MicroRNAs in Vitis vinifera cv. Chardonnay Are Differentially Expressed in Response to Diaporthe Species
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Fungal Isolates
2.3. Plant Inoculation
2.4. RNA Extraction and Quality Control
2.5. Library Preparation and Sequencing
2.6. Bioinformatics and Data Evaluation
2.7. miRNA Target Prediction and Functional Annotation
2.8. Validation of miRNA Expression Profiles by Real-Time RT-qPCR
3. Results
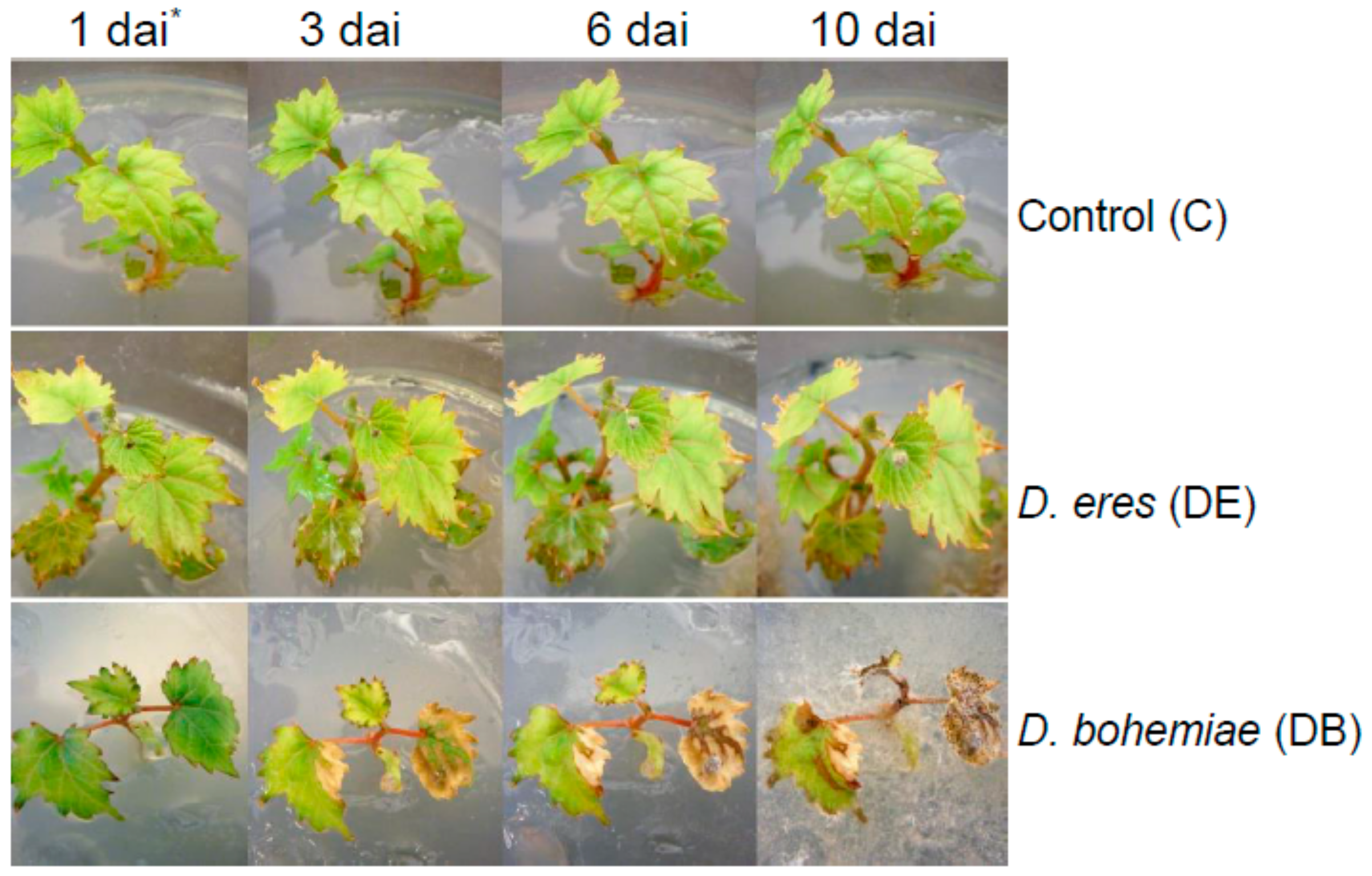
3.1. Plant Inoculation
3.2. The Abundance of sRNAs in Grapevines in Vitro
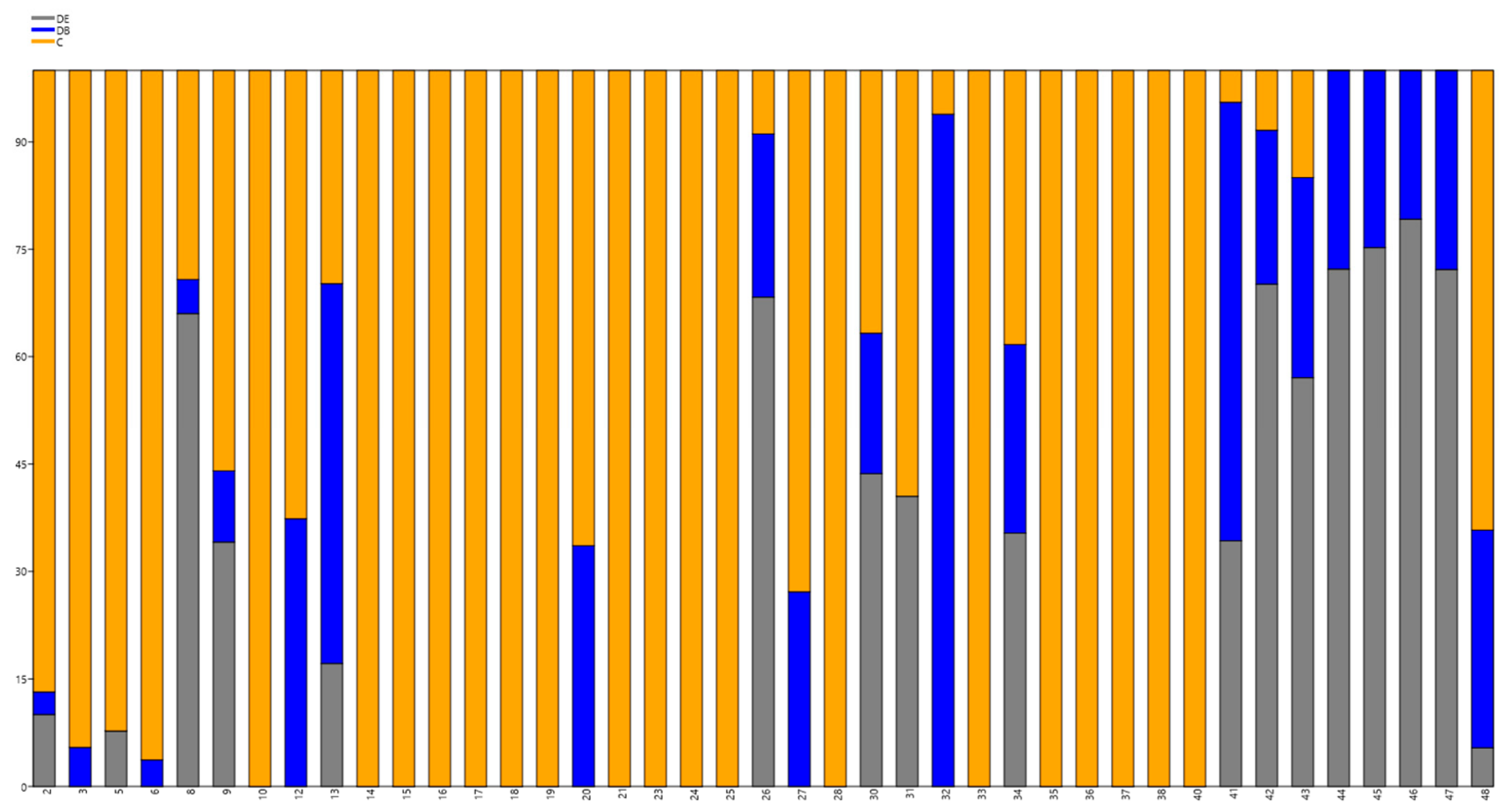
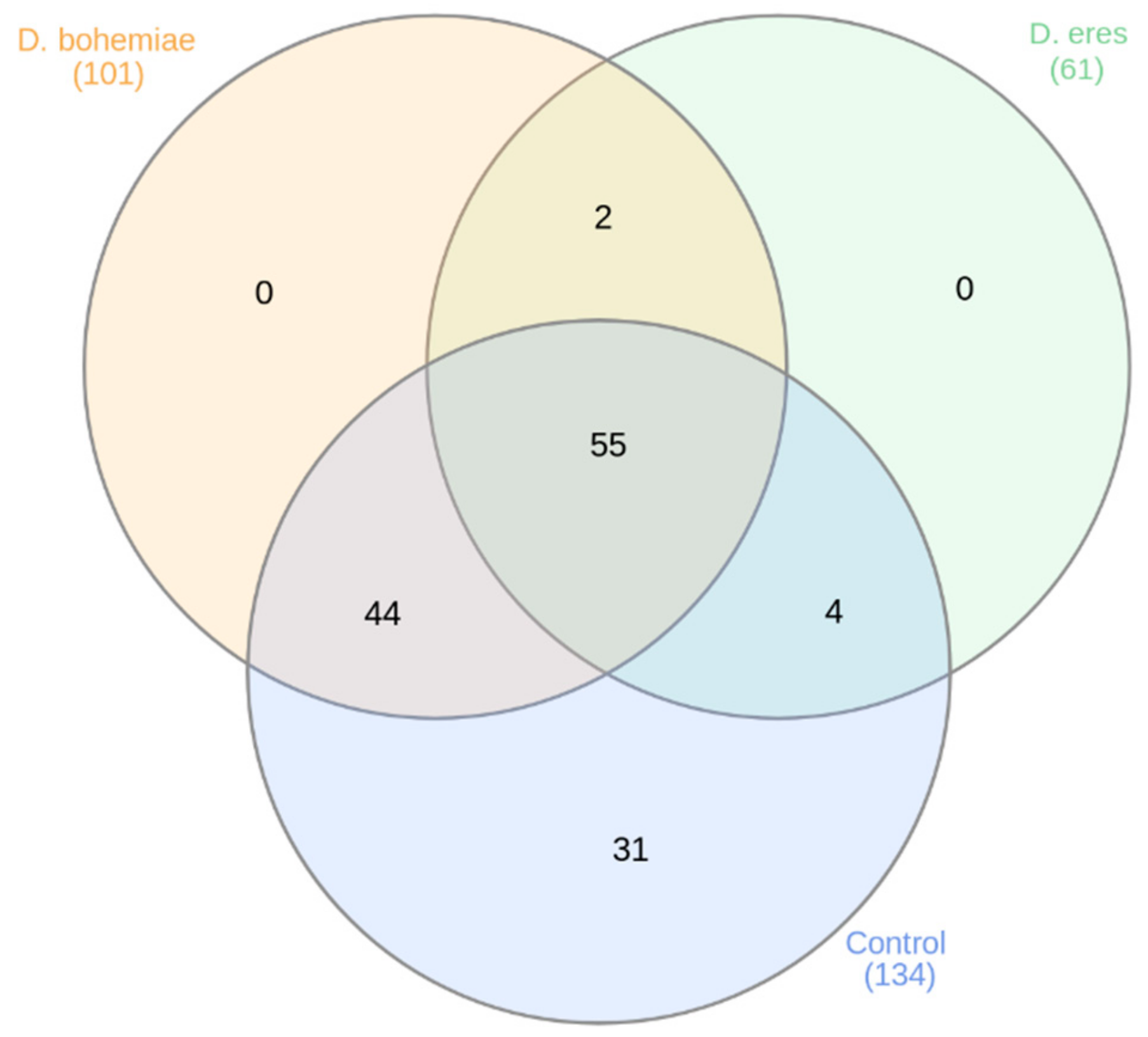
3.3. New and Conserved miRNAs Identified in Grapevine cv. Chardonnay in Vitro
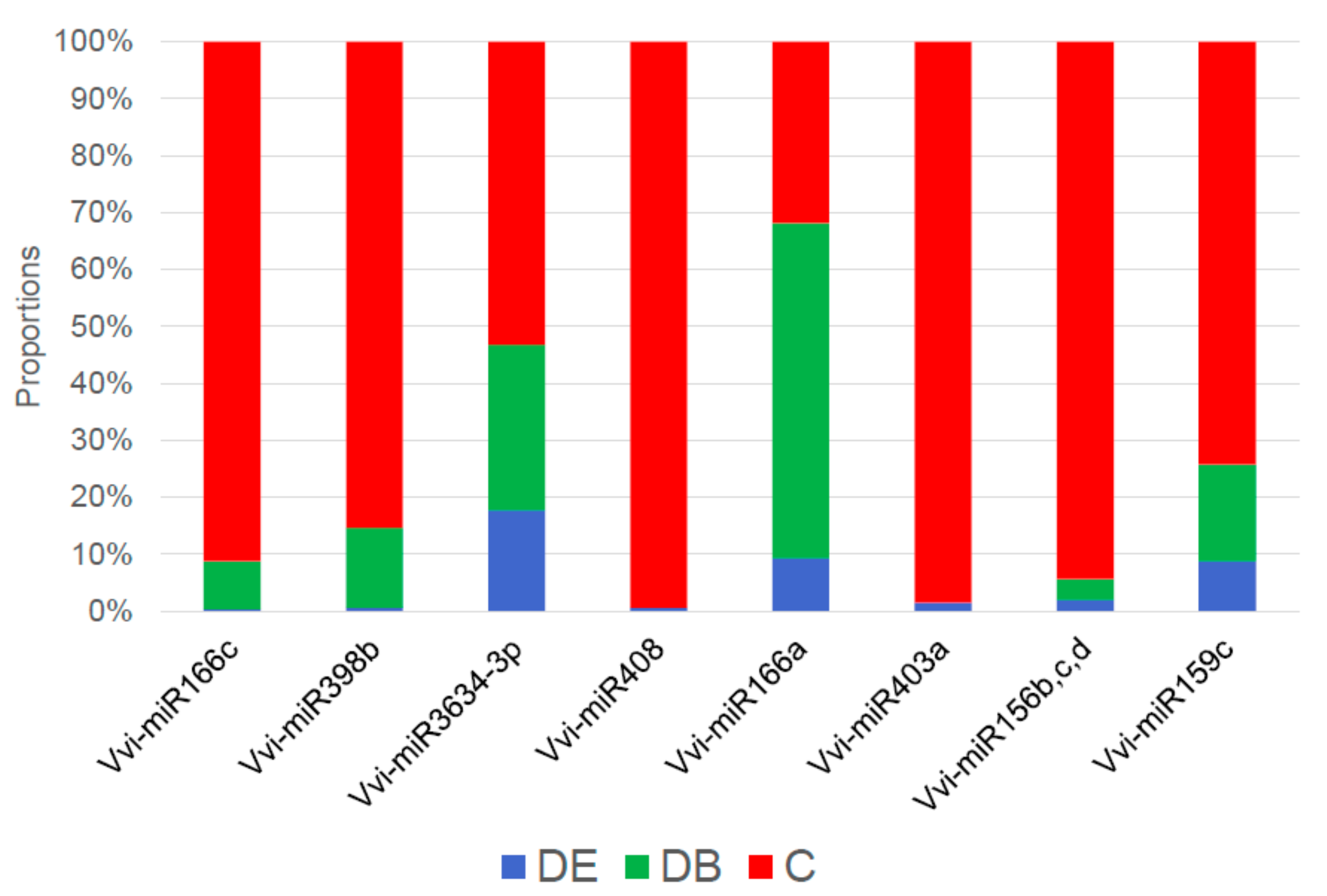
3.4. miRNA Expression Profiles by Real-Time RT-qPCR
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Muralli, T.S.; Suryanarayanan, T.S.; Geeta, R. Endophytic Phomopsis species: Host range and implications for diversity estimates. Can. J. Microbiol. 2006, 52, 673–680. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Reyne, A.; López-Medrano, F.; Morales, J.M.; García Esteban, C.; Martín, I.; Eraña, I.; Meije, Y.; Lalueza, A.; Alastruey-Izquierdo, A.; Rodríguez-Tudela, J.L.; et al. Cutaneous infection by Phomopsis longicolla in a renal transplant recipient from Guinea: First report of human infection by this fungus. Transpl. Infect. Dis. 2011, 13, 204–207. [Google Scholar] [CrossRef] [PubMed]
- Santos, J.M.; Vrandecčicć, K.; CĆosicć, J.; Duvnjak, T.; Phillips, A. Resolving the Diaporthe species occurring on soybean in Croatia. Persoonia 2011, 27, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Guarnaccia, V.; Groenewald, J.Z.; Woodhall, J.; Armengol, J.; Cinelli, T.; Eichmeier, A.; Ezra, D.; Fontaine, F.; Gramaje, D.; Gutierrez-Aguirregabiria, A.; et al. Diaporthe diversity and pathogenicity revealed from a broad survey of grapevine diseases in Europe. Pers. Mol. Phylogeny Evol. Fungi 2018, 40, 135–153. [Google Scholar] [CrossRef] [PubMed]
- Gramaje, D.; Úrbez-Torres, J.R.; Sosnowski, M.R. Managing grapevine trunk diseases with respect to etiology and epidemiology: Current strategies and future prospects. Plant Dis. 2018, 102, 12–39. [Google Scholar] [CrossRef]
- Van Rensburg, J.C.J.; Lamprecht, S.C.; Groenewald, J.Z.; Castlebury, L.A.; Crous, P.W.; Van Rensburg, J.C.J. Characterization of Phomopsis spp. associated with die-back of rooibos (Aspalathus linearis) in South Africa. Stud. Mycol. 2006, 55, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Santos, J.M.; Phillips, A.J.L. Resolving the complex of Diaporthe (Phomopsis) species occurring on Foeniculum vulgare in Portugal. Fungal Divers. 2009, 34, 111–125. [Google Scholar]
- Crous, P.; Groenewald, J.; Shivas, R.; Edwards, J.; Seifert, K.; Alfenas, A.; Alfenas, R.; Burgess, T.; Carnegie, A.; Hardy, G.; et al. Fungal Planet description sheets: 69–91. Persoonia 2011, 26, 108–156. [Google Scholar] [CrossRef]
- Crous, P.W.; Summerell, B.A.; Swart, L.; Denman, S.; Taylor, J.E.; Bezuidenhout, C.M.; Palm, M.E.; Marincowitz, S.; Groenewald, J.Z. Fungal pathogens of Proteaceae. Persoonia 2011, 27, 20–45. [Google Scholar] [CrossRef]
- Thompson, S.; Tan, Y.; Young, A.; Neate, S.; Aitken, E.; Shivas, R. Stem cankers on sunflower (Helianthus annuus) in Australia reveal a complex of pathogenic Diaporthe (Phomopsis) species. Persoonia 2011, 27, 80–89. [Google Scholar] [CrossRef]
- Gramaje, D.; Agustí-Brisach, C.; Pérez-Sierra, A.; Moralejo, E.; Olmo, D.; Mostert, L.; Damm, U.; Armengol, J. Fungal trunk pathogens associated with wood decay of almond trees on Mallorca (Spain). Pers. Mol. Phylogeny Evol. Fungi 2012, 28, 1. [Google Scholar] [CrossRef] [PubMed]
- Grasso, F.M.; Marini, M.; Vitale, A.; Firrao, G.; Granata, G. Canker and dieback on Platanus × acerifolia caused by Diaporthe scabra. For. Pathol. 2012, 42, 510–513. [Google Scholar] [CrossRef]
- Huang, F.; Hou, X.; Dewdney, M.M.; Fu, Y.; Chen, G.; Hyde, K.D.; Li, H. Diaporthe species occurring on citrus in China. Fungal Diversity 2013, 61, 237–250. [Google Scholar] [CrossRef]
- Lombard, L.; Van Leeuwen, G.C.M.; Guarnaccia, V.; Polizzi, Z.; Rijswick, P.C.J.V.; Rosendahl, K.C.H.M.V.; Gabler, J.; Crous, P.W. Diaporthe species associated with Vaccinium, with specific reference to Europe. Phytopathol. Mediterr. 2014, 53, 287–299. [Google Scholar] [CrossRef]
- Gao, Y.H.; Liu, F.; Cai, L. Unravelling Diaporthe species associated with Camellia. Syst. Biodivers. 2016, 14, 102–117. [Google Scholar] [CrossRef]
- Udayanga, D.; Castlebury, L.A.; Rossman, A.Y.; Chukeatirote, E.; Hyde, K.D. The Diaporthe sojae species complex: Phylogenetic re-assessment of pathogens associated with soybean, cucurbits and other field crops. Fungal Biol. 2015, 119, 383–407. [Google Scholar] [CrossRef] [PubMed]
- Guarnaccia, V.; Vitale, A.; Cirvilleri, G.; Aiello, D.; Susca, A.; Epifani, F.; Perrone, G.; Polizzi, G. Characterisation and pathogenicity of fungal species associated with branch cankers and stem-end rot of avocado in Italy. Eur. J. Plant Pathol. 2016, 146, 963–976. [Google Scholar] [CrossRef]
- Guarnaccia, V.; Crous, P.W. Emerging citrus diseases in Europe caused by Diaporthe spp. IMA Fungus 8 2017, 317–334. [Google Scholar] [CrossRef]
- Mostert, L.; Crous, P.W.; Kang, J.-C.; Phillips, A.J.L. Species of Phomopsis and a Libertella sp. occurring on grapevines with specific reference to South Africa: Morphological, cultural, molecular and pathological characterization. Mycologia 2001, 93, 146–167. [Google Scholar] [CrossRef]
- Mostert, L.; Kang, J.C.; Crous, P.W.; Denman, S. Phomopsis saccharata sp. nov., causing a canker and die-back disease of Protea repens in South Africa. Sydowia 2001, 53, 227–235. [Google Scholar]
- Udayanga, D.; Liu, X.; McKenzie, E.H.C.; Chukeatirote, E.; Bahkali, A.H.A.; Hyde, K.D. The genus Phomopsis: Biology, applications, species concepts and names of common phytopathogens. Fungal Divers. 2011, 50, 189–225. [Google Scholar] [CrossRef]
- Santos, L.; Alves, A.; Alves, R. Evaluating multi-locus phylogenies for species boundaries determination in the genus Diaporthe. PeerJ 2017, 5, e3120. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Fan, X.-L.; Guarnaccia, V.; Tian, C.-M. High diversity of Diaporthe species associated with dieback diseases in China, with twelve new species described. MycoKeys 2018, 39, 97–149. [Google Scholar] [CrossRef] [PubMed]
- Van Niekerk, J.M.; Groenewald, J.Z.; Farr, D.F.; Fourie, P.H.; Halleen, F.; Crous, P.W.; Halleer, F. Reassessment of Phomopsis species on grapevines. Australas. Plant Pathol. 2005, 34, 27–39. [Google Scholar] [CrossRef]
- Gomes, R.R.; Glienke, C.; Videira, S.; Lombard, L.; Groenewald, J.Z.; Crous, P.W. Diaporthe: A genus of endophytic, saprobic and plant pathogenic fungi. Persoonia 2013, 31, 1–41. [Google Scholar] [CrossRef]
- Úrbez-Torres, J.R.; Peduto, F.; Smith, R.J.; Gubler, W.D. Phomopsis dieback: A grapevine trunk disease caused by Phomopsis viticola in California. Plant Dis. 2013, 97, 1571–1579. [Google Scholar] [CrossRef]
- Dissanayake, A.J.; Liu, M.; Zhang, W.; Chen, Z.; Udayanga, D.; Chukeatirote, E.; Li, X.; Yan, J.; Hyde, K.D. Morphological and molecular characterisation of Diaporthe species associated with grapevine trunk disease in China. Fungal Biol. 2015, 119, 283–294. [Google Scholar] [CrossRef] [PubMed]
- Calvo, A.M.; Wilson, R.A.; Bok, J.W.; Keller, N.P. Relationship between secondary metabolism and fungal development. Microbiol. Mol. Biol. Rev. 2002, 66, 447–459. [Google Scholar] [CrossRef]
- Chaves, M.M.; Zarrouk, O.; Francisco, R.; Costa, J.M.; Santos, T.; Regalado, A.P.; Rodrigues, M.L.; Lopes, C.M. Grapevine under deficit irrigation: Hints from physiological and molecular data. Ann. Bot. 2010, 105, 661–676. [Google Scholar] [CrossRef]
- Abdelrahman, M.; Suzumura, N.; Mitoma, M.; Matsuo, S.; Ikeuchi, T.; Mori, M.; Murakami, K.; Ozaki, Y.; Matsumoto, M.; Uragami, A.; et al. Comparative de novo transcriptome profiles in Asparagus officinali s and A. kiusianus during the early stage of Phomopsis asparagi infection. Sci. Rep. 2017, 7, 2608. [Google Scholar] [CrossRef]
- Zhou, J.; Li, X.; Chen, Y.; Dai, C. De novo transcriptome assembly of Phomopsis liquidambari provides insights into genes associated with different lifestyles in rice (Oryza sativa L.). Front. Plant Sci. 2017, 8, 121. [Google Scholar] [CrossRef] [PubMed]
- Abdelrahman, M.; Mitoma, M.; Ikeuchi, T.; Mori, M.; Murakami, K.; Ozaki, Y.; Matsumoto, M.; Uragami, A.; Kanno, A. Differential gene expression analysis and SNP/InDel marker discovery in resistant wild Asparagus kiusianus and susceptible A. officinalis in response to Phomopsis asparagi infection. Data Brief 2018, 21, 2117–2121. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Ferrer, V.; Voinnet, O. Roles of plant small RNAs in biotic stress responses. Annu. Rev. Plant Biol. 2009, 60, 485–510. [Google Scholar] [CrossRef] [PubMed]
- Sunkar, R.; Li, Y.F.; Jagadeeswaran, G. Functions of microRNAs in plant stress responses. Trends Plant Sci. 2012, 17, 196–203. [Google Scholar] [CrossRef]
- Voinnet, O. Origin, biogenesis, and activity of plant microRNAs. Cell 2009, 136, 669–687. [Google Scholar] [CrossRef]
- Allen, E.; Xie, Z.; Gustafson, A.M.; Carrington, J.C. microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 2005, 121, 207–221. [Google Scholar] [CrossRef]
- Vaucheret, H. Post-transcriptional small RNA pathways in plants: Mechanisms and regulations. Genes Dev. 2006, 20, 759–771. [Google Scholar] [CrossRef]
- Budak, H.; Akpinar, B.A. Plant miRNAs: Biogenesis, organization and origins. Funct. Integr. Genom. 2015, 15, 523–531. [Google Scholar] [CrossRef]
- Eichmeier, A.; Komínková, M.; Komínek, P.; Baránek, M. Comprehensive virus detection using next generation sequencing in grapevine vascular tissues of plants obtained from the wine regions of Bohemia and Moravia (Czech Republic). PLoS ONE 2016, 11, e0167966. [Google Scholar] [CrossRef]
- Baldrich, P.; Campo, S.; Wu, M.T.; Liu, T.T.; Hsing, Y.I.C.; Segundo, B.S. MicroRNA-mediated regulation of gene expression in the response of rice plants to fungal elicitors. RNA Biol. 2015, 12, 847–863. [Google Scholar] [CrossRef]
- Soto-Suárez, M.; Baldrich, P.; Weigel, D.; Rubio-Somoza, I.; San Segundo, B. The Arabidopsis miR396 mediates pathogen-associated molecular pattern-triggered immune responses against fungal pathogens. Sci. Rep. 2017, 7, 44898. [Google Scholar] [CrossRef] [PubMed]
- Hua, C.; Zhao, J.H.; Guo, H.S. Trans-kingdom RNA silencing in plant–fungal pathogen interactions. Mol. Plant 2018, 11, 235–244. [Google Scholar] [CrossRef] [PubMed]
- The French–Italian Public Consortium for Grapevine Genome Characterization; Jaillon, O.; Aury, J.-M.; Noel, B.; Policriti, A.; Clepet, C.; Casagrande, A.; Choisne, N.; Aubourg, S.; Vitulo, N.; et al. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 2007, 449, 463. [Google Scholar] [CrossRef]
- Velasco, R.; Zharkikh, A.; Troggio, M.; Cartwright, D.A.; Cestaro, A.; Pruss, D.; Pindo, M.; Fitzgerald, L.M.; Vezzulli, S.; Reid, J.; et al. A high quality draft consensus sequence of the genome of a heterozygous grapevine variety. PLoS ONE 2007, 2, e1326. [Google Scholar] [CrossRef]
- Pantaleo, V.; Szittya, G.; Moxon, S.; Miozzi, L.; Moulton, V.; Dalmay, T.; Burgyan, J. Identification of grapevine microRNAs and their targets using high-throughput sequencing and degradome analysis. Plant J. 2010, 62, 960–976. [Google Scholar] [CrossRef]
- Mica, E.; Piccolo, V.; Delledonne, M.; Ferrarini, A.; Pezzotti, M.; Casati, C.; Del Fabbro, C.; Valle, G.; Policriti, A.; Morgante, M.; et al. High throughput approaches reveal splicing of primary microRNA transcripts and tissue specific expression of mature microRNAs in Vitis vinifera. BMC Genomics 2009, 10, 558. [Google Scholar] [CrossRef]
- Kozomara, A.; Griffiths-Jones, S. miRBase: Integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res. 2010, 39 (Suppl. 1), D152–D157. [Google Scholar] [CrossRef]
- Baránek, M.; Křižan, B.; Ondrušíková, E.; Pidra, M. DNA-methylation changes in grapevine somaclones following in vitro culture and thermotherapy. Plant Celltissue Organ Cult. 2010, 101, 11–22. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A quality control tool for high throughput sequence data. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 5 November 2019).
- Dai, X.; Zhao, P.X. psRNATarget: A plant small RNA target analysis server. Nucleic Acids Res. 2011, 39 (Suppl. 2), W155–W159. [Google Scholar] [CrossRef]
- Dai, X.; Zhao, P.X. pssRNAMiner: A plant short small RNA regulatory cascade analysis server. Nucleic Acids Res. 2008, 36 (Suppl. 2), W114–W118. [Google Scholar] [CrossRef]
- Chen, C.; Ridzon, D.A.; Broomer, A.J.; Zhou, Z.; Lee, D.H.; Nguyen, J.T.; Barbisin, M.; Xu, N.L.; Mahuvakar, V.R.; Andersen, M.R.; et al. Real-time quantification of microRNAs by stem–loop RT–PCR. Nucleic Acids Res. 2005, 33, e179. [Google Scholar] [CrossRef] [PubMed]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]
- Hellemans, J.; Mortier, G.; De Paepe, A.; Speleman, F.; Vandesompele, J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol. 2007, 8, R19. [Google Scholar] [CrossRef]
- Fan, J.; Liu, J.; Culty, M.; Papadopoulos, V. Acyl-coenzyme A binding domain containing 3 (ACBD3; PAP7; GCP60): An emerging signaling molecule. Prog. Lipid Res. 2010, 49, 218–234. [Google Scholar] [CrossRef] [PubMed]
- OIV. OIV FOCUS 2017 Distribution of the World’s Grapevine Varieties; International Organisation of Vine and Wine: Paris, France, 2017; p. 54. ISBN 979-10-91799-89-8. [Google Scholar]
- Liu, W.; Cheng, C.; Chen, F.; Ni, S.; Lin, Y.; Lai, Z. High-throughput sequencing of small RNAs revealed the diversified cold-responsive pathways during cold stress in the wild banana (Musa itinerans). BMC Plant Biol. 2018, 18, 308. [Google Scholar] [CrossRef]
- Ma, X.; Bologna, N.; Palma-Guerrero, J. Small RNA bidirectional crosstalk during the interaction between wheat and Zymoseptoria tritici. bioRxiv 2018, 501593. [Google Scholar] [CrossRef]
- Marchi, G. Susceptibility to esca of various grapevine (Vitis vinifera) cultivars grafted on different rootstocks in a vineyard in the province of Siena (Italy). Phytopathol. Mediterr. 2001, 40, 27–36. [Google Scholar] [CrossRef]
- Murolo, S.; Romanazzi, G. Effects of grapevine cultivar, rootstock and clone on esca disease. Australas. Plant Pathol. 2014, 43, 215–221. [Google Scholar] [CrossRef]
- Rajagopalan, R.; Vaucheret, H.; Trejo, J.; Bartel, D.P. A diverse and evolutionarily fluid set of microRNAs in Arabidopsis thaliana. Genes Dev. 2006, 20, 3407–3425. [Google Scholar] [CrossRef]
- Fahlgren, N.; Howell, M.D.; Kasschau, K.D.; Chapman, E.J.; Sullivan, C.M.; Cumbie, J.S.; Givan, S.A.; Law, T.F.; Grant, S.R.; Dangl, J.L.; et al. High-throughput sequencing of Arabidopsis microRNAs: Evidence for frequent birth and death of MIRNA genes. PLoS ONE 2007, 2, e219. [Google Scholar] [CrossRef] [PubMed]
- Moxon, S.; Jing, R.; Szittya, G.; Schwach, F.; Pilcher, R.L.R.; Moulton, V.; Dalmay, T. Deep sequencing of tomato short RNAs identifies microRNAs targeting genes involved in fruit ripening. Genome Res. 2008, 18, 1602–1609. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.; Johansen, L.K.; Gustafson, A.M.; Kasschau, K.D.; Lellis, A.D.; Zilberman, D.; Jacobsen, S.E.; Carrington, J.C. Genetic and functional diversification of small RNA pathways in plants. PLoS Biol. 2004, 2, e104. [Google Scholar] [CrossRef] [PubMed]
- Chan, S.W.L.; Henderson, I.R.; Jacobsen, S.E. Gardening the genome: DNA methylation in Arabidopsis thaliana. Nat. Rev. Genet. 2005, 6, 351. [Google Scholar] [CrossRef]
- Pantaleo, V.; Vitali, M.; Boccacci, P.; Miozzi, L.; Cuozzo, D.; Chitarra, W.; Mannini, F.; Lovisolo, C.; Gambino, G. Novel functional microRNAs from virus-free and infected Vitis vinifera plants under water stress. Sci. Rep. 2016, 6, 20167. [Google Scholar] [CrossRef]
- Raabe, C.A.; Tang, T.H.; Brosius, J.; Rozhdestvensky, T.S. Biases in small RNA deep sequencing data. Nucleic Acids Res. 2013, 42, 1414–1426. [Google Scholar] [CrossRef]
- Baev, V.; Milev, I.; Naydenov, M.; Apostolova, E.; Minkov, G.; Minkov, I.; Yahubyan, G. Implementation of a de novo genome-wide computational approach for updating Brachypodium miRNAs. Genomics 2011, 97, 282–293. [Google Scholar] [CrossRef]
- Axtell, M.J.; Snyder, J.A.; Bartel, D.P. Common functions for diverse small RNAs of land plants. Plant Cell 2007, 19, 1750–1769. [Google Scholar] [CrossRef]
- Bittner-Eddy, P.D.; Beynon, J.L. The Arabidopsis downy mildew resistance gene, RPP13-Nd, functions independently of NDR1 and EDS1 and does not require the accumulation of salicylic acid. Mol. Plant Microbe Interact. 2007, 14, 416–421. [Google Scholar] [CrossRef]
- Bhattarai, K.; Wang, W.; Cao, Z.; Deng, Z. Comparative analysis of impatiens leaf transcriptomes reveal candidate genes for resistance to downy mildew caused by Plasmopara obducens. Int. J. Mol. Sci. 2018, 19, 2057. [Google Scholar] [CrossRef]
- Rhoades, M.W.; Reinhart, B.J.; Lim, L.P.; Burge, C.B.; Bartel, B.; Bartel, D.P. Prediction of plant microRNA targets. Cell 2002, 110, 513–520. [Google Scholar] [CrossRef]
- Zhong, R.; Ye, Z.H. Regulation of HD-ZIP III genes by microRNA 165. Plant Signal. Behav. 2007, 2, 351–353. [Google Scholar] [CrossRef] [PubMed]
- Du, Q.; Wang, H. The role of HD-ZIP III transcription factors and miR165/166 in vascular development and secondary cell wall formation. Plant Signal. Behav. 2015, 10, e1078955. [Google Scholar] [CrossRef] [PubMed]
- Ohashi-Ito, K.; Kubo, M.; Demura, T.; Fukuda, H. Class III homeodomain leucine-zipper proteins regulate xylem cell differentiation. Plant Cell Physiol. 2005, 46, 1646–1656. [Google Scholar] [CrossRef] [PubMed]
- Prigge, M.J.; Otsuga, D.; Alonso, J.M.; Ecker, J.R.; Drews, G.N.; Clark, S.E. Class III homeodomain-leucine zipper gene family members have overlapping, antagonistic, and distinct roles in Arabidopsis development. Plant Cell 2005, 17, 61–76. [Google Scholar] [CrossRef]
- Jung, J.H.; Park, C.M. MIR166/165 genes exhibit dynamic expression patterns in regulating shoot apical meristem and floral development in Arabidopsis. Planta 2007, 225, 1327–1338. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Jung, J.-H.; Reyes, J.L.; Kim, Y.-S.; Kim, S.-Y.; Chung, K.-S.; Kim, J.A.; Lee, M.; Lee, Y.; Kim, V.N.; et al. microRNA-directed cleavage of ATHB15 mRNA regulates vascular development in Arabidopsis inflorescence stems. Plant J. 2005, 42, 84–94. [Google Scholar] [CrossRef] [PubMed]
- Dugas, D.V.; Bartel, B. Sucrose induction of Arabidopsis miR398 represses two Cu/Zn superoxide dismutases. Plant Mol. Biol. 2008, 67, 403–417. [Google Scholar] [CrossRef]
- Chitarra, W.; Pagliarani, C.; Abbà, S.; Boccacci, P.; Birello, G.; Rossi, M.; Palmano, S.; Marzachì, C.; Perrone, I.; Gambino, G. miRVIT: A novel miRNA database and its application to uncover vitis responses to flavescence dorée infection. Front. Plant Sci. 2018, 9, 1034. [Google Scholar] [CrossRef]
- Abdel-Ghany, S.E.; Pilon, M. MicroRNA-mediated systemic down-regulation of copper protein expression in response to low copper availability in Arabidopsis. J. Biol. Chem. 2008, 283, 15932–15945. [Google Scholar] [CrossRef]
- Zhang, X.; Zhao, H.; Gao, S.; Wang, W.-C.; Katiyar-Agarwal, S.; Huang, H.-D.; Raikhel, N.; Jin, H. Arabidopsis Argonaute 2 regulates innate immunity via miRNA393∗-mediated silencing of a Golgi-localized SNARE gene, MEMB12. Mol. Cell 2011, 42, 356–366. [Google Scholar] [CrossRef]
- Zeng, R.F.; Zhou, J.J.; Liu, S.R.; Gan, Z.M.; Zhang, J.Z.; Hu, C.G. Genome-Wide Identification and Characterization of SQUAMOSA—Promoter-Binding Protein (SBP) Genes Involved in the Flowering Development of Citrus Clementina. Biomolecules 2019, 9, 66. [Google Scholar] [CrossRef] [PubMed]
- Weigel, D.; Alvarez, J.; Smyth, D.R.; Yanofsky, M.F.; Meyerowitz, E.M. LEAFY controls floral meristem identity in Arabidopsis. Cell 1992, 69, 843–859. [Google Scholar] [CrossRef]
- Bettiga, L.J. Grape Pest Management; University of California Agriculture and Natural Resources (UCANR): Santa Maria, CA, USA, 2013. [Google Scholar]


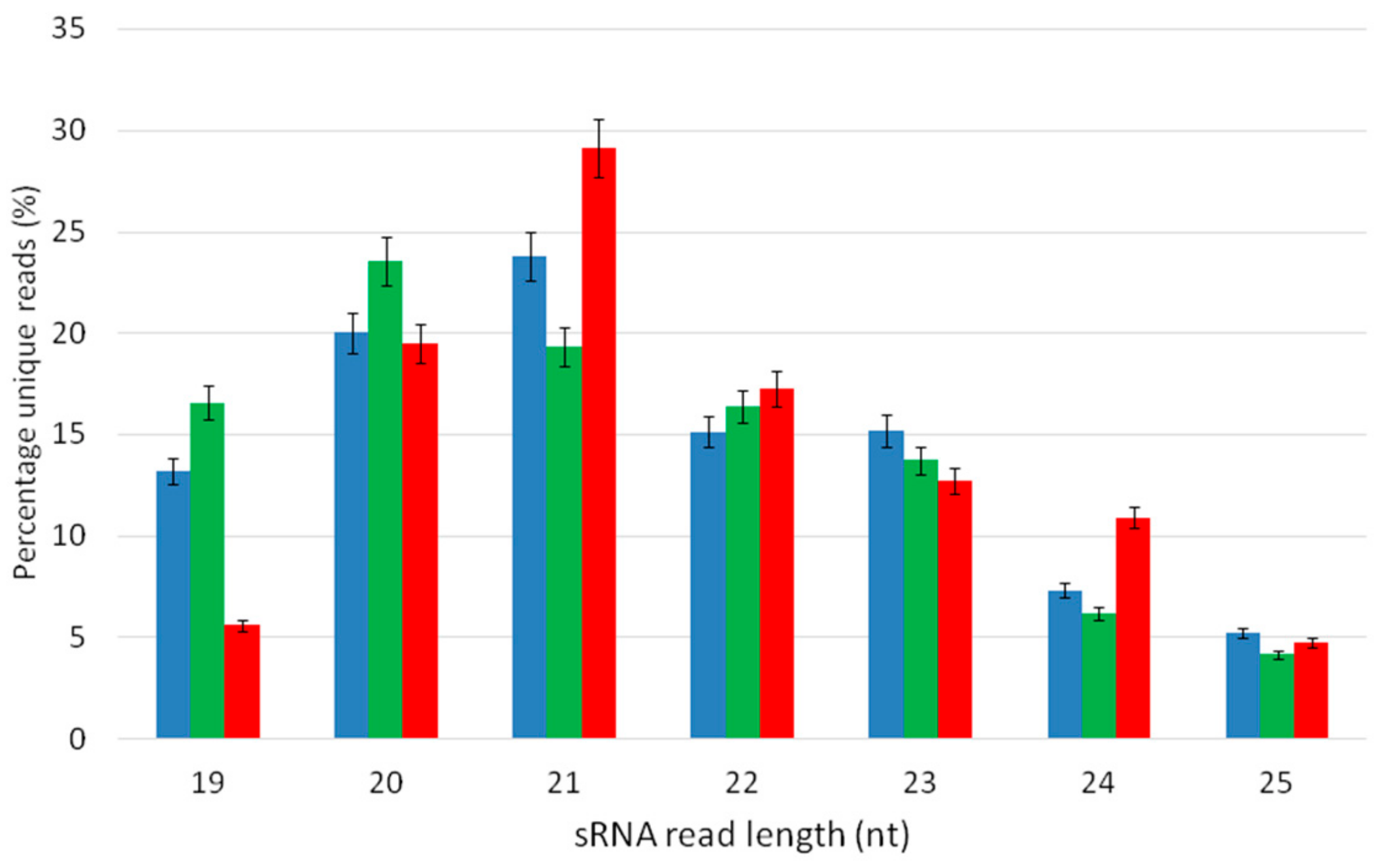
| 19 | 20 | 21 | 22 | 23 | 24 | 25 | Total | |
|---|---|---|---|---|---|---|---|---|
| DE | 287,897 | 436,557 | 518,754 | 330,042 | 331,108 | 159,806 | 113,975 | 2,178,138 |
| DB | 563,942 | 801,579 | 656,600 | 556,699 | 467,013 | 211,409 | 141,876 | 3,399,118 |
| C | 104,300 | 362,066 | 540,728 | 320,848 | 236,014 | 202,736 | 88,881 | 1,855,572 |
| miRNA Name | miRNA Sequence | Putative Target Identified Using NCBI | Target Acc. Based on the Highest Expectations (E), psRNA Target | Inhibition |
|---|---|---|---|---|
| 2 | CCCAGUCCCGAACCCGUCGGC | similar to aspartate aminotransferase; similar to Aspartate aminotransferase 2, transcript variant X9, misc_RNA, importin α isoform 9 | chr11.gff3_MRNA_VIT_11s0149g00200.t01 | Cleavage |
| 3 | AGUUACUAAUUCAUGAUCUGGC | importin α isoform 9 | chr2.gff3_MRNA_VIT_02s0033g00980.t01 | Cleavage |
| 5 | CCAGUCCCGAACCCGUCGGC | Vitis vinifera contig VV78X128415.10, whole genome shotgun sequence | chr7_random.gff3_MRNA_VIT_07s0151g00980.t01 | Cleavage |
| 6 | UCUCGGACCAGGCUUCAUUCC | Vitis vinifera microRNA MIR166a (MIR166A), microRNA, http://www.mirbase.org/cgi-bin/mature.pl?mature_acc=MIMAT0020658 | chr18.gff3_MRNA_VIT_18s0075g00480.t01 | Translocation |
| 8 | GGUGGCUGUAGUUUAGUGGU | Vitis vinifera contig VV78X038801.3, whole genome shotgun sequence | chr18.gff3_MRNA_VIT_18s0001g12770.t01 | Cleavage |
| 9 | CGGUGGACUGCUCGAGCUGC | Vitis vinifera contig VV78X196950.19, whole genome shotgun sequence | chr15.gff3_MRNA_VIT_15s0048g02810.t01 | Translocation |
| 10 | CUAACAGACCGGUAGACUUGAAC | Vitis vinifera contig VV78X130314.7, whole genome shotgun sequence | chr15.gff3_MRNA_VIT_15s0048g02810.t01 | Translation |
| 12 | CCCAGUCCCGAACCCGUCGGCU | Vitis vinifera contig VV78X156561.10, whole genome shotgun sequence | chr11.gff3_MRNA_VIT_11s0149g00200.t01 | Cleavage |
| 13 | GCGCCUGUAGCUCAGUGGA | Vitis vinifera contig VV78X046944.3, whole genome shotgun sequence | chr8.gff3_MRNA_VIT_08s0007g07620.t01 | Cleavage |
| 14 | UUCAUGGACGUUGAUAAGAUCCU | Vitis vinifera subsp. sylvestris chloroplast DNA, complete genome | chr7.gff3_MRNA_VIT_07s0005g00750.t01 | Cleavage |
| 15 | UAACAGACCGGUAGACUUGAAC | PREDICTED: Vitis vinifera pentatricopeptide repeat-containing protein At5g50990 (LOC100247459) | chr18.gff3_MRNA_VIT_18s0001g09480.t01 | Cleavage |
| 16 | UGCACUGCCUCUUCCCUGGCU | Vitis vinifera microRNA MIR408 gene, complete sequence, http://www.mirbase.org/cgi-bin/mirna_entry.pl?acc=MI0005917 | chr18.gff3_MRNA_VIT_18s0001g15240.t01 | Cleavage |
| 17 | CCUAACAGACCGGUAGACUUGAAC | PREDICTED: Vitis vinifera ATP synthase subunit α, chloroplastic-like (LOC109124299), mRNA | chr18.gff3_MRNA_VIT_18s0001g11300.t01 | Cleavage |
| 18 | UCCUAACAGACCGGUAGACUUGAAC | Vitis vinifera subsp. sylvestris chloroplast DNA, complete genome | chr18.gff3_MRNA_VIT_18s0001g11300.t01 | Cleavage |
| 19 | UCCUAACAGACCGGUAGACUUGAAC | Vitis vinifera subsp. sylvestris chloroplast DNA, complete genome | chr18.gff3_MRNA_VIT_18s0001g11300.t01 | Cleavage |
| 20 | UUAGAUGAUCAUCAACAAACU | Vitis vinifera ankyrin repeat-containing protein NPR4-like (LOC100260982), transcript variant X10, mRNA | chr7.gff3_MRNA_VIT_07s0005g02430.t01 | Cleavage |
| 21 | CAGACCGGUAGACUUGAAC | PREDICTED: Vitis vinifera ATP synthase subunit α, chloroplastic-like (LOC109124299), mRNA | chr19.gff3_MRNA_VIT_19s0090g01480.t01 | Cleavage |
| 23 | ACAGACCGGUAGACUUGAAC | PREDICTED: Vitis vinifera ATP synthase subunit α, chloroplastic-like (LOC109124299), mRNA | chr18.gff3_MRNA_VIT_18s0001g09480.t01 | Cleavage |
| 24 | UUCCACAGCUUUCUUGAACUU | Vitis vinifera microRNA MIR396b (MIR396B), microRNA | chr15.gff3_MRNA_VIT_15s0021g02580.t01 | Cleavage |
| 25 | AACAGACCGGUAGACUUGAAC | PREDICTED: Vitis vinifera ATP synthase subunit α, chloroplastic-like (LOC109124299), mRNA | chr18.gff3_MRNA_VIT_18s0001g09480.t01 | Cleavage |
| 26 | CCGGCGAUGCGCUCCUGGCC | Vitis vinifera contig VV78X197078.6, whole genome shotgun sequence | chr12.gff3_MRNA_VIT_12s0034g02480.t01 | Cleavage |
| 27 | CAGUCCCGAACCCGUCGGC | Vitis vinifera contig VV78X156561.10, whole genome shotgun sequence | chr11.gff3_MRNA_VIT_11s0149g00200.t01 | Cleavage |
| 28 | UGUUGAGCUCACCUUGUACCC | PREDICTED: Vitis vinifera kinase-interacting family protein (LOC100246194), transcript variant X1, mRNA | chr9.gff3_MRNA_VIT_09s0002g03120.t01 | Translation |
| 30 | CGGUGGACUGCUCGAGCUGCU | Vitis vinifera contig VV78X156561.10, whole genome shotgun sequence | chr15.gff3_MRNA_VIT_15s0048g02810.t01 | Translation |
| 31 | GUUGAGCUCACCUUGUACCCA | PREDICTED: Vitis vinifera kinase-interacting family protein (LOC100246194), transcript variant X1, mRNA | chr9.gff3_MRNA_VIT_09s0002g03120.t01 | Translation |
| 32 | CCCAGUCCCGAACCCGUCGG | Vitis vinifera contig VV78X128415.10, whole genome shotgun sequence, mRNA sequence acyl CoA binding protein domain containing protein 3 which is a Golgi protein involved in several signalling events | chr6.gff3_MRNA_VIT_06s0004g04740.t01 | Cleavage |
| 33 | UGAAGGUCCAAGGCCGAGGCU | PREDICTED: Vitis vinifera uncharacterized LOC100855078 (LOC100855078), ncRNA | chr14.gff3_MRNA_VIT_14s0006g03100.t01 | Cleavage |
| 34 | GGGAUGGGUCGACCGGUCC | Vitis vinifera contig VV78X071755.8, whole genome shotgun sequence | chr12.gff3_MRNA_VIT_12s0034g01520.t01 | Cleavage |
| 35 | UCGGAUAAAGGGUUAUACAUC | PREDICTED: Vitis vinifera uncharacterized LOC100853315 (LOC100853315), transcript variant X1, ncRNA | chr6.gff3_MRNA_VIT_06s0009g03800.t01 | Cleavage |
| 36 | UGCACUGCCUCUUCCCUGGC | Vitis vinifera microRNA MIR408 (MIR408), microRNA | chr18.gff3_MRNA_VIT_18s0001g15240.t01 | Cleavage |
| 37 | CUGGAUUAUGACUGAACGCCU | PREDICTED: Vitis vinifera lysosomal Pro-X carboxypeptidase (LOC100244772), transcript variant X2, mRNA | chr4.gff3_MRNA_VIT_04s0210g00160.t01 | Cleavage |
| 38 | UUCCACAGCUUUCUUGAACU | Vitis vinifera microRNA MIR396c (MIR396C), microRNA | chr15.gff3_MRNA_VIT_15s0021g02580.t01 | Cleavage |
| 40 | AGUUACUAAUUCAUGAUCUGGCC | PREDICTED: Vitis vinifera scopoletin glucosyltransferase (LOC100260498), mRNA | chr2.gff3_MRNA_VIT_02s0033g00980.t01 | Cleavage |
| 41 | CCGGCGAUGCGCUCCUGGCC | mRNA sequence with expression of RPP13-like protein 1, potential disease resistance protein | chr12.gff3_MRNA_VIT_12s0034g02480.t01 | Cleavage |
| 42 | CCGGCGAUGCGCUCCUGGCCU | PREDICTED: Vitis vinifera putative disease resistance RPP13-like protein 1 (LOC100258269), transcript variant X4, mRNA | chr12.gff3_MRNA_VIT_12s0034g02480.t01 | Cleavage |
| 43 | ACCGGCGAUGCGCUCCUGGCCU | PREDICTED: Vitis vinifera auxin efflux carrier component 3 (LOC100268124), mRNA | chr1.gff3_MRNA_VIT_01s0011g01820.t01 | Cleavage |
| 44 | GCCCGUGGAGACGUCGUCGCCUCG | PREDICTED: Vitis vinifera oxalate--CoA ligase (LOC100256632), mRNA | chr1.gff3_MRNA_VIT_01s0011g00770.t01 | Cleavage |
| 45 | CGCCGUCCGAAUUGUAGUCUGGA | PREDICTED: Vitis vinifera uncharacterized LOC109123385 (LOC109123385), mRNA | chr12.gff3_MRNA_VIT_12s0134g00450.t01 | Cleavage |
| 46 | UCGGGUUAACAUUCCUGAACCGGGA | PREDICTED: Vitis vinifera AUGMIN subunit 7 (LOC100243653), transcript variant X1, mRNA | chr1.gff3_MRNA_VIT_01s0011g01130.t01 | Cleavage |
| 47 | CGGUGGACUGCUCGAGCUGCU | PREDICTED: Vitis vinifera non-specific lipid-transfer protein-like protein At5g64080 (LOC100247017), mRNA | chr15.gff3_MRNA_VIT_15s0048g02810.t01 | Translation |
| 48 | CCAGUCCCGAACCCGUCGGC | PREDICTED: Vitis vinifera acyl-CoA-binding domain-containing protein 3 (LOC100268114), transcript variant X6, mRNA | chr7_random.gff3_MRNA_VIT_07s0151g00980.t01 | Cleavage |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Eichmeier, A.; Kiss, T.; Penazova, E.; Pecenka, J.; Berraf-Tebbal, A.; Baranek, M.; Pokluda, R.; Cechova, J.; Gramaje, D.; Grzebelus, D. MicroRNAs in Vitis vinifera cv. Chardonnay Are Differentially Expressed in Response to Diaporthe Species. Genes 2019, 10, 905. https://doi.org/10.3390/genes10110905
Eichmeier A, Kiss T, Penazova E, Pecenka J, Berraf-Tebbal A, Baranek M, Pokluda R, Cechova J, Gramaje D, Grzebelus D. MicroRNAs in Vitis vinifera cv. Chardonnay Are Differentially Expressed in Response to Diaporthe Species. Genes. 2019; 10(11):905. https://doi.org/10.3390/genes10110905
Chicago/Turabian StyleEichmeier, Ales, Tomas Kiss, Eliska Penazova, Jakub Pecenka, Akila Berraf-Tebbal, Miroslav Baranek, Robert Pokluda, Jana Cechova, David Gramaje, and Dariusz Grzebelus. 2019. "MicroRNAs in Vitis vinifera cv. Chardonnay Are Differentially Expressed in Response to Diaporthe Species" Genes 10, no. 11: 905. https://doi.org/10.3390/genes10110905
APA StyleEichmeier, A., Kiss, T., Penazova, E., Pecenka, J., Berraf-Tebbal, A., Baranek, M., Pokluda, R., Cechova, J., Gramaje, D., & Grzebelus, D. (2019). MicroRNAs in Vitis vinifera cv. Chardonnay Are Differentially Expressed in Response to Diaporthe Species. Genes, 10(11), 905. https://doi.org/10.3390/genes10110905








