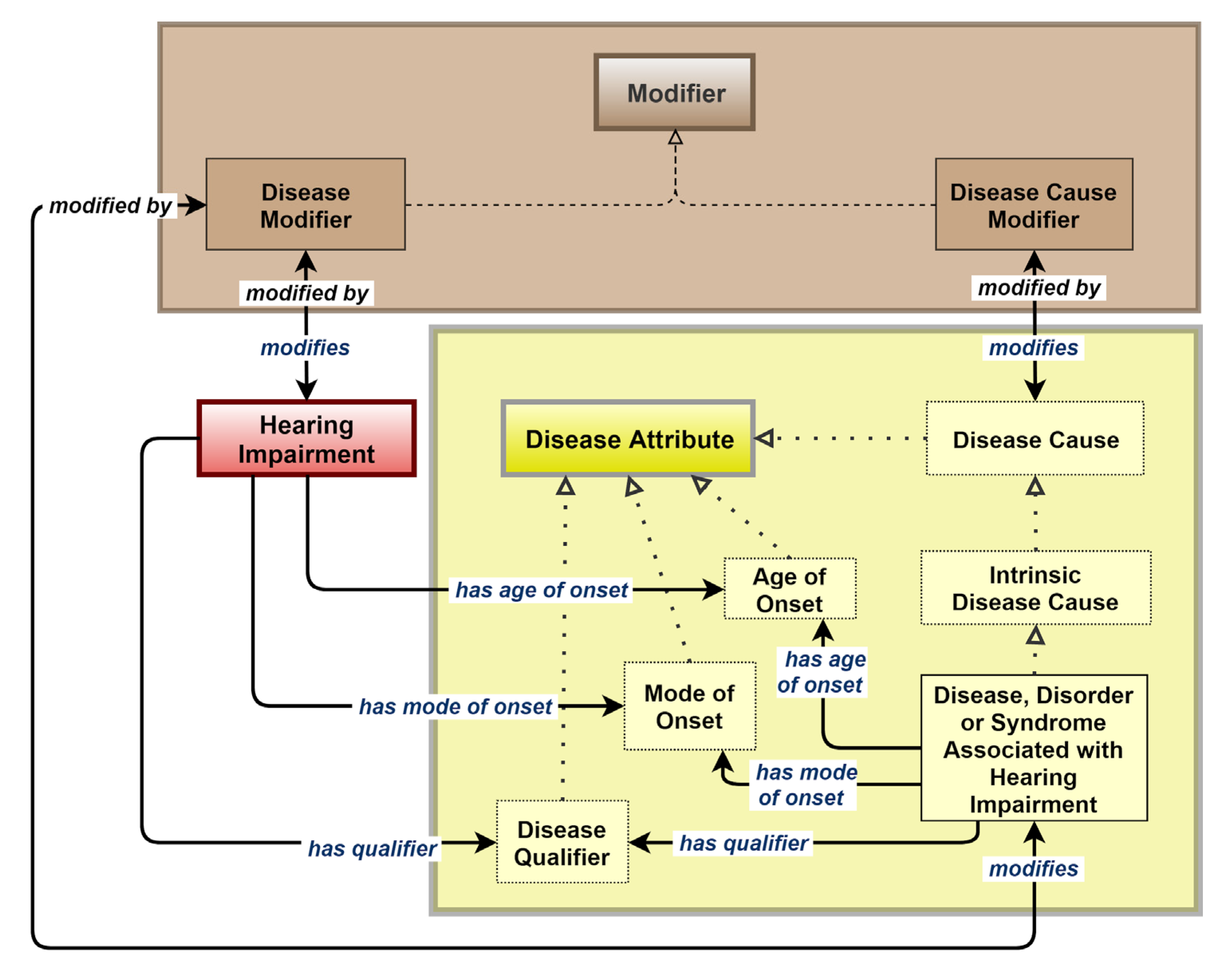
2.1. SCDO Model-Based HIO Development
2.2. Building Different HIO Objects
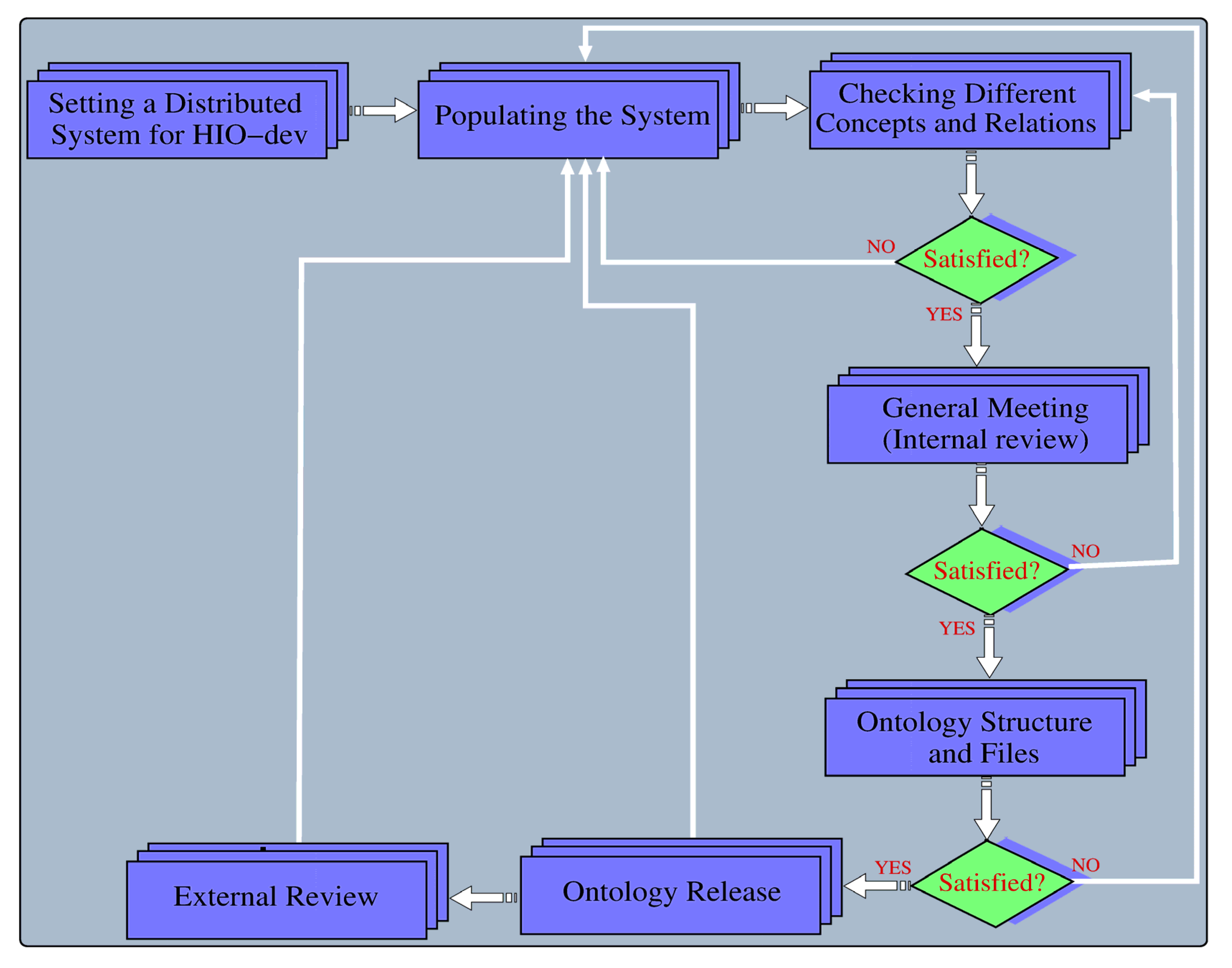
2.3. Distributed Model-Based HIO Design
2.4. HIO File Refinement and Evolution
3. Results
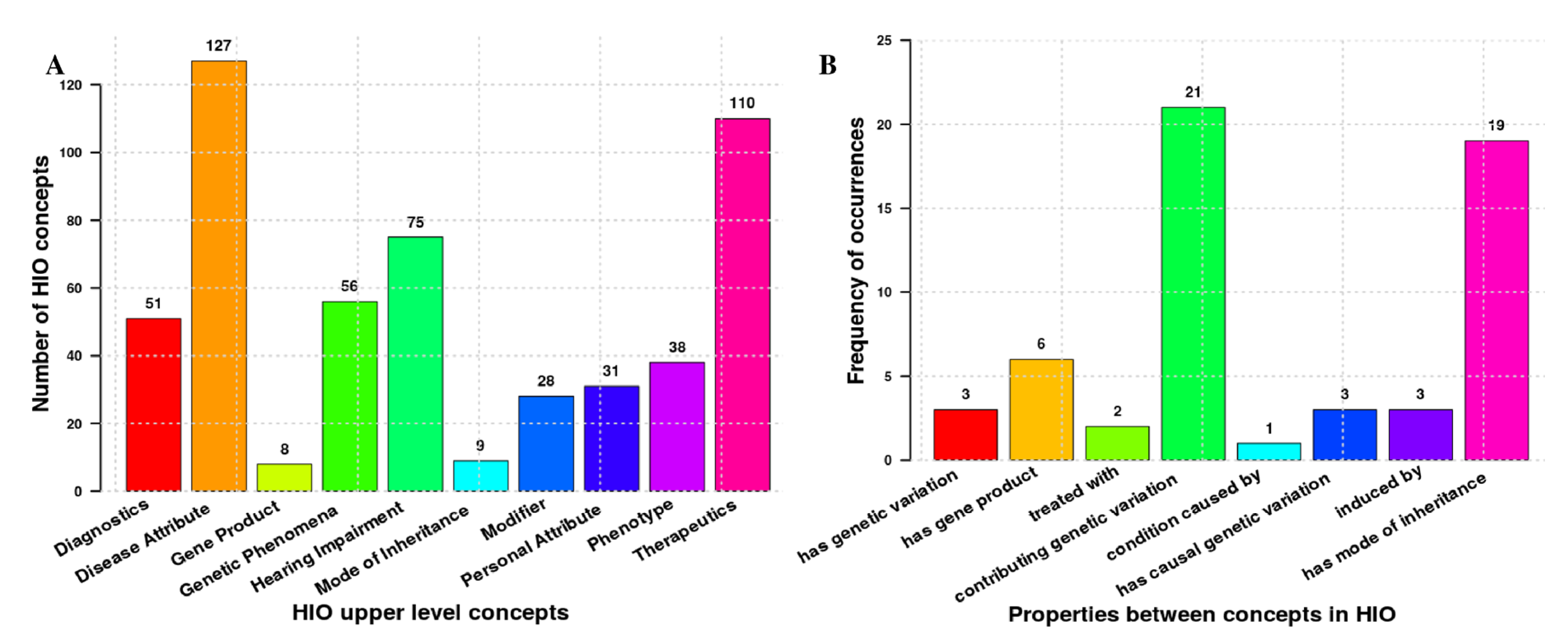
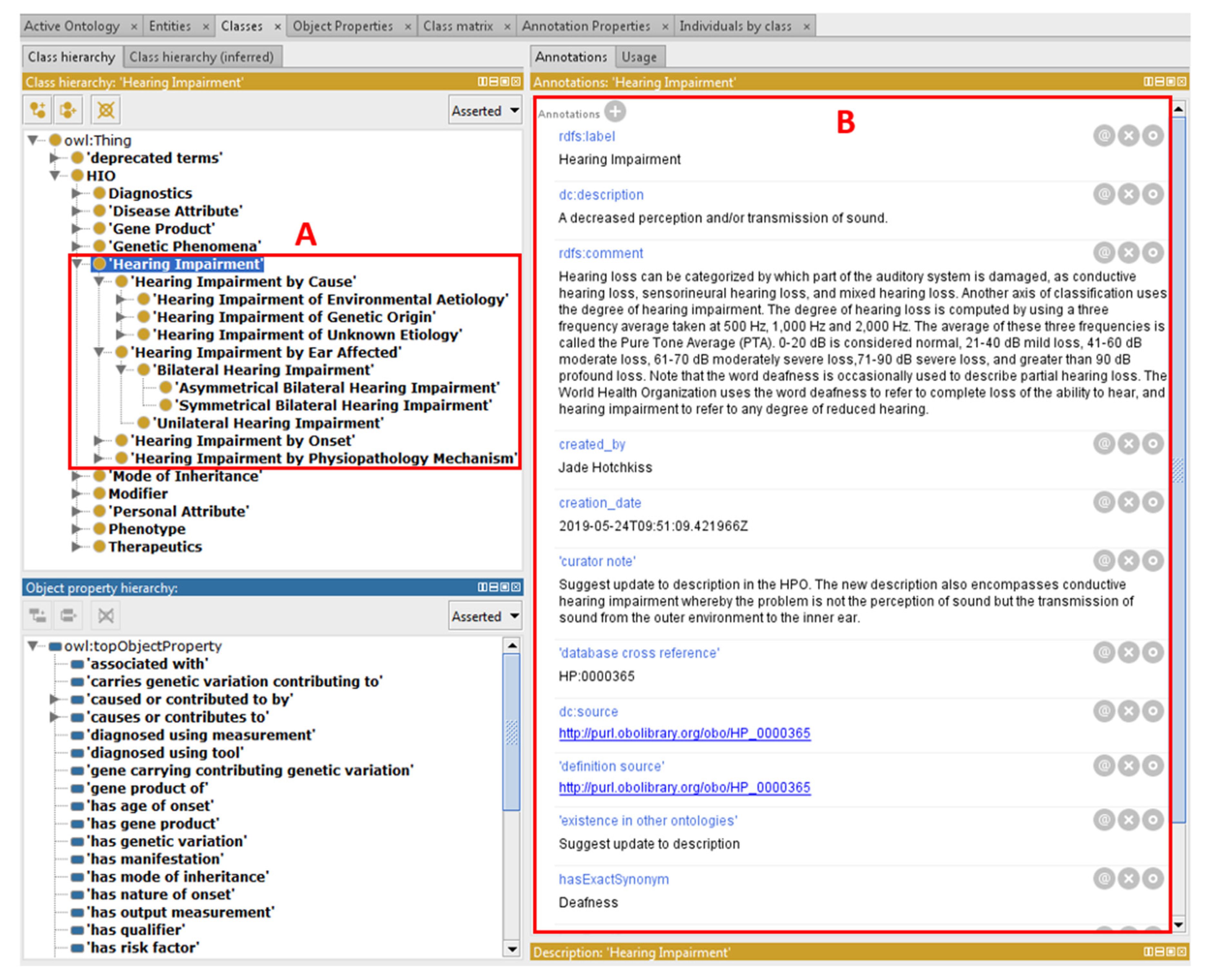
3.1. HIO General Description
3.2. Assessing the Relevance of HIO
3.3. HIO Release and License
3.4. Different HIO Access Platforms
4. Discussion
4.1. HIO Structure, Other Disease Ontologies and HI Online Datasets
4.2. HIO Potential Future Applications
4.3. Challenges and Future Direction
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Chadha, S.; Cieza, A.; Reyes, K. Public health approach to hearing across the life course: A call-for-papers. Bull. World Health Organ. 2018, 96, 592. [Google Scholar] [CrossRef] [PubMed]
- Murray, E.; McCabe, P.; Heard, R.; Ballard, K.J. Differential diagnosis of children with suspected childhood apraxia of speech. J. Speech Lang. Hear. Res. 2015, 58, 43–60. [Google Scholar] [CrossRef] [PubMed]
- Vos, T.; Allen, C.; Arora, M.; Barber, R.M.; Bhutta, Z.A.; Brown, A.; Carter, A.; Casey, D.C.; Charlson, F.J.; Chen, A.Z.; et al. Global, regional, and national incidence, prevalence, and years lived with disability for 310 diseases and injuries, 1990–2015: A systematic analysis for the global burden of disease study 2015. Lancet 2016, 388, 1545–1602. [Google Scholar] [CrossRef]
- Global Estimates on Hearing Loss; The World Health Organization: Geneva, Switzerland, 2018; Available online: http://www.who.int/pbd/deafness/estimates/en/ (accessed on 27 June 2019).
- Graydon, K.; Waterworth, C.; Miller, H.; Gunasekera, H. Global burden of hearing impairment and ear disease. J. Laryngol. Otol. 2019, 133, 18–25. [Google Scholar] [CrossRef] [PubMed]
- WHO. Global Costs of Unaddressed Hearing Loss and Cost-Effectiveness of Intervention; The World Health Organization: Geneva, Switzerland, 2017. [Google Scholar]
- The SCDO Working Group. The Sickle Cell Disease Ontology: Enabling universal sickle cell-based knowledge representation. Database 2019, in press. [Google Scholar]
- Löhler, J.; Walther, L.E.; Hansen, F.; Kapp, P.; Meerpohl, J.; Wollenberg, B.; Schönweiler, R.; Schmucker, C. The prevalence of hearing loss and use of hearing aids among adults in Germany: A systematic review. Eur. Arch. Otorhinolaryngol. 2019, 276, 945–956. [Google Scholar] [CrossRef]
- Martinez-Cruz, C.; Blanco, I.J.; Vila, M.A. Ontologies versus relational databases: Are they so different? A comparison. Artif. Intell. Rev. 2011, 38, 271–290. [Google Scholar] [CrossRef]
- Gruber, T.R. Toward principles for the design of ontologies used for knowledge sharing? Int. J. Hum-Comput. St. 1995, 43, 907–928. [Google Scholar] [CrossRef]
- Mazandu, G.K.; Mulder, N.J. A topology-based metric for measuring term similarity in the Gene Ontology. Adv. Bioinform. 2012, 2012, 17. [Google Scholar] [CrossRef]
- Mazandu, G.K.; Chimusa, E.R.; Mulder, N.J. Gene ontology semantic similarity tools: Survey on features and challenges for biological knowledge discovery. Brief. Bioinform. 2018, 18, 886–901. [Google Scholar] [CrossRef]
- Kibbe, W.A.; Arze, C.; Felix, V.; Mitraka, E.; Bolton, E.; Fu, G.; Mungall, C.J.; Binder, J.X.; Malone, J.; Vasant, D.; et al. Disease Ontology 2015 update: An expanded and updated database of human diseases for linking biomedical knowledge through disease data. Nucleic Acids Res. 2015, 43, D1071–D1078. [Google Scholar] [CrossRef] [PubMed]
- Köhler, S.; Vasilevsky, N.A.; Engelstad, M.; Foster, E.; McMurry, J.; Aymé, S.; Baynam, G.; Bello, S.M.; Boerkoel, C.F.; Boycott, K.M.; et al. The Human Phenotype Ontology in 2017. Nucleic Acids Res. 2017, 45, D865–D876. [Google Scholar] [CrossRef] [PubMed]
- Köhler, S.; Carmody, L.; Vasilevsky, N.; Jacobsen, J.O.B.; Danis, D.; Gourdine, J.P.; Gargano, M.; Harris, N.L.; Matentzoglu, N.; McMurry, J.A.; et al. Expansion of the Human Phenotype Ontology (HPO) knowledge base and resources. Nucleic Acids Res. 2019, 47, D1018–D1027. [Google Scholar] [CrossRef] [PubMed]
- Matentzoglu, N.; Malone, J.; Mungall, C.; Stevens, R. MIRO: Guidelines for minimum information for the reporting of an ontology. J. Biomed. Semant. 2018, 9, 6. [Google Scholar] [CrossRef]
- Horridge, M.; Gonçalves, R.S.; Nyulas, C.; Musen, M.A. WebProtégé: A Cloud-Based Ontology Editor. arXiv 2019, arXiv:1902.08251. [Google Scholar]
- Jackson, R.C.; Balhoff, J.P.; Douglass, E.; Harris, N.L.; Mungall, C.J.; Overton, J.A. ROBOT: A Tool for Automating Ontology Workflows. BMC Bioinform. 2019, 20, 407. [Google Scholar] [CrossRef]
- Nance, W.E. The genetics of deafness. Ment. Retard. Dev. Disabil. Res. Rev. 2003, 9, 109–119. [Google Scholar] [CrossRef]
- Adadey, S.M.; Awandare, G.; Amedofu, G.K.; Wonkam, A. Public Health Burden of Hearing Impairment and the Promise of Genomics and Environmental Research: A Case Study in Ghana, Africa. OMICS 2017, 21, 638–646. [Google Scholar] [CrossRef]
- Fook, L.; Morgan, R. Hearing impairment in older people: A review. Postgrad. Med. J. 2000, 76, 537–541. [Google Scholar] [CrossRef]
- Szyfter, W.; Wróbel, M.J.; Szyfter-Harris, J.; Greczka, G. Hearing impairment in Polish infants. Epidemiology 2013, 24, 333. [Google Scholar] [CrossRef]
- Cunningham, L.L.; Tucci, D.L. Hearing Loss in Adults. N. Engl. J. Med. 2017, 377, 2465–2473. [Google Scholar] [CrossRef] [PubMed]
- Rudman, J.R.; Kabahuma, R.I.; Bressler, S.E.; Feng, Y.; Blanton, S.H.; Yan, D.; Liu, X.Z. The genetic basis of deafness in populations of African descent. J. Genet. Genom. 2017, 44, 285–294. [Google Scholar] [CrossRef] [PubMed]
- Angela, S.; Lin, X.; Liu, X.Z. Genetics of Hearing and Deafness. Anat. Rec. 2012, 295, 1812–1829. [Google Scholar] [CrossRef] [PubMed]
- Lustig, L.R. Drug-Induced Ototoxicity. 2018. Available online: https://www.msdmanuals.com/professional/ear,-nose,-and-throat-disorders/inner-ear-disorders/drug-induced-ototoxicity (accessed on 28 June 2019).
- Shen, J.; Scheffer, D.I.; Kwan, K.Y.; Corey, D.P. SHIELD: An integrative gene expression database for inner ear research. Database 2015, 2015, bav071. [Google Scholar] [CrossRef] [PubMed]
- Azaiez, H.; Booth, K.T.; Ephraim, S.S.; Crone, B.; Black-Ziegelbein, E.A.; Marini, R.J.; Shearer, A.E.; Sloan-Heggen, C.M.; Kolbe, D.; Casavant, T.; et al. Genomic Landscape and Mutational Signatures of Deafness-Associated Genes. Am. J. Hum. Genet. 2018, 103, 484–497. [Google Scholar] [CrossRef]
- Fokkema, I.F.; Taschner, P.E.; Schaafsma, G.C.; Celli, J.; Laros, J.F.; den Dunnen, J.T. LOVD v.2.0: The next generation in gene variant databases. Hum. Mutat. 2011, 32, 557–563. [Google Scholar] [CrossRef]
- Amberger, J.S.; Bocchini, C.A.; Scott, A.F.; Hamosh, A. OMIM.org: Leveraging knowledge across phenotype-gene relationships. Nucleic Acids Res. 2019, 47, D1038–D1043. [Google Scholar] [CrossRef]
- Mazandu, G.K.; Kyomugisha, I.; Geza, E.; Seuneu, M.; Bah, B.; Chimusa, E.R. Designing Data-Driven Learning Algorithms: A Necessity to Ensure Effective Post-Genomic Medicine and Biomedical Research, Artificial Intelligence—Applications in Medicine and Biology; IntechOpen: London, UK, 2019; Available online: https://www.intechopen.com/books/artificial-intelligence-applications-in-medicine-and-biology/designing-data-driven-learning-algorithms-a-necessity-to-ensure-effective-post-genomic-medicine-and- (accessed on 16 September 2019).
- Zhao, Y.; Li, J.; Zhang, M.; Lu, Y.; Xie, H.; Tian, Y.; Qiu, W. Machine Learning Models for the Hearing Impairment Prediction in Workers Exposed to Complex Industrial Noise: A Pilot Study. Ear Hear. 2019, 40, 690–699. [Google Scholar] [CrossRef]

| Existence Status | Explanation of Status | No. Terms | % Terms |
|---|---|---|---|
| Sufficient | Exists in other ontology and has appropriate description | 284 | 71.2 |
| Suggest update to description | Used term from existing ontology but will suggest they update their description to ours | 27 | 6.8 |
| Suggest update to label | Used term from existing ontology but will suggest they update their label to ours | 0 | 0 |
| Suggest update to label and description | Used term from existing ontology but will suggest they update their label and description to ours | 0 | 0 |
| Few but definitions not available | Term exists in a few ontologies but has not been given a description in any | 3 | 0.8 |
| Few but definitions not freely available | Term exists in a few ontologies but the description is not freely available | 8 | 2.0 |
| Few but definitions not specific enough | Term exists in a few ontologies but the definitions are not specific enough for the HIO’s needs | 9 | 2.3 |
| Not relevant to context of hearing impairment | Term exists in other ontologies but the definitions are not relevant to the HI field | 3 | 0.8 |
| Negligible | No description or outdated ontology | 2 | 0.5 |
| None | Not in any existing ontology | 63 | 15.8 |
| Term Label | Term ID | Term Description |
|---|---|---|
| Symmetrical Bilateral Hearing Impairment | HIO:0000365 | When the severity and configuration of hearing impairment is approximately the same in both ears. |
| Asymmetrical Bilateral Hearing Impairment | HIO:0000366 | When each ear has a different severity and configuration of hearing impairment. |
| Postlingual Hearing Impairment | HIO:0000475 | Hearing impairment which develops after the acquisition of speech and language, usually after the age of six. |
| Prelingual Hearing Impairment | HIO:0000476 | Hearing impairment which is either congenital or develops before the acquisition of speech and language, usually before the age of 6. |
| Temporal Bone Fracture with Otic Capsule Involvement | HIO:0000287 | Traumatic injury to the temporal bone in which the continuity of the bone is broken and violation of the otic capsule is involved. |
| Temporal Bone Fracture without Otic Capsule Involvement | HIO:0000288 | Traumatic injury to the temporal bone in which the continuity of the bone is broken and violation of the otic capsule is not involved. |
| Pseudo-Dominant Inheritance | HIO:0000228 | When the inheritance of a recessive trait mimics a dominant pattern of inheritance. |
| Cisplatin-Induced Hearing Impairment | HIO:0000215 | Hearing loss caused by cisplatin (a chemotherapeutic agent) ototoxicity. |
| Neomycin-Induced Hearing Impairment | HIO:0000285 | Partial or complete loss of hearing following ingestion of neomycin. |
| Maternal Medical History | HIO:0000362 | A record of a patient’s biological mother’s background regarding health and the occurrence of disease events of the mother. |
| Hearing Impairment based on Immaturity | HIO:0000514 | Hearing impairment that occurs due to premature birth (birth at or before 37 weeks of gestational age). |
| Scheme | Description | Types | URL | Reference |
|---|---|---|---|---|
| HHL | Hereditary Hearing Loss Homepage | An up-to-date overview of the genetics of hereditary hearing impairment for researchers and clinicians working in the field. | https://hereditaryhearingloss.org/ | - |
| SHIELD | The Shared Harvard Inner Ear Laboratory Database | An integrative gene expression database for inner ear research | https://shield.hms.harvard.edu | [27] |
| DVD | Deafness Variation Database | A comprehensive resource integrating available genetic, genomic, and clinical data together with expert curation to generate a single classification for each variant in 152 genes implicated in syndromic and non-syndromic deafness. | http://deafnessvariationdatabase.org/ | [28] |
| LOVD | Leiden Open Variation Databse | Retinal and hearing impairment genetic variant database | https://databases.lovd.nl/shared/genes/OTOF | [29] |
| NIDCD | National Institute on Deafness and Other Communication Disorders | A resource providing knowledge about Hearing, Ear Infections, and Deafness Diseases and Conditions. It also provides NIDCD Temporal Bone Registry at https://www.tbregistry.org/, a resource for learning about the pathology and pathophysiology of otologic disorders, which serves as a resource for scientists to analyze data from a collection of more than 12,000 temporal bone specimens. | https://www.nidcd.nih.gov/health/hearing-ear-infections-deafness | - |
| gEAR | Gene Expression Analysis Resource | Visualization and analysis of multiomic data both in public and private domains. | https://umgear.org/ | - |
| OMIM | Online Mendelian Inheritance in Man | An Online Catalog of Human Genes and Genetic Disorders | https://www.omim.org | [30] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hotchkiss, J.; Manyisa, N.; Mawuli Adadey, S.; Oluwole, O.G.; Wonkam, E.; Mnika, K.; Yalcouye, A.; Nembaware, V.; Haendel, M.; Vasilevsky, N.; et al. The Hearing Impairment Ontology: A Tool for Unifying Hearing Impairment Knowledge to Enhance Collaborative Research. Genes 2019, 10, 960. https://doi.org/10.3390/genes10120960
Hotchkiss J, Manyisa N, Mawuli Adadey S, Oluwole OG, Wonkam E, Mnika K, Yalcouye A, Nembaware V, Haendel M, Vasilevsky N, et al. The Hearing Impairment Ontology: A Tool for Unifying Hearing Impairment Knowledge to Enhance Collaborative Research. Genes. 2019; 10(12):960. https://doi.org/10.3390/genes10120960
Chicago/Turabian StyleHotchkiss, Jade, Noluthando Manyisa, Samuel Mawuli Adadey, Oluwafemi Gabriel Oluwole, Edmond Wonkam, Khuthala Mnika, Abdoulaye Yalcouye, Victoria Nembaware, Melissa Haendel, Nicole Vasilevsky, and et al. 2019. "The Hearing Impairment Ontology: A Tool for Unifying Hearing Impairment Knowledge to Enhance Collaborative Research" Genes 10, no. 12: 960. https://doi.org/10.3390/genes10120960
APA StyleHotchkiss, J., Manyisa, N., Mawuli Adadey, S., Oluwole, O. G., Wonkam, E., Mnika, K., Yalcouye, A., Nembaware, V., Haendel, M., Vasilevsky, N., Mulder, N. J., Jupp, S., Wonkam, A., & K. Mazandu, G. (2019). The Hearing Impairment Ontology: A Tool for Unifying Hearing Impairment Knowledge to Enhance Collaborative Research. Genes, 10(12), 960. https://doi.org/10.3390/genes10120960





