Enhancing Genetic Medicine: Rapid and Cost-Effective Molecular Diagnosis for a GJB2 Founder Mutation for Hearing Impairment in Ghana
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethical Approvals
2.2. Study Participants
2.3. Molecular Analyses
2.4. Data Analysis
3. Results
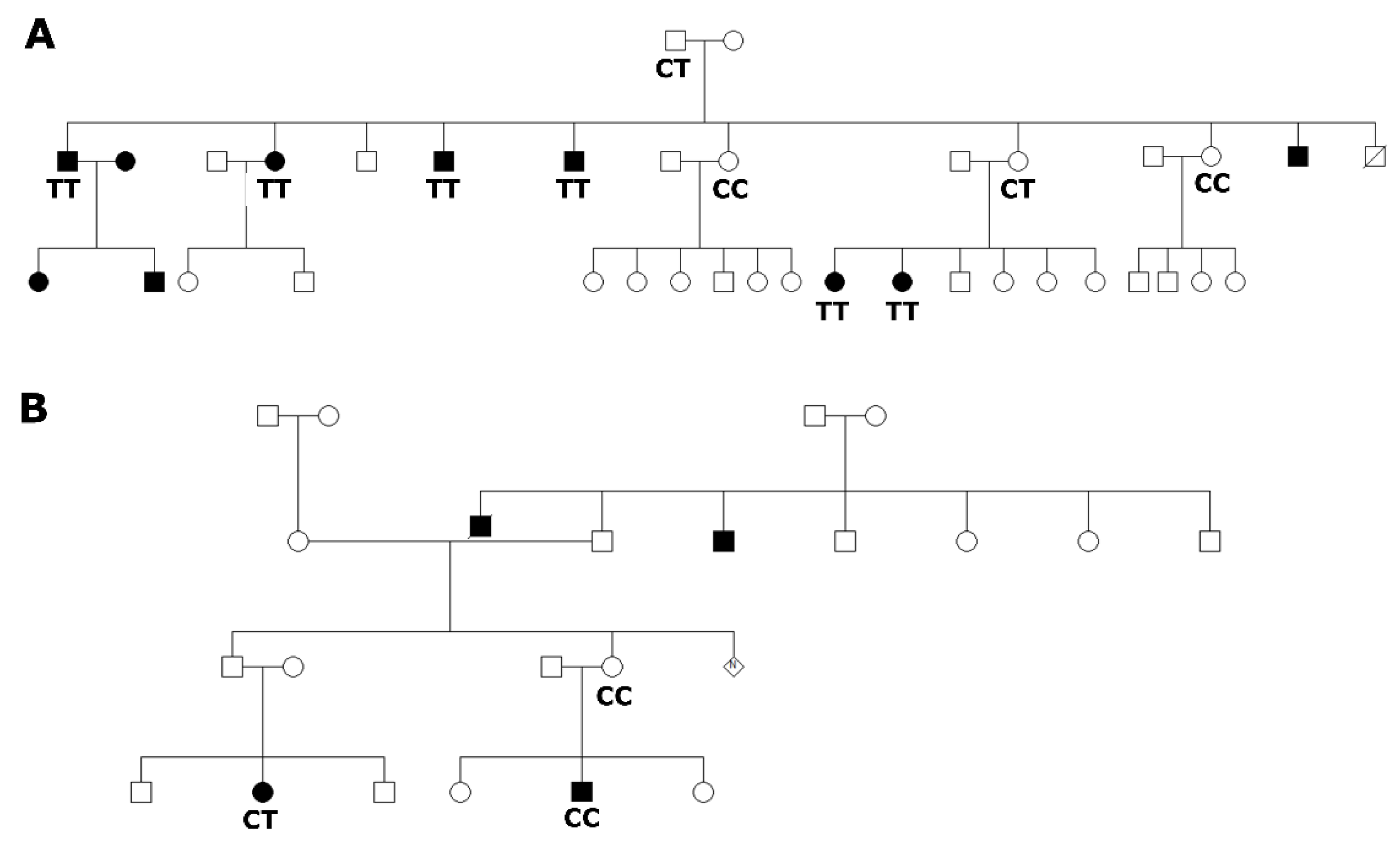
3.1. Selected Families Segregating Hearing Impairment from Adamorobe Village, Ghana
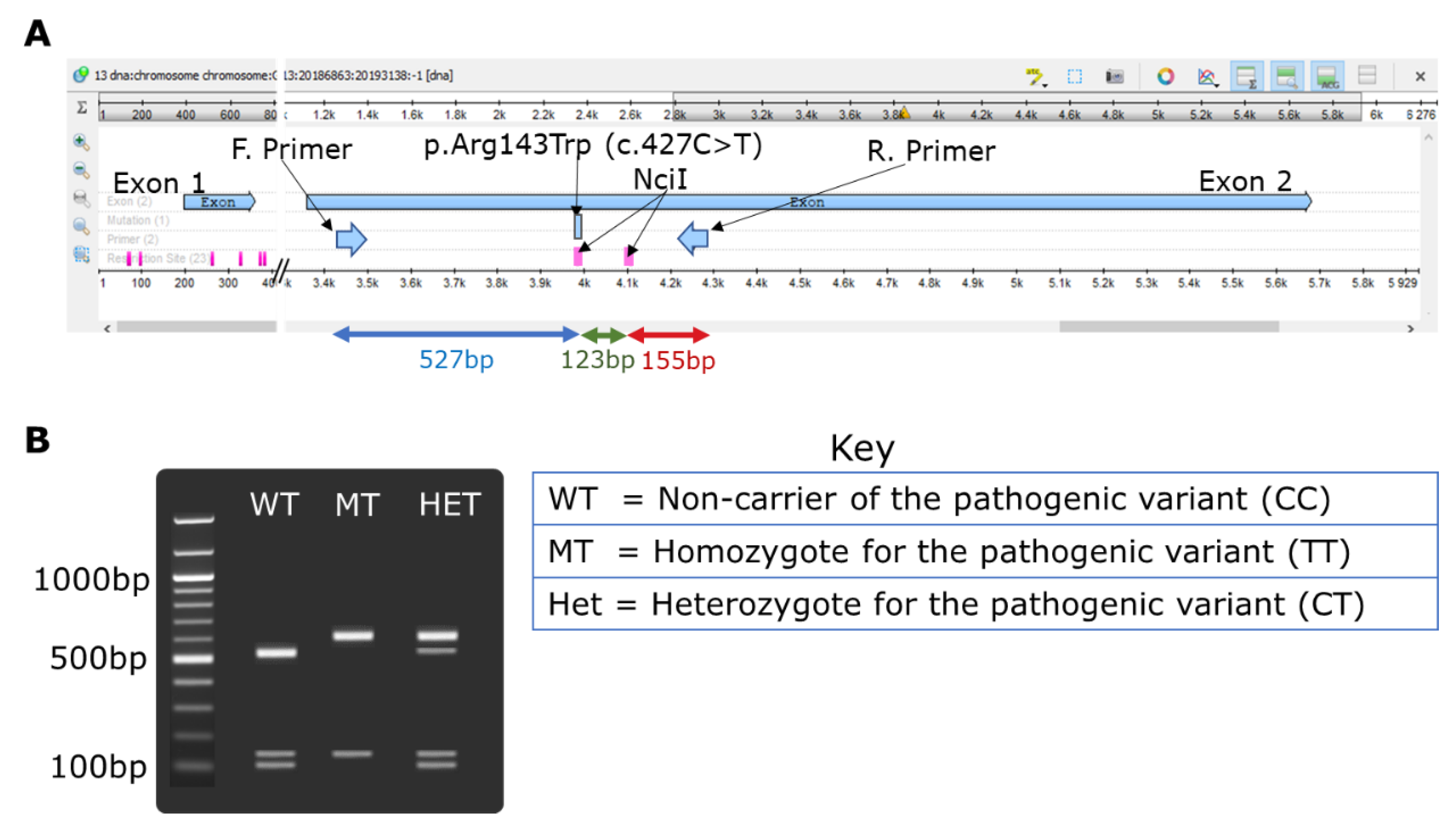
3.2. Restriction Fragment Polymorphism Design for GJB2-p.Arg143Trp
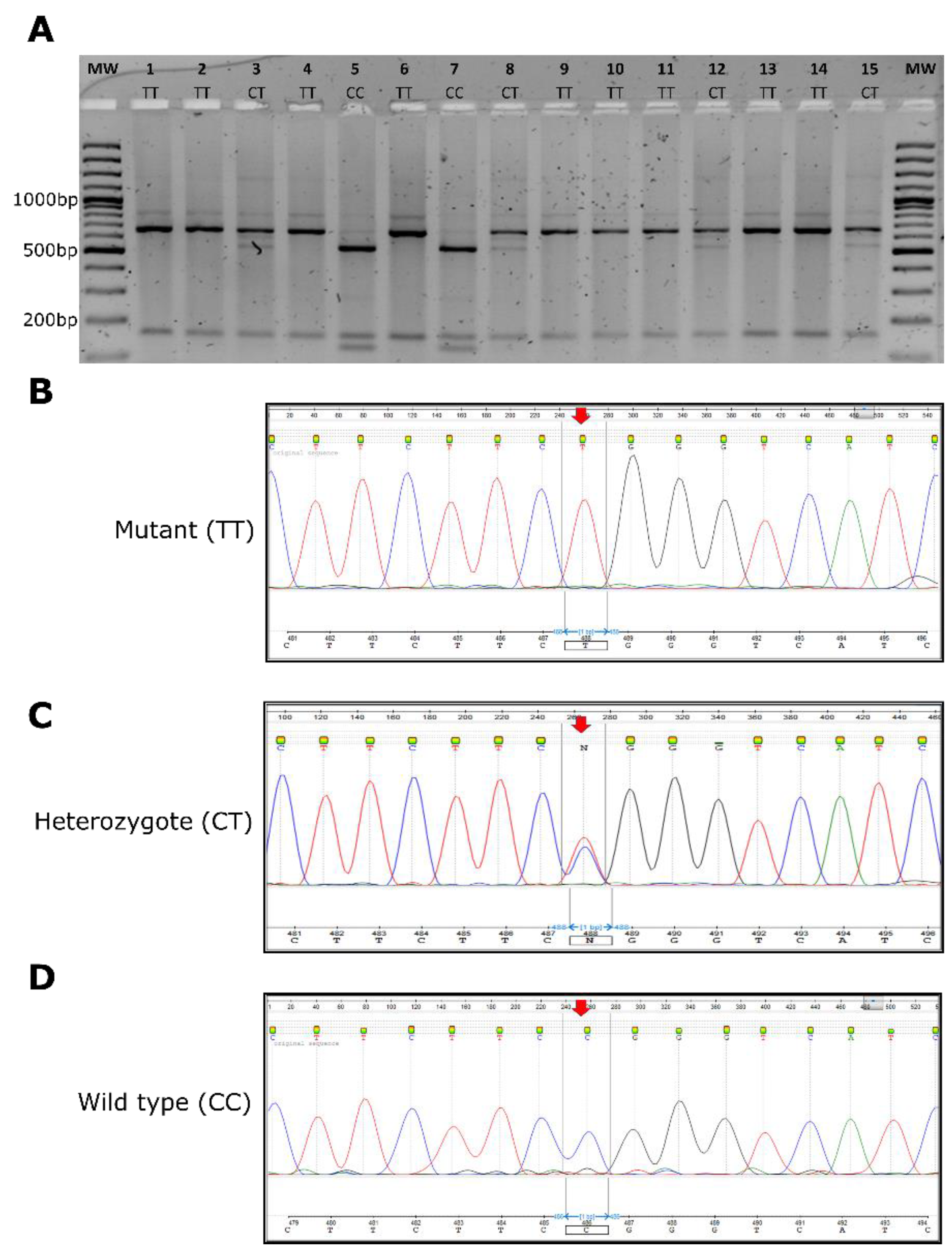
3.3. GJB2-p.Arg143Trp NciI-Restriction Fragment Polymorphism Investigations
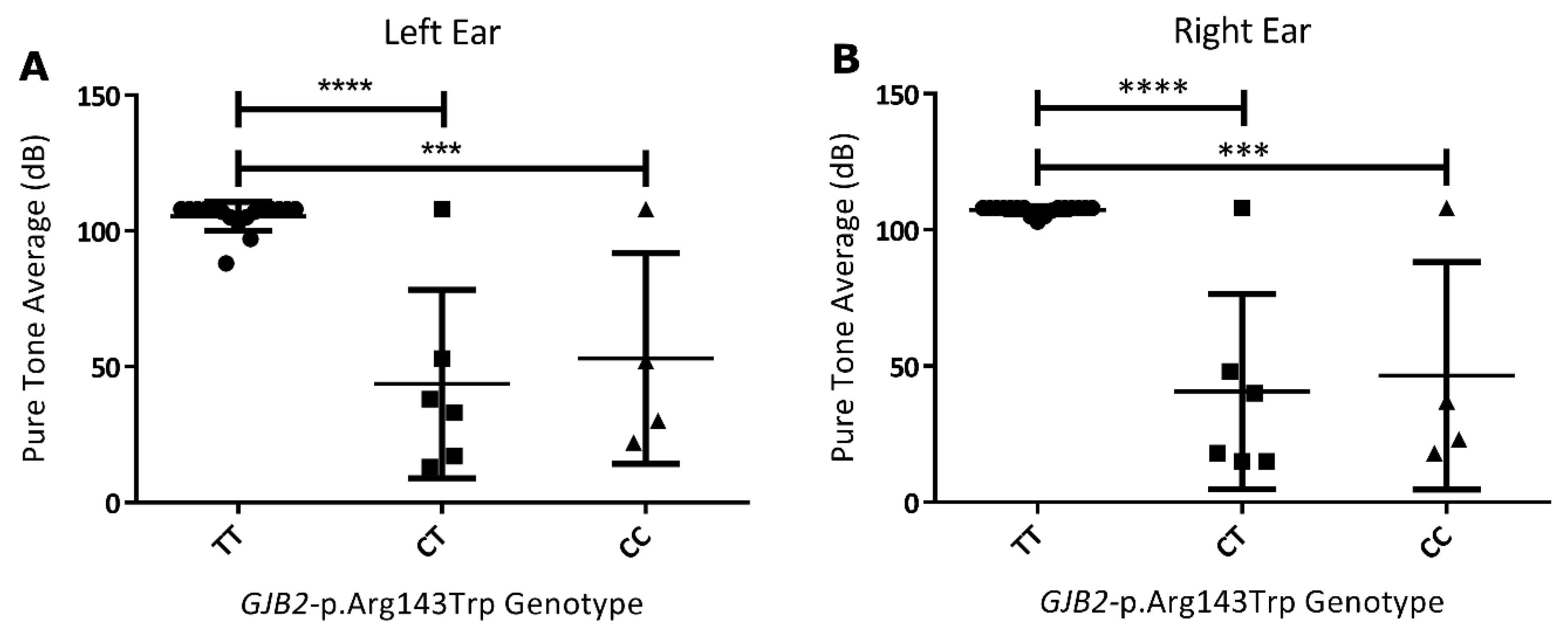
3.4. Genotype to Phenotype Correlations
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rudman, J.R.; Kabahuma, R.I.; Bressler, S.E.; Feng, Y.; Blanton, S.H.; Yan, D.; Liu, X.-Z. The genetic basis of deafness in populations of African descent. J. Genet. Genom. 2017, 44, 285–294. [Google Scholar] [CrossRef] [PubMed]
- WHO. Deafness and Hearing Loss. Available online: https://www.who.int/news-room/fact-sheets/detail/deafness-and-hearing-loss (accessed on 10 August 2019).
- Van Camp, G.; Smith, R. Hereditary Hearing Loss Homepage. Available online: https://hereditaryhearingloss.org/ (accessed on 9 September 2019).
- Ahmad, S.; Tang, W.; Chang, Q.; Qu, Y.; Hibshman, J.; Li, Y.; Söhl, G.; Willecke, K.; Chen, P.; Lin, X. Restoration of connexin26 protein level in the cochlea completely rescues hearing in a mouse model of human connexin30-linked deafness. Proc. Natl. Acad. Sci. USA 2007, 104, 1337–1341. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Paris, J.; Schrijver, I. The digenic hypothesis unraveled: The GJB6 del(GJB6-D13S1830) mutation causes allele-specific loss of GJB2 expression in cis. Biochem. Biophys. Res. Commun. 2009, 389, 354–359. [Google Scholar] [CrossRef] [PubMed]
- Distefano, M.T.; On Behalf of the ClinGen Hearing Loss Clinical Domain Working Group; Hemphill, S.E.; Oza, A.M.; Siegert, R.K.; Grant, A.R.; Hughes, M.Y.; Cushman, B.J.; Azaiez, H.; Booth, K.T.; et al. ClinGen expert clinical validity curation of 164 hearing loss gene–disease pairs. Genet. Med. 2019, 21, 2239–2247. [Google Scholar] [CrossRef] [PubMed]
- Wonkam, A. Letter to the editor regarding “GJB2, GJB6 or GJA1 genes should not be investigated in routine in non syndromic deafness in people of sub-Saharan African descent”. Int. J. Pediatr. Otorhinolaryngol. 2015, 79, 632–633. [Google Scholar] [CrossRef]
- Wonkam, A.; Bosch, J.; Noubiap, J.J.N.; Lebeko, K.; Makubalo, N.; Dandara, C. No evidence for clinical utility in investigating the connexin genes GJB2, GJB6 and GJA1 in non-syndromic hearing loss in black Africans. S. Afr. Med. J. 2015, 105, 23–26. [Google Scholar] [CrossRef]
- Gazzaz, B.; Weil, M.; Raïs, L.; Akhyat, O.; Azeddoug, H.; Nadifi, S. Autosomal recessive and sporadic deafness in Morocco: High frequency of the 35delG GJB2 mutation and absence of the 342-kb GJB6 variant. Hear. Res. 2005, 210, 80–84. [Google Scholar] [CrossRef]
- Ratbi, I.; Hajji, S.; Ouldim, K.; Aboussair, N.; Feldmann, D.; Sefiani, A. The mutation 35delG of the gene of the connexin 26 is a frequent cause of autosomal-recessive non-syndromic hearing loss in Morocco. Arch. Pediatr. 2007, 14, 450–453. [Google Scholar] [CrossRef]
- Gasmelseed, N.M.; Schmidt, M.; Magzoub, M.M.; Macharia, M.; Elmustafa, O.M.; Ototo, B.; Winkler, E.; Ruge, G.; Horstmann, R.D.; Meyer, C.G. Low frequency of deafness-associatedGJB2 variants in Kenya and Sudan and novelGJB2 variants. Hum. Mutat. 2004, 23, 206–207. [Google Scholar] [CrossRef]
- Adadey, S.M.; Manyisa, N.; Mnika, K.; De Kock, C.; Nembaware, V.; Quaye, O.; Amedofu, G.K.; Awandare, G.A.; Wonkam, A. GJB2 and GJB6 Mutations in Non-Syndromic Childhood Hearing Impairment in Ghana. Front. Genet. 2019, 10, 841. [Google Scholar] [CrossRef]
- Brobby, G.W.; Horstmann, R.D.; Müller-Myhsok, B. Connexin 26 R143W Mutation Associated with Recessive Nonsyndromic Sensorineural Deafness in Africa. N. Engl. J. Med. 1998, 338, 548–550. [Google Scholar] [CrossRef] [PubMed]
- Hamelmann, C.; Amedofu, G.K.; Albrecht, K.; Muntau, B.; Gelhaus, A.; Brobby, G.W.; Horstmann, R.D. Pattern of connexin 26 (GJB2) mutations causing sensorineural hearing impairment in Ghana. Hum. Mutat. 2001, 18, 84–85. [Google Scholar] [CrossRef] [PubMed]
- Nyst, V.A.S. A Descriptive Analysis of Adamorobe Sign Language (Ghana); LOT: Utrecht, The Netherlands, 2007. [Google Scholar]
- Kusters, A. Being a deaf white anthropologist in Adamorobe: Some ethical and methodological issues. In Village Sign Languages: Anthropological and Linguistic Insights; Ishara Press & Mouton: UK, 2012; pp. 27–52. Available online: https://www.degruyter.com/downloadpdf/books/9781614511496/9781614511496.27/9781614511496.27.pdf (accessed on 22 January 2020).
- de Freitas Cordeiro-Silva, M.; Barbosa, A.; Santiago, M.; Provetti, M.; Dettogni, R.S.; Tovar, T.T.; Rabbi-Bortolini, E.; Louro, I.D. Mutation analysis of GJB2 and GJB6 genes in Southeastern Brazilians with hereditary nonsyndromic deafness. Mol. Biol. Rep. 2011, 38, 1309–1313. [Google Scholar] [CrossRef] [PubMed]
- Sloan-Heggen, C.M.; Bierer, A.O.; Shearer, A.E.; Kolbe, D.L.; Nishimura, C.J.; Frees, K.L.; Ephraim, S.S.; Shibata, S.B.; Booth, K.T.; Campbell, C.A.; et al. Comprehensive genetic testing in the clinical evaluation of 1119 patients with hearing loss. Qual. Life Res. 2016, 135, 441–450. [Google Scholar] [CrossRef] [PubMed]
- Shearer, A.E.; DeLuca, A.P.; Hildebrand, M.S.; Taylor, K.R.; Gurrola, J.; Scherer, S.; Scheetz, T.E.; Smith, R.J.H. Comprehensive genetic testing for hereditary hearing loss using massively parallel sequencing. Proc. Natl. Acad. Sci. USA 2010, 107, 21104–21109. [Google Scholar] [CrossRef]
- Gu, X.; Guo, L.; Ji, H.; Sun, S.; Chai, R.; Wang, L.; Li, H. Genetic testing for sporadic hearing loss using targeted massively parallel sequencing identifies 10 novel mutations. Clin. Genet. 2015, 87, 588–593. [Google Scholar] [CrossRef]
- Tayoun, A.N.A.; Mason-Suares, H.; Frisella, A.L.; Bowser, M.; Duffy, E.; Mahanta, L.; Funke, B.; Rehm, H.L.; Amr, S.S. Targeted droplet-digital PCR as a tool for novel deletion discovery at the DFNB1 locus. Hum. Mutat. 2016, 37, 119–126. [Google Scholar] [CrossRef]
- Schade, G.; Kothe, C.; Ruge, G.; Hess, M.; Meyer, C.G. Non-invasive screening for GJB2 mutations in buccal smears for the diagnosis of inherited hearing impairment. Laryngo-rhino-Otol. 2003, 82, 397–401. [Google Scholar] [CrossRef]
- Schrauwen, I.; Sommen, M.; Corneveaux, J.J.; Reiman, R.A.; Hackett, N.J.; Claes, C.; Claes, K.; Bitner-Glindzicz, M.; Coucke, P.; Van Camp, G. A sensitive and specific diagnostic test for hearing loss using a microdroplet PCR-based approach and next generation sequencing. Am. J. Med. Genet. Part A 2013, 161, 145–152. [Google Scholar] [CrossRef]
- Antoniadi, T.; Pampanos, A.; Petersen, M.B. Prenatal diagnosis of prelingual deafness: Carrier testing and prenatal diagnosis of the common GJB2 35delG mutation. Prenat. Diagn. 2001, 21, 10–13. [Google Scholar] [CrossRef]
- Lucotte, G.; Bathelier, C.; Champenois, T. PCR test for diagnosis of the common GJB2 (connexin 26) 35delG mutation on dried blood spots and determination of the carrier frequency in France. Mol. Cell. Probes 2001, 15, 57–59. [Google Scholar] [CrossRef] [PubMed]
- BIAP. Classification Audiométrique Des Déficiences Auditives. Available online: http://www.biap.org/index.php?option=com_content&view=article&id=5%3Arecommandation-biap-021-bis&catid=65%3Act-2-classification-des-surdites&Itemid=19&lang=en (accessed on 25 February 2019).
- Wonkam, A.; Noubiap, J.J.N.; Djomou, F.; Fieggen, K.; Njock, R.; Toure, G.B. Aetiology of childhood hearing loss in Cameroon (sub-Saharan Africa). Eur. J. Med. Genet. 2013, 56, 20–25. [Google Scholar] [CrossRef] [PubMed]
- Bosch, J.; Noubiap, J.J.N.; Dandara, C.; Makubalo, N.; Wright, G.; Entfellner, J.-B.D.; Tiffin, N.; Wonkam, A. Sequencing of GJB2 in Cameroonians and Black South Africans and comparison to 1000 Genomes Project Data Support Need to Revise Strategy for Discovery of Nonsyndromic Deafness Genes in Africans. OMICS 2014, 18, 705–710. [Google Scholar] [CrossRef] [PubMed]
- Del Castillo, I.; Villamar, M.; Moreno-Pelayo, M.A.; Del Castillo, F.J.; Álvarez, A.; Tellería, D.; Menendez, I.; Moreno, F. A Deletion Involving the Connexin 30 Gene in Nonsyndromic Hearing Impairment. N. Engl. J. Med. 2002, 346, 243–249. [Google Scholar] [CrossRef] [PubMed]
- Baratloo, A.; Hosseini, M.; Negida, A.; El Ashal, G. Part 1: Simple Definition and Calculation of Accuracy, Sensitivity and Specificity. Arch. Acad. Emerg. Med. 2015, 3, 48–49. [Google Scholar]
- Okonechnikov, K.G.O.; Fursov, M. The_UGENE_Team Unipro UGENE: A Unified Bioinformatics Toolkit, Version 33. Bioinformatics 2012, 28, 1166–1167. [Google Scholar] [CrossRef]
- Gao, X.; Dai, P. Impact of next-generation sequencing on molecular diagnosis of inherited non-syndromic hearing loss. J. Otol. 2014, 9, 122–125. [Google Scholar] [CrossRef]
- Lebeko, K.; Bosch, J.; Noubiap, J.J.N.; Dandara, C.; Wonkam, A. Genetics of hearing loss in Africans: Use of next generation sequencing is the best way forward. Pan Afr. Med. J. 2015, 20, 383. [Google Scholar] [CrossRef]
- Lebeko, K.; Sloan-Heggen, C.M.; Noubiap, J.J.N.; Dandara, C.; Kolbe, D.L.; Ephraim, S.S.; Booth, K.T.; Azaiez, H.; Santos-Cortez, R.L.P.; Leal, S.M.; et al. Targeted genomic enrichment and massively parallel sequencing identifies novel nonsyndromic hearing impairment pathogenic variants in Cameroonian families. Clin. Genet. 2016, 90, 288–290. [Google Scholar] [CrossRef]
- Pinto, Y.M.; Elliott, P.M.; Arbustini, E.; Adler, Y.; Anastasakis, A.; Böhm, M.; Duboc, D.; Gimeno, J.; De Groote, P.; Imazio, M.; et al. Proposal for a revised definition of dilated cardiomyopathy, hypokinetic non-dilated cardiomyopathy, and its implications for clinical practice: A position statement of the ESC working group on myocardial and pericardial diseases. Eur. Heart J. 2016, 37, 1850–1858. [Google Scholar] [CrossRef]
- Calistri, A.; Palù, G. Editorial Commentary: Unbiased Next-Generation Sequencing and New Pathogen Discovery: Undeniable Advantages and Still-Existing Drawbacks; Oxford University Press: Oxford, UK, 2015. [Google Scholar]
- Abe, S.; Nishio, S.-Y.; Yokota, Y.; Moteki, H.; Kumakawa, K.; Usami, S.-I. Diagnostic pitfalls for GJB2-related hearing loss: A novel deletion detected by Array-CGH analysis in a Japanese patient with congenital profound hearing loss. Clin. Case Rep. 2018, 6, 2111–2116. [Google Scholar] [CrossRef] [PubMed]
- Brown, K.K.; Rehm, H.L. Molecular Diagnosis of Hearing Loss. Curr. Protoc. Hum. Genet. 2012. [Google Scholar] [CrossRef] [PubMed]
- Yan, D.; Xiang, G.X.; Chai, X.P.; Qing, J.; Shang, H.Q.; Zou, B.; Mittal, R.; Shen, J.; Smith, R.J.H.; Fan, Y.S.; et al. Screening of deafness-causing DNA variants that are common in patients of European ancestry using a microarray-based approach. PLoS ONE 2017, 12, e0169219. [Google Scholar] [CrossRef] [PubMed]
- Šimundić, A.-M. Measures of diagnostic accuracy: Basic definitions. Ejifcc 2009, 19, 203–211. [Google Scholar] [PubMed]
| Familial Cases from Adamorobe | ||||
| Sanger Sequencing | ||||
| Genotype | TT | CT | CC | |
| GJB2-p.Arg143Trp NciI-RFLP | TT | 12 | 0 | 0 |
| CT | 0 | 6 | 0 | |
| CC | 0 | 0 | 2 | |
| Nation-Wide Isolated/Non-Familial Cases | ||||
| Sanger Sequencing | ||||
| Genotype | TT | CT | CC | |
| GJB2-p.Arg143Trp NciI-RFLP | TT | 6 | 0 | 0 |
| CT | 0 | 1 | 0 | |
| CC | 0 | 0 | 104 | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
M. Adadey, S.; Tingang Wonkam, E.; Twumasi Aboagye, E.; Quansah, D.; Asante-Poku, A.; Quaye, O.; K. Amedofu, G.; A. Awandare, G.; Wonkam, A. Enhancing Genetic Medicine: Rapid and Cost-Effective Molecular Diagnosis for a GJB2 Founder Mutation for Hearing Impairment in Ghana. Genes 2020, 11, 132. https://doi.org/10.3390/genes11020132
M. Adadey S, Tingang Wonkam E, Twumasi Aboagye E, Quansah D, Asante-Poku A, Quaye O, K. Amedofu G, A. Awandare G, Wonkam A. Enhancing Genetic Medicine: Rapid and Cost-Effective Molecular Diagnosis for a GJB2 Founder Mutation for Hearing Impairment in Ghana. Genes. 2020; 11(2):132. https://doi.org/10.3390/genes11020132
Chicago/Turabian StyleM. Adadey, Samuel, Edmond Tingang Wonkam, Elvis Twumasi Aboagye, Darius Quansah, Adwoa Asante-Poku, Osbourne Quaye, Geoffrey K. Amedofu, Gordon A. Awandare, and Ambroise Wonkam. 2020. "Enhancing Genetic Medicine: Rapid and Cost-Effective Molecular Diagnosis for a GJB2 Founder Mutation for Hearing Impairment in Ghana" Genes 11, no. 2: 132. https://doi.org/10.3390/genes11020132
APA StyleM. Adadey, S., Tingang Wonkam, E., Twumasi Aboagye, E., Quansah, D., Asante-Poku, A., Quaye, O., K. Amedofu, G., A. Awandare, G., & Wonkam, A. (2020). Enhancing Genetic Medicine: Rapid and Cost-Effective Molecular Diagnosis for a GJB2 Founder Mutation for Hearing Impairment in Ghana. Genes, 11(2), 132. https://doi.org/10.3390/genes11020132





