Chromatin-Mediated Regulation of Genome Plasticity in Human Fungal Pathogens
Abstract
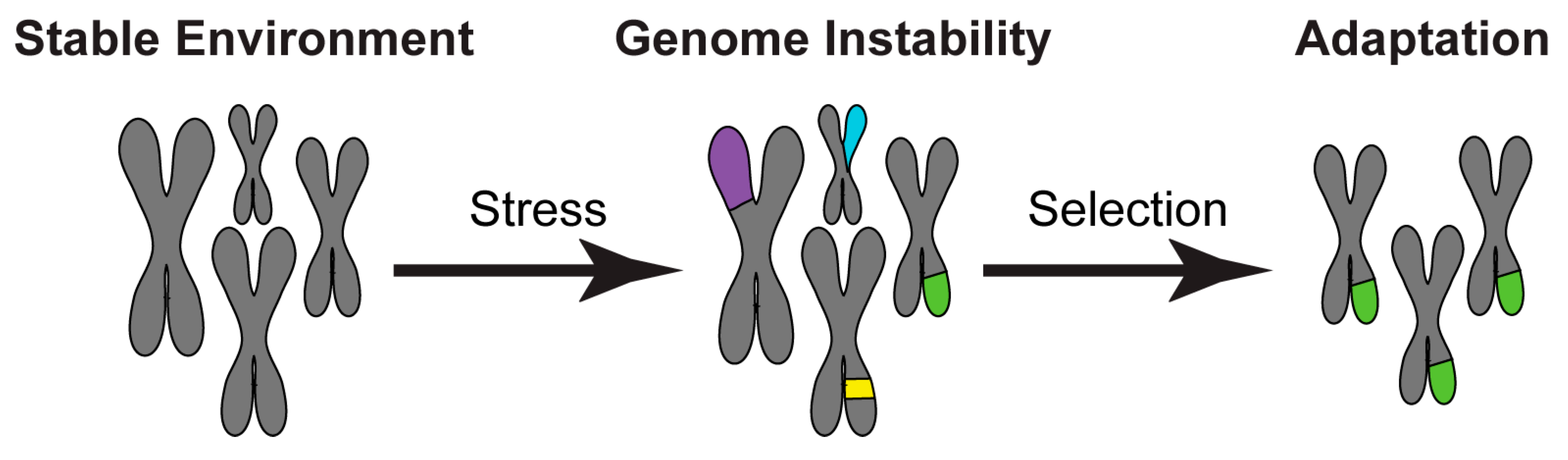
1. Introduction
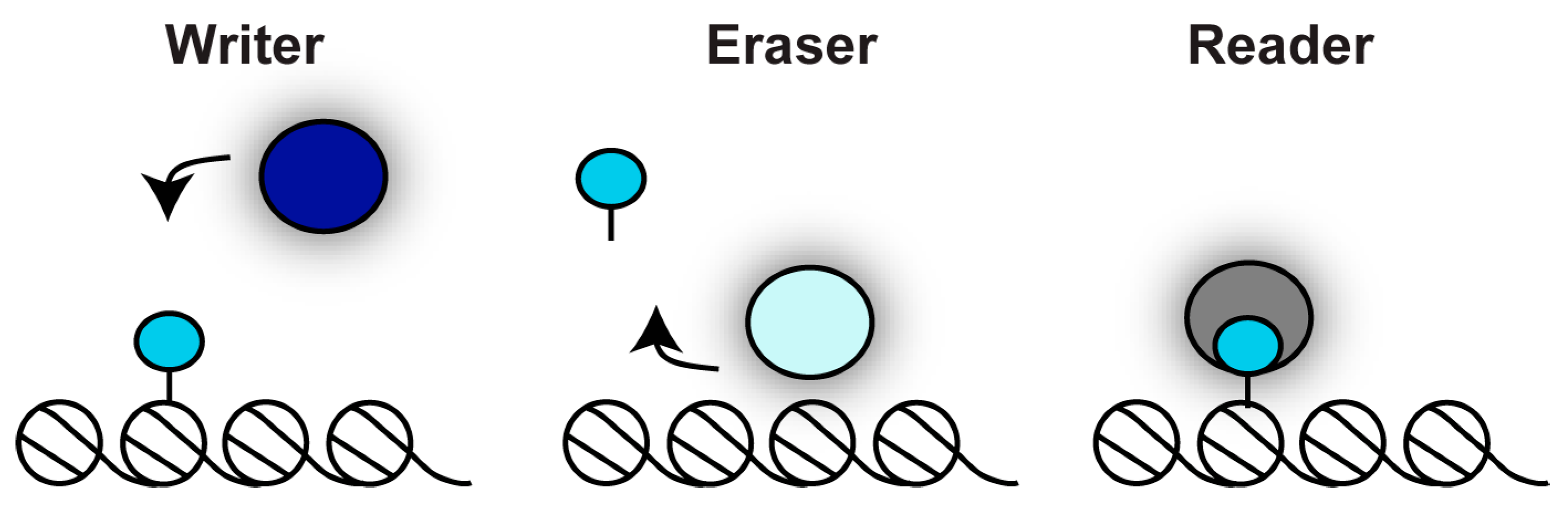
2. Regulation of Chromatin Structure by Post Translation Modification of Histone Proteins
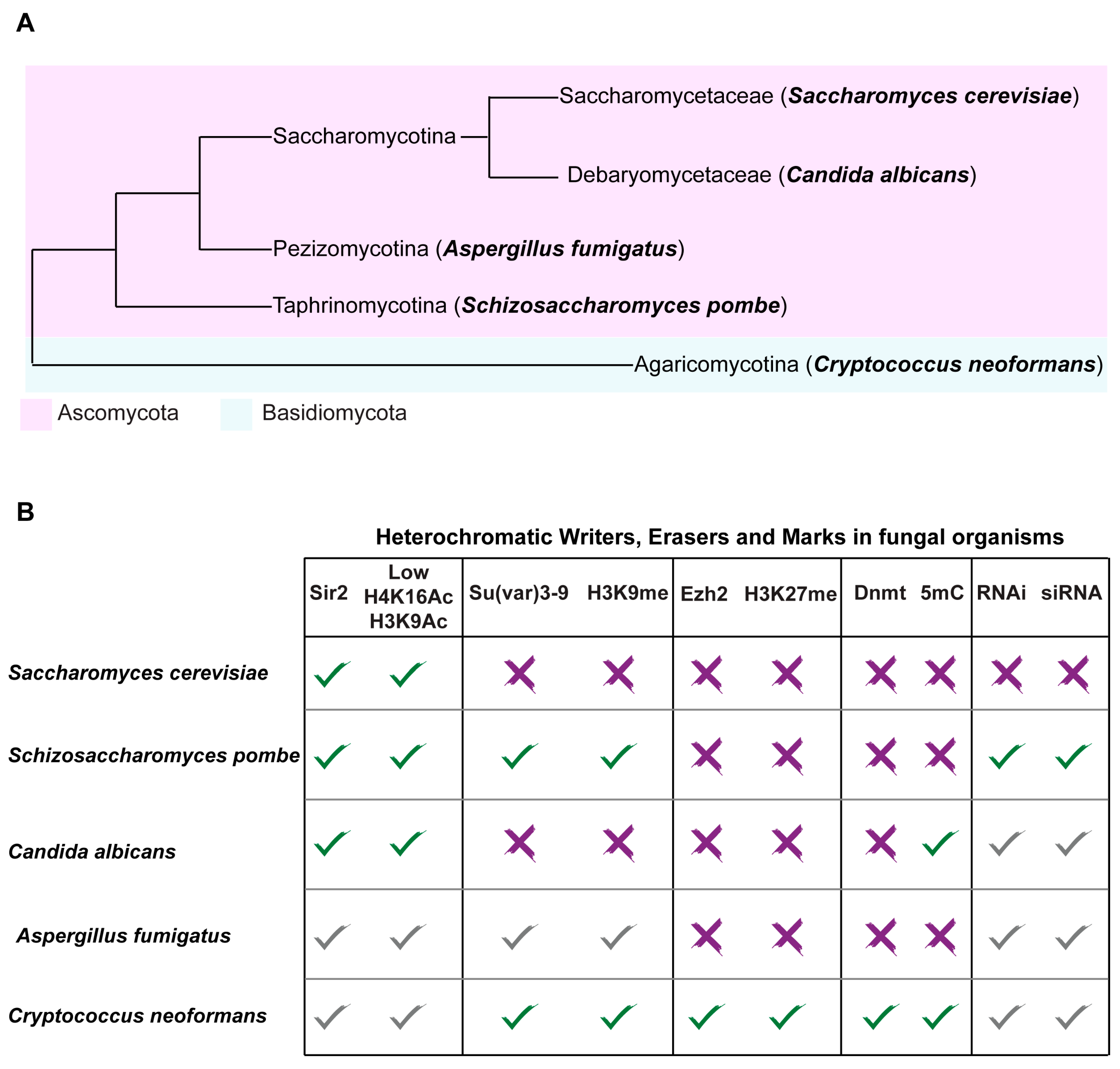
3. Fungal Model Systems as a Road Map for Understanding Heterochromatin Structure and its Function in Promoting Genome Stability
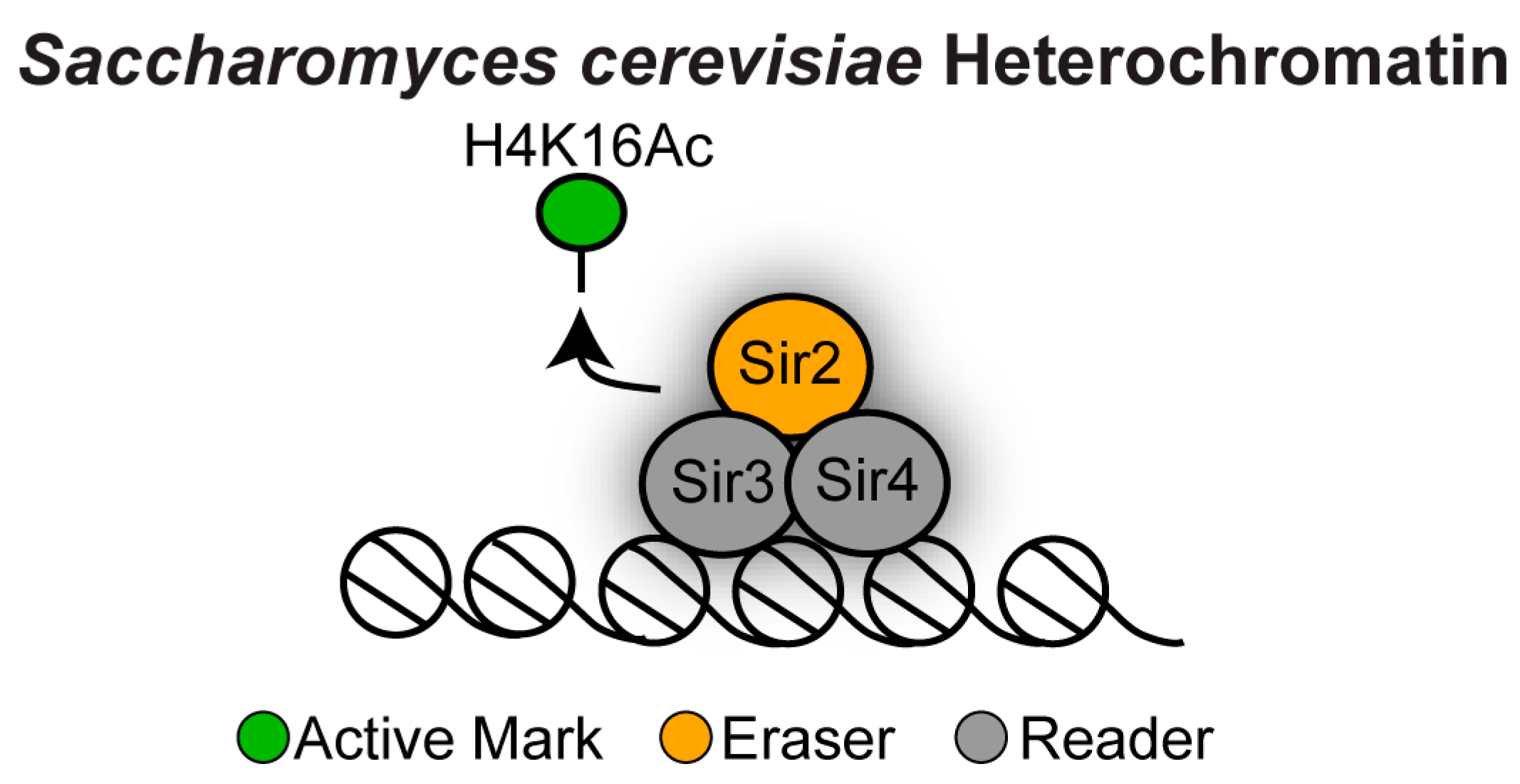
3.1. Sir2-Dependent Heterochromatin in Saccharomyces cerevisiae
3.1.1. Heterochromatin at the S. cerevisiae the MAT Locus
3.1.2. Heterochromatin at the S. cerevisiae Subtelomeres
3.1.3. Heterochromatin at the S. cerevisiae rDNA Locus
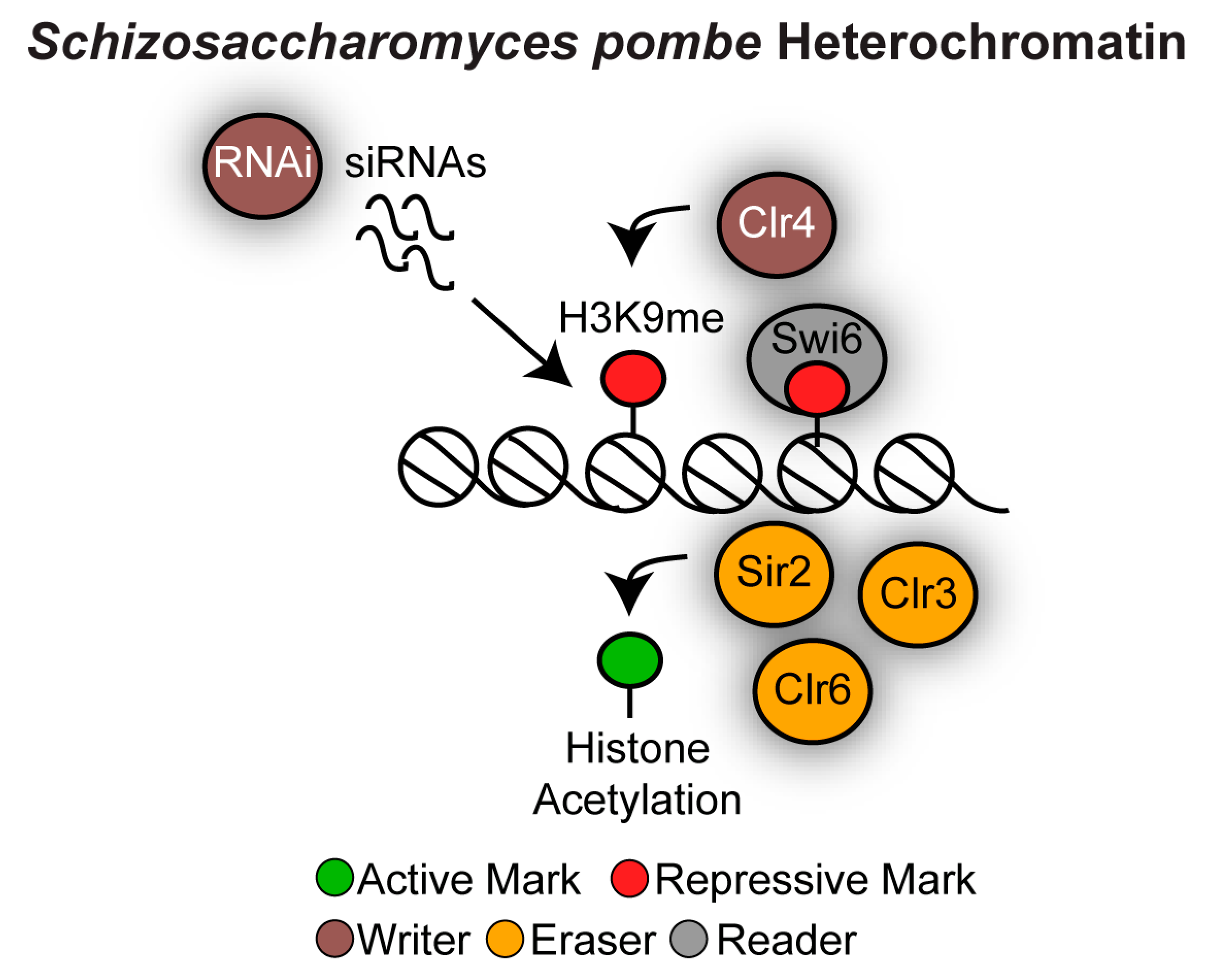
3.2. Heterochromatin in Schizosaccharomyces pombe
3.2.1. Heterochromatin at S. pombe Pericentromeres
3.2.2. S. pombe Heterochromatin at Other Genomic Locations
4. Heterochromatin Structure and Function in Human Fungal Pathogens
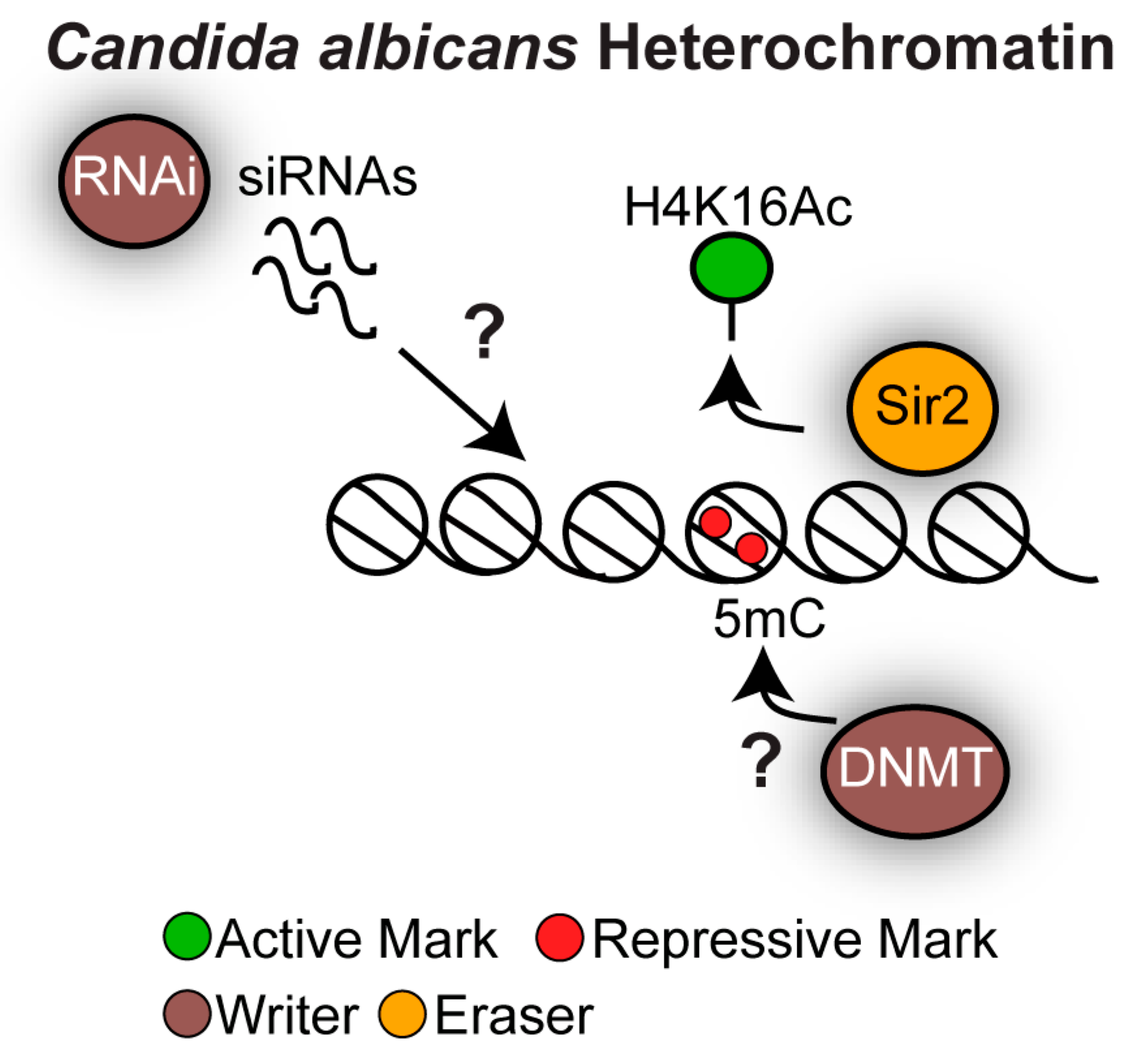
4.1. Distinct Chromatin States Mark Different Repetitive Elements in Candida albicans
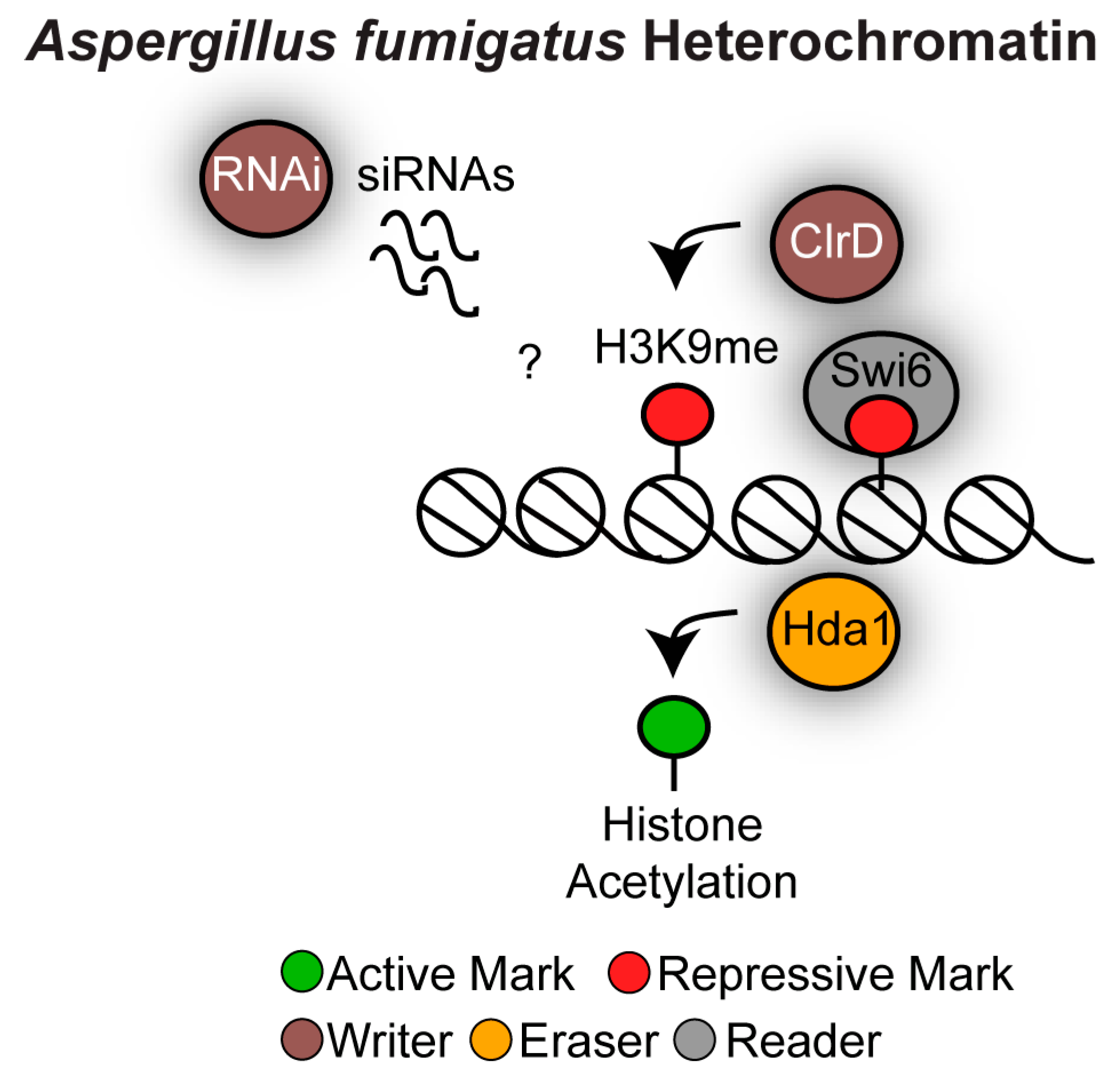
4.2. Heterochromatin as a Regulator of Secondary Metabolite Gene Clusters in Aspergillus fumigatus
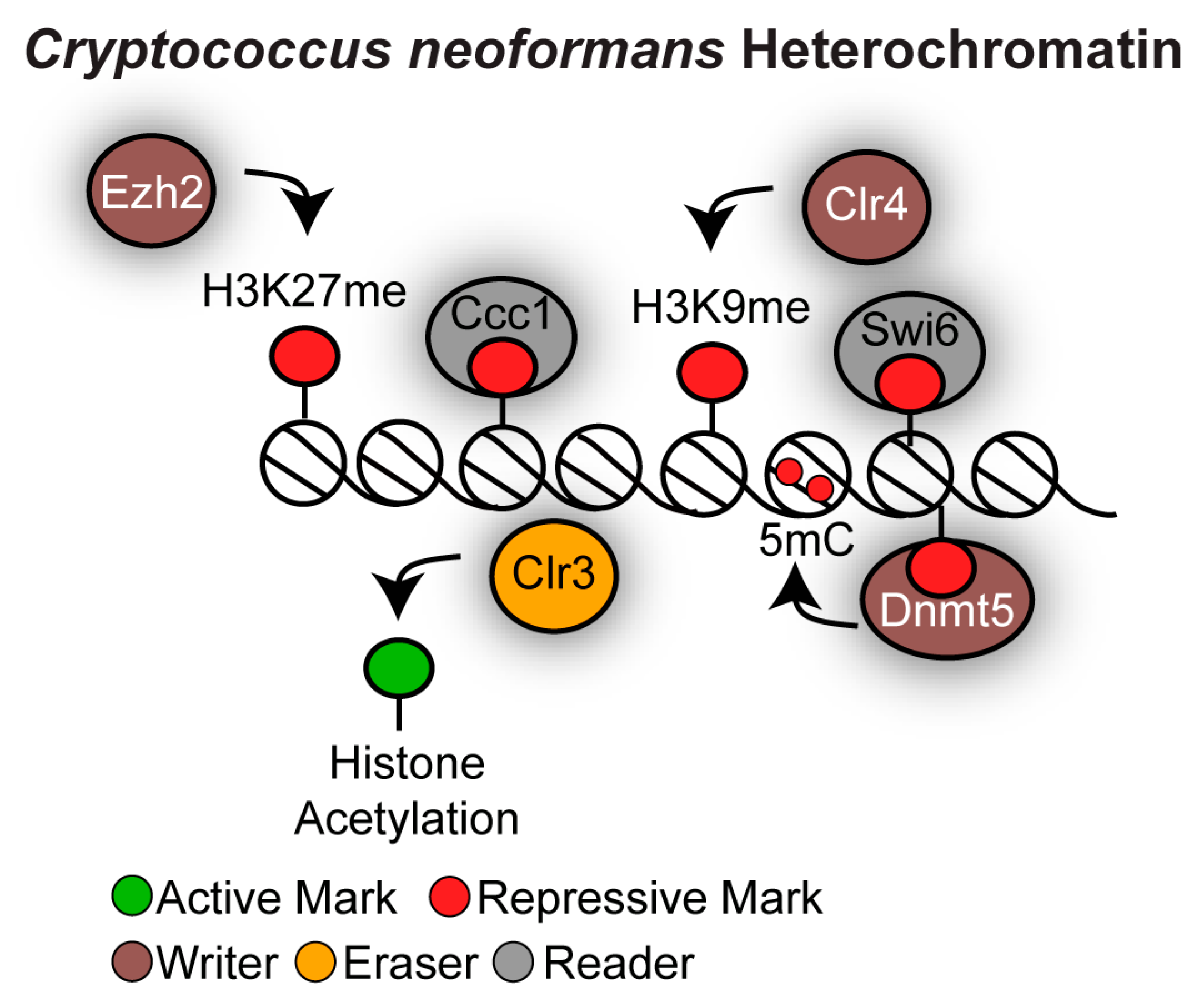
4.3. Distinct Chromatin States Associated With Cryptococcus Neoformans Repetitive Elements
5. Advancing Our Understanding of Heterochromatin Structure and Function in Human Fungal Pathogens
6. Concluding Remarks
Funding
Acknowledgments
Conflicts of Interest
References
- Brown, G.D.; Denning, D.W.; Gow, N.A.R.; Levitz, S.M.; Netea, M.G.; White, T.C. Hidden Killers: Human Fungal Infections. Sci. Transl. Med. 2012, 4, 165rv13. [Google Scholar] [CrossRef] [PubMed]
- Fisher, M.C.; Hawkins, N.J.; Sanglard, D.; Gurr, S.J. Worldwide emergence of resistance to antifungal drugs challenges human health and food security. Science 2018, 360, 739–742. [Google Scholar] [CrossRef] [PubMed]
- Erwig, L.P.; Gow, N.A.R. Interactions of fungal pathogens with phagocytes. Nat. Rev. Genet. 2016, 14, 163–176. [Google Scholar] [CrossRef] [PubMed]
- Galhardo, R.S.; Hastings, P.J.; Rosenberg, S.M. Mutation as a stress response and the regulation of evolvability. Crit. Rev. Biochem. Mol. Biol. 2007, 42, 399–435. [Google Scholar] [CrossRef] [PubMed]
- Ormerod, K.L.; Fraser, J.A. Balancing Stability and Flexibility within the Genome of the Pathogen Cryptococcus neoformans. PLoS Pathog. 2013, 9, e1003764. [Google Scholar] [CrossRef][Green Version]
- Hirakawa, M.P.; Martinez, D.A.; Sakthikumar, S.; Anderson, M.Z.; Berlin, A.; Gujja, S.; Zeng, Q.; Zisson, E.; Wang, J.M.; Greenberg, J.M.; et al. Genetic and phenotypic intra-species variation in Candida albicans. Genome Res. 2015, 25, 413–425. [Google Scholar] [CrossRef]
- Beale, M.A.; Sabiiti, W.; Robertson, E.J.; Fuentes-Cabrejo, K.M.; O’Hanlon, S.J.; Jarvis, J.N.; Loyse, A.; Meintjes, G.; Harrison, T.S.; May, R.C.; et al. Genotypic Diversity Is Associated with Clinical Outcome and Phenotype in Cryptococcal Meningitis across Southern Africa. PLoS Negl. Trop. Dis. 2015, 9, e0003847. [Google Scholar] [CrossRef]
- Selmecki, A.; Forche, A.; Berman, J. Genomic Plasticity of the Human Fungal Pathogen Candida albicans. Eukaryot. Cell 2010, 9, 991–1008. [Google Scholar] [CrossRef]
- Forche, A.; Abbey, D.; Pisithkul, T.; Weinzierl, M.A.; Ringstrom, T.; Bruck, D.; Petersen, K.; Berman, J. Stress Alters Rates and Types of Loss of Heterozygosity in Candida albicans. mBio 2011, 2, e00129-11. [Google Scholar] [CrossRef]
- Freire-Benéitez, V.; Gourlay, S.; Berman, J.; Buscaino, A. Sir2 regulates stability of repetitive domains differentially in the human fungal pathogen Candida albicans. Nucleic Acids Res. 2016, 44, 9166–9179. [Google Scholar]
- Yang, F.; Teoh, F.; Tan, A.S.M.; Cao, Y.; Pavelka, N.; Berman, J. Aneuploidy Enables Cross-Adaptation to Unrelated Drugs. Mol. Biol. Evol. 2019, 36, 1768–1782. [Google Scholar] [CrossRef] [PubMed]
- Selmecki, A.M.; Dulmage, K.; Cowen, L.E.; Anderson, J.B.; Berman, J. Acquisition of Aneuploidy Provides Increased Fitness during the Evolution of Antifungal Drug Resistance. PLoS Genet. 2009, 5, e1000705. [Google Scholar] [CrossRef] [PubMed]
- Selmecki, A.; Forche, A.; Berman, J. Aneuploidy and isochromosome formation in drug-resistant Candida albicans. Science 2006, 313, 367–370. [Google Scholar] [CrossRef] [PubMed]
- Chow, E.W.; Morrow, C.A.; Djordjevic, J.T.; Wood, I.A.; Fraser, J.A. Microevolution of Cryptococcus neoformans Driven by Massive Tandem Gene Amplification. Mol. Biol. Evol. 2012, 29, 1987–2000. [Google Scholar] [CrossRef] [PubMed]
- Gerstein, A.C.; Fu, M.S.; Mukaremera, L.; Li, Z.; Ormerod, K.L.; Fraser, J.A.; Berman, J.; Nielsen, K. Polyploid Titan Cells Produce Haploid and Aneuploid Progeny To Promote Stress Adaptation. mBio 2015, 6, 01340-15. [Google Scholar] [CrossRef] [PubMed]
- Ene, I.V.; Farrer, R.A.; Hirakawa, M.P.; Agwamba, K.; Cuomo, C.A.; Bennett, R.J. Global analysis of mutations driving microevolution of a heterozygous diploid fungal pathogen. Proc. Natl. Acad. Sci. USA 2018, 115, 201806002. [Google Scholar] [CrossRef]
- Todd, R.T.; Wikoff, T.D.; Forche, A.; Selmecki, A. Genome plasticity in Candida albicans is driven by long repeat sequences. eLife 2019, 8, 8. [Google Scholar] [CrossRef]
- Allshire, R.C.; Madhani, H.D. Ten principles of heterochromatin formation and function. Nat. Rev. Mol. Cell Biol. 2017, 19, 229–244. [Google Scholar] [CrossRef]
- Peng, J.C.; Karpen, G.H. Epigenetic regulation of heterochromatic DNA stability. Curr. Opin. Genet. Dev. 2008, 18, 204–211. [Google Scholar] [CrossRef]
- Luger, K.; Mäder, A.W.; Richmond, R.K.; Sargent, D.F.; Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 1997, 389, 251–260. [Google Scholar] [CrossRef]
- Allis, C.D.; Jenuwein, T. The molecular hallmarks of epigenetic control. Nat. Rev. Genet. 2016, 17, 487–500. [Google Scholar] [CrossRef] [PubMed]
- Janssen, A.; Colmenares, S.U.; Karpen, G.H. Heterochromatin: Guardian of the Genome. Annu. Rev. Cell Dev. Biol. 2018, 34, 265–288. [Google Scholar] [CrossRef] [PubMed]
- Rea, S.; Eisenhaber, F.; O’Carroll, D.; Strahl, B.D.; Sun, Z.-W.; Schmid, M.; Opravil, S.; Mechtler, K.; Ponting, C.P.; Allis, C.D.; et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 2000, 406, 593–599. [Google Scholar] [CrossRef] [PubMed]
- Rose, N.R.; Klose, R.J. Understanding the relationship between DNA methylation and histone lysine methylation. Biochim. Biophys. Acta (BBA) Bioenerg. 2014, 1839, 1362–1372. [Google Scholar] [CrossRef] [PubMed]
- Kuzmichev, A.; Nishioka, K.; Erdjument-Bromage, H.; Tempst, P.; Reinberg, D. Histone methyltransferase activity associated with a human multiprotein complex containing the Enhancer of Zeste protein. Genome Res. 2002, 16, 2893–2905. [Google Scholar] [CrossRef]
- Kellis, M.; Birren, B.W.; Lander, E.S. Proof and evolutionary analysis of ancient genome duplication in the yeast Saccharomyces cerevisiae. Nature 2004, 428, 617–624. [Google Scholar] [CrossRef]
- Wolfe, K.H.; Shields, D.C. Molecular evidence for an ancient duplication of the entire yeast genome. Nature 1997, 387, 708–713. [Google Scholar] [CrossRef]
- Pfaller, M.A.; Diekema, D.J. Epidemiology of Invasive Mycoses in North America. Crit. Rev. Microbiol. 2010, 36, 1–53. [Google Scholar] [CrossRef]
- Freitag, M. Histone Methylation by SET Domain Proteins in Fungi. Annu. Rev. Microbiol. 2017, 71, 413–439. [Google Scholar] [CrossRef]
- Hecht, A.; Laroche, T.; Strahl-Bolsinger, S.; Gasser, S.M.; Grunstein, M. Histone H3 and H4 N-termini interact with SIR3 and SIR4 proteins: A molecular model for the formation of heterochromatin in yeast. Cell 1995, 80, 583–592. [Google Scholar] [CrossRef]
- Onishi, M.; Liou, G.-G.; Buchberger, J.R.; Walz, T.; Moazed, D. Role of the Conserved Sir3-BAH Domain in Nucleosome Binding and Silent Chromatin Assembly. Mol. Cell 2007, 28, 1015–1028. [Google Scholar] [CrossRef] [PubMed]
- Szilard, R.K.; Jacques, P.-É.; Laramée, L.; Cheng, B.; Galicia, S.; Bataille, A.R.; Yeung, M.; Mendez, M.; Bergeron, M.; Robert, F.; et al. Systematic identification of fragile sites via genome-wide location analysis of gamma-H2AX. Nat. Struct. Mol. Biol. 2010, 17, 299–305. [Google Scholar] [CrossRef] [PubMed]
- Kitada, T.; Schleker, T.; Sperling, A.S.; Xie, W.; Gasser, S.M.; Grunstein, M. γH2A is a component of yeast heterochromatin required for telomere elongation. Cell Cycle 2011, 10, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Iacovoni, J.S.; Caron, P.; Lassadi, I.; Nicolas, E.; Massip, L.; Trouche, D.; Legube, G. High-resolution profiling of gammaH2AX around DNA double strand breaks in the mammalian genome. EMBO J. 2010, 29, 1446–1457. [Google Scholar] [CrossRef]
- Sherman, J.M.; Stone, E.M.; Freeman-Cook, L.L.; Brachmann, C.B.; Boeke, J.D.; Pillus, L. The Conserved Core of a Human SIR2 Homologue Functions in Yeast Silencing. Mol. Biol. Cell 1999, 10, 3045–3059. [Google Scholar] [CrossRef]
- Ellahi, A.; Thurtle, D.M.; Rine, J. The Chromatin and Transcriptional Landscape of Native Saccharomyces cerevisiaeTelomeres and Subtelomeric Domains. Genetics 2015, 200, 505–521. [Google Scholar] [CrossRef]
- Loo, S.; Rine, J. Silencers and domains of generalized repression. Science 1994, 264, 1768–1771. [Google Scholar] [CrossRef]
- Pryde, F.E.; Louis, E.J. Limitations of silencing at native yeast telomeres. EMBO J 1999, 18, 2538. [Google Scholar] [CrossRef]
- Rine, J.; Strathern, J.N.; Hicks, J.B.; Herskowitz, I. A suppressor of mating-type locus mutations in saccharomyces cerevisiae: Evidence FOr and identification of cryptic mating-type lOCI. Genetics 1979, 93, 877–901. [Google Scholar]
- Rine, J.; Herskowitz, I. Four Genes Responsible for a Position Effect on Expression From HML and HMR in Saccharomyces cerevisiae. Genetics 1987, 116, 9–22. [Google Scholar]
- Kobayashi, T.; Horiuchi, T.; Tongaonkar, P.; Vu, L.; Nomura, M. SIR2 regulates recombination between different rDNA repeats, but not recombination within individual rRNA genes in yeast. Cell 2004, 117, 441–453. [Google Scholar] [CrossRef]
- Haber, J.E. Mating-Type Genes and MAT Switching in Saccharomyces cerevisiae. Genetics 2012, 191, 33–64. [Google Scholar] [CrossRef] [PubMed]
- Connolly, B.; White, C.I.; Haber, J.E. Physical monitoring of mating type switching in Saccharomyces cerevisiae. Mol. Cell. Biol. 1988, 8, 2342–2349. [Google Scholar] [CrossRef] [PubMed]
- Brown, C.A.; Murray, A.W.; Verstrepen, K.J. Rapid Expansion and Functional Divergence of Subtelomeric Gene Families in Yeasts. Curr. Biol. 2010, 20, 895–903. [Google Scholar] [CrossRef]
- Louis, E.J.; Haber, J.E. The Structure and Evolution of Subtelomeric Y’ Repeats in Saccharomyces Cerevisiae. Genetics 1992, 131, 559–574. [Google Scholar]
- Liti, G.; Carter, D.M.; Moses, A.M.; Warringer, J.; Parts, L.; James, S.A.; Davey, R.P.; Roberts, I.N.; Burt, A.; Koufopanou, V.; et al. Population genomics of domestic and wild yeasts. Nature 2009, 458, 337–341. [Google Scholar] [CrossRef]
- Kobayashi, T.; Nomura, M.; Horiuchi, T. Identification of DNA cis Elements Essential for Expansion of Ribosomal DNA Repeats in Saccharomyces cerevisiae. Mol. Cell. Biol. 2001, 21, 136–147. [Google Scholar] [CrossRef]
- Kobayashi, T.; Ganley, A.R.D. Recombination Regulation by Transcription-Induced Cohesin Dissociation in rDNA Repeats. Science 2005, 309, 1581–1584. [Google Scholar] [CrossRef]
- Prieto, M.; Wedin, M. Dating the Diversification of the Major Lineages of Ascomycota (Fungi). PLoS ONE 2013, 8, e65576. [Google Scholar] [CrossRef]
- Erlendson, A.A.; Friedman, S.; Freitag, M. A matter of scale and dimensions: Chromatin of chromosome landmarks in the fungi. Microbiol. Spectr. 2017, 5. [Google Scholar] [CrossRef]
- Volpe, T.A.; Kidner, C.; Hall, I.M.; Teng, G.; Grewal, S.I.S.; Martienssen, R.A. Regulation of Heterochromatic Silencing and Histone H3 Lysine-9 Methylation by RNAi. Science 2002, 297, 1833–1837. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, J.-I.; Rice, J.C.; Strahl, B.D.; Allis, C.D.; Grewal, S.I.S. Role of Histone H3 Lysine 9 Methylation in Epigenetic Control of Heterochromatin Assembly. Science 2001, 292, 110–113. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, J.; Klar, A.J.; Grewal, S.I. A chromodomain protein, Swi6, performs imprinting functions in fission yeast during mitosis and meiosis. Cell 2000, 101, 307–317. [Google Scholar] [CrossRef]
- Zhang, K.; Mosch, K.; Fischle, W.; Grewal, S.I.S. Roles of the Clr4 methyltransferase complex in nucleation, spreading and maintenance of heterochromatin. Nat. Struct. Mol. Biol. 2008, 15, 381–388. [Google Scholar] [CrossRef]
- Buscaino, A.; Lejeune, E.; Audergon, P.; Hamilton, G.; Pidoux, A.; Allshire, R.C. Distinct roles for Sir2 and RNAi in centromeric heterochromatin nucleation, spreading and maintenance. EMBO J. 2013, 32, 1250–1264. [Google Scholar] [CrossRef]
- Wirén, M.; Silverstein, R.A.; Sinha, I.; Walfridsson, J.; Lee, H.-M.; Laurenson, P.; Pillus, L.; Robyr, D.; Grunstein, M.; Ekwall, K. Genomewide analysis of nucleosome density histone acetylation and HDAC function in fission yeast. EMBO J. 2005, 24, 2906–2918. [Google Scholar] [CrossRef]
- Sugiyama, T.; Cam, H.P.; Sugiyama, R.; Noma, K.-I.; Zofall, M.; Kobayashi, R.; Grewal, S.I. SHREC, an Effector Complex for Heterochromatic Transcriptional Silencing. Cell 2007, 129, 1227. [Google Scholar] [CrossRef][Green Version]
- Buscaino, A.; Allshire, R.; Pidoux, A. Building centromeres: Home sweet home or a nomadic existence? Curr. Opin. Genet. Dev. 2010, 20, 118–126. [Google Scholar] [CrossRef]
- Takahashi, K.; Chen, E.S.; Yanagida, M. Requirement of Mis6 Centromere Connector for Localizing a CENP-A-Like Protein in Fission Yeast. Science 2000, 288, 2215–2219. [Google Scholar] [CrossRef]
- Partridge, J.F.; Borgstrøm, B.; Allshire, R.C. Complex centromere Distinct protein interaction domains and protein spreading in a complex centromere. Genes Dev. 2000, 14, 783–791. [Google Scholar] [CrossRef]
- Bernard, P.; Maure, J.-F.; Partridge, J.F.; Genier, S.; Javerzat, J.-P.; Allshire, R.C. Requirement of Heterochromatin for Cohesion at Centromeres. Science 2001, 294, 2539–2542. [Google Scholar] [CrossRef] [PubMed]
- Pidoux, A.L.; Uzawa, S.; Perry, P.E.; Cande, W.Z.; Allshire, R.C. Live analysis of lagging chromosomes during anaphase and their effect on spindle elongation rate in fission yeast. J. Cell Sci. 2000, 113, 4177–4191. [Google Scholar] [PubMed]
- Zaratiegui, M.; Castel, S.E.; Irvine, D.V.; Kloc, A.; Ren, J.; Li, F.; De Castro, E.; Marín, L.; Chang, A.-Y.; Goto, D.; et al. RNAi promotes heterochromatic silencing through replication-coupled release of RNA Pol II. Nature 2011, 479, 135–138. [Google Scholar] [CrossRef] [PubMed]
- Ellermeier, C.; Higuchi, E.C.; Phadnis, N.; Holm, L.; Geelhood, J.L.; Thon, G.; Smith, G.R. RNAi and heterochromatin repress centromeric meiotic recombination. Proc. Natl. Acad. Sci. USA 2010, 107, 8701–8705. [Google Scholar] [CrossRef] [PubMed]
- Nimmo, E.; Cranston, G.; Allshire, R. Telomere-associated chromosome breakage in fission yeast results in variegated expression of adjacent genes. EMBO J. 1994, 13, 3801–3811. [Google Scholar] [CrossRef]
- Jain, D.; Hebden, A.K.; Nakamura, T.M.; Miller, K.M.; Cooper, J.P. HAATI survivors replace canonical telomeres with blocks of generic heterochromatin. Nature 2010, 467, 223–227. [Google Scholar] [CrossRef]
- Zofall, M.; Yamanaka, S.; Reyes-Turcu, F.E.; Zhang, K.; Rubin, C.; Grewal, S.I.S. RNA elimination machinery targeting meiotic mRNAs promotes facultative heterochromatin formation. Science 2011, 335, 96–100. [Google Scholar] [CrossRef]
- Gallagher, P.S.; Larkin, M.; Thillainadesan, G.; Dhakshnamoorthy, J.; Balachandran, V.; Xiao, H.; Wellman, C.; Chatterjee, R.; Wheeler, D.; Grewal, S.I.S. Iron homeostasis regulates facultative heterochromatin assembly in adaptive genome control. Nat. Struct. Mol. Biol. 2018, 25, 372–383. [Google Scholar] [CrossRef]
- Krassowski, T.; Coughlan, A.Y.; Shen, X.-X.; Zhou, X.; Kominek, J.; Opulente, D.A.; Riley, R.; Grigoriev, I.V.; Maheshwari, N.; Shields, D.C.; et al. Evolutionary instability of CUG-Leu in the genetic code of budding yeasts. Nat. Commun. 2018, 9, 1887. [Google Scholar] [CrossRef]
- Mühlhausen, S.; Kollmar, M. Molecular Phylogeny of Sequenced Saccharomycetes Reveals Polyphyly of the Alternative Yeast Codon Usage. Genome Biol. Evol. 2014, 6, 3222–3237. [Google Scholar] [CrossRef]
- Antinori, S.; Milazzo, L.; Sollima, S.; Galli, M.; Corbellino, M. Candidemia and invasive candidiasis in adults: A narrative review. Eur. J. Intern. Med. 2016, 34, 21–28. [Google Scholar] [CrossRef] [PubMed]
- Forche, A.; Cromie, G.; Gerstein, A.C.; Solis, N.V.; Pisithkul, T.; Srifa, W.; Jeffery, E.; Abbey, D.; Filler, S.G.; Dudley, A.M.; et al. Rapid Phenotypic and Genotypic Diversification After Exposure to the Oral Host Niche in Candida albicans. Genetics 2018, 209, 725–741. [Google Scholar] [PubMed]
- Forche, A.; Solis, N.V.; Swidergall, M.; Thomas, R.; Guyer, A.; Beach, A.; Cromie, G.A.; Le, G.T.; Lowell, E.; Pavelka, N.; et al. Selection of Candida albicans trisomy during oropharyngeal infection results in a commensal-like phenotype. PLoS Genet. 2019, 15, e1008137. [Google Scholar] [CrossRef] [PubMed]
- Forche, A.; Alby, K.; Schaefer, D.; Johnson, A.D.; Berman, J.; Bennett, R.J. The parasexual cycle in Candida albicans provides an alternative pathway to meiosis for the formation of recombinant strains. PLoS Biol. 2008, 6, e110. [Google Scholar] [CrossRef]
- van het Hoog, M.; Rast, T.J.; Martchenko, M.; Grindle, S.; Dignard, D.; Hogues, H.; Cuomo, C.; Berriman, M.; Scherer, S.; Magee, B.B.; et al. Assembly of the Candida albicans genome into sixteen supercontigs aligned on the eight chromosomes. Genome Biol. 2007, 8, R52. [Google Scholar] [CrossRef]
- Poulter, R.T.; Goodwin, T.J. Multiple LTR-Retrotransposon Families in the Asexual Yeast Candida albicans. Genome Res. 2000, 10, 174–191. [Google Scholar]
- Anderson, M.Z.; Baller, J.A.; Dulmage, K.; Wigen, L.; Berman, J. The Three Clades of the Telomere-Associated TLO Gene Family of Candida albicans Have Different Splicing, Localization, and Expression Features. Eukaryot. Cell 2012, 11, 1268–1275. [Google Scholar] [CrossRef]
- Chibana, H.; Magee, P.T. The enigma of the major repeat sequence ofCandida albicans. Future Microbiol. 2009, 4, 171–179. [Google Scholar] [CrossRef]
- Hull, C.M. Identification of a Mating Type-Like Locus in the Asexual Pathogenic Yeast Candida albicans. Science 1999, 285, 1271–1275. [Google Scholar] [CrossRef]
- Drinnenberg, I.A.; Weinberg, D.E.; Xie, K.T.; Mower, J.P.; Wolfe, K.H.; Fink, G.R.; Bartel, D.P. RNAi in budding yeast. Science 2009, 326, 544–550. [Google Scholar] [CrossRef]
- Staab, J.F.; White, T.C.; Marr, K.A. Hairpin dsRNA does not trigger RNA interference in Candida albicans cells. Yeast 2011, 28, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Freire-Benéitez, V.; Price, R.J.; Tarrant, D.; Berman, J.; Buscaino, A. Candida albicans repetitive elements display epigenetic diversity and plasticity. Sci. Rep. 2016, 6, 22989. [Google Scholar] [CrossRef] [PubMed]
- Price, R.J.; Weindling, E.; Berman, J.; Buscaino, A. Chromatin Profiling of the Repetitive and Nonrepetitive Genomes of the Human Fungal Pathogen Candida albicans. mBio 2019, 10, e01376-19. [Google Scholar] [CrossRef] [PubMed]
- Anderson, M.Z.; Gerstein, A.C.; Wigen, L.; Baller, J.A.; Berman, J. Silencing Is Noisy: Population and Cell Level Noise in Telomere-Adjacent Genes Is Dependent on Telomere Position and Sir2. PLoS Genet. 2014, 10, e1004436. [Google Scholar] [CrossRef]
- Mishra, P.K.; Baum, M.; Carbon, J. DNA methylation regulates phenotype-dependent transcriptional activity in Candida albicans. Proc. Natl. Acad. Sci. USA 2011, 108, 11965–11970. [Google Scholar] [CrossRef]
- Chindamporn, A.; Chibana, H.; Dois, M.; Tanaka, K. Repetitive sequences (RPSs) in the chromosomes of Candida albicans are sandwiched between two novel stretches, HOK and RB2, common to each chromosome. Microbiology 1998, 144, 849–857. [Google Scholar] [CrossRef][Green Version]
- Beimforde, C.; Feldberg, K.; Nylinder, S.; Rikkinen, J.; Tuovila, H.; Dörfelt, H.; Gube, M.; Jackson, D.J.; Reitner, J.; Seyfullah, L.J.; et al. Estimating the Phanerozoic history of the Ascomycota lineages: Combining fossil and molecular data. Mol. Phylogenetics Evol. 2014, 78, 386–398. [Google Scholar] [CrossRef]
- Morgan, J.; Wannemuehler, K.A.; Marr, K.A.; Hadley, S.; Kontoyiannis, D.P.; Walsh, T.J.; Fridkin, S.K.; Pappas, P.G.; Warnock, D.W. Incidence of invasive aspergillosis following hematopoietic stem cell and solid organ transplantation: Interim results of a prospective multicenter surveillance program. Med. Mycol. 2005, 43, 49–58. [Google Scholar] [CrossRef]
- Garcia-Rubio, R.; Cuenca-Estrella, M.; Mellado, E. Triazole Resistance in Aspergillus Species: An Emerging Problem. Drugs 2017, 77, 599–613. [Google Scholar] [CrossRef]
- O’Gorman, C.M.; Fuller, H.T.; Dyer, P.S. Discovery of a sexual cycle in the opportunistic fungal pathogen Aspergillus fumigatus. Nature 2009, 457, 471–474. [Google Scholar] [CrossRef]
- Perrin, R.M.; Fedorova, N.D.; Bok, J.W.; Cramer, R.A.; Wortman, J.R.; Kim, H.S.; Nierman, W.C.; Keller, N.P. Transcriptional Regulation of Chemical Diversity in Aspergillus fumigatus by LaeA. PLoS Pathog. 2007, 3, e50. [Google Scholar] [CrossRef] [PubMed]
- Bok, J.W.; Balajee, S.A.; Marr, K.A.; Andes, D.; Nielsen, K.F.; Frisvad, J.C.; Keller, N.P. LaeA, a Regulator of Morphogenetic Fungal Virulence Factors. Eukaryot. Cell 2005, 4, 1574–1582. [Google Scholar] [CrossRef] [PubMed]
- McDonagh, A.; Fedorova, N.D.; Crabtree, J.; Yu, Y.; Kim, S.; Chen, D.; Loss, O.; Cairns, T.; Goldman, G.; Armstrong-James, D.; et al. Sub-Telomere Directed Gene Expression during Initiation of Invasive Aspergillosis. PLOS Pathog. 2008, 4, e1000154. [Google Scholar] [CrossRef] [PubMed]
- Lind, A.L.; Wisecaver, J.H.; Lameiras, C.; Wiemann, P.; Palmer, J.M.; Keller, N.P.; Rodrigues, F.; Goldman, G.H.; Rokas, A. Drivers of genetic diversity in secondary metabolic gene clusters within a fungal species. PLoS Biol. 2017, 15, e2003583. [Google Scholar] [CrossRef] [PubMed]
- Nie, X.; Li, B.; Wang, S. Epigenetic and Posttranslational Modifications in Regulating the Biology of Aspergillus Species. Adv. Appl. Microbiol. 2018, 105, 191–226. [Google Scholar] [PubMed]
- Gowher, H.; Ehrlich, K.C.; Jeltsch, A. DNA from Aspergillus flavus contains 5-methylcytosine. FEMS Microbiol. Lett. 2001, 205, 151–155. [Google Scholar] [CrossRef]
- Liu, S.-Y.; Lin, J.-Q.; Wu, H.-L.; Wang, C.-C.; Huang, S.-J.; Luo, Y.-F.; Sun, J.-H.; Zhou, J.-X.; Yan, S.-J.; He, J.-G.; et al. Bisulfite Sequencing Reveals That Aspergillus flavus Holds a Hollow in DNA Methylation. PLoS ONE 2012, 7, e30349. [Google Scholar] [CrossRef]
- Yang, K.; Liang, L.; Ran, F.; Liu, Y.; Li, Z.; Lan, H.; Gao, P.; Zhuang, Z.; Zhang, F.; Nie, X.; et al. The DmtA methyltransferase contributes to Aspergillus flavus conidiation, sclerotial production, aflatoxin biosynthesis and virulence. Sci. Rep. 2016, 6, 23259. [Google Scholar] [CrossRef]
- Hammond, T.M.; Keller, N.P. RNA Silencing in Aspergillus nidulans Is Independent of RNA-Dependent RNA Polymerases. Genetics 2005, 169, 607–617. [Google Scholar] [CrossRef]
- Hammond, T.M.; Andrewski, M.D.; Roossinck, M.J.; Keller, N.P. Aspergillus Mycoviruses Are Targets and Suppressors of RNA Silencing. Eukaryot. Cell 2008, 7, 350–357. [Google Scholar] [CrossRef]
- Reyes-Dominguez, Y.; Bok, J.W.; Berger, H.; Shwab, E.K.; Basheer, A.; Gallmetzer, A.; Scazzocchio, C.; Keller, N.; Strauss, J. Heterochromatic marks are associated with the repression of secondary metabolism clusters in Aspergillus nidulans. Mol. Microbiol. 2010, 76, 1376–1386. [Google Scholar] [CrossRef] [PubMed]
- Lee, I.; Oh, J.-H.; Shwab, E.K.; Dagenais, T.R.T.; Andes, D.; Keller, N.P. HdaA, a class 2 histone deacetylase of Aspergillus fumigatus, affects germination and secondary metabolite production. Fungal Genet. Biol. 2009, 46, 782–790. [Google Scholar] [CrossRef] [PubMed]
- Palmer, J.M.; Perrin, R.M.; Dagenais, T.R.T.; Keller, N.P. H3K9 Methylation Regulates Growth and Development in Aspergillus fumigatus. Eukaryot. Cell 2008, 7, 2052–2060. [Google Scholar] [CrossRef] [PubMed]
- Park, B.J.; Wannemuehler, K.A.; Marston, B.J.; Govender, N.; Pappas, P.G.; Chiller, T.M. Estimation of the current global burden of cryptococcal meningitis among persons living with HIV/AIDS. AIDS 2009, 23, 525–530. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.; Kim, S.-H. A genome Tree of Life for the Fungi kingdom. Proc. Natl. Acad. Sci. USA 2017, 114, 9391–9396. [Google Scholar] [CrossRef]
- Kwon-Chung, K.J.; Fraser, J.A.; Doering, T.L.; Wang, Z.; Janbon, G.; Idnurm, A.; Bahn, Y.-S. Cryptococcus neoformans and Cryptococcus gattii, the Etiologic Agents of Cryptococcosis. Cold Spring Harb. Perspect. Med. 2014, 4, a019760. [Google Scholar] [CrossRef]
- Zaragoza, O.; Nielsen, K. Titan cells in Cryptococcus neoformans: Cells with a giant impact. Curr. Opin. Microbiol. 2013, 16, 409–413. [Google Scholar] [CrossRef]
- Hommel, B.; Mukaremera, L.; Cordero, R.J.B.; Coelho, C.; Desjardins, C.A.; Sturny-Leclère, A.; Janbon, G.; Perfect, J.R.; Fraser, J.A.; Casadevall, A.; et al. Titan cells formation in Cryptococcus neoformans is finely tuned by environmental conditions and modulated by positive and negative genetic regulators. PLoS Pathog. 2018, 14, e1006982. [Google Scholar] [CrossRef]
- Dambuza, I.M.; Drake, T.; Chapuis, A.; Zhou, X.; Correia, J.; Taylor-Smith, L.; Legrave, N.; Rasmussen, T.; Fisher, M.C.; Bicanic, T.; et al. The Cryptococcus neoformans Titan cell is an inducible and regulated morphotype underlying pathogenesis. PLoS Pathog. 2018, 14, e1006978. [Google Scholar] [CrossRef]
- Trevijano-Contador, N.; De Oliveira, H.C.; García-Rodas, R.; Rossi, S.A.; Llorente, I.; Zaballos, A.; Janbon, G.; Arino, J.; Zaragoza, O. Cryptococcus neoformans can form titan-like cells in vitro in response to multiple signals. PLoS Pathog. 2018, 14, e1007007. [Google Scholar] [CrossRef]
- Okagaki, L.H.; Nielsen, K. Titan Cells Confer Protection from Phagocytosis in Cryptococcus neoformans Infections. Eukaryot. Cell 2012, 11, 820–826. [Google Scholar] [CrossRef] [PubMed]
- Sionov, E.; Lee, H.; Chang, Y.C.; Kwon-Chung, K.J. Cryptococcus neoformans overcomes stress of azole drugs by formation of disomy in specific multiple chromosomes. PLoS Pathog. 2010, 6, e1000848. [Google Scholar] [CrossRef] [PubMed]
- Stone, N.R.; Rhodes, J.; Fisher, M.C.; Mfinanga, S.; Kivuyo, S.; Rugemalila, J.; Segal, E.S.; Needleman, L.; Molloy, S.F.; Kwon-Chung, J.; et al. Dynamic ploidy changes drive fluconazole resistance in human cryptococcal meningitis. J. Clin. Investig. 2019, 129, 999–1014. [Google Scholar] [CrossRef] [PubMed]
- Huff, J.T.; Zilberman, D. Dnmt1-independent CG methylation contributes to nucleosome positioning in diverse eukaryotes. Cell 2014, 156, 1286–1297. [Google Scholar] [CrossRef]
- Catania, S.; Dumesic, P.A.; Pimentel, H.; Nasif, A.; Stoddard, C.I.; Burke, J.E.; Diedrich, J.K.; Cook., S.; Shea, T.; Geinger, E.; et al. Evolutionary persistence of DNA methylation for millions of years after ancient loss of a de novo methyltransferase. bioRxiv 2019. [Google Scholar] [CrossRef]
- Janbon, G.; Maeng, S.; Yang, D.-H.; Ko, Y.-J.; Jung, K.-W.; Moyrand, F.; Floyd, A.; Heitman, J.; Bahn, Y.-S. Characterizing the role of RNA silencing components in Cryptococcus neoformans. Fungal. Genet. Biol. 2010, 47, 1070–1080. [Google Scholar] [CrossRef]
- Hsueh, Y.-P.; Heitman, J. Orchestration of sexual reproduction and virulence by the fungal mating-type locus. Curr. Opin. Microbiol. 2008, 11, 517–524. [Google Scholar] [CrossRef]
- Janbon, G.; Ormerod, K.L.; Paulet, D.; Byrnes, E.J.; Yadav, V.; Chatterjee, G.; Mullapudi, N.; Hon, C.-C.; Billmyre, R.B.; Brunel, F.; et al. Analysis of the Genome and Transcriptome of Cryptococcus neoformans var. grubii Reveals Complex RNA Expression and Microevolution Leading to Virulence Attenuation. PLoS Genet. 2014, 10, e1004261. [Google Scholar] [CrossRef]
- Yadav, V.; Sun, S.; Billmyre, R.B.; Thimmappa, B.C.; Shea, T.; Lintner, R.; Bakkeren, G.; Cuomo, C.A.; Heitman, J.; Sanyal, K. RNAi is a critical determinant of centromere evolution in closely related fungi. Proc. Natl. Acad. Sci. USA 2018, 115, 3108–3113. [Google Scholar] [CrossRef]
- Yadav, V.; Sreekumar, L.; Guin, K.; Sanyal, K. Five pillars of centromeric chromatin in fungal pathogens. PLoS Pathog. 2018, 14, e1007150. [Google Scholar] [CrossRef]
- Wang, X.; Hsueh, Y.-P.; Li, W.; Floyd, A.; Skalsky, R.; Heitman, J. Sex-induced silencing defends the genome of Cryptococcus neoformans via RNAi. Genes Dev. 2010, 24, 2566–2582. [Google Scholar] [CrossRef] [PubMed]
- Dumesic, P.A.; Homer, C.M.; Moresco, J.J.; Pack, L.R.; Shanle, E.K.; Coyle, S.M.; Strahl, B.D.; Fujimori, D.G.; Yates, J.R.; Madhani, H.D. Product binding enforces the genomic specificity of a yeast polycomb repressive complex. Cell 2015, 160, 204–218. [Google Scholar] [CrossRef] [PubMed]
- Cam, H.P.; Sugiyama, T.; Chen, E.S.; Chen, X.; Fitzgerald, P.C.; Grewal, S.I.S. Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nat. Genet. 2005, 37, 809–819. [Google Scholar] [CrossRef] [PubMed]
- Pomraning, K.R.; Smith, K.M.; Freitag, M. Genome-wide high throughput analysis of DNA methylation in eukaryotes. Methods 2009, 47, 142–150. [Google Scholar] [CrossRef]
- Wierer, M.; Mann, M. Proteomics to study DNA-bound and chromatin-associated gene regulatory complexes. Hum. Mol. Genet. 2016, 25, R106–R114. [Google Scholar] [CrossRef]
- Cam, H.P.; Whitehall, S. Reporter Gene Silencing Assays in Fission Yeast. Cold Spring Harb. Protoc. 2016, 2016, pdb.prot091512. [Google Scholar] [CrossRef]
- Jalan, M.; Oehler, J.; Morrow, C.A.; Osman, F.; Whitby, M.C. Factors affecting template switch recombination associated with restarted DNA replication. eLife 2019, 8, 8. [Google Scholar] [CrossRef]
© 2019 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Buscaino, A. Chromatin-Mediated Regulation of Genome Plasticity in Human Fungal Pathogens. Genes 2019, 10, 855. https://doi.org/10.3390/genes10110855
Buscaino A. Chromatin-Mediated Regulation of Genome Plasticity in Human Fungal Pathogens. Genes. 2019; 10(11):855. https://doi.org/10.3390/genes10110855
Chicago/Turabian StyleBuscaino, Alessia. 2019. "Chromatin-Mediated Regulation of Genome Plasticity in Human Fungal Pathogens" Genes 10, no. 11: 855. https://doi.org/10.3390/genes10110855
APA StyleBuscaino, A. (2019). Chromatin-Mediated Regulation of Genome Plasticity in Human Fungal Pathogens. Genes, 10(11), 855. https://doi.org/10.3390/genes10110855




