Addition of High Molecular Weight Hyaluronic Acid to Fibroblast-Like Stromal Cells Modulates Endogenous Hyaluronic Acid Metabolism and Enhances Proteolytic Processing and Secretion of Versican
Abstract
1. Introduction
2. Materials and Methods
2.1. (Fibroblast-Like-Stromal Cell) FLSC Cultures
2.2. Treatment of Cultures with LPS and HMW HA
2.3. RTqPCR Assays
2.4. Confocal Microscopy
2.5. Solubilization of Cell Layer Associated ACAN, VCAN, and PRG4
2.6. Purification of Aggrecan, Versican and PRG4 from Culture Medium
2.7. Western Blotting
3. Results
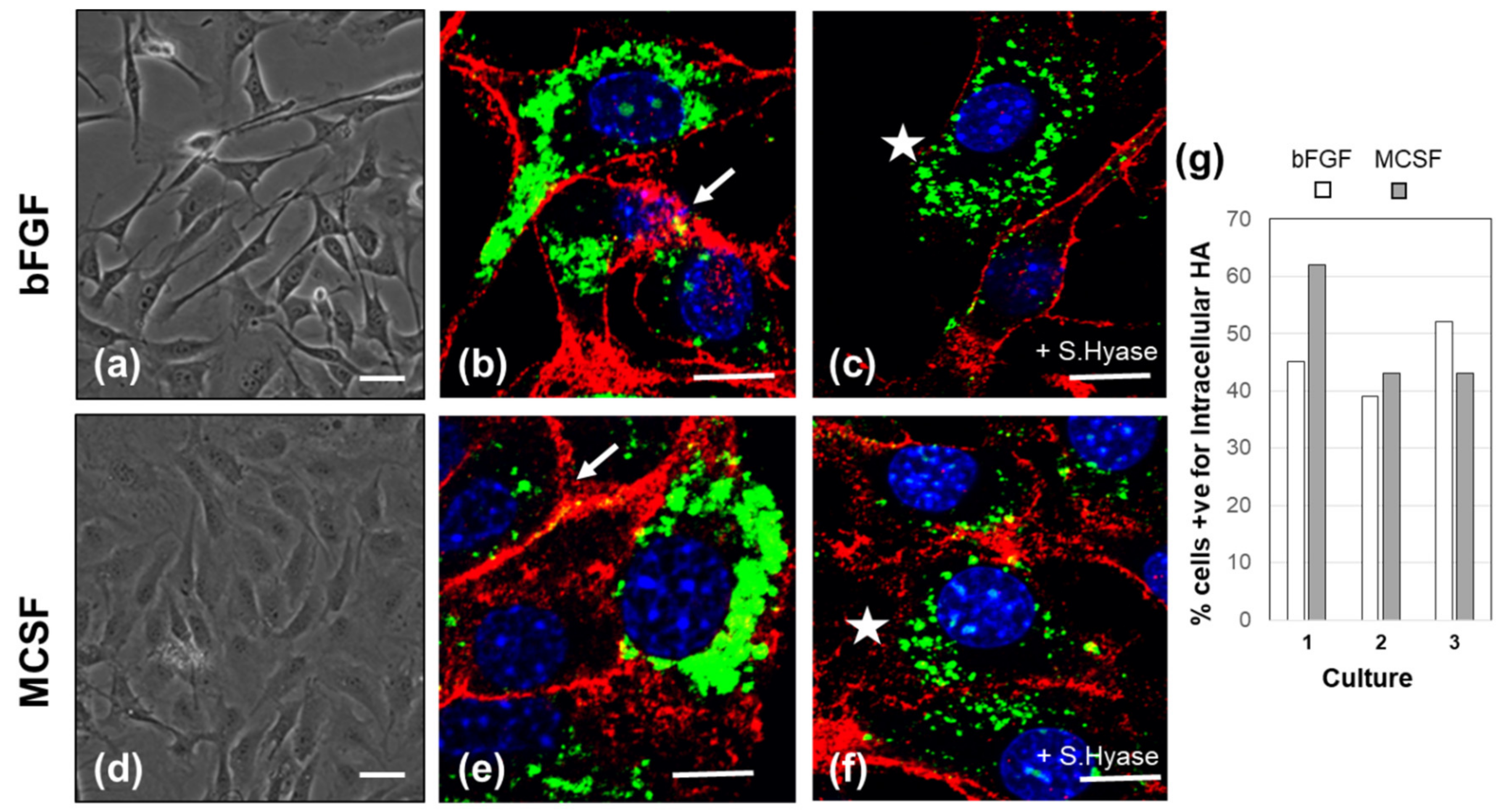
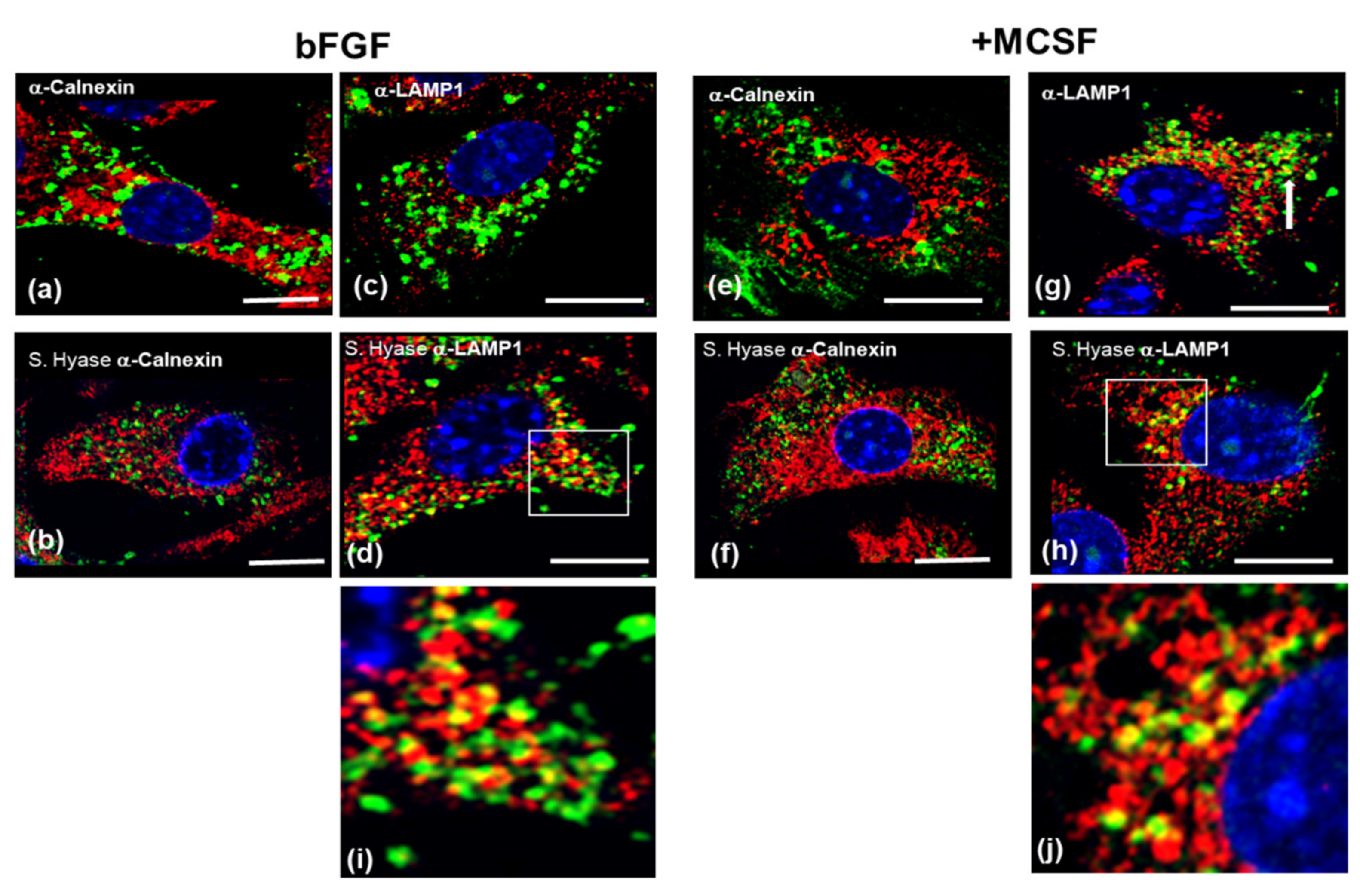
3.1. HA Metabolism in FLSC Cultures Expanded in bFGF and Treated with MCSF to Enhance a Macrophage Phenotype
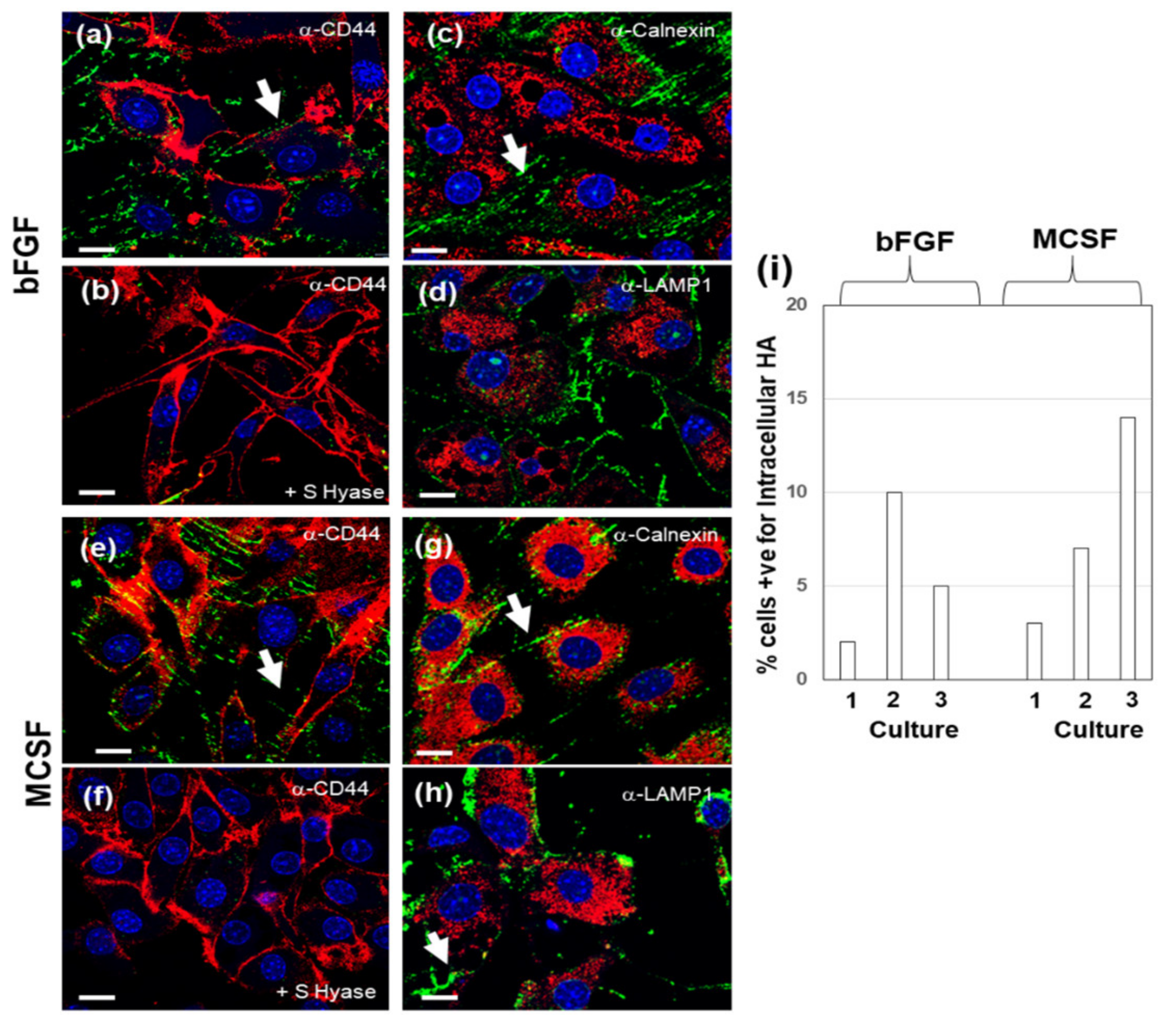
3.2. Effect of LPS on Cell-Associated HA in FLSCs Expanded in bFGF and Treated with MCSF
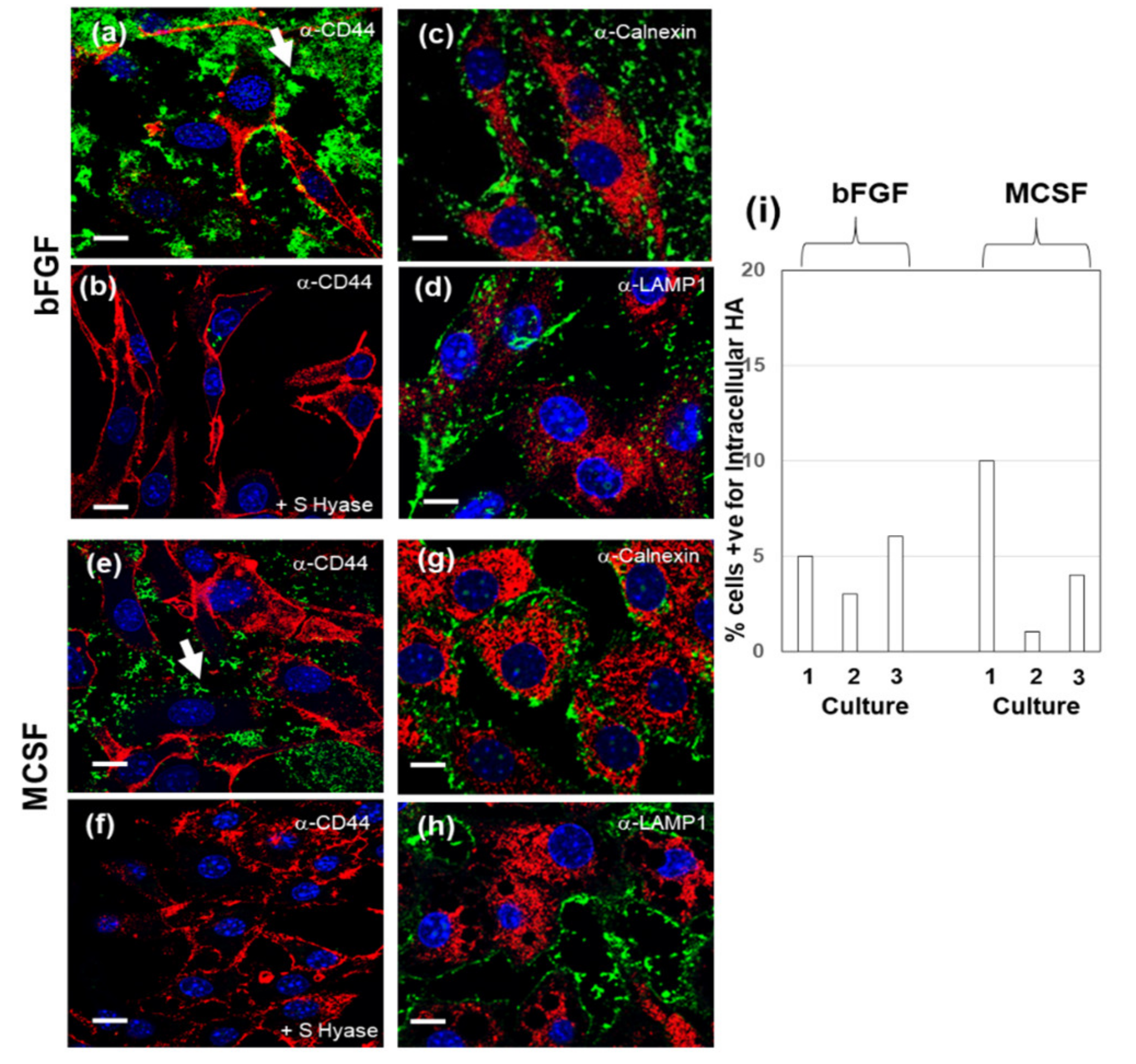
3.3. Effect of Addition of HMW HA on Endogenous HA Metabolism in Basal or LPS Stimulated FLSC Cultures
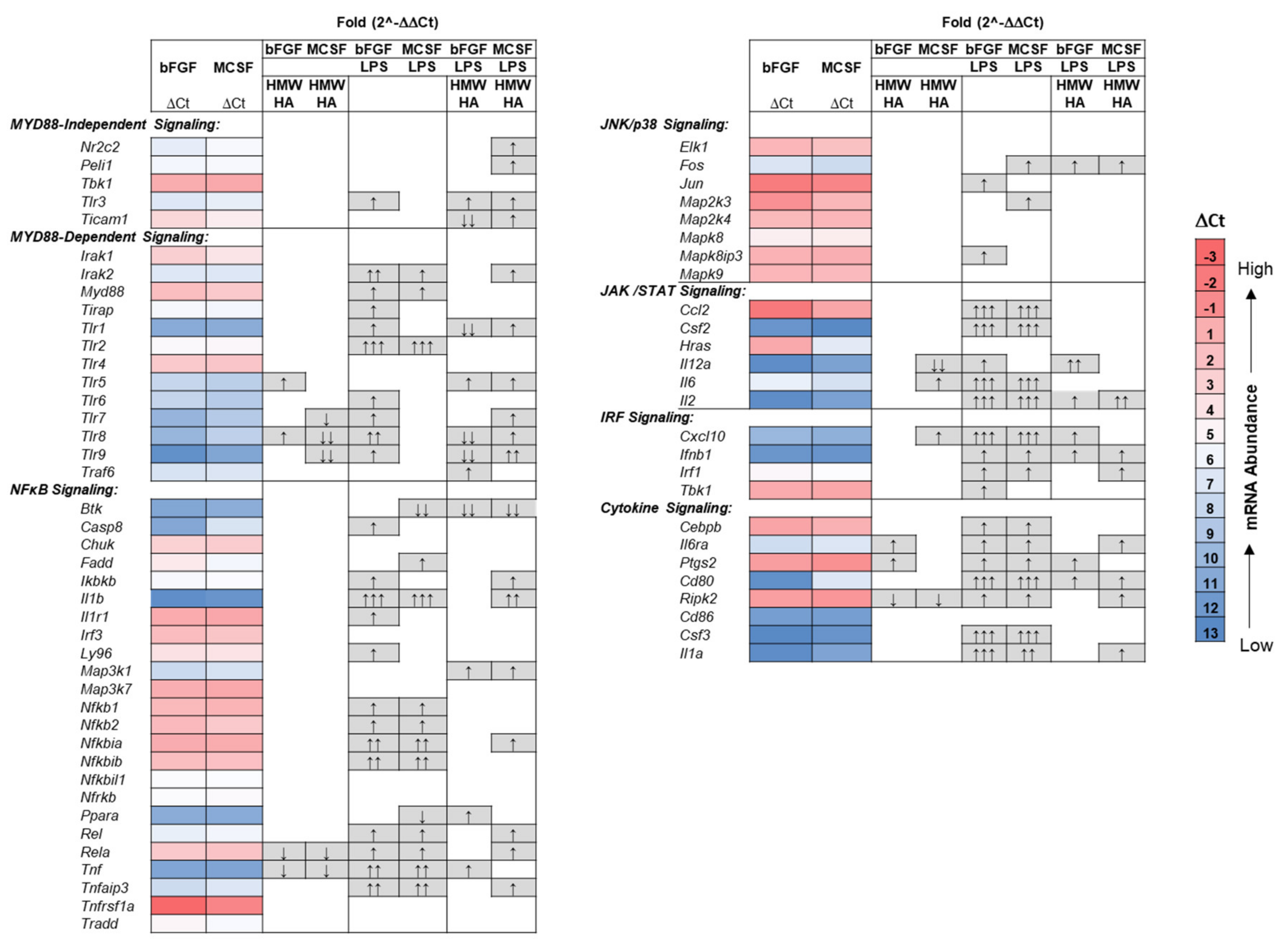
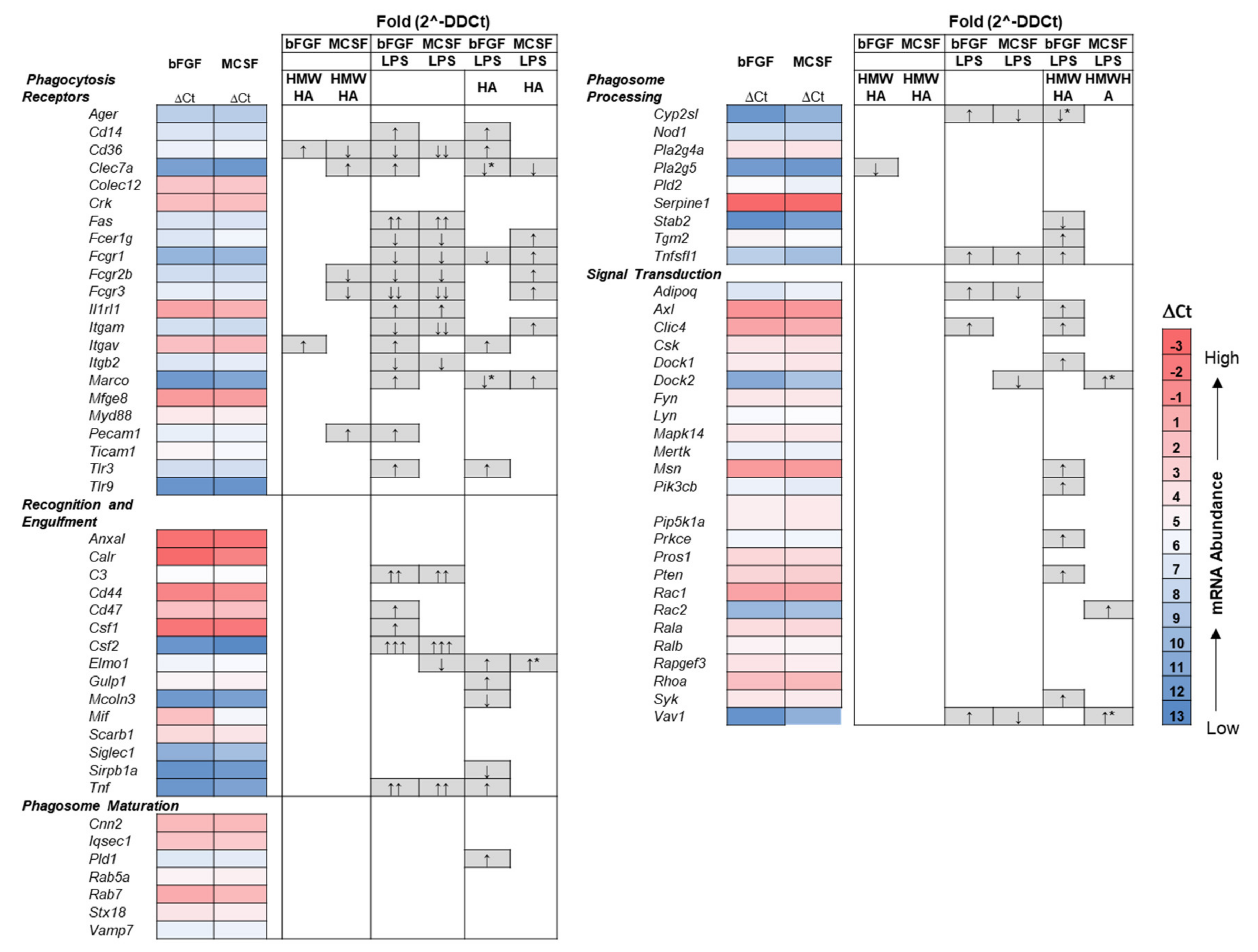
3.4. Effect of Addition of Exogenous HMW HA on TLR and Phagocytosis Pathway Genes in Basal and LPS Stimulated Cultures
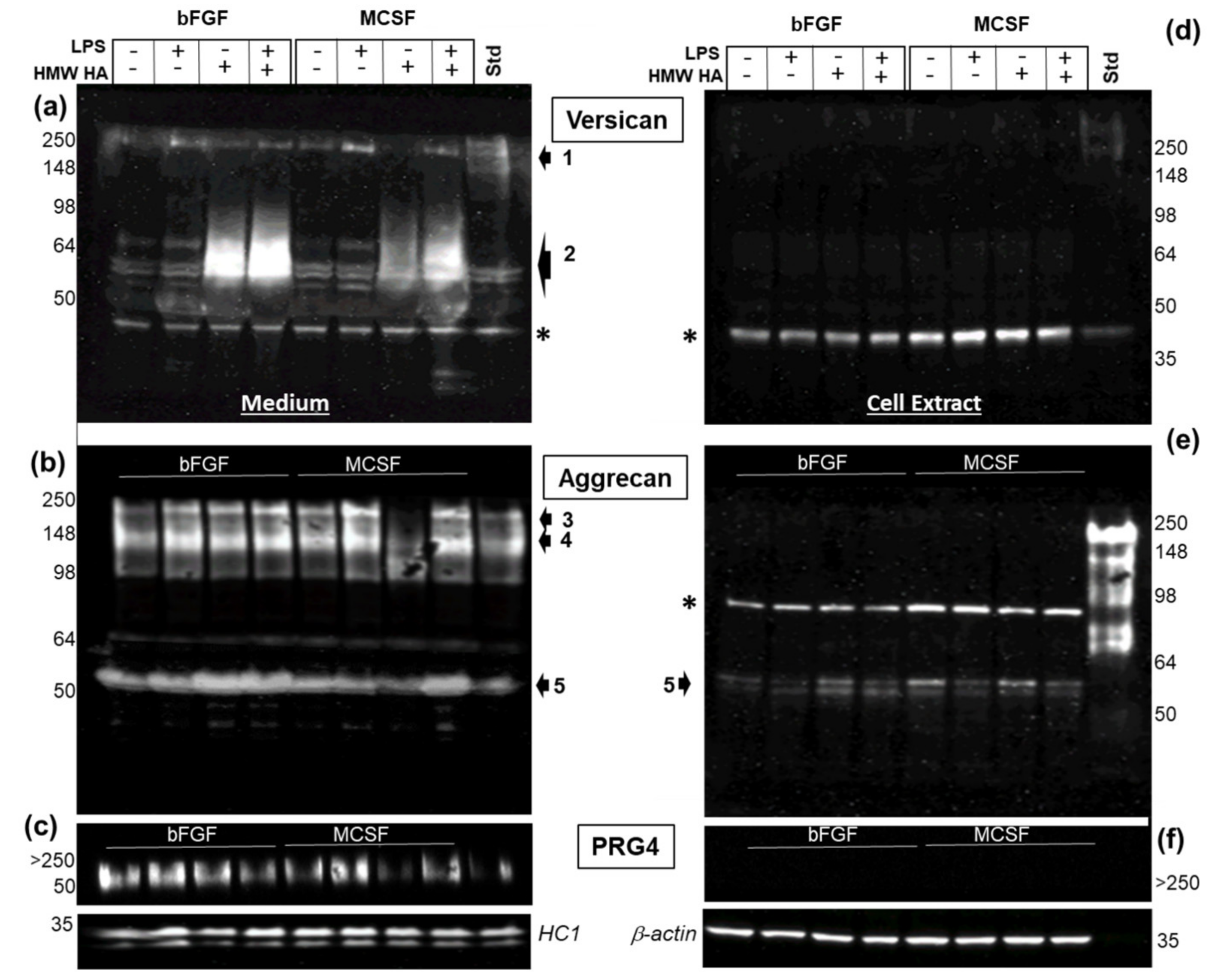
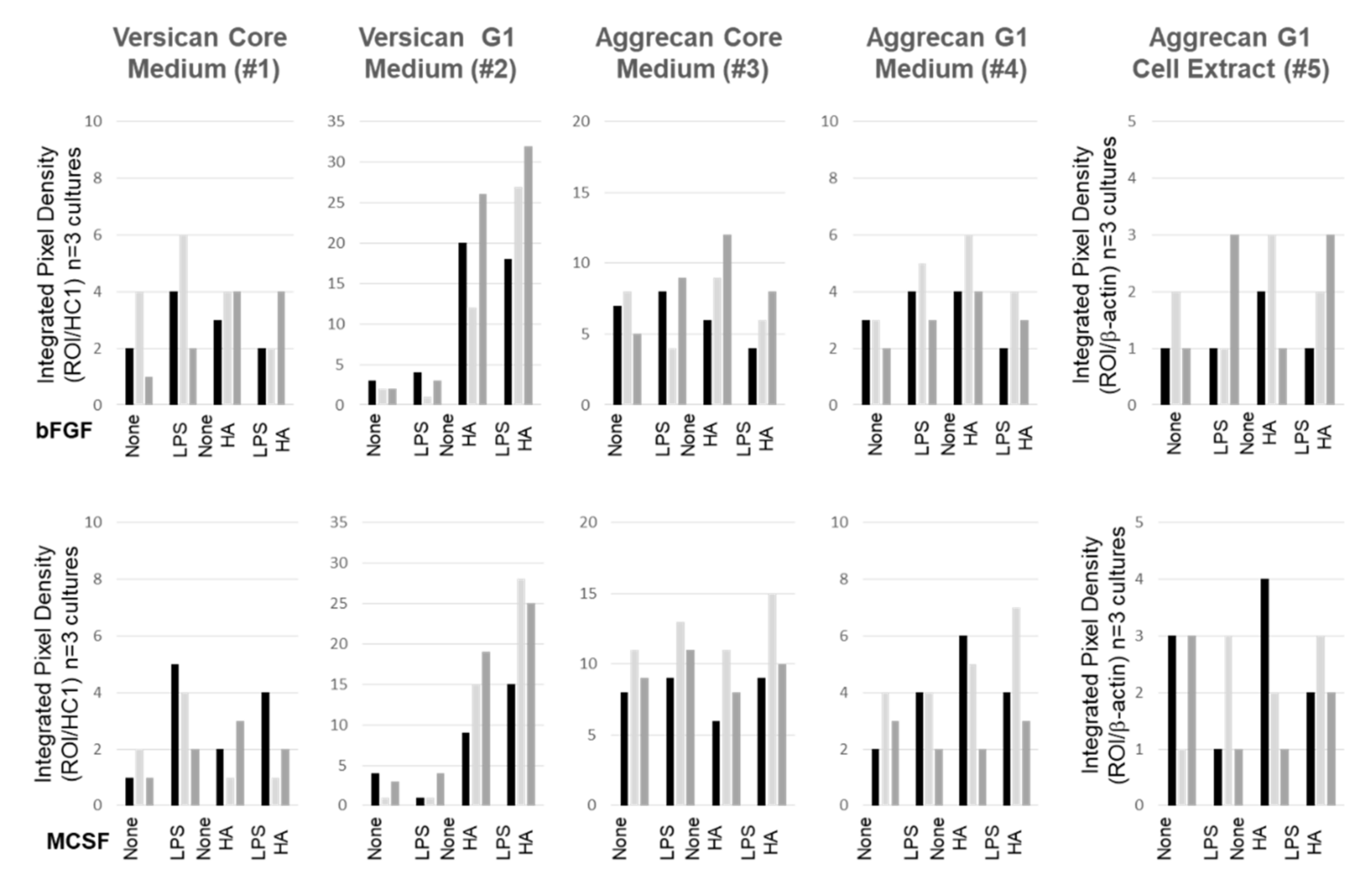
3.5. Effect of LPS and Exogenous HMW HA on VCAN, ACAN, and PRG4 Gene Expression and Protein Levels
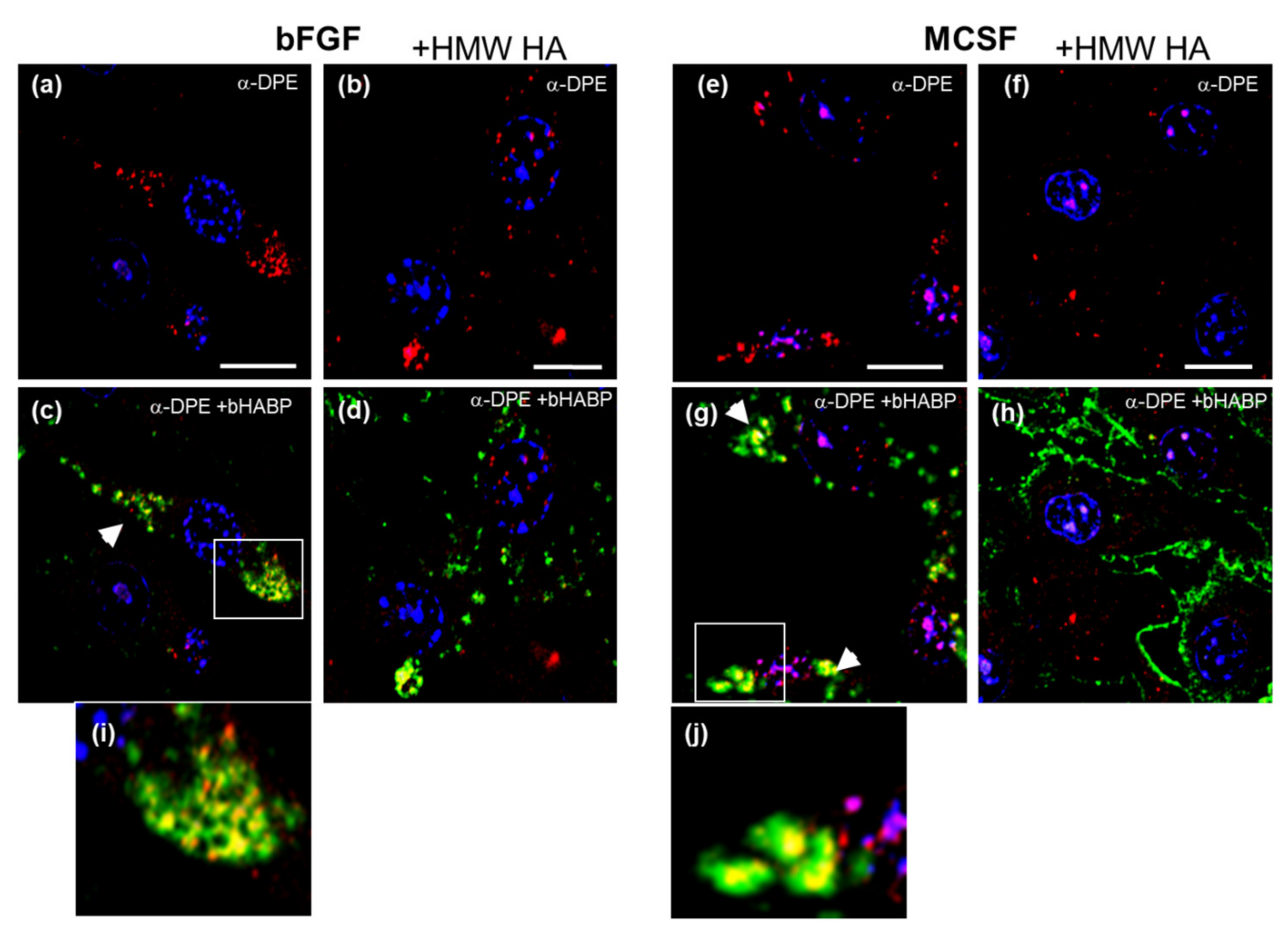
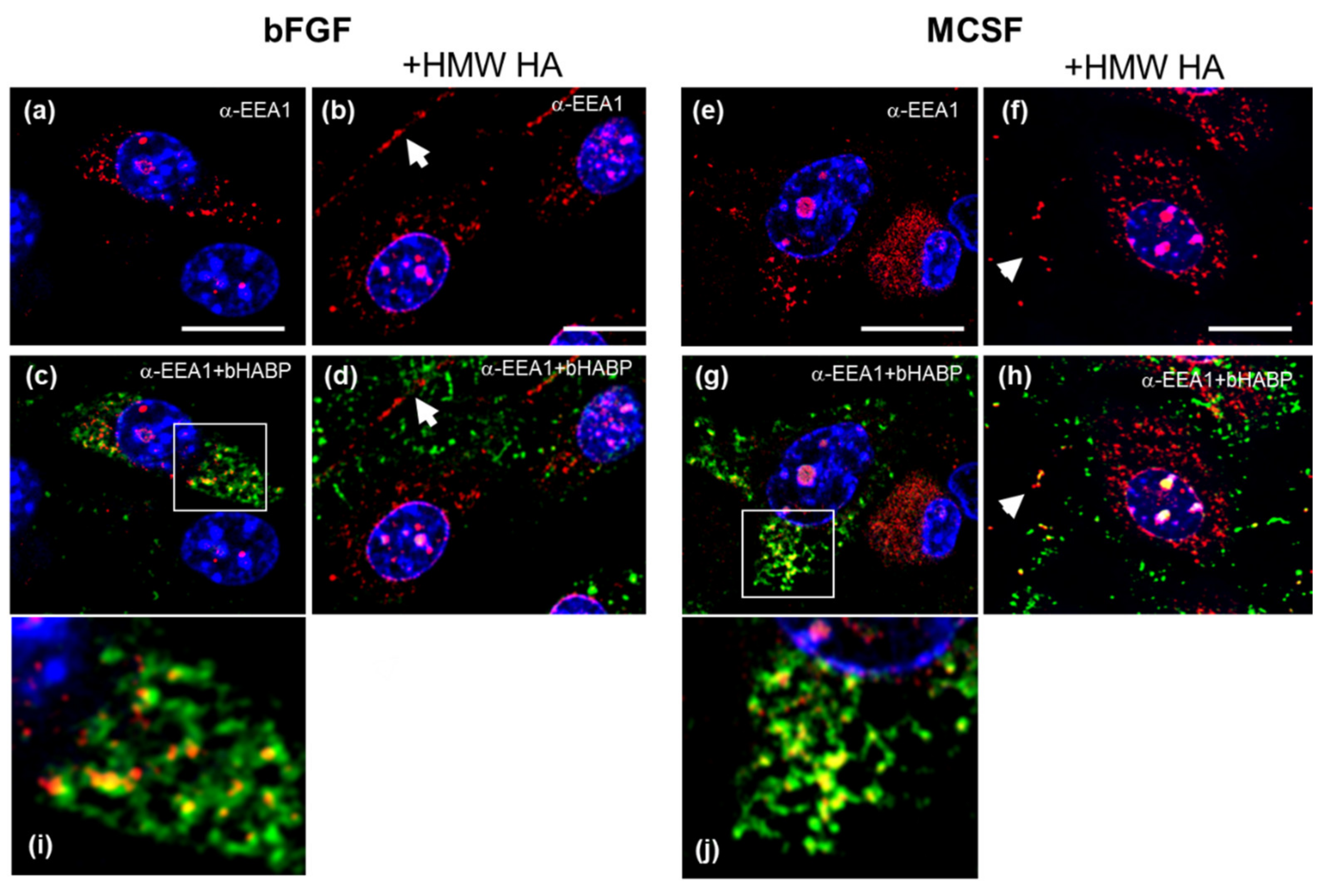
3.6. Confocal Localization of VCAN and ACAN before and after Addition of Exogenous HMW HA
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Woodell-May, J.E.; Sommerfeld, S.D. Role of Inflammation and the Immune System in the Progression of Osteoarthritis. J. Orthop. Res. 2019, 38, 253–257. [Google Scholar] [CrossRef] [PubMed]
- Carrión, M.; Frommer, K.; García, S.P.; Müller-Ladner, U.; Gomariz, R.P.; Neumann, E. The Adipokine Network in Rheumatic Joint Diseases. Int. J. Mol. Sci. 2019, 20, 4091. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Fan, X.; Xing, L.; Tian, F.-M. Wnt signaling: A promising target for osteoarthritis therapy. Cell Commun. Signal. 2019, 17, 97–114. [Google Scholar] [CrossRef] [PubMed]
- Tang, C.; Chen, Y.; Huang, J.; Zhao, K.; Chen, X.; Yin, Z.; Heng, B.C.; Chen, W.; Shen, W. The roles of inflammatory mediators and immunocytes in tendinopathy. J. Orthop. Transl. 2018, 14, 23–33. [Google Scholar] [CrossRef]
- Morita, W.; Dakin, S.G.; Snelling, S.; Carr, A.J. Cytokines in tendon disease. Bone Jt. Res. 2018, 6, 656–664. [Google Scholar] [CrossRef]
- Tarbell, J.M.; Cancel, L.M. The glycocalyx and its significance in human medicine. J. Intern. Med. 2016, 280, 97–113. [Google Scholar] [CrossRef]
- Uchimido, R.; Schmidt, E.P.; Shapiro, N.I. The glycocalyx: A novel diagnostic and therapeutic target in sepsis. Crit. Care 2019, 23, 16. [Google Scholar] [CrossRef]
- Sieve, I.; Münster-Kühnel, A.K.; Hilfiker-Kleiner, D. Regulation and function of endothelial glycocalyx layer in vascular diseases. Vasc. Pharmacol. 2018, 100, 26–33. [Google Scholar] [CrossRef]
- He, H.; Kuriyan, A.E.; Su, C.-W.; Mahabole, M.; Zhang, Y.; Zhu, Y.-T.; Flynn, H.W.; Parel, J.-M.; Tseng, S.C.G. Inhibition of Proliferation and Epithelial Mesenchymal Transition in Retinal Pigment Epithelial Cells by Heavy Chain-Hyaluronan/Pentraxin 3. Sci. Rep. 2017, 7, 43736. [Google Scholar] [CrossRef]
- Johnson, P.; Arif, A.A.; Lee-Sayer, S.S.M.; Dong, Y. Hyaluronan and Its Interactions With Immune Cells in the Healthy and Inflamed Lung. Front. Immunol. 2018, 9. [Google Scholar] [CrossRef]
- Grandoch, M.; Bollyky, P.L.; Fischer, J.W. Hyaluronan: A Master Switch between Vascular Homeostasis and Inflammation. Circ. Res. 2018, 122, 1341–1343. [Google Scholar] [CrossRef] [PubMed]
- Petrey, A.C.; De La Motte, C.A. Hyaluronan in inflammatory bowel disease: Cross-linking inflammation and coagulation. Matrix Boil. 2019, 79, 314–323. [Google Scholar] [CrossRef] [PubMed]
- Day, A.J.; Milner, C. TSG-6: A multifunctional protein with anti-inflammatory and tissue-protective properties. Matrix Boil. 2019, 79, 60–83. [Google Scholar] [CrossRef]
- Lamas, A.; Marshburn, J.; Stober, V.P.; Donaldson, S.H.; Garantziotis, S. Effects of inhaled high-molecular weight hyaluronan in inflammatory airway disease. Respir. Res. 2016, 17, 123. [Google Scholar] [CrossRef]
- Cantor, J.; Ma, S.; Turino, G. A pilot clinical trial to determine the safety and efficacy of aerosolized hyaluronan as a treatment for COPD. Int. J. Chronic Obstr. Pulm. Dis. 2017, 12, 2747–2752. [Google Scholar] [CrossRef] [PubMed]
- Richardson, C.; Plaas, A.; Block, J.A. Intra-articular Hyaluronan Therapy for Symptomatic Knee Osteoarthritis. Rheum. Dis. Clin. N. Am. 2019, 45, 439–451. [Google Scholar] [CrossRef] [PubMed]
- Altman, R.D.; Manjoo, A.; Fierlinger, A.; Niazi, F.; Nicholls, M. The mechanism of action for hyaluronic acid treatment in the osteoarthritic knee: A systematic review. Bmc Musculoskelet. Disord. 2015, 16, 321. [Google Scholar] [CrossRef]
- Ohtsuki, T.; Asano, K.; Inagaki, J.; Shinaoka, A.; Kumagishi-Shinaoka, K.; Cilek, M.Z.; Hatipoglu, O.F.; Oohashi, T.; Nishida, K.; Komatsubara, I.; et al. High molecular weight hyaluronan protects cartilage from degradation by inhibiting aggrecanase expression. J. Orthop. Res. 2018, 36, 3247–3255. [Google Scholar] [CrossRef]
- Ikeuchi, M.; Izumi, M.; Aso, K.; Sugimura, N.; Kato, T.; Tani, T. Effects of intra-articular hyaluronic acid injection on immunohistochemical characterization of joint afferents in a rat model of knee osteoarthritis. Eur. J. Pain 2015, 19, 334–340. [Google Scholar] [CrossRef]
- Muramatsu, Y.; Sasho, T.; Saito, M.; Yamaguchi, S.; Akagi, R.; Mukoyama, S.; Akatsu, Y.; Katsuragi, J.; Fukawa, T.; Endo, J.; et al. Preventive effects of hyaluronan from deterioration of gait parameters in surgically induced mice osteoarthritic knee model. Osteoarthr. Cartil. 2014, 22, 831–835. [Google Scholar] [CrossRef]
- Li, J.; Gorski, D.J.; Anemaet, W.; Velasco, J.; Takeuchi, J.; Sandy, J.D.; Plaas, A. Hyaluronan injection in murine osteoarthritis prevents TGFbeta 1-induced synovial neovascularization and fibrosis and maintains articular cartilage integrity by a CD44-dependent mechanism. Arthritis Res. Ther. 2012, 14, R151. [Google Scholar] [CrossRef] [PubMed]
- Ariyoshi, W.; Okinaga, T.; Knudson, C.B.; Knudson, W.; Nishihara, T. High molecular weight hyaluronic acid regulates osteoclast formation by inhibiting receptor activator of NF-κB ligand through Rho kinase. Osteoarthr. Cartil. 2014, 22, 111–120. [Google Scholar] [CrossRef]
- Yasuda, T. Hyaluronan inhibits cytokine production by lipopolysaccharide-stimulated U937 macrophages through down-regulation of NF-κB via ICAM-1. Inflamm. Res. 2007, 56, 246–253. [Google Scholar] [CrossRef] [PubMed]
- Sheehan, K.M.; De Lott, L.; Day, S.M.; DeHeer, D.H. Hyalgan® has a dose-dependent differential effect on macrophage proliferation and cell death. J. Orthop. Res. 2003, 21, 744–751. [Google Scholar] [CrossRef]
- Yasuda, T. Hyaluronan Inhibits Prostaglandin E2 Production via CD44 in U937 Human Macrophages. Tohoku J. Exp. Med. 2010, 220, 229–235. [Google Scholar] [CrossRef] [PubMed]
- Rayahin, J.E.; Buhrman, J.S.; Zhang, Y.; Koh, T.J.; Gemeinhart, R.A. High and Low Molecular Weight Hyaluronic Acid Differentially Influence Macrophage Activation. Acs Biomater. Sci. Eng. 2015, 1, 481–493. [Google Scholar] [CrossRef]
- Kumari, A.; Silakari, O.; Singh, R.K. Recent advances in colony stimulating factor-1 receptor/c-FMS as an emerging target for various therapeutic implications. Biomed. Pharmacother. 2018, 103, 662–679. [Google Scholar] [CrossRef]
- Celik, I.; Krack, W.; Zeiler, T.; Kretschmer, V.; Solinas, S.; Gajek, H.; Lorenz, W. Plasma histamine levels during plasmapheresis: Difficult interpretation of adverse reactions to plasma substitutes. Inflamm. Res. 2001, 50, 65–67. [Google Scholar] [CrossRef]
- Ota, T.; Urakawa, H.; Kozawa, E.; Ikuta, K.; Hamada, S.; Tsukushi, S.; Shimoyama, Y.; Ishiguro, N.; Nishida, Y. Expression of colony-stimulating factor 1 is associated with occurrence of osteochondral change in pigmented villonodular synovitis. Tumor Boil. 2015, 36, 5361–5367. [Google Scholar] [CrossRef]
- Millerand, M.; Berenbaum, F.; Jacques, C. Danger signals and inflammaging in osteoarthritis. Clin. Exp. Rheumatol 2019, 120, 48–56. [Google Scholar]
- Scanzello, C.R.; Goldring, S.R. The role of synovitis in osteoarthritis pathogenesis. Bone 2012, 51, 249–257. [Google Scholar] [CrossRef]
- Neuenschwander, H.M.; Moreira, J.J.; Vendruscolo, C.P.; Fülber, J.; Seidel, S.R.T.; Michelacci, Y.M.; Baccarin, R.Y.A. Hyaluronic acid has chondroprotective and joint-preserving effects on LPS-induced synovitis in horses. J. Ve. Sci. 2019, 20. [Google Scholar] [CrossRef]
- Kim, S.E.; Lee, J.Y.; Shim, K.-S.; Lee, S.; Min, K.; Bae, J.-H.; Kim, H.J.; Park, K.; Song, H.-R. Attenuation of inflammation and cartilage degradation by sulfasalazine-containing hyaluronic acid on osteoarthritis rat model. Int. J. Boil. Macromol. 2018, 114, 341–348. [Google Scholar] [CrossRef] [PubMed]
- Scanzello, C.R.; Plaas, A.; Crow, M.K. Innate immune system activation in osteoarthritis: Is osteoarthritis a chronic wound? Curr. Opin. Rheumatol. 2008, 20, 565–572. [Google Scholar] [CrossRef] [PubMed]
- Gorski, D.J.; Xiao, W.; Li, J.; Luo, W.; Lauer, M.; Kisiday, J.; Plaas, A.; Sandy, J. Deletion of ADAMTS5 does not affect aggrecan or versican degradation but promotes glucose uptake and proteoglycan synthesis in murine adipose derived stromal cells. Matrix Boil. 2015, 47, 66–84. [Google Scholar] [CrossRef]
- Plaas, A.; Osborn, B.; Yoshihara, Y.; Bai, Y.; Bloom, T.; Nelson, F.; Mikecz, K.; Sandy, J. Aggrecanolysis in human osteoarthritis: Confocal localization and biochemical characterization of ADAMTS5–hyaluronan complexes in articular cartilages. Osteoarthr. Cartil. 2007, 15, 719–734. [Google Scholar] [CrossRef][Green Version]
- Schmidt, T.A.; Plaas, A.H.; Sandy, J.D. Disulfide-bonded multimers of proteoglycan 4 (PRG4) are present in normal synovial fluids. Biochim. Et Biophys. Acta (Bba) – Gen. Subj. 2009, 1790, 375–384. [Google Scholar] [CrossRef] [PubMed]
- Yoshihara, Y.; Plaas, A.; Osborn, B.; Margulis, A.; Nelson, F.; Stewart, M.; Rugg, M.; Milner, C.; Day, A.J.; Nemoto, K.; et al. Superficial zone chondrocytes in normal and osteoarthritic human articular cartilages synthesize novel truncated forms of inter-alpha-trypsin inhibitor heavy chains which are attached to a chondroitin sulfate proteoglycan other than bikunin. Osteoarthr. Cartil. 2008, 16, 1343–1355. [Google Scholar] [CrossRef] [PubMed]
- Skandalis, S.S.; Karalis, T.; Heldin, P. Intracellular hyaluronan: Importance for cellular functions. Semin. Cancer Boil. 2020, 62, 20–30. [Google Scholar] [CrossRef]
- Wang, A.; Ren, J.; Wang, C.P.; Hascall, V.C. Heparin Prevents Intracellular Hyaluronan Synthesis and Autophagy Responses in Hyperglycemic Dividing Mesangial Cells and Activates Synthesis of an Extensive Extracellular Monocyte-adhesive Hyaluronan Matrix after Completing Cell Division. J. Boil. Chem. 2014, 289, 9418–9429. [Google Scholar] [CrossRef]
- Knudson, W.; Chow, G.; Knudson, C.B. CD44-mediated uptake and degradation of hyaluronan. Matrix Boil. 2002, 21, 15–23. [Google Scholar] [CrossRef]
- Shenkman, M.; Lederkremer, G. Compartmentalization and Selective Tagging for Disposal of Misfolded Glycoproteins. Trends Biochem. Sci. 2019, 44, 827–836. [Google Scholar] [CrossRef] [PubMed]
- Fraser, J.R.E.; Laurent, T.C. Hyaluronan: Its nature, distribution, functions and turnover. J. Intern. Med. 1997, 242, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Petrey, A.C.; De La Motte, C.A. Thrombin Cleavage of Inter-α-inhibitor Heavy Chain 1 Regulates Leukocyte Binding to an Inflammatory Hyaluronan Matrix. J. Boil. Chem. 2016, 291, 24324–24334. [Google Scholar] [CrossRef] [PubMed]
- Ferrandez, E.; Gutierrez, O.; Segundo, D.S.; Fernández-Luna, J.L. NFκB activation in differentiating glioblastoma stem-like cells is promoted by hyaluronic acid signaling through TLR4. Sci. Rep. 2018, 8, 6341. [Google Scholar] [CrossRef]
- Jiang, D.; Liang, J.; Fan, J.; Yu, S.; Chen, S.; Luo, Y.; Prestwich, G.D.; Mascarenhas, M.M.; Garg, H.G.; A Quinn, D.; et al. Regulation of lung injury and repair by Toll-like receptors and hyaluronan. Nat. Med. 2005, 11, 1173–1179. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Potts-Kant, E.N.; Garantziotis, S.; Foster, W.M.; Hollingsworth, J.W. Hyaluronan Signaling during Ozone-Induced Lung Injury Requires TLR4, MyD88, and TIRAP. PLoS ONE 2011, 6, e27137. [Google Scholar] [CrossRef]
- Lv, J.; He, X.; Wang, H.; Wang, Z.; Kelly, G.T.; Wang, X.; Chen, Y.; Wang, T.; Qian, Z. TLR4-NOX2 axis regulates the phagocytosis and killing of Mycobacterium tuberculosis by macrophages. Bmc Pulm. Med. 2017, 17, 194. [Google Scholar] [CrossRef]
- Blander, J.M. Signalling and phagocytosis in the orchestration of host defence. Cell. Microbiol. 2007, 9, 290–299. [Google Scholar] [CrossRef]
- Merrilees, M.; Zuo, N.; Evanko, S.P.; Day, A.J.; Wight, T.N. G1 Domain of Versican Regulates Hyaluronan Organization and the Phenotype of Cultured Human Dermal Fibroblasts. J. Histochem. Cytochem. 2016, 64, 353–363. [Google Scholar] [CrossRef]
- Velasco, J.; Li, J.; DiPietro, L.; Stepp, M.A.; Sandy, J.D.; Plaas, A. Adamts5 Deletion Blocks Murine Dermal Repair through CD44-mediated Aggrecan Accumulation and Modulation of Transforming Growth Factor β1 (TGFβ1) Signaling. J. Boil. Chem. 2011, 286, 26016–26027. [Google Scholar] [CrossRef] [PubMed]
- Ali, L.; Flowers, S.A.; Jin, C.; Bennett, E.P.; Ekwall, A.-K.H.; Karlsson, N.G. The O-glycomap of lubricin, a novel mucin responsible for joint lubrication, identified by site-specific glycopeptide analysis. Mol. Cell. Proteom. 2014, 13, 3396–3409. [Google Scholar] [CrossRef] [PubMed]
- Raposo, G.; Stoorvogel, W. Extracellular vesicles: Exosomes, microvesicles, and friends. J. Cell Boil. 2013, 200, 373–383. [Google Scholar] [CrossRef]
- Chan, D.; Xiao, W.; Li, J.; De La Motte, C.A.; Sandy, J.D.; Plaas, A. Deficiency of hyaluronan synthase 1 (Has1) results in chronic joint inflammation and widespread intra-articular fibrosis in a murine model of knee joint cartilage damage. Osteoarthr. Cartil. 2015, 23, 1879–1889. [Google Scholar] [CrossRef]
- Macchi, V.; Stocco, E.; Stecco, C.; Belluzzi, E.; Favero, M.; Porzionato, A.; De Caro, R. The infrapatellar fat pad and the synovial membrane: An anatomo-functional unit. J. Anat. 2018, 233, 146–154. [Google Scholar] [CrossRef]
- Polly, S.S.; Nichols, A.E.C.; Donnini, E.; Inman, D.J.; Scott, T.J.; Apple, S.M.; Werre, S.R.; Dahlgren, L.A. Adipose-Derived Stromal Vascular Fraction and Cultured Stromal Cells as Trophic Mediators for Tendon Healing. J. Orthop. Res. 2019, 37, 1429–1439. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Chen, S.; Pei, M. Comparative advantages of infrapatellar fat pad: An emerging stem cell source for regenerative medicine. Rheumatology 2018, 57, 2072–2086. [Google Scholar] [CrossRef]
- Hindle, P.; Khan, N.; Biant, L.; Péault, B. The Infrapatellar Fat Pad as a Source of Perivascular Stem Cells with Increased Chondrogenic Potential for Regenerative Medicine. Stem Cells Transl. Med. 2016, 6, 77–87. [Google Scholar] [CrossRef]
- Kouroupis, D.; Bowles, A.C.; Willman, M.A.; Orfei, C.P.; Colombini, A.; Best, T.M.; Kaplan, L.D.; Correa, D. Infrapatellar fat pad-derived MSC response to inflammation and fibrosis induces an immunomodulatory phenotype involving CD10-mediated Substance P degradation. Sci. Rep. 2019, 9, 10864–10916. [Google Scholar] [CrossRef]
- Sivasubramaniyan, K.; Koevoet, W.J.; Hakimiyan, A.A.; Sande, M.; Farrell, E.; Hoogduijn, M.J.; Verhaar, J.A.; Chubinskaya, S.; Bühring, H.-J.; Van Osch, G.J. Cell-surface markers identify tissue resident multipotential stem/stromal cell subsets in synovial intimal and sub-intimal compartments with distinct chondrogenic properties. Osteoarthr. Cartil. 2019, 27, 1831–1840. [Google Scholar] [CrossRef]
- Affan, A.; Al-Jezani, N.; Railton, P.; Powell, J.N.; Krawetz, R. Multiple mesenchymal progenitor cell subtypes with distinct functional potential are present within the intimal layer of the hip synovium. Bmc Musculoskelet. Disord. 2019, 20, 125. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Zheng, H.; Seol, D.; Yu, Y.; Martin, J.A. Gene expression profiles reveal that chondrogenic progenitor cells and synovial cells are closely related. J. Orthop. Res. 2014, 32, 981–988. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Newton, P.T.; Bouderlique, T.; Sejnohova, M.; Zikmund, T.; Kozhemyakina, E.; Xie, M.; Krivanek, J.; Kaiser, J.; Qian, H.; et al. Superficial cells are self-renewing chondrocyte progenitors, which form the articular cartilage in juvenile mice. Faseb J. 2016, 31, 1067–1084. [Google Scholar] [CrossRef] [PubMed]
- Chun, S.Y.; Lim, J.O.; Lee, E.H.; Han, M.-H.; Ha, Y.-S.; Lee, J.N.; Kim, B.S.; Park, M.J.; Yeo, M.; Jung, B.; et al. Preparation and Characterization of Human Adipose Tissue-Derived Extracellular Matrix, Growth Factors, and Stem Cells: A Concise Review. Tissue Eng. Regen. Med. 2019, 16, 385–393. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Tang, Y.; Song, B.; Yu, M.; Li, Q.; Zhang, C.; Hou, J.; Yang, R. Nomenclature clarification: Synovial fibroblasts and synovial mesenchymal stem cells. Stem Cell Res. Ther. 2019, 10, 260–267. [Google Scholar] [CrossRef]
- Jones, A.R.C.; Flannery, C.R. Bioregulation of lubricin expression by growth factors and cytokines. Eur. Cell Mater. 2007, 13, 40–45. [Google Scholar] [CrossRef]
- Alquraini, A.; Jamal, M.; Zhang, L.; Schmidt, T.A.; Jay, G.D.; Elsaid, K.A. The autocrine role of proteoglycan-4 (PRG4) in modulating osteoarthritic synoviocyte proliferation and expression of matrix degrading enzymes. Arthritis Res. 2017, 19, 89. [Google Scholar] [CrossRef]
- Zhou, D.; Huang, C.; Lin, Z.; Zhan, S.; Kong, L.; Fang, C.; Li, J. Macrophage polarization and function with emphasis on the evolving roles of coordinated regulation of cellular signaling pathways. Cell. Signal. 2014, 26, 192–197. [Google Scholar] [CrossRef]
- Barboza, E.; Hudson, J.; Chang, W.-P.; Kovats, S.; Towner, R.A.; Silasi, R.; Lupu, F.; Kent, C.; Griffin, T.M. Profibrotic Infrapatellar Fat Pad Remodeling Without M1 Macrophage Polarization Precedes Knee Osteoarthritis in Mice With Diet-Induced Obesity. Arthritis Rheumatol. 2017, 69, 1221–1232. [Google Scholar] [CrossRef] [PubMed]
- Abu-Hakmeh, A.E.; Fleck, A.K.M.; Wan, L.Q. Temporal effects of cytokine treatment on lubricant synthesis and matrix metalloproteinase activity of fibroblast-like synoviocytes. J. Tissue Eng. Regen. Med. 2018, 13, 87–98. [Google Scholar] [CrossRef]
- Olivotto, E.; Merli, G.; Assirelli, E.; Cavallo, C.; Belluzzi, E.; Ramonda, R.; Favero, M.; Filardo, G.; Roffi, A.; Kon, E.; et al. Cultures of a human synovial cell line to evaluate platelet-rich plasma and hyaluronic acid effects. J. Tissue Eng. Regen. Med. 2018, 12, 1835–1842. [Google Scholar] [CrossRef] [PubMed]
- Qiu, B.; Xu, X.; Deng, R.; Xia, G.; Shang, X.; Zhou, P.-H. Hyaluronic acid-chitosan nanoparticles encoding CrmA attenuate interleukin-1β induced inflammation in synoviocytes in vitro. Int. J. Mol. Med. 2018, 43. [Google Scholar] [CrossRef] [PubMed]
- Kilborne, A.H.; Hussein, H.; Bertone, A.L. Effects of hyaluronan alone or in combination with chondroitin sulfate and N. -acetyl-d.-glucosamine on lipopolysaccharide challenge-exposed equine fibroblast-like synovial cells. Am. J. Veter- Res. 2017, 78, 579–588. [Google Scholar] [CrossRef]
- Vigetti, D.; Viola, M.; Karousou, E.; Deleonibus, S.; Karamanou, K.; De Luca, G.; Passi, A. Epigenetics in extracellular matrix remodeling and hyaluronan metabolism. Febs J. 2014, 281, 4980–4992. [Google Scholar] [CrossRef]
- Vigetti, D.; Karousou, E.; Viola, M.; Deleonibus, S.; De Luca, G.; Passi, A. Hyaluronan: Biosynthesis and signaling. Biochim. Et Biophys. Acta (Bba) – Gen. Subj. 2014, 1840, 2452–2459. [Google Scholar] [CrossRef] [PubMed]
- Tammi, M.I.; Oikari, S.; Pasonen-Seppänen, S.; Rilla, K.; Auvinen, P.; Tammi, R.H. Activated hyaluronan metabolism in the tumor matrix-Causes and consequences. Matrix Boil. 2019, 79, 147–164. [Google Scholar] [CrossRef]
- Tammi, R.; Rilla, K.; Pienimäki, J.-P.; Maccallum, D.K.; Hogg, M.; Luukkonen, M.; Hascall, V.C.; Tammi, M. Hyaluronan Enters Keratinocytes by a Novel Endocytic Route for Catabolism. J. Boil. Chem. 2001, 276, 35111–35122. [Google Scholar] [CrossRef]
- Knudson, W.; Ishizuka, S.; Terabe, K.; Askew, E.B.; Knudson, C.B. The pericellular hyaluronan of articular chondrocytes. Matrix Boil. 2019, 79, 32–46. [Google Scholar] [CrossRef]
- Weigel, P.H. Discovery of the Liver Hyaluronan Receptor for Endocytosis (HARE) and Its Progressive Emergence as the Multi-Ligand Scavenger Receptor Stabilin-2. Biomolecules 2019, 9, 454. [Google Scholar] [CrossRef]
- Nakamura, S.; Yoshimori, T. New insights into autophagosome–lysosome fusion. J. Cell Sci. 2017, 130, 1209–1216. [Google Scholar] [CrossRef]
- Naslavsky, N.; Caplan, S. The enigmatic endosome-sorting the ins and outs of endocytic trafficking. J. Cell Sci. 2018, 131, jcs216499. [Google Scholar] [CrossRef] [PubMed]
- Kuznetsova, S.A.; Issa, P.; Perruccio, E.M.; Zeng, B.; Sipes, J.M.; Ward, Y.; Seyfried, N.T.; Fielder, H.L.; Day, A.J.; Wight, T.N.; et al. Versican-thrombospondin-1 binding in vitro and colocalization in microfibrils induced by inflammation on vascular smooth muscle cells. J. Cell Sci. 2006, 119, 4499–4509. [Google Scholar] [CrossRef]
- De La Motte, C.A.; Hascall, V.C.; Drazba, J.; Bandyopadhyay, S.K.; Strong, S.A. Mononuclear Leukocytes Bind to Specific Hyaluronan Structures on Colon Mucosal Smooth Muscle Cells Treated with Polyinosinic Acid:Polycytidylic Acid. Am. J. Pathol. 2003, 163, 121–133. [Google Scholar] [CrossRef]
- Wight, T.N.; Kang, I.; Merrilees, M.J. Versican and the control of inflammation. Matrix Boil. 2014, 35, 152–161. [Google Scholar] [CrossRef] [PubMed]
- Evanko, S.P.; Potter-Perigo, S.; Bollyky, P.L.; Nepom, G.T.; Wight, T.N. Hyaluronan and versican in the control of human T-lymphocyte adhesion and migration. Matrix Boil. 2011, 31, 90–100. [Google Scholar] [CrossRef] [PubMed]
- Salwowska, N.M.; Bebenek, K.A.; Żądło, D.A.; Wcisło-Dziadecka, D. Physiochemical properties and application of hyaluronic acid A systematic review. J. Cosmet. Dermatol. 2016, 15, 520–526. [Google Scholar] [CrossRef]
- Martin-Alarcon, L.; Schmidt, T.A. Rheological effects of macromolecular interactions in synovial fluid. Biorheology 2016, 53, 49–67. [Google Scholar] [CrossRef]
- McDonald, J.N.; Levick, J.R. Effect of intra-articular hyaluronan on pressure-flow relation across synovium in anaesthetized rabbits. J. Physiol. 1995, 485, 179–193. [Google Scholar] [CrossRef]
- Takeda, K. Toll-like receptors in innate immunity. Int. Immunol. 2004, 17, 1–14. [Google Scholar] [CrossRef]
- Sánchez-De-Diego, C.; Valer, J.A.; Pimenta-Lopes, C.; Rosa, J.; Ventura, F. Interplay between BMPs and Reactive Oxygen Species in Cell Signaling and Pathology. Biomolecules 2019, 9, 534. [Google Scholar] [CrossRef]
- Bourguignon, L.Y.W.; Singleton, P.A.; Diedrich, F.; Stern, R.; Gilad, E. CD44 Interaction with Na+-H+Exchanger (NHE1) Creates Acidic Microenvironments Leading to Hyaluronidase-2 and Cathepsin B Activation and Breast Tumor Cell Invasion. J. Boil. Chem. 2004, 279, 26991–27007. [Google Scholar] [CrossRef] [PubMed]
- Parolini, I.; Federici, C.; Raggi, C.; Lugini, L.; Palleschi, S.; De Milito, A.; Coscia, C.; Iessi, E.; Logozzi, M.; Molinari, A.; et al. Microenvironmental pH Is a Key Factor for Exosome Traffic in Tumor Cells*. J. Boil. Chem. 2009, 284, 34211–34222. [Google Scholar] [CrossRef] [PubMed]
- Savina, A.; Furlán, M.; Vidal, M.; Colombo, M.I.; Gamel-Didelon, K.; Kunz, L.; Föhr, K.J.; Gratzl, M.; Mayerhofer, A. Exosome Release Is Regulated by a Calcium-dependent Mechanism in K562 Cells. J. Boil. Chem. 2003, 278, 20083–20090. [Google Scholar] [CrossRef]
- A Crawford, S.; Diamond, D.; Brustolon, L.; Penarreta, R. Effect of Increased Extracellular Ca++ on Microvesicle Production and Tumor Spheroid Formation. Cancer Microenviron. 2010, 4, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Van Niel, G.; D’Angelo, G.; Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Boil. 2018, 19, 213–228. [Google Scholar] [CrossRef]
- Kim, Y.; West, G.A.; Ray, G.; Kessler, S.P.; Petrey, A.C.; Fiocchi, C.; McDonald, C.; Longworth, M.S.; Nagy, L.E.; De La Motte, C.A. Layilin is critical for mediating hyaluronan 35 kDa-induced intestinal epithelial tight junction protein ZO-1 in vitro and in vivo. Matrix Boil. 2018, 66, 93–109. [Google Scholar] [CrossRef]
- Tolg, C.; McCarthy, J.B.; Yazdani, A.; Turley, E.A. Hyaluronan and RHAMM in Wound Repair and the “Cancerization” of Stromal Tissues. Biomed. Res. Int. 2014, 2014, 1–18. [Google Scholar] [CrossRef]
- Hessvik, N.P.; Llorente, A. Current knowledge on exosome biogenesis and release. Cell. Mol. Life Sci. 2017, 75, 193–208. [Google Scholar] [CrossRef]
- Rilla, K.; Mustonen, A.-M.; Arasu, U.T.; Härkönen, K.; Matilainen, J.; Nieminen, P. Extracellular vesicles are integral and functional components of the extracellular matrix. Matrix Boil. 2019, 76, 201–219. [Google Scholar] [CrossRef]


| Gene | 4 h + bFGF | 4 h + MCSF | 36 h + bFGF | 36 h + MCSF |
|---|---|---|---|---|
| ⚜Ct * | ⚜Ct * | ⚜Ct * | ⚜Ct * | |
| Il6 | 10.88 (± 2.17) | 11.28 (± 2.54) | 12.78 (±1.55) | 13.15 (± 2.00) |
| Nos2 | ND | ND | ND | ND |
| Has 1 | 7.64 (± 1.01) | 8.11 (± 1.92) | 8.83 (±0.15) | 9.22 (± 0.93) |
| Has2 | 8.33 (± 1.48) | 8.09 (± 2.31) | 8.52 (±0.92) | 7.65 (± 1.32) |
| Ptx3 | 8.19 (± 2.56) | 6.07 (± 3.21) | 8.35 (±0.68) | 8.92 (± 1.29) |
| Tnfaip6 | 9.96 (± 2.01) | 9.28 (± 1.89) | 12.68 (±1.49) | 13.13 (± 2.19) |
| Cd44 | 1.14 (± 0.31) | 1.67 (± 0.52) | 1.26 (±0.41) | 1.41 (± 0.31) |
| Hyla1 | 9.05 (± 2.11) | 8.52 (± 2.42) | 7.33 (±0.46) | 7.12 (± 0.68) |
| Hyal2 | 5.39 (± 1.11) | 4.94 (± 0.76) | 4.34 (±0.28) | 3.97 (± 0.63) |
| Cemip | 8.81 (± 2.25) | 6.82 (± 1.13) | 7.53 (±0.52) | 7.82 (± 0.67) |
| Tmem2 | 6.26 (± 0.99) | 6.05 (± 1.26) | 9.84 (±0.69) | 8.17 (± 0.88) |
| 4 h bFGF + LPS | 4 h MCSF + LPS | +16 h bFGF | +16 h MCSF | |
|---|---|---|---|---|
| Gene | * FOLD | FOLD | FOLD | FOLD |
| Il6 | 101.5 (± 0.92) | 93.4 (± 3.44) | 155.1 (± 22.13) | 76.64 (± 11.11) |
| Nos2 | ** >>100 | >>100 | >>100 | >>100 |
| Has1 | 1.98 (± 0.21) | 2.90 (± 0.87) | 4.85 (± 0.58) | 8.90 (± 0.13) |
| Has2 | 1.62 (± 0.03) | 1.65 (± 0.11) | 0.69 (± 0.11) | 0.36 (± 0.08) |
| Ptx3 | 12.40 (± 2.33) | 5.32 (± 0.82) | 66.67 (± 4.88) | 132.5 (± 25.3) |
| Tnfaip6 | 6.09 (± 1.20) | 12.62 (± 2.11) | 4.97 (± 0.81) | 7.89 (± 0.39) |
| Cd44 | 1.04 (± 0.32) | 1.49 (± 0.46) | 0.95 (± 0.28) | 1.23 (± 0.41) |
| Hyla1 | 0.86 (± 0.14) | 0.59 (± 0.32) | 0.45 (± 0.12) | 0.28 (± 0.03) |
| Hyal2 | 0.85 (± 0.29) | 0.43 (± 0.08) | 0.28 (± 0.11) | 0.11 (± 0.06) |
| Cemip | 1.57 (± 0.22) | 0.88 (± 0.71) | 1.30 (± 0.21) | 0.78 (± 0.39) |
| Tmem2 | 1.50 (± 0.41) | 1.14 (± 0.15) | 0.24 (± 0.08) | 0.30 (± 0.06) |
| 4 h Basal Medium | 4 h LPS Treatment | |||
|---|---|---|---|---|
| 16 h bFGF + HA | 16 h MCSF + HA | 16 h bFGF + HA | 16 h MCSF + HA | |
| Gene | * FOLD | FOLD | FOLD | FOLD |
| Il6 | # 2.58 (± 1.07) | # 2.29 (± 0.84) | ### 21.4 (± 4.5) | ### 44.3 (± 5.99) |
| Nos2 | ND | ND | ### 0.03 (± 0.06) | ### 0.14 (± 0.03) |
| Has1 | 0.86 (± 0.41) | 1.38 (± 0.51) | 1.06 (± 0.21) | 1.04 (± 0.23) |
| Has2 | 1.08 (± 0.33) | 1.44 (± 0.59) | # 1.97 (± 0.31) | # 1.87 (± 0.29) |
| Ptx3 | 0.87 (± 0.64) | 1.01 (± 0.14) | 1.08 (± 0.09) | 1.13 (± 0.25) |
| Tnfaip6 | 0.58 (± 0.44) | 1.10 (± 0.61) | 1.33 (± 0.26) | 1.25(± 0.44) |
| Cd44 | 1.23 (± 0.29) | 1.14 (± 0.17) | 0.97 (± 0.31) | 1.19 (± 0.29) |
| Hyal1 | 1.12 (± 0.44) | 1.13 (± 0.22) | # 2.01 (± 0.14) | ## 2.68(± 0.27) |
| Hyal2 | 0.60 (± 0.71) | 1.27 (± 0.41) | # 1.95 (± 0.54) | ## 3.29 (± 0.98) |
| Cemip | 1.09 (± 0.32) | 0.97 (± 0.31) | 0.96 (± 0.33) | 1.15 (± 0.66) |
| Tmem2 | 0.68 (± 0.58) | 0.74 (± 0.29) | # 1.78 (± 0.11) | ## 2.32 (± 0.28) |
| Gene | 4 h + bFGF | 4 h + MCSF | 36 h + bFGF | 36 h + MCSF |
|---|---|---|---|---|
| ⚜Ct * | ⚜Ct * | ⚜Ct * | ⚜Ct * | |
| Acan | 9.12 (± 1.1) | 6.93 (± 0.82) | 8.97 (± 1.6) | 6.42 (± 0.61) |
| Vcan | 6.91 (± 0.48) | 4.76 (± 1.01) | 6.29 (± 0.57) | 4.74 (± 2.01) |
| Prg4 | 3.55 (± 0.51) | 3.92 (± 0.21) | 3.67 (± 0.86) | 3.19 (± 0.20) |
| Gene | 4 h bFGF + LPS | 4 h MCSF + LPS | + 16 h bFGF | + 16 h MCSF |
|---|---|---|---|---|
| * FOLD | FOLD | FOLD | FOLD | |
| Acan | ## 0.24 (± 0.11) | ## 0.29 (± 0.03) | ## 0.31 (± 0.08) | ## 0.09 (± 0.03) |
| Vcan | 0.84 (± 0.37) | 1.44 (± 0.38) | # 2.44 (± 0.41) | 1.03 (± 0.38) |
| Prg4 | 0.74 (± 0.18) | 0.78 (± 0.34) | # 2.07 (± 4.88) | 0.94 (± 25.3) |
| Gene | bFGF + HA | MCSF + HA | bFGF + LPS + HA | MCSF + LPS + HA |
|---|---|---|---|---|
| * FOLD | FOLD | FOLD | FOLD | |
| Acan | 1.64 (± 0.22) | 0.62 (± 0.62) | 1.56 (± 0.51) | 2.26 (± 0.61) |
| Vcan | 1.76 (± 0.21) | 2.59 (± 0.11) | 0.78 (± 0.32) | 0.40 (± 0.63) |
| Prg4 | 1.44 (± 0.25) | 1.56 (± 0.31) | 1.04 (± 0.08) | 1.69 (± 0.21) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xue, J.; Chen, J.; Shen, Q.; Chan, D.; Li, J.; Tanguay, A.P.; Schmidt, T.A.; Niazi, F.; Plaas, A. Addition of High Molecular Weight Hyaluronic Acid to Fibroblast-Like Stromal Cells Modulates Endogenous Hyaluronic Acid Metabolism and Enhances Proteolytic Processing and Secretion of Versican. Cells 2020, 9, 1681. https://doi.org/10.3390/cells9071681
Xue J, Chen J, Shen Q, Chan D, Li J, Tanguay AP, Schmidt TA, Niazi F, Plaas A. Addition of High Molecular Weight Hyaluronic Acid to Fibroblast-Like Stromal Cells Modulates Endogenous Hyaluronic Acid Metabolism and Enhances Proteolytic Processing and Secretion of Versican. Cells. 2020; 9(7):1681. https://doi.org/10.3390/cells9071681
Chicago/Turabian StyleXue, Jiapeng, Jinnan Chen, Quan Shen, Deva Chan, Jun Li, Adam P. Tanguay, Tannin A. Schmidt, Faizan Niazi, and Anna Plaas. 2020. "Addition of High Molecular Weight Hyaluronic Acid to Fibroblast-Like Stromal Cells Modulates Endogenous Hyaluronic Acid Metabolism and Enhances Proteolytic Processing and Secretion of Versican" Cells 9, no. 7: 1681. https://doi.org/10.3390/cells9071681
APA StyleXue, J., Chen, J., Shen, Q., Chan, D., Li, J., Tanguay, A. P., Schmidt, T. A., Niazi, F., & Plaas, A. (2020). Addition of High Molecular Weight Hyaluronic Acid to Fibroblast-Like Stromal Cells Modulates Endogenous Hyaluronic Acid Metabolism and Enhances Proteolytic Processing and Secretion of Versican. Cells, 9(7), 1681. https://doi.org/10.3390/cells9071681






