Comprehensive Insight into Gibberellin- and Jasmonate-Mediated Stamen Development
Abstract
1. Introduction
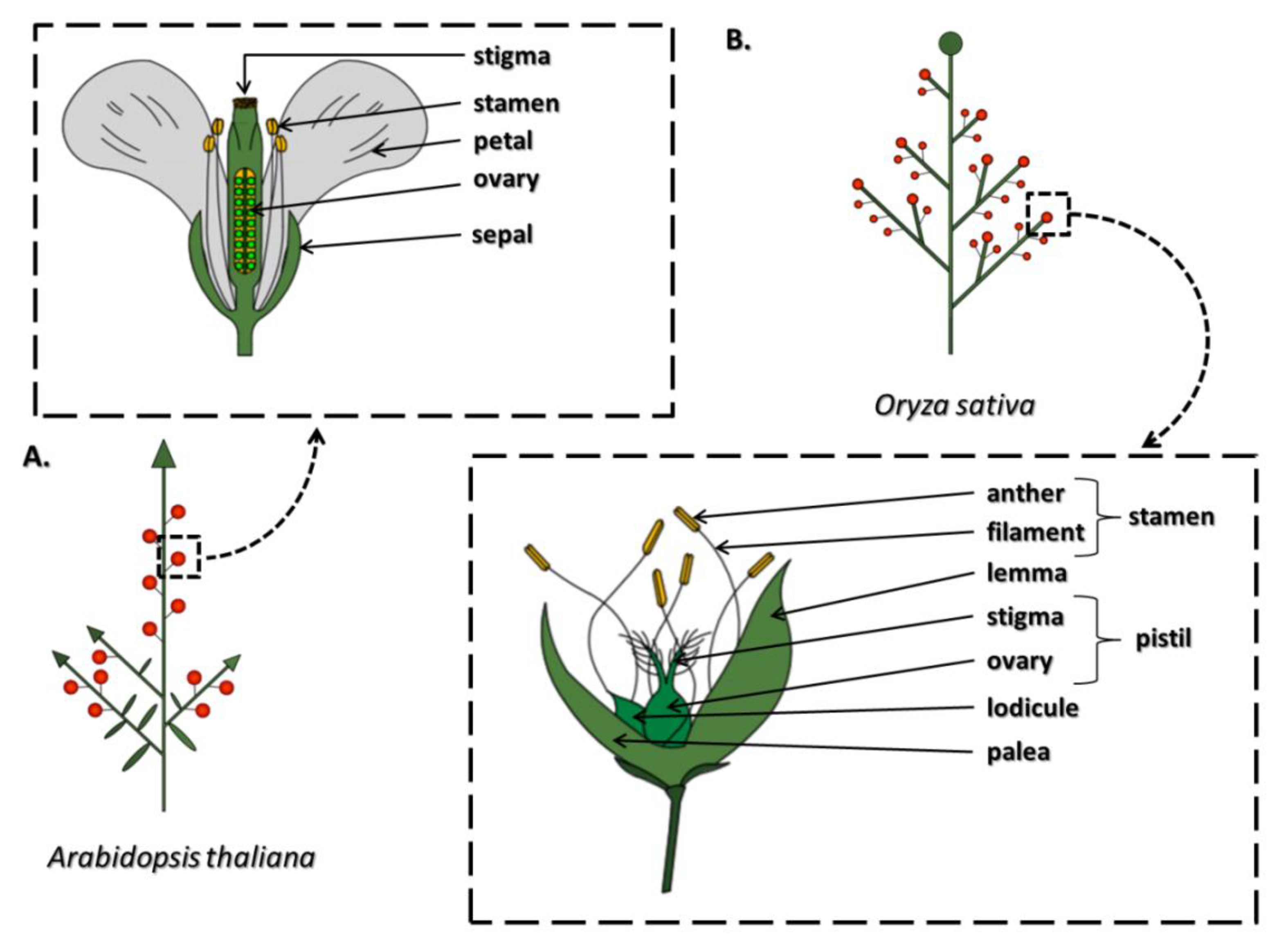
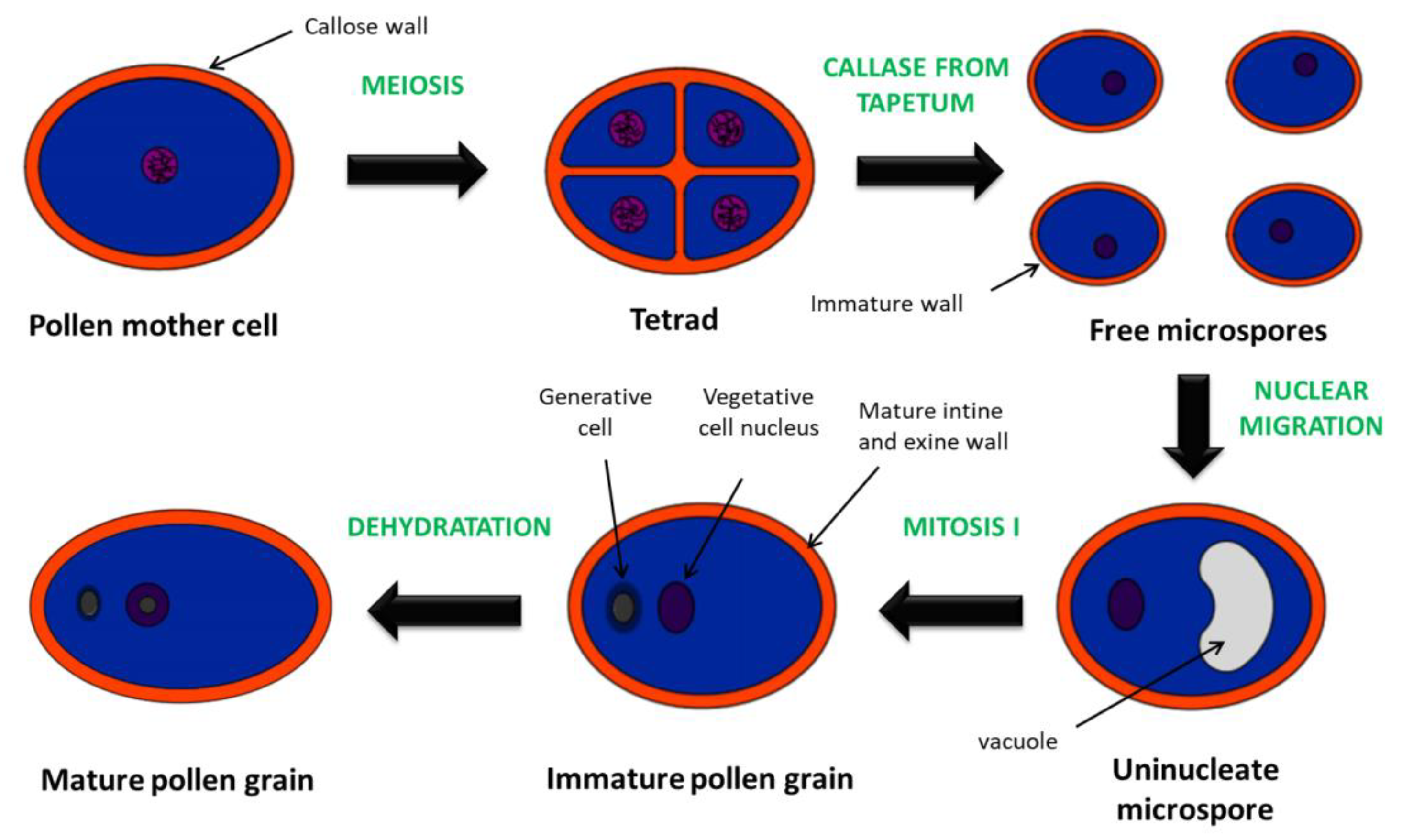
2. Structure, Function and Development of Stamens
3. Gibberellin-Mediated Stamen Development and Functioning
3.1. GA Biosynthesis Is Crucial for Correct Stamen Development and Functioning in Various Plants
3.1.1. GA Biosynthesis Pathway—General Information
3.1.2. Early Stages of GA Biosynthesis
3.1.3. Late Stages of GA Biosynthesis
3.2. Perception, Signal Transduction and Action of GAs during Stamen Development
3.2.1. GA Signaling Pathway—General Information
3.2.2. Receptor—Dependent Signaling
3.2.3. The Role of DELLA Proteins
3.2.4. Events Downstream of DELLAs during Filament Elongation and Anther Development
4. Jasmonate-Dependent Stamen Development and Functioning
4.1. The Importance of JA Biosynthesis in Proper Stamen Development
4.1.1. JA Metabolism—General Information
4.1.2. Studies on JA Biosynthesis Mutants
4.2. Perception, Signal Transduction and Action of JAs during Stamen Development
4.2.1. JA Signaling Pathway—General Information
4.2.2. JA Signaling Dependent on the COI Receptor
4.2.3. The Pathway Downstream of the JAZ Repressor
5. Gibberellin-Jasmonate Interactions in the Regulation of Stamen Development
6. Hormonal Transport in Stamen Development
7. Summary
Author Contributions
Funding
Conflicts of Interest
References
- Plackett, A.R.G.; Thomas, S.G.; Wilson, Z.A.; Hedden, P. Gibberellin control of stamen development: A fertile field. Trends Plant Sci. 2011, 16, 568–578. [Google Scholar] [CrossRef] [PubMed]
- Turczyn, M. ABC of flower architecture. Post Biol. Kom. 2011, 38, 673–684. [Google Scholar]
- Wilson, Z.A.; Zhang, D.-B. From Arabidopsis to rice: Pathways in pollen development. J. Exp. Bot. 2009, 60, 1479–1492. [Google Scholar] [CrossRef] [PubMed]
- Teo, Z.W.N.; Song, S.; Wang, Y.-Q.; Liu, J.; Yu, H. New insights into the regulation of inflorescence architecture. Trends Plant Sci. 2014, 19, 158–165. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.; Sun, B.; Looi, L.S.; Xu, Y.; Gan, E.S.; Huang, J.; Ito, T. Co-ordination of flower development through epigenetic regulation in two model species: Rice and Arabidopsis. Plant Cell Physiol. 2015, 56, 830–842. [Google Scholar] [CrossRef] [PubMed]
- Coen, E.S.; Meyerowitz, E.M. The war of the whorls: Genetic interactions controlling flower development. Nature 1991, 353, 31–37. [Google Scholar] [CrossRef] [PubMed]
- Theißen, G. Development of floral organ identity: Stories from the MADS house. Curr. Opin. Plant Biol. 2001, 4, 75–85. [Google Scholar] [CrossRef]
- Theißen, G.; Melzer, R.; Rümpler, F. MADS-domain transcription factors and the floral quartet model of flower development: Linking plant development and evolution. Development 2016, 143, 3259–3271. [Google Scholar] [CrossRef]
- Song, S.; Qi, T.; Huang, H.; Xie, D. Regulation of stamen development by coordinated actions of jasmonate, auxin, and gibberellin in Arabidopsis. Mol. Plant 2013, 6, 1065–1073. [Google Scholar] [CrossRef]
- Chandler, J.W. The hormonal regulation of flower development. J. Plant Growth Regul. 2011, 30, 242–254. [Google Scholar] [CrossRef]
- De Jong, A.W.; Bruinsma, J. Pistil development in Cleome flowers III. Effects of growth-regulating substances on flower buds of Cleome iberidella Welw. ex Oliv. Grown in vitro. Zeitschrift für Pflanzenphysiologie 1974, 73, 142–151. [Google Scholar] [CrossRef]
- Pharis, R.P.; Ross, S.D.; McMullan, E. Promotion of flowering in the Pinaceae by gibberellins III. Seedlings of Douglas fir. Physiol. Plant 1980, 50, 119–126. [Google Scholar] [CrossRef]
- Wilmowicz, E.; Frankowski, K.; Glazińska, P.; Sidłowska, M.; Marciniak, K.; Kopcewicz, J. The role of gibberellins in the regulation of flowering in plants. Kosmos 2011, 60, 129–140. [Google Scholar]
- Marciniak, K.; Turowski, T.; Wilmowicz, E.; Frankowski, K.; Kęsy, J.; Kopcewicz, J. Ubiquitin ligases in auxin, jasmontate and gibberellin signal transduction pathways. Post. Biol. Kom. 2010, 37, 409–432. [Google Scholar]
- Marciniak, K.; Wilmowicz, E.; Kućko, A.; Kęsy, J.; Kopcewicz, J. Protein kinases in plant hormone signaling pathways. Post. Biol. Kom. 2013, 40, 253–294. [Google Scholar]
- Cardarelli, M.; Cecchetti, V. Auxin polar transport in stamen formation and development: How many actors? Front. Plant Sci. 2014, 5, 333. [Google Scholar] [CrossRef]
- Goldberg, R.B.; Beals, T.P.; Sanders, P.M. Anther development: Basic principles and practical applications. Plant Cell 1993, 5, 1217–1229. [Google Scholar]
- Scott, R.J.; Spielman, M.; Dickinson, H.G. Stamen structure and function. Plant Cell 2004, 16, 46–60. [Google Scholar] [CrossRef]
- Sanders, P.M.; Bui, A.Q.; Weterings, K.; McIntire, K.N.; Hsu, Y.C.; Lee, P.Y.; Truong, M.T.; Beals, T.P.; Goldberg, R.B. Anther developmental defects in Arabidopsis thaliana male-sterile mutants. Sex Plant Reprod. 1999, 11, 297–322. [Google Scholar] [CrossRef]
- Wilson, Z.A.; Song, J.; Taylor, B.; Yang, C. The final split: The regulation of anther dehiscence. J. Exp. Bot. 2011, 62, 1633–1649. [Google Scholar] [CrossRef]
- Smyth, D.R.; Bowman, J.L.; Meyerowitz, E.M. Early flower development in Arabidopsis. Plant Cell 1990, 2, 755–767. [Google Scholar] [PubMed]
- Bowman, J.L. Arabidopsis: An Atlas of Morphology and Development; Springer: New York, NY, USA, 1994. [Google Scholar]
- Gómez, J.F.; Talle, B.; Wilson, Z.A. Anther and pollen development: A conserved developmental pathway. J. Integr. Plant Biol. 2015, 57, 876–891. [Google Scholar] [CrossRef] [PubMed]
- McCormick, S. Male gametophyte development. Plant Cell 1993, 5, 1265–1275. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.B.; Wilson, Z.A. Stamen specification and anther development in rice. Chin. Sci Bull 2009, 54, 2342–2353. [Google Scholar] [CrossRef]
- Chen, C.-B.; Xu, Y.-Y.; Ma, H.; Chong, K. Cell biological characterization of male meiosis and pollen development in rice. J. Int. Plant Biol. 2005, 47, 734–744. [Google Scholar] [CrossRef]
- Itoh, J.; Nonomura, K.; Ikeda, K.; Yamaki, S.; Inukai, Y.; Yamagishi, H.; Kitano, H.; Nagato, Y. Rice plant development: From zygote to spikelet. Plant Cell Physiol. 2005, 46, 23–47. [Google Scholar] [CrossRef]
- Wilson, R.N.; Heckman, J.W.; Somerville, C.R. Gibberellin is required for flowering in Arabidopsis thaliana under short days. Plant Physiol. 1992, 100, 403–408. [Google Scholar] [CrossRef]
- Marciniak, K.; Wilmowicz, E.; Kućko, A.; Kopcewicz, J. Photoperiod and ethylene-dependent expression of gibberellin metabolism gene InEKO1 during flower induction of Ipomoea nil. Biol. Plant 2018, 62, 194–199. [Google Scholar] [CrossRef]
- Mutasa-Göttgens, E.; Hedden, P. Gibberellin as a factor in floral regulatory networks. J. Exp. Bot. 2009, 60, 1979–1989. [Google Scholar] [CrossRef]
- Kulikowska-Gulewska, H.; Majewska, M.; Kopcewicz, J. Gibberellins in the control of photoperiodic flower transition in Pharbitis nil. Physiol. Plant. 2000, 108, 202–207. [Google Scholar] [CrossRef]
- Marciniak, K.; Kućko, A.; Wilmowicz, E.; Świdziński, M.; Kęsy, J.; Kopcewicz, J. Photoperiodic flower induction in Ipomoea nil is accompanied by decreasing content of gibberellins. Plant Growth Regul. 2018, 84, 395–400. [Google Scholar] [CrossRef]
- Yang, Y.Y.; Yamaguchi, I.; Takeno-wada, K.; Suzuki, Y.; Murofushi, N. Metabolism and translocation of gibberellins in seedlings of Pharbitis nil. (I) Effect of photoperiod on stem elongation and endogenous gibberellins in cotyledons and their phloem exudates. Plant Cell Physiol. 1995, 36, 221–227. [Google Scholar] [CrossRef]
- Takeno, K.; Tsuruta, T.; Maeda, T. Gibberellins are not essential for photoperiodic flower induction of Pharbitis nil. Physiol. Plant 1996, 97, 397–401. [Google Scholar] [CrossRef]
- Griffiths, J.; Murase, K.; Rieu, I.; Zentella, R.; Zhang, Z.L.; Powers, S.J.; Gong, F.; Phillips, A.L.; Hedden, P.; Sun, T.P.; et al. Genetic characterization and functional analysis of the GID1 gibberellin receptors in Arabidopsis. Plant Cell 2006, 18, 3399–3414. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.H.; Mitchum, M.G.; Barnaby, N.; Ayele, B.T.; Ogawa, M.; Nam, E.; Lai, W.C.; Hanada, A.; Alonso, J.M.; Ecker, J.R.; et al. Potential sites of bioactive gibberellin production during reproductive growth in Arabidopsis. Plant Cell 2008, 20, 320–336. [Google Scholar] [CrossRef] [PubMed]
- Rieu, I.; Ruiz-Rivero, O.; Fernandez-Garcia, N.; Griffiths, J.; Powers, S.J.; Gong, F.; Phillips, A.L. The gibberellin biosynthetic genes AtGA20ox1 and AtGA20ox2 act, partially redundantly, to promote growth and development throughout the Arabidopsis life cycle. Plant J. 2008, 53, 488–504. [Google Scholar] [CrossRef]
- Goto, N.; Pharis, R.P. Role of gibberellins in the development of floral organs of the gibberellin-deficient mutant, ga1-1, of Arabidopsis thaliana. Can. J. Bot. 1999, 77, 944–954. [Google Scholar]
- Nester, J.E.; Zeevaart, J.A.D. Flower development in normal tomato and a gibberellin-deficient (ga-2) mutant. Am. J. Bot. 1988, 75, 45–55. [Google Scholar] [CrossRef]
- Cheng, H.; Qin, L.; Lee, S.; Fu, X.; Richards, D.E.; Cao, D.; Luo, D.; Harberd, N.P.; Peng, J. Gibberellin regulates Arabidopsis floral development via suppression of DELLA protein function. Development 2004, 131, 1055–1064. [Google Scholar] [CrossRef]
- Marciniak, K.; Grzegorzewska, W.; Kęsy, J.; Szmidt-Jaworska, A.; Tretyn, A.; Kopcewicz, J. Regulation of gibberellin metabolism in plants. Kosmos 2012, 61, 213–232. [Google Scholar]
- Marciniak, K.; Kęsy, J.; Tretyn, A.; Kopcewicz, J. Gibberellins–Structure, biosynthesis and deactivation in plants. Post. Biochem. 2012, 58, 14–25. [Google Scholar]
- Hedden, P.; Phillips, A.L. Gibberellin metabolism: New insights revealed by the genes. Trends Plant Sci. 2000, 5, 523–530. [Google Scholar] [CrossRef]
- Hedden, P.; Thomas, S.G. Gibberellin biosynthesis and its regulation. Biochem. J. 2012, 444, 11–25. [Google Scholar] [CrossRef] [PubMed]
- Davière, J.M.; Achard, P. Gibberellin signaling in plants. Development 2013, 140, 1147–1151. [Google Scholar] [CrossRef]
- Peng, J. Gibberellin and jasmonate crosstalk during stamen development. J. Integr. Plant Biol. 2009, 51, 1064–1070. [Google Scholar] [CrossRef]
- Jacobsen, S.E.; Olszewski, N.E. Characterization of the arrest in another development associated with gibberellin deficiency of the gib-1 mutant of tomato. Plant Physiol. 1991, 97, 409–414. [Google Scholar] [CrossRef]
- Aya, K.; Ueguchi-Tanaka, M.; Kondo, M.; Hamada, K.; Yano, K.; Nishimura, M.; Matsuoka, M. Gibberellin modulates anther development in rice via the transcriptional regulation of GAMYB. Plant Cell 2009, 21, 1453–1472. [Google Scholar] [CrossRef]
- Chhun, T.; Aya, K.; Asano, K.; Yamamoto, E.; Morinaka, Y.; Watanabe, M.; Kitano, H.; Ashikari, M.; Matsuoka, M.; Ueguchi-Tanaka, M. Gibberellin regulates pollen viability and pollen tube growth in rice. Plant Cell 2007, 19, 3876–3888. [Google Scholar] [CrossRef]
- Mitchum, M.G.; Yamaguchi, S.; Hanada, A.; Kuwahara, A.; Yoshioka, Y.; Kato, T.; Tabata, S.; Kamiya, Y.; Sun, T.P. Distinct and overlapping roles of two gibberellin 3-oxidases in Arabidopsis development. Plant J. 2006, 45, 804–818. [Google Scholar] [CrossRef]
- Hirano, K.; Aya, K.; Hobo, T.; Sakakibara, H.; Kojima, M.; Shim, R.A.; Hasegawa, Y.; Ueguchi-Tanaka, M.; Matsuoka, M. Comprehensive transcriptome analysis of phytohormone biosynthesis and signaling genes in microspore/pollen and tapetum of rice. Plant Cell Physiol. 2008, 49, 1429–1450. [Google Scholar] [CrossRef]
- Silverstone, A.L.; Chang, C.-W.; Krol, E.; Sun, T.P. Developmental regulation of the gibberellin biosynthetic gene GA1 in Arabidopsis thaliana. Plant J. 1997, 12, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Plackett, A.R.G.; Powers, S.J.; Fernandez-Garcia, N.; Urbanova, T.; Takebayashi, Y.; Seo, M.; Jikumaru, Y.; Benlloch, R.; Nilsson, O.; Ruiz-Rivero, O.; et al. Analysis of the developmental roles of the Arabidopsis gibberellin 20-oxidases demonstrates that GA20ox1, -2, and -3 are the dominant paralogs. Plant Cell 2012, 24, 941–960. [Google Scholar] [CrossRef] [PubMed]
- Ueguchi-Tanaka, M.; Ashikari, M.; Nakajima, M.; Itoh, H.; Katoh, E.; Kobayashi, M.; Chow, T.Y.; Hsing, Y.I.; Kitano, H.; Yamaguchi, I.; et al. GIBBERELLIN INSENSITIVE DWARF1 encodes a soluble receptor for gibberellin. Nature 2005, 437, 693–698. [Google Scholar] [CrossRef] [PubMed]
- Hirano, K.; Ueguchi-Tanaka, M.; Matsuoka, M. GID1-mediated gibberellin signaling in plants. Trends Plant Sci. 2008, 13, 192–199. [Google Scholar] [CrossRef]
- Marciniak, K.; Świeżawska, B.; Kęsy, J.; Tretyn, A.; Kopcewicz, J. Gibberellins–Perception and signal transduction in plants. Post. Biol. Kom. 2012, 39, 25–48. [Google Scholar]
- Huang, S.; Cerny, R.E.; Qi, Y.; Bhat, D.; Aydt, C.M.; Hanson, D.D.; Malloy, K.P.; Ness, L.A. Transgenic studies on the involvement of cytokinin and gibberellin in male development. Plant Physiol. 2003, 131, 1270–1282. [Google Scholar] [CrossRef]
- Hynes, L.W.; Peng, J.; Richards, D.E.; Harberd, N.P. Transgenic expression of the Arabidopsis DELLA proteins GAI and gai confers altered gibberellin response in tobacco. Transgenic Res. 2003, 12, 707–714. [Google Scholar] [CrossRef]
- Ikeda, A.; Ueguchi-Tanaka, M.; Sonoda, Y.; Kitano, H.; Koshioka, M.; Futsuhara, Y.; Matsuoka, M.; Yamaguchi, J. Slender rice, a constitutive gibberellin response mutant, is caused by a null mutation of the SLR1 gene, an ortholog of the height-regulating gene GAI/RGA/RHT/D8. Plant Cell 2001, 13, 999–1010. [Google Scholar] [CrossRef]
- Chandler, P.M.; Marion-Poll, A.; Ellis, M.; Gubler, F. Mutants at the Slender1 locus of barley cv Himalaya. Molecular and physiological characterization. Plant Physiol. 2002, 129, 181–190. [Google Scholar] [CrossRef]
- Dill, A.; Sun, T. Synergistic derepression of gibberellin signaling by removing RGA and GAI function in Arabidopsis thaliana. Genetics 2001, 159, 777–785. [Google Scholar]
- King, K.E.; Moritz, T.; Harberd, N.P. Gibberellins are not required for stem growth in Arabidopsis thaliana in the absence of GAI and RGA. Genetics 2001, 159, 767–776. [Google Scholar] [PubMed]
- Lee, S.; Cheng, H.; King, K.E.; Wang, W.; He, Y.; Hussain, A.; Lo, J.; Harberd, N.P.; Peng, J. Gibberellin regulates Arabidopsis seed germination via RGL2, a GAI/RGA-like gene whose expression is up-regulated following imbibition. Genes Dev. 2002, 16, 646–658. [Google Scholar] [CrossRef] [PubMed]
- Cao, D.; Cheng, H.; Wu, W.; Soo, H.M.; Peng, J. Gibberellin mobilizes distinct DELLA-dependent transcriptomes to regulate seed germination and floral development in Arabidopsis. Plant Physiol. 2006, 142, 509–525. [Google Scholar] [CrossRef] [PubMed]
- Cao, D.N.; Hussain, A.; Cheng, H.; Peng, J.R. Loss of function of four DELLA genes leads to light- and gibberellin-independent seed germination in Arabidopsis. Planta 2005, 223, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Hou, X.L.; Hu, W.W.; Shen, L.S.; Lee, L.Y.C.; Tao, Z.; Han, J.H.; Yu, H. Global identification of DELLA target genes during Arabidopsis flower development. Plant Physiol. 2008, 147, 1126–1142. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Song, S.S.; Xiao, L.T.; Soo, H.M.; Cheng, Z.W.; Xie, D.X.; Peng, J. Gibberellin acts through jasmonate to control the expression of MYB21, MYB24 and MYB57 to promote stamen filament growth in Arabidopsis. PLoS Genet. 2009, 5, e1000440. [Google Scholar] [CrossRef]
- Stracke, R.; Werber, M.; Weisshaar, B. The R2R3-MYB gene family in Arabidopsis thaliana. Curr. Opin. Plant Biol. 2001, 4, 447–456. [Google Scholar] [CrossRef]
- Kranz, H.D.; Denekamp, M.; Greco, R.; Jin, H.; Leyva, A.; Meissner, R.C.; Petroni, K.; Urzainqui, A.; Bevan, M.; Martin, C.; et al. Towards functional characterisation of the members of the R2R3-MYB gene family from Arabidopsis thaliana. Plant J. 1998, 16, 263–276. [Google Scholar] [CrossRef]
- Millar, A.A.; Gubler, F. The Arabidopsis GAMYB-like genes, MYB33 and MYB65, are microRNA-regulated genes that redundantly facilitate anther development. Plant Cell 2005, 17, 705–721. [Google Scholar] [CrossRef]
- Zhang, D.; Liang, W.; Yin, C.; Zong, J.; Gu, F.; Zhang, D. OsC6, encoding a lipid transfer protein, is required for postmeiotic anther development in rice. Plant Physiol. 2010, 154, 149–162. [Google Scholar] [CrossRef]
- Li, N.; Zhang, D.S.; Liu, H.S.; Yin, C.S.; Li, X.X.; Liang, W.Q.; Yuan, Z.; Xu, B.; Chu, H.W.; Wang, J.; et al. The rice Tapetum Degeneration Retardation gene is required for tapetum degradation and anther development. Plant Cell 2006, 18, 2999. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Yang, C.; Yuan, Z.; Zhang, D.; Gondwe, M.Y.; Ding, Z.; Liang, W.; Zhang, D.; Wilson, Z.A. The ABORTED MICROSPORES regulatory network is required for postmeiotic male reproductive development in Arabidopsis thaliana. Plant Cell 2010, 22, 91–107. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Sun, Y.; Timofejeva, L.; Chen, C.; Grossniklaus, U.; Ma, H. Regulation of Arabidopsis tapetum development and function by Dysfunctional Tapetum1 (DYT1) encoding a putative bHLH transcription factor. Development 2006, 133, 3085–3095. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Liu, B.; Liu, L.; Song, S. Jasmonate action in plant growth and development. J. Exp. Bot. 2017, 68, 1349–1359. [Google Scholar] [CrossRef] [PubMed]
- Avanci, N.C.; Luche, D.D.; Goldman, G.H.; Goldman, N.H. Jasmonates are phytohormones with multiple functions, including plant defense and reproduction. Genet. Mol. Res. 2010, 9, 484–505. [Google Scholar] [CrossRef] [PubMed]
- Liechti, R.; Farmer, E.E. Jasmonate biochemical pathway. Sci. Signal. 2006, 2006, cm18. [Google Scholar] [CrossRef]
- Ishiguro, S.; Kawai-Oda, A.; Ueda, J.; Nishida, I.; Okada, K. The defective in anther dehiscience gene encodes a novel phospholipase A1 catalyzing the initial step of jasmonic acid biosynthesis, which synchronizes pollen maturation, anther dehiscence, and flower opening in Arabidopsis. Plant Cell 2001, 13, 2191–2209. [Google Scholar] [CrossRef]
- Koo, A.J.K.; Cooke, T.F.; Howe, G.A. Cytochrome P450 CYP94B3 mediates catabolism and inactivation of the plant hormone jasmonoyl-L-isoleucine. Proc. Natl. Acad. Sci. USA 2011, 108, 9298–9303. [Google Scholar] [CrossRef]
- Cheong, J.J.; Choi, Y.D. Methyl jasmonate as a vital substance in plants. Trends Genet. 2003, 19, 409–413. [Google Scholar] [CrossRef]
- Yu, Z.Q.; Zhu, J.; Gao, J.F.; Yang, Z.N. Functional analysis of rice P0491E01 gene regulating anther development. Fen Zi Xi Bao Sheng Wu Xue Bao 2006, 39, 467–472. [Google Scholar]
- Caldelari, D.; Wang, G.; Farmer, E.E.; Dong, X. Arabidopsis lox3 lox4 double mutants are male sterile and defective in global proliferative arrest. Plant Mol. Biol. 2011, 75, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Von Malek, B.; van der Graaff, E.; Schneitz, K.; Keller, B. The Arabidopsis male-sterile mutant dde2-2 is defective in the Allene Oxide Synthase gene encoding one of the key enzymes of the jasmonic acid biosynthesis pathway. Planta 2002, 216, 187–192. [Google Scholar] [CrossRef] [PubMed]
- Sanders, P.M.; Lee, P.Y.; Biesgen, C.; Boone, J.D.; Bealsm, T.P.; Weiler, E.W.; Goldberg, R.B. The Arabidopsis Delayed Dehiscence1 gene encodes an enzyme in the jasmonic acid synthesis pathway. Plant Cell 2000, 12, 1041–1061. [Google Scholar] [CrossRef] [PubMed]
- Stintzi, A.; Browse, J. The Arabidopsis male-sterile mutant, opr3, lacks the 12-oxophytodienoic acid reductase required for jasmonate synthesis. Proc. Natl. Acad. Sci. USA 2000, 97, 10625–10630. [Google Scholar] [CrossRef]
- Wasternack, C.; Song, S. Jasmonates: Biosynthesis, metabolism, and signaling by proteins activating and repressing transcription. J. Exp. Bot. 2017, 68, 1303–1321. [Google Scholar] [CrossRef]
- Jewell, J.B.; Browse, J. Epidermal jasmonate perception is sufficient for all aspects of jasmonate-mediated male fertility in Arabidopsis. Plant J. 2016, 85, 634–647. [Google Scholar] [CrossRef]
- Wang, Z.; Dai, L.; Jiang, Z.; Peng, W.; Zhang, L.; Wang, G.; Xie, D. GmCOI1, a soybean F-Box protein gene, shows ability to mediate jasmonate-regulated plant defense and fertility in Arabidopsis. Mol. Plant Microbe Interact 2005, 18, 1285–1295. [Google Scholar] [CrossRef]
- Lee, H.Y.; Seo, J.S.; Cho, J.H.; Jung, H.; Kim, J.K.; Lee, J.S.; Do Choi, Y. Oryza sativa COI homologues restore jasmonate signal transduction in Arabidopsis coi1-1 mutants. PLoS ONE 2013, 8, e52802. [Google Scholar] [CrossRef]
- Li, L.; Zhao, Y.; McCaig, B.C.; Wingerd, B.A.; Wang, J.; Whalon, M.E.; Pichersky, E.; Howe, G.A. The tomato homolog of Coronatine-Insensitive1 is required for the maternal control of seed maturation, jasmonate-signaled defense responses, and glandular trichome development. Plant Cell 2004, 16, 126–143. [Google Scholar] [CrossRef]
- Mandaokar, A.; Thines, B.; Shin, B.; Lange, B.M.; Choi, G.; Koo, Y.J.; Yoo, Y.J.; Choi, Y.D.; Browse, J. Transcriptional regulators of stamen development in Arabidopsis identified by transcriptional profiling. Plant J. 2006, 46, 984–1008. [Google Scholar] [CrossRef]
- Song, S.; Qi, T.; Huang, H.; Ren, Q.; Wu, D.; Chang, C.; Peng, W.; Liu, Y.; Peng, J.; Xie, D. The jasmonate-ZIM domain proteins interact with the R2R3-MYB transcription factors MYB21 and MYB24 to affect jasmonate-regulated stamen development in Arabidopsis. Plant Cell 2011, 23, 1000–1013. [Google Scholar] [CrossRef] [PubMed]
- Mandaokar, A.; Browse, J. MYB108 acts together with MYB24 to regulate jasmonate-mediated stamen maturation in Arabidopsis. Plant Physiol. 2009, 149, 851–862. [Google Scholar] [CrossRef] [PubMed]
- Qi, T.; Huang, H.; Song, S.; Xie, D. Regulation of jasmonate-mediated stamen development and seed production by a bHLH-MYB complex in Arabidopsis. Plant Cell 2015, 27, 1620–1633. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhao, H.; Luo, T.; Liu, Y.; Nie, X.; Li, H. Characteristics and expression pattern of MYC genes in Triticum aestivum, Oryza sativa, and Brachypodium distachyon. Plants 2019, 8, 274. [Google Scholar] [CrossRef]
- Pires, N.; Dolan, L. Origin and diversification of basic-helix-loop-helix proteins in plants. Mol. Biol. Evol. 2010, 27, 862–874. [Google Scholar] [CrossRef]
- Reeves, P.H.; Ellis, C.M.; Ploense, S.E.; Wu, M.F.; Yadav, V.; Tholl, D.; Chételat, A.; Haupt, I.; Kennerley, B.J.; Hodgens, C.; et al. A regulatory network for coordinated flower maturation. PLoS Genet. 2012, 8, e1002506. [Google Scholar] [CrossRef]
- Chen, X.; Huang, H.; Qi, T.; Liu, B.; Song, S. New perspective of the bHLH-MYB complex in jasmonate-regulated plant fertility in Arabidopsis. Plant Signal Behav. 2016, 11, e1135280. [Google Scholar] [CrossRef]
- Cai, Q.; Yuan, Z.; Chen, M.; Yin, C.; Luo, Z.; Zhao, X.; Liang, W.; Hu, J.; Zhang, D. Jasmonic acid regulates spikelet development in rice. Nat. Commun. 2014, 5, 3476. [Google Scholar] [CrossRef]
- Uji, Y.; Taniguchi, S.; Tamaoki, D.; Shishido, H.; Akimitsu, K.; Gomi, K. Overexpression of OsMYC2 Results in the Up-Regulation of Early JA-Rresponsive Genes and Bacterial Blight Resistance in Rice. Plant Cell Physiol. 2016, 57, 1814–1827. [Google Scholar] [CrossRef]
- Feys, J.F.; Benedetti, C.E.; Penfold, C.N.; Turner, J.G. Arabidopsis mutants selected for resistance to the phytotoxin coronatine are male sterile, insensitive to methyl jasmonate, and resistant to a bacterial pathogen. Plant Cell 1994, 6, 751–759. [Google Scholar] [CrossRef]
- Xie, D.X.; Feys, B.F.; James, S.; Nieto-Rostro, M.; Turner, J.G. COI1: An Arabidopsis gene required for jasmonate-regulated defense and fertility. Science 1998, 280, 1091–1094. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Ito, T.; Wellmer, F.; Meyerowitz, E.M. Repression of AGAMOUS-LIKE 24 is a crucial step in promoting flower development. Nat. Genet. 2004, 368, 157–161. [Google Scholar] [CrossRef] [PubMed]
- Ito, T.; Ng, K.H.; Lim, T.S.; Yu, H.; Meyerowitz, E.M. The homeotic protein AGAMOUS controls late stamen development by regulating a jasmonate biosynthetic gene in Arabidopsis. Plant Cell 2007, 198, 3516–3529. [Google Scholar] [CrossRef] [PubMed]
- Grant, M.R.; Jones, J.D.G. Hormone (dis)harmony moulds plant health and disease. Science 2009, 324, 750–752. [Google Scholar] [CrossRef]
- Hong, G.J.; Xue, X.Y.; Mao, Y.B.; Wang, L.J.; Chen, X.Y. Arabidopsis MYC2 interacts with DELLA proteins in regulating sesquiterpene synthase gene expression. Plant Cell 2012, 24, 2635–2648. [Google Scholar] [CrossRef]
- Park, J.; Lee, Y.; Martinoia, E.; Geisler, M. Plant hormone transporters: What we know and what we would like to know. BMC Biol. 2017, 15, 93. [Google Scholar] [CrossRef]
- Kanno, Y.; Hanada, A.; Chiba, Y.; Ichikawa, T.; Nakazawa, M.; Matsui, M.; Koshiba, T.; Kamiya, Y.; Seo, M. Identification of an abscisic acid transporter by functional screening using the receptor complex as a sensor. Proc. Natl. Acad. Sci. USA 2012, 109, 9653–9658. [Google Scholar] [CrossRef]
- Nour-Eldin, H.H.; Andersen, T.G.; Burow, M.; Madsen, S.R.; Jørgensen, M.E.; Olsen, C.E.; Dreyer, I.; Hedrich, R.; Geiger, D.; Halkier, B.A. NRT/PTR transporters are essential for translocation of glucosinolate defence compounds to seeds. Nature 2012, 488, 531–534. [Google Scholar] [CrossRef]
- Saito, H.; Oikawa, T.; Hamamoto, S.; Ishimaru, Y.; Kanamori-Sato, M.; Sasaki-Sekimoto, Y.; Utsumi, Y.; Chen, J.; Kanno, Y.; Masuda, S.; et al. The jasmonate-responsive GTR1 transporter is required for gibberellin-mediated stamen development in Arabidopsis. Nature 2015, 6, 6095. [Google Scholar] [CrossRef]
- Kanno, Y.; Jikumaru, Y.; Hanada, A.; Nambara, E.; Abrams, S.R.; Kamiya, Y.; Seo, M. Comprehensive hormone profiling in developing Arabidopsis seeds: Examination of the site of ABA biosynthesis, ABA transport and hormone interactions. Plant Cell Physiol. 2010, 51, 1988–2001. [Google Scholar] [CrossRef]




| Gene | Mutant | Phenotype | Species | Refs |
|---|---|---|---|---|
| AtCPS | ga1-3 | Male-sterile phenotype, which can be reversed by GA application | Arabidopsis | [40] |
| Anther and pollen development is blocked after meiosis but prior to mitosis | ||||
| Pollen sacs expansion arrest | ||||
| Inability to release microspores | ||||
| Tapetum remains at the vacuolated stage and degenerates together with the microspores | ||||
| Inhibition of filament elongation by a reduction of the length, not the number of cells | ||||
| Altered ratio of stamen-pistil length in the flowers of mature mutant | ||||
| LeCPS | gib-1 | Initiation of floral meristems and development of all floral organs proceeds normally up to a certain point, but then normal development ceases and flower buds eventually abort | tomato | [47] |
| Microsporogenesis is blocked before meiosis | ||||
| Anthers of developmentally arrested buds contain PMCs that are in the G1 phase of premeiotic interphase. Following treatment of mutant buds with GA3, premeiotic DNA synthesis and callose accumulation in PMCs are evident by 48 h posttreatment, and within 66 h, prophase I of meiosis and meiosis-related changes in tapetum development are observable | ||||
| OsCPS1 | oscps1-1 | One of the most severe GA-deficient mutant | rice | [48] |
| GA treatment rescues the defect in stamen development | ||||
| Abnormal enlargement of tapetal cells, to the point of almost filling the locule space | ||||
| Collapse of microspores | ||||
| LeKAO | ga-2 | Flower buds are initiated, but do not develop to maturity and eventually abort | tomato | [39] |
| Cells of the sporogenous layer are initiated, but growth is arrested and cells eventually degenerate | ||||
| Inhibition of microsporogenesis occurs before meiosis | ||||
| Stamen do not elongate | ||||
| OsKAO | rpe1 | Intermediate severity GA-deficient mutant | rice | [49] |
| The mutant develops typical flowers with normal pistils and stamens | ||||
| Pollen viability and the number of mature pollen grains in mutant are similar to those of the WT plant | ||||
| Impaired pollen germination and elongation | ||||
| GA20ox | ga20ox1 ga20ox2 | Semidwarf, semifertile phenotype, with early flowers failing to set seed | Arabidopsis | [37] |
| Self-rescue of seed set occurs in later flowers, although the mechanism remains undetermined | ||||
| Normal tapetum degradation | ||||
| Fully viable pollen | ||||
| Delayed or inhibited anther dehiscence | ||||
| Disturbance of filament elongation | ||||
| ga20ox1 ga20ox2 ga20ox3-1 | For many phenotypic characters, the triple mutant is not significantly different from the ga1-3 | [53] | ||
| Postmeiotic arrest in stamen development | ||||
| Defect in tapetum degeneration. Tapetum layer fails to degenerate completely and remains in anther locules | ||||
| Inhibited anther dehiscence | ||||
| Do not undergo late-stage stamen acceleration, with growth and development instead halting | ||||
| Shorter stamens at flower opening | ||||
| GA3ox | ga3ox1 ga3ox3 | The epidermal layer of the anther remains intact, although the tapetum layer disappears, suggesting that anther development is arrested around stages 11 and 12 | Arabidopsis | [36] |
| Defective pollen after its maturation | ||||
| Delayed or inhibited anther dehiscence | ||||
| Disturbances in filament elongation | ||||
| All defects gradually decrease in the later flowers |
| Gene | Mutant | Phenotype | Species | Refs |
|---|---|---|---|---|
| AtGID1 | gid1a-1 gid1b-1 gid1c-1 | Complete infertility and unresponsiveness to GA treatment | Arabidopsis | [35] |
| The triple mutant exhibits more pronounced disturbances in stamen development than ga1-3 | ||||
| Anther development in this mutant has not been described | ||||
| Dramatic reduction in length of filaments | ||||
| OsGID1 | gid1-4 | Anther-wide developmental arrest to occur either just prior to or during meiosis | rice | [48] |
| PMCs are condensed and do not form tetrads | ||||
| Abnormal stamens with shrunken and whitened anthers | ||||
| Slightly enlarged tapetal cells that nearly fill the locule and contain the degraded meiocyte | ||||
| Middle layer of cells does not degrade | ||||
| Failure in epidermal cell expansion | ||||
| GAI RGA RGL1 RGL2 | ga1-3 gai-t6 rga-t2 rgl1-1 rgl2-1 | Penta mutant can produce fully developed fertile flowers as the WT control | Arabidopsis | [40] |
| ga1-3 gai-t6 rgl1-1 rgl2-1 ga1-3 gai-t6 rgl1-1 rga-t2 | Those quadruple mutants with expression of only RGA or RGL2 are completely sterile | |||
| Mutants are effective in inhibiting the expression of MYB21, MYB24 and MYB57 | ||||
| ga1-3 rga-t2 rgl1-1 rgl2-1 ga1-3 gai-t6 rga-t2 rgl2-1 | Those quadruple mutants with expression of only GAI or RGL1 are fully fertile | |||
| Mutants are ineffective in inhibiting the expression of MYB21, MYB24 and MYB57 | ||||
| SLR1 | Slr1-d3 | Constitutive GA responce mutant is semifertile, even though it develops normal flowers with morphologically normal stamens and pistils | rice | [49] |
| The anthers appear normal and produce a similar number of pollen grains as WT plants | ||||
| High frequency of nonviable pollen | ||||
| slr1-1 | Sterile phenotype | [59] | ||
| Impaired floral development | ||||
| SLN1 | sln | Infertility due to impaired floral development | barley | [60] |
| MYB21 MYB24 MYB57 | myb21-t1 myb24-t1 myb57-t1 | Pollen is partial viable | Arabidopsis | [67] |
| Short stamens are the main cause of the mutant sterility | ||||
| MYB33 MYB65 | myb33 myb65 | Anthers are smaller than those in the WT plants and fail to produce pollen. The block in pollen development appears to be premeiotic occurring between anther stages 5 and 6 | Arabidopsis | [70] |
| During sixth stage of anther development when the PMCs begin to separate in a clearly defined locule and the tapetum begins to vacuolate, the mutant is similar, except that the tapetum begins to enlarge. Next, the tapetum expand to such an extent that there is no locule, and the PMCs have an irregular shape. Whereas microspores form in the locule of WT anthers and eventually form mature pollen, the tapetum of the mutant continues to expand until the contents collapse and degenerate. The expansion of the tapetum appears to be due to an increase in cell size, not in cell number | ||||
| Stamens shorter than their WT counterparts and fail to fully extend to the pistil | ||||
| Other than sterility and the associated characteristics of sterile plants, mutant shows no obvious morphological differences from WT plants |
| Gene | Mutant | Phenotype | Species | Refs |
|---|---|---|---|---|
| PLA1/ DAD1 | dad1 | WT phenotype can be rescued by the JA application | Arabidopsis | [76] |
| Developmental delay of flower bud opening | ||||
| Before flower opening, all cell types are normally developed in mutant anthers, similar to all structural features | ||||
| Pollen grains develop normally up to the trinucleate stage | ||||
| A defect in pollen grains occurs at the final stage of their maturation | ||||
| Defective in anther dehiscence | ||||
| P0491E01 | Normal anther development at the initial stages | rice | [81] | |
| Microspores development into mature pollen grains is impaired | ||||
| LOX3 LOX4 | lox3 lox4 | Male sterile. JA application restored fertility | Arabidopsis | [82] |
| Abnormal anther maturation | ||||
| Pollen is not viable | ||||
| Defective dehiscence | ||||
| Shorter filaments | ||||
| AOS | dde2-2 | Male-sterile phenotype which can be rescued by Me-JA application | Arabidopsis | [83] |
| Impaired anther dehiscence and filament elongation | ||||
| DDE1/ OPR3 | dde1/opr3 | WT phenotype can be rescued by the MeJA application | Arabidopsis | [84,85] |
| Floral organs develop normally within the closed bud | ||||
| The anther locules do not dehisce at the time of flower opening | ||||
| Pollen develops to the trinucleate stage | ||||
| Pollen grains are predominantly inviable | ||||
| The filaments do not elongate sufficiently to position the locules above the stigma at anthesis |
| Gene | Mutant | Phenotype | Species | Refs |
|---|---|---|---|---|
| COI1 | coi-1 | Delayed anther dehiscence | Arabidopsis | [101,102] |
| Reduced pollen viability in the 13th phase of flower development | ||||
| Abnormal filament elongation | ||||
| MYC2 MYC3 MYC4 MYC5 | myc2 myc3 myc4 myc5 | Pollen grains do not germinate in vitro | [94] | |
| The anthers dehisce and release viable pollen at floral stage 15 | ||||
| The filament does not elongate normally at floral stage 13 | ||||
| MYB21 MYB24 | myb21 myb24 | Greatly reduced male fertility. Restore the WT phenotype via JA application | [91] | |
| Delayed anther dehiscence | ||||
| Very short filaments | ||||
| MYB108 | myb108 | Reduced male fertility | [93] | |
| Delayed anther dehiscence | ||||
| Reduced pollen viability |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marciniak, K.; Przedniczek, K. Comprehensive Insight into Gibberellin- and Jasmonate-Mediated Stamen Development. Genes 2019, 10, 811. https://doi.org/10.3390/genes10100811
Marciniak K, Przedniczek K. Comprehensive Insight into Gibberellin- and Jasmonate-Mediated Stamen Development. Genes. 2019; 10(10):811. https://doi.org/10.3390/genes10100811
Chicago/Turabian StyleMarciniak, Katarzyna, and Krzysztof Przedniczek. 2019. "Comprehensive Insight into Gibberellin- and Jasmonate-Mediated Stamen Development" Genes 10, no. 10: 811. https://doi.org/10.3390/genes10100811
APA StyleMarciniak, K., & Przedniczek, K. (2019). Comprehensive Insight into Gibberellin- and Jasmonate-Mediated Stamen Development. Genes, 10(10), 811. https://doi.org/10.3390/genes10100811




