Emerging Roles of Post-Translational Modifications in Nucleotide Excision Repair
Abstract
1. Introduction
2. Ubiquitylation-Mediated Processes During Nucleotide Excision Repair
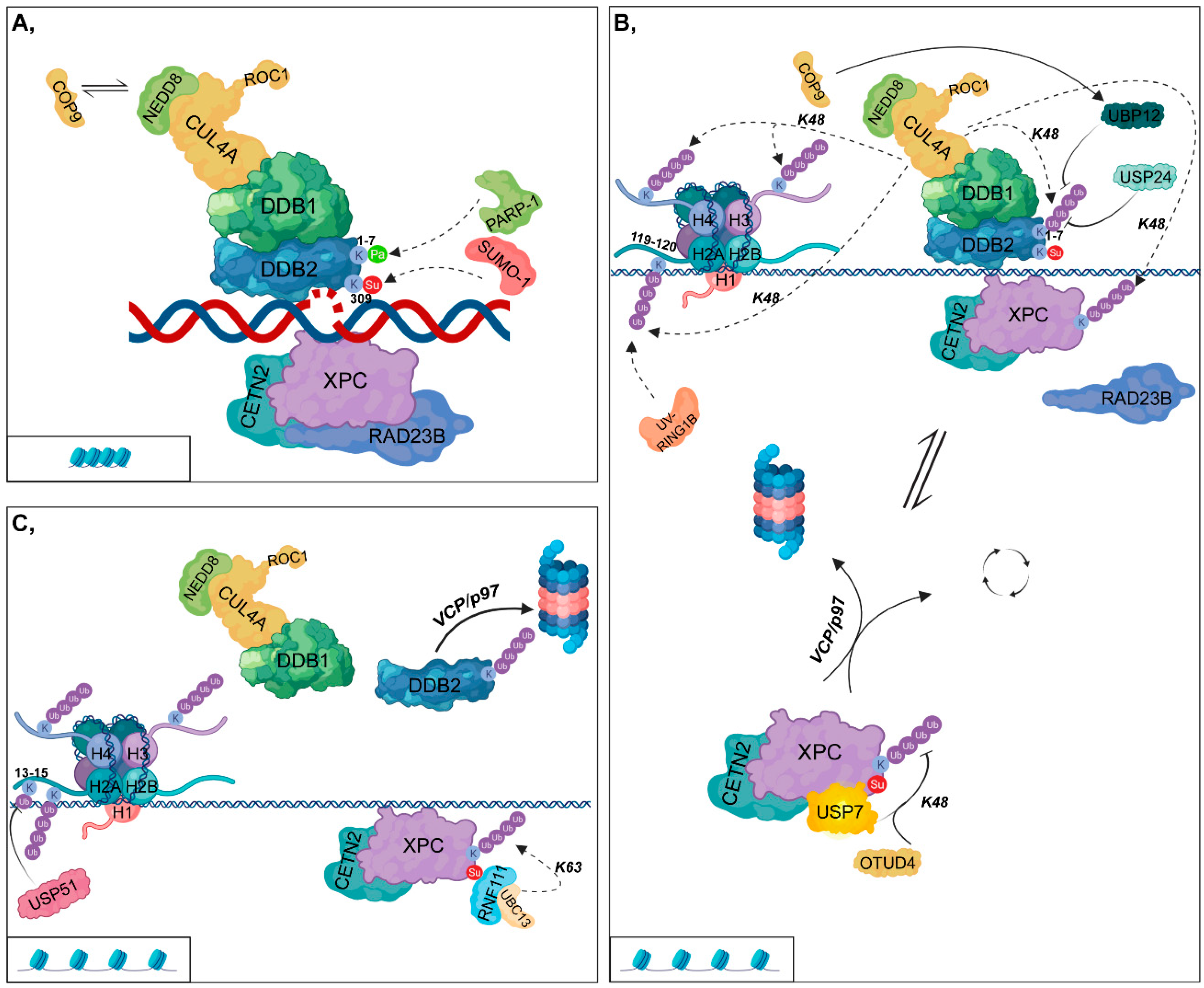
2.1. Global Genomic-Nucleotide Excision Repair
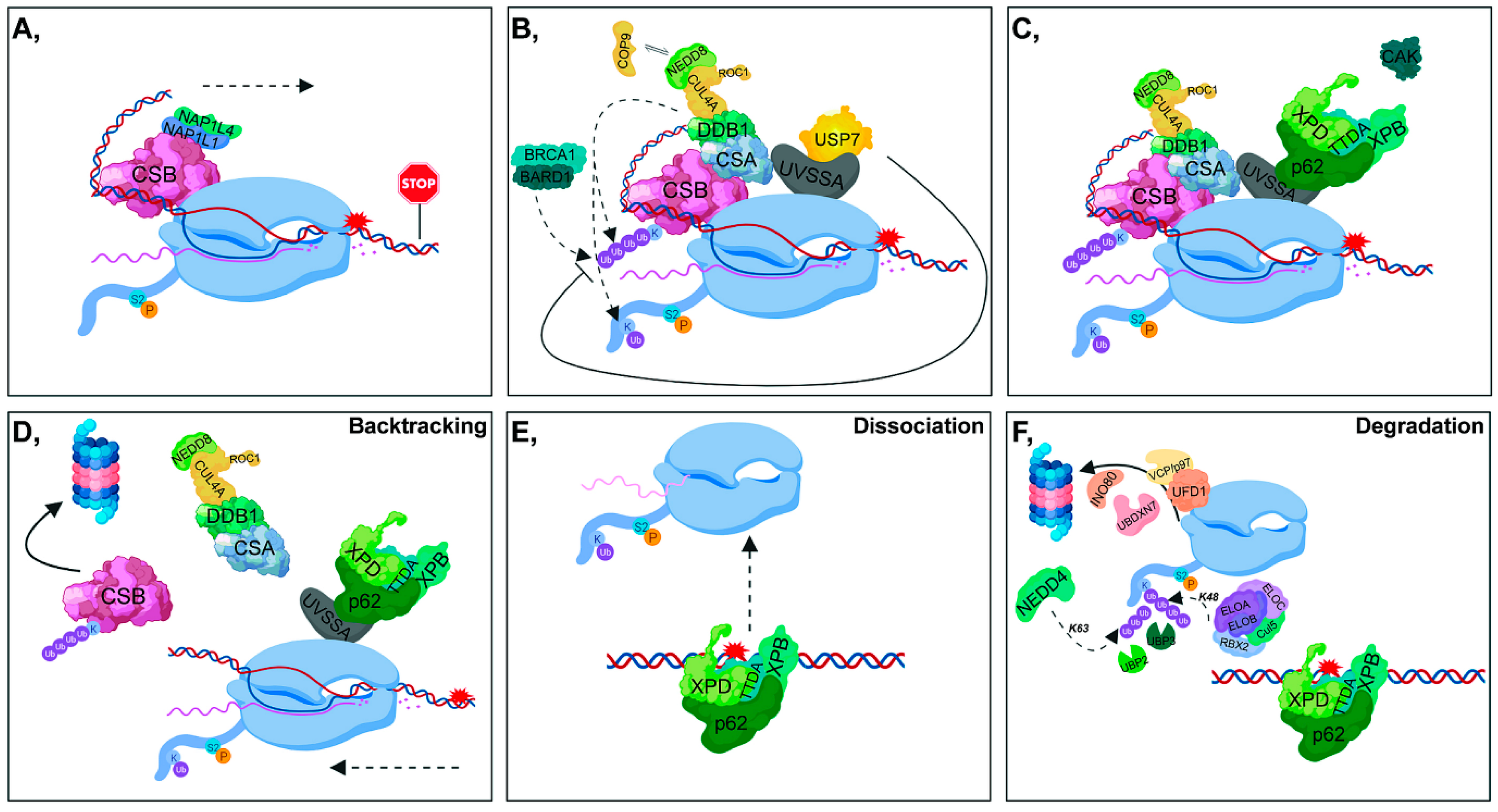
2.2. Transcription-Coupled-Nucleotide Excision Repair
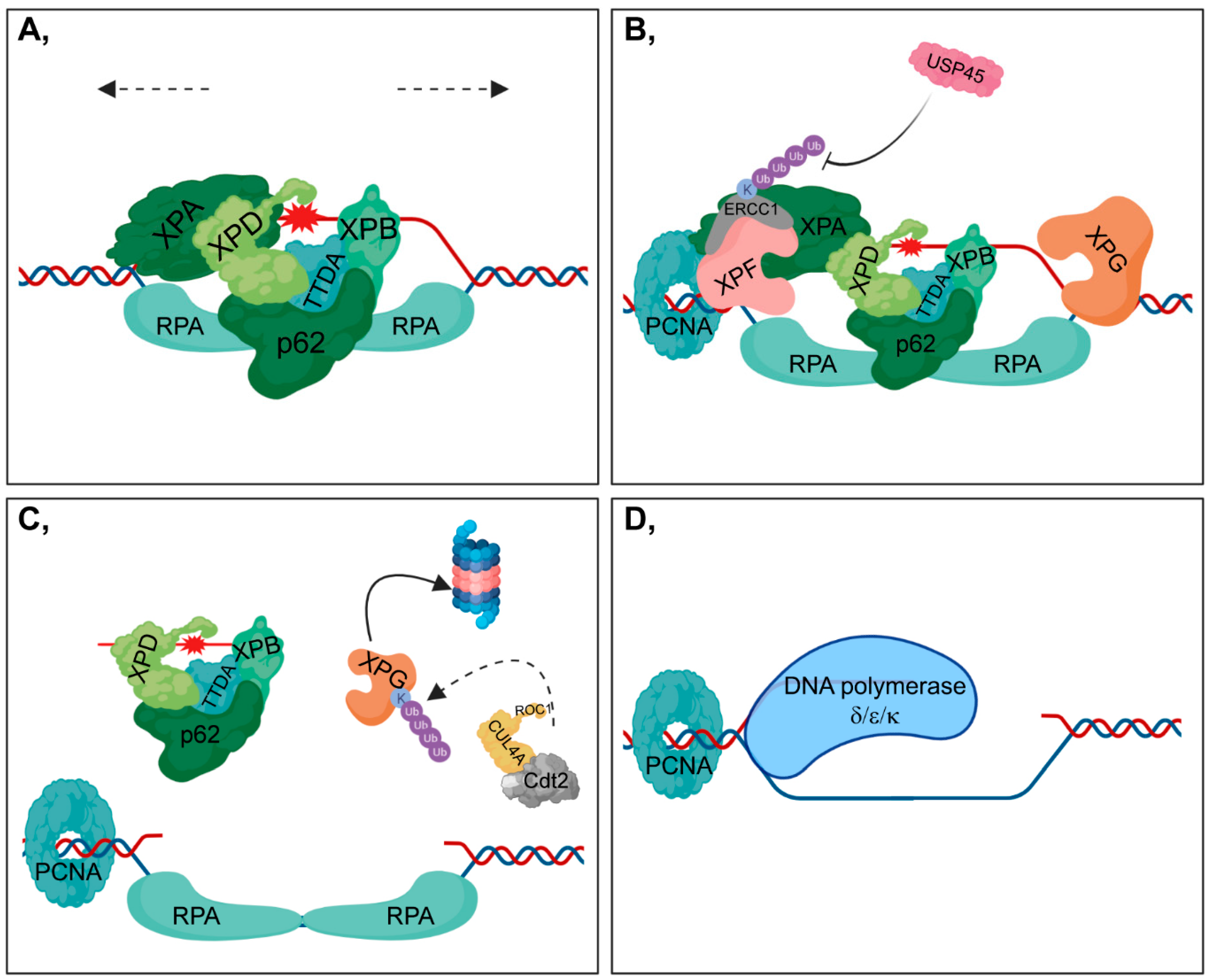
2.3. DNA Damage Verification and Repair During NER
3. Precise Coordination of Ubiquitin-Mediated Removal of RNAPII upon Transcription Blockage
4. Concluding Remarks and Future Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Bohr, V. DNA repair in an active gene: Removal of pyrimidine dimers from the DHFR gene of CHO cells is much more efficient than in the genome overall. Cell 1985, 40, 359–369. [Google Scholar] [CrossRef]
- Mellon, I. Selective removal of transcription-blocking DNA damage from the transcribed strand of the mammalian DHFR gene. Cell 1987, 51, 241–249. [Google Scholar] [CrossRef]
- Hershko, A.; Ciechanover, A. THE UBIQUITIN SYSTEM. Annu. Rev. Biochem. 1998, 67, 425–479. [Google Scholar] [CrossRef] [PubMed]
- Kulathu, Y.; Komander, D. Atypical ubiquitylation — The unexplored world of polyubiquitin beyond Lys48 and Lys63 linkages. Nat. Rev. Mol. Cell Boil. 2012, 13, 508–523. [Google Scholar] [CrossRef]
- Wei, L.; Lan, L.; Yasui, A.; Tanaka, K.; Saijo, M.; Matsuzawa, A.; Kashiwagi, R.; Maseki, E.; Hu, Y.; Parvin, J.D.; et al. BRCA1 contributes to transcription-coupled repair of DNA damage through polyubiquitination and degradation of Cockayne syndrome B protein. Cancer Sci. 2011, 102, 1840–1847. [Google Scholar] [CrossRef]
- Harreman, M.; Taschner, M.; Sigurdsson, S.; Anindya, R.; Reid, J.; Somesh, B.; Kong, S.E.; Banks, C.A.S.; Conaway, R.C.; Conaway, J.W.; et al. Distinct ubiquitin ligases act sequentially for RNA polymerase II polyubiquitylation. Proc. Natl. Acad. Sci. USA 2009, 106, 20705–20710. [Google Scholar] [CrossRef] [PubMed]
- Anindya, R.; Aygün, O.; Svejstrup, J.Q. Damage-Induced Ubiquitylation of Human RNA Polymerase II by the Ubiquitin Ligase Nedd4, but Not Cockayne Syndrome Proteins or BRCA1. Mol. Cell 2007, 28, 386–397. [Google Scholar] [CrossRef]
- Wilson, M.D.; Harreman, M.; Svejstrup, J.Q. Ubiquitylation and degradation of elongating RNA polymerase II: The last resort. Biochim. Biophys. Acta Bioenerg. 2013, 1829, 151–157. [Google Scholar] [CrossRef]
- Hottiger, M.O. Nuclear ADP-Ribosylation and Its Role in Chromatin Plasticity, Cell Differentiation, and Epigenetics. Annu. Rev. Biochem. 2015, 84, 227–263. [Google Scholar] [CrossRef]
- D’Amours, D.; Desnoyers, S.; D’Silva, I.; Poirier, G.G. Poly(adp-ribosyl)ation reactions in the regulation of nuclear functions. Biochem. J. 1999, 342, 249–268. [Google Scholar] [CrossRef]
- Juarez-Salinas, H.; Levi, V.; Jacobson, E.L.; Jacobson, M.K. Poly(ADP-ribose) has a branched structure in vivo. J. Boil. Chem. 1982, 257, 607–609. [Google Scholar]
- Bai, P.; Canto, C.; Oudart, H.; Brunyánszki, A.; Cen, Y.; Thomas, C.; Yamamoto, H.; Huber, A.; Kiss, B.; Houtkooper, R.H.; et al. PARP-1 Inhibition Increases Mitochondrial Metabolism through SIRT1 Activation. Cell Metab. 2011, 13, 461–468. [Google Scholar] [CrossRef]
- Bai, P.; Nagy, L.; Fodor, T.; Liaudet, L.; Pacher, P. Poly(ADP-ribose) polymerases as modulators of mitochondrial activity. Trends Endocrinol. Metab. 2015, 26, 75–83. [Google Scholar] [CrossRef] [PubMed]
- Slade, D.; Dunstan, M.S.; Barkauskaite, E.; Weston, R.; Lafite, P.; Dixon, N.; Ahel, M.; Leys, D.; Ahel, I. The structure and catalytic mechanism of a poly(ADP-ribose) glycohydrolase. Nature 2011, 477, 616–620. [Google Scholar] [CrossRef] [PubMed]
- Mukhopadhyay, D.; Dasso, M. Modification in reverse: The SUMO proteases. Trends Biochem. Sci. 2007, 32, 286–295. [Google Scholar] [CrossRef]
- Gong, L.; Li, B.; Millas, S.; Yeh, E.T. Molecular cloning and characterization of human AOS1 and UBA2, components of the sentrin-activating enzyme complex. FEBS Lett. 1999, 448, 185–189. [Google Scholar] [CrossRef]
- Poulsen, S.L.; Hansen, R.K.; Wagner, S.A.; Van Cuijk, L.; Van Belle, G.J.; Streicher, W.; Wikström, M.; Choudhary, C.; Houtsmuller, A.B.; Marteijn, J.A.; et al. RNF111/Arkadia is a SUMO-targeted ubiquitin ligase that facilitates the DNA damage response. J. Cell Boil. 2013, 201, 797–807. [Google Scholar] [CrossRef] [PubMed]
- Masutani, C.; Sugasawa, K.; Yanagisawa, J.; Sonoyama, T.; Ui, M.; Enomoto, T.; Takio, K.; Tanaka, K.; Van Der Spek, P.; Bootsma, D. Purification and cloning of a nucleotide excision repair complex involving the xeroderma pigmentosum group C protein and a human homologue of yeast RAD23. EMBO J. 1994, 13, 1831–1843. [Google Scholar] [CrossRef]
- Nishi, R.; Okuda, Y.; Watanabe, E.; Mori, T.; Iwai, S.; Masutani, C.; Sugasawa, K.; Hanaoka, F. Centrin 2 Stimulates Nucleotide Excision Repair by Interacting with Xeroderma Pigmentosum Group C Protein. Mol. Cell. Boil. 2005, 25, 5664–5674. [Google Scholar] [CrossRef]
- Reardon, J.T.; Sancar, A. Recognition and repair of the cyclobutane thymine dimer, a major cause of skin cancers, by the human excision nuclease. Genome Res. 2003, 17, 2539–2551. [Google Scholar] [CrossRef]
- Groisman, S.; Polanowska, J.; Kuraoka, I.; Sawada, J.-I.; Saijo, M.; Drapkin, R.; Kisselev, A.F.; Tanaka, K.; Nakatani, Y. The Ubiquitin Ligase Activity in the DDB2 and CSA Complexes Is Differentially Regulated by the COP9 Signalosome in Response to DNA Damage. Cell 2003, 113, 357–367. [Google Scholar] [CrossRef]
- Scrima, A.; Koníčková, R.; Czyzewski, B.K.; Kawasaki, Y.; Jeffrey, P.D.; Groisman, S.; Nakatani, Y.; Iwai, S.; Pavletich, N.P.; Thomä, N.H. Structural Basis of UV DNA-Damage Recognition by the DDB1–DDB2 Complex. Cell 2008, 135, 1213–1223. [Google Scholar] [CrossRef] [PubMed]
- Han, C.; Zhao, R.; Kroger, J.; He, J.; Wani, G.; Wang, Q.-E.; A Wani, A. UV radiation-induced SUMOylation of DDB2 regulates nucleotide excision repair. Carcinogenesis 2017, 38, 976–985. [Google Scholar] [CrossRef]
- Pines, A.; Vrouwe, M.G.; Marteijn, J.A.; Typas, D.; Luijsterburg, M.S.; Cansoy, M.; Hensbergen, P.J.; Deelder, A.; De Groot, A.; Matsumoto, S.; et al. PARP1 promotes nucleotide excision repair through DDB2 stabilization and recruitment of ALC1. J. Cell Boil. 2012, 199, 235–249. [Google Scholar] [CrossRef]
- Bai, P. Biology of Poly(ADP-Ribose) Polymerases: The Factotums of Cell Maintenance. Mol. Cell 2015, 58, 947–958. [Google Scholar] [CrossRef]
- Zhang, L.; Lubin, A.; Chen, H.; Sun, Z.; Gong, F. The deubiquitinating protein USP24 interacts with DDB2 and regulates DDB2 stability. Cell Cycle 2012, 11, 4378–4384. [Google Scholar] [CrossRef]
- Hannah, J.; Zhou, P. Regulation of DNA damage response pathways by the cullin-RING ubiquitin ligases. DNA Repair 2009, 8, 536–543. [Google Scholar] [CrossRef]
- Zhou, C.; Wee, S.; Rhee, E.; Naumann, M.; Dubiel, W.; Wolf, D.A. Fission yeast COP9/signalosome suppresses cullin activity through recruitment of the deubiquitylating enzyme Ubp12p. Mol. Cell 2003, 11, 927–938. [Google Scholar] [CrossRef]
- Bergink, S.; Toussaint, W.; Luijsterburg, M.S.; Dinant, C.; Alekseev, S.; Hoeijmakers, J.H.J.; Dantuma, N.P.; Houtsmuller, A.B.; Vermeulen, W. Recognition of DNA damage by XPC coincides with disruption of the XPC–RAD23 complex. J. Cell Boil. 2012, 196, 681–688. [Google Scholar] [CrossRef]
- Wang, Q.-E.; Zhu, Q.; Wani, G.; El-Mahdy, M.A.; Li, J.; Wani, A.A. DNA repair factor XPC is modified by SUMO-1 and ubiquitin following UV irradiation. Nucleic Acids Res. 2005, 33, 4023–4034. [Google Scholar] [CrossRef]
- Sugasawa, K.; Okuda, Y.; Saijo, M.; Nishi, R.; Matsuda, N.; Chu, G.; Mori, T.; Iwai, S.; Tanaka, K.; Tanaka, K.; et al. UV-Induced Ubiquitylation of XPC Protein Mediated by UV-DDB-Ubiquitin Ligase Complex. Cell 2005, 121, 387–400. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Zhu, Q.; Wani, G.; Sharma, N.; Han, C.; Qian, J.; Pentz, K.; Wang, Q.-E.; Wani, A.A. Ubiquitin-specific Protease 7 Regulates Nucleotide Excision Repair through Deubiquitinating XPC Protein and Preventing XPC Protein from Undergoing Ultraviolet Light-induced and VCP/p97 Protein-regulated Proteolysis*. J. Boil. Chem. 2014, 289, 27278–27289. [Google Scholar] [CrossRef] [PubMed]
- Lubin, A.; Zhang, L.; Chen, H.; White, V.M.; Gong, F. A Human XPC Protein Interactome—A Resource. Int. J. Mol. Sci. 2013, 15, 141–158. [Google Scholar] [CrossRef] [PubMed]
- Riedl, T.; Hanaoka, F.; Egly, J.M. The comings and goings of nucleotide excision repair factors on damaged DNA. EMBO J. 2003, 22, 5293–5303. [Google Scholar] [CrossRef] [PubMed]
- Kapetanaki, M.G.; Guerrero-Santoro, J.; Bisi, D.C.; Hsieh, C.L.; Rapić-Otrin, V.; Levine, A.S. The DDB1-CUL4ADDB2 ubiquitin ligase is deficient in xeroderma pigmentosum group E and targets histone H2A at UV-damaged DNA sites. Proc. Natl. Acad. Sci. USA 2006, 103, 2588–2593. [Google Scholar] [CrossRef]
- Lan, L.; Nakajima, S.; Kapetanaki, M.G.; Hsieh, C.L.; Fagerburg, M.; Thickman, K.; Rodriguez-Collazo, P.; Leuba, S.; Levine, A.S.; Rapić-Otrin, V. Monoubiquitinated Histone H2A Destabilizes Photolesion-containing Nucleosomes with Concomitant Release of UV-damaged DNA-binding Protein E3 Ligase. J. Boil. Chem. 2012, 287, 12036–12049. [Google Scholar] [CrossRef]
- Wang, H.; Zhai, L.; Xu, J.; Joo, H.-Y.; Jackson, S.; Erdjument-Bromage, H.; Tempst, P.; Xiong, Y.; Zhang, Y. Histone H3 and H4 Ubiquitylation by the CUL4-DDB-ROC1 Ubiquitin Ligase Facilitates Cellular Response to DNA Damage. Mol. Cell 2006, 22, 383–394. [Google Scholar] [CrossRef]
- Gracheva, E.; Chitale, S.; Wilhelm, T.; Rapp, A.; Byrne, J.; Stadler, J.; Medina, R.; Cardoso, M.C.; Richly, H. ZRF1 mediates remodeling of E3 ligases at DNA lesion sites during nucleotide excision repair. J. Cell Boil. 2016, 213, 185–200. [Google Scholar] [CrossRef]
- Richly, H.; Rocha-Viegas, L.; Ribeiro, J.D.; Demajo, S.; Gundem, G.; Lopez-Bigas, N.; Nakagawa, T.; Rospert, S.; Ito, T.; Di Croce, L. Transcriptional activation of polycomb-repressed genes by ZRF1. Nature 2010, 468, 1124–1128. [Google Scholar] [CrossRef]
- Papadopoulou, T.; Richly, H. On-site remodeling at chromatin: How multiprotein complexes are rebuilt during DNA repair and transcriptional activation. BioEssays 2016, 38, 1130–1140. [Google Scholar] [CrossRef]
- Wang, Z.; Zhang, H.; Liu, J.; Cheruiyot, A.; Lee, J.-H.; Ordog, T.; Lou, Z.; You, Z.; Zhang, Z. USP51 deubiquitylates H2AK13,15ub and regulates DNA damage response. Genes Dev. 2016, 30, 946–959. [Google Scholar] [CrossRef] [PubMed]
- Cho, I.; Tsai, P.-F.; Lake, R.J.; Basheer, A.; Fan, H.-Y. ATP-Dependent Chromatin Remodeling by Cockayne Syndrome Protein B and NAP1-Like Histone Chaperones Is Required for Efficient Transcription-Coupled DNA Repair. PLoS Genet. 2013, 9, e1003407. [Google Scholar] [CrossRef] [PubMed]
- Fousteri, M.; Vermeulen, W.; Van Zeeland, A.A.; Mullenders, L.H. Cockayne Syndrome A and B Proteins Differentially Regulate Recruitment of Chromatin Remodeling and Repair Factors to Stalled RNA Polymerase II In Vivo. Mol. Cell 2006, 23, 471–482. [Google Scholar] [CrossRef] [PubMed]
- Groisman, S.; Kuraoka, I.; Chevallier, O.; Gaye, N.; Magnaldo, T.; Tanaka, K.; Kisselev, A.F.; Harel-Bellan, A.; Nakatani, Y. CSA-dependent degradation of CSB by the ubiquitin–proteasome pathway establishes a link between complementation factors of the Cockayne syndrome. Genome Res. 2006, 20, 1429–1434. [Google Scholar] [CrossRef] [PubMed]
- Anindya, R.; Mari, P.-O.; Kristensen, U.; Kool, H.; Giglia-Mari, G.; Mullenders, L.H.; Fousteri, M.; Vermeulen, W.; Egly, J.-M.; Svejstrup, J.Q. A Ubiquitin-Binding Domain in Cockayne Syndrome B Required for Transcription-Coupled Nucleotide Excision Repair. Mol. Cell 2010, 38, 637–648. [Google Scholar] [CrossRef]
- Nakazawa, Y.; Sasaki, K.; Mitsutake, N.; Matsuse, M.; Shimada, M.; Nardo, T.; Takahashi, Y.; Ohyama, K.; Ito, K.; Mishima, H.; et al. Mutations in UVSSA cause UV-sensitive syndrome and impair RNA polymerase IIo processing in transcription-coupled nucleotide-excision repair. Nat. Genet. 2012, 44, 586–592. [Google Scholar] [CrossRef] [PubMed]
- Okuda, M.; Nakazawa, Y.; Guo, C.; Ogi, T.; Nishimura, Y. Common TFIIH recruitment mechanism in global genome and transcription-coupled repair subpathways. Nucleic Acids Res. 2017, 45, 13043–13055. [Google Scholar] [CrossRef]
- Higa, M.; Zhang, X.; Tanaka, K.; Saijo, M. Stabilization of Ultraviolet (UV)-stimulated Scaffold Protein A by Interaction with Ubiquitin-specific Peptidase 7 Is Essential for Transcription-coupled Nucleotide Excision Repair*. J. Boil. Chem. 2016, 291, 13771–13779. [Google Scholar] [CrossRef]
- Higa, M.; Tanaka, K.; Saijo, M. Inhibition of UVSSA ubiquitination suppresses transcription-coupled nucleotide excision repair deficiency caused by dissociation from USP 7. FEBS J. 2018, 285, 965–976. [Google Scholar] [CrossRef]
- Woudstra, E.C.; Gilbert, C.; Fellows, J.; Jansen, L.E.; Brouwer, J.; Erdjument-Bromage, H.; Tempst, P.; Svejstrup, J.Q. A Rad26–Def1 complex coordinates repair and RNA pol II proteolysis in response to DNA damage. Nature 2002, 415, 929–933. [Google Scholar] [CrossRef]
- Svejstrup, J.Q. Rescue of arrested RNA polymerase II complexes. J. Cell Sci. 2003, 116, 447–451. [Google Scholar] [CrossRef] [PubMed]
- Charlet-Berguerand, N.; Feuerhahn, S.; E Kong, S.; Ziserman, H.; Conaway, J.W.; Conaway, R.; Egly, J.M. RNA polymerase II bypass of oxidative DNA damage is regulated by transcription elongation factors. EMBO J. 2006, 25, 5481–5491. [Google Scholar] [CrossRef] [PubMed]
- Li, C.-L.; Golebiowski, F.M.; Onishi, Y.; Samara, N.L.; Sugasawa, K.; Yang, W. Tripartite DNA Lesion Recognition and Verification by XPC, TFIIH, and XPA in Nucleotide Excision Repair. Mol. Cell 2015, 59, 1025–1034. [Google Scholar] [CrossRef]
- Lafon, A.; Taranum, S.; Pietrocola, F.; Dingli, F.; Loew, D.; Brahma, S.; Bartholomew, B.; Papamichos-Chronakis, E. INO80 Chromatin Remodeler Facilitates Release of RNA Polymerase II from Chromatin for Ubiquitin-Mediated Proteasomal Degradation. Mol. Cell 2015, 60, 784–796. [Google Scholar] [CrossRef]
- Dinant, C.; Ampatziadis-Michailidis, G.; Lans, H.; Tresini, M.; Lagarou, A.; Grosbart, M.; Theil, A.F.; Van Cappellen, W.A.; Kimura, H.; Bartek, J.; et al. Enhanced Chromatin Dynamics by FACT Promotes Transcriptional Restart after UV-Induced DNA Damage. Mol. Cell 2013, 51, 469–479. [Google Scholar] [CrossRef]
- Oksenych, V.; Zhovmer, A.; Ziani, S.; Mari, P.-O.; Eberova, J.; Nardò, T.; Stefanini, M.; Giglia-Mari, G.; Egly, J.-M.; Coin, F. Histone Methyltransferase DOT1L Drives Recovery of Gene Expression after a Genotoxic Attack. PLoS Genet. 2013, 9, e1003611. [Google Scholar] [CrossRef]
- Adam, S.; Polo, S.; Almouzni, G. Transcription Recovery after DNA Damage Requires Chromatin Priming by the H3.3 Histone Chaperone HIRA. Cell 2013, 155, 94–106. [Google Scholar] [CrossRef] [PubMed]
- Gaillard, P.H.L.; Martini, E.M.-D.; Kaufman, P.D.; Stillman, B.; Moustacchi, E.; Almouzni, G. Chromatin Assembly Coupled to DNA Repair: A New Role for Chromatin Assembly Factor I. Cell 1996, 86, 887–896. [Google Scholar] [CrossRef]
- Green, C.; Almouzni, G. Local action of the chromatin assembly factor CAF-1 at sites of nucleotide excision repair in vivo. EMBO J. 2003, 22, 5163–5174. [Google Scholar] [CrossRef]
- Polo, S.; Roche, D.; Almouzni, G. New Histone Incorporation Marks Sites of UV Repair in Human Cells. Cell 2006, 127, 481–493. [Google Scholar] [CrossRef]
- Oksenych, V.; De Jesus, B.B.; Zhovmer, A.; Egly, J.-M.; Coin, F. Molecular insights into the recruitment of TFIIH to sites of DNA damage. EMBO J. 2009, 28, 2971–2980. [Google Scholar] [CrossRef] [PubMed]
- Coin, F.; Oksenych, V.; Mocquet, V.; Groh, S.; Blattner, C.; Egly, J.M. Nucleotide Excision Repair Driven by the Dissociation of CAK from TFIIH. Mol. Cell 2008, 31, 9–20. [Google Scholar] [CrossRef]
- Coin, F.; Oksenych, V.; Egly, J.M. Distinct Roles for the XPB/p52 and XPD/p44 Subcomplexes of TFIIH in Damaged DNA Opening during Nucleotide Excision Repair. Mol. Cell 2007, 26, 245–256. [Google Scholar] [CrossRef] [PubMed]
- Winkler, S.; Araujo, S.J.; Fiedler, U.; Vermeulen, W.; Coin, F.; Egly, J.-M.; Hoeijmakers, J.H.J.; Wood, R.D.; Timmers, H.T.M.; Weeda, G. TFIIH with Inactive XPD Helicase Functions in Transcription Initiation but Is Defective in DNA Repair. J. Boil. Chem. 2000, 275, 4258–4266. [Google Scholar] [CrossRef]
- Camenisch, U.; Dip, R.; Schumacher, S.B.; Schuler, B.; Naegeli, H. Recognition of helical kinks by xeroderma pigmentosum group A protein triggers DNA excision repair. Nat. Struct. Mol. Boil. 2006, 13, 278–284. [Google Scholar] [CrossRef] [PubMed]
- Gervais, V.; Lamour, V.; Jawhari, A.; Frindel, F.; Wasielewski, E.; Dubaele, S.; Egly, J.-M.; Thierry, J.-C.; Kieffer, B.; Poterszman, A. TFIIH contains a PH domain involved in DNA nucleotide excision repair. Nat. Struct. Mol. Boil. 2004, 11, 616–622. [Google Scholar] [CrossRef]
- Wold, M. REPLICATION PROTEIN A:A Heterotrimeric, Single-Stranded DNA-Binding Protein Required for Eukaryotic DNA Metabolism. Annu. Rev. Biochem. 1997, 66, 61–92. [Google Scholar] [CrossRef] [PubMed]
- De Laat, W.; Appeldoorn, E.; Sugasawa, K.; Weterings, E.; Jaspers, N.G.; Hoeijmakers, J. DNA-binding polarity of human replication protein A positions nucleases in nucleotide excision repair. Genome Res. 1998, 12, 2598–2609. [Google Scholar] [CrossRef]
- O’Donovan, A.; Davies, A.A.; Moggs, J.G.; West, S.; Wood, R.D. XPG endonuclease makes the 3′ incision in human DNA nucleotide excision repair. Nature 1994, 371, 432–435. [Google Scholar] [CrossRef]
- Sijbers, A.M.; De Laat, W.; Ariza, R.R.; Biggerstaff, M.; Wei, Y.-F.; Moggs, J.G.; Carter, K.C.; Shell, B.K.; Evans, E.; De Jong, M.C.; et al. Xeroderma Pigmentosum Group F Caused by a Defect in a Structure-Specific DNA Repair Endonuclease. Cell 1996, 86, 811–822. [Google Scholar] [CrossRef]
- Tsodikov, O.V.; Enzlin, J.H.; Schärer, O.D.; Ellenberger, T. Crystal structure and DNA binding functions of ERCC1, a subunit of the DNA structure-specific endonuclease XPF–ERCC1. Proc. Natl. Acad. Sci. USA 2005, 102, 11236–11241. [Google Scholar] [CrossRef] [PubMed]
- Tripsianes, K.; Folkers, G.E.; Ab, E.; Das, D.; Odijk, H.; Jaspers, N.G.; Hoeijmakers, J.H.; Kaptein, R.; Boelens, R. The Structure of the Human ERCC1/XPF Interaction Domains Reveals a Complementary Role for the Two Proteins in Nucleotide Excision Repair. Structure 2005, 13, 1849–1858. [Google Scholar] [CrossRef] [PubMed]
- Biggerstaff, M.; Szymkowski, D.; Wood, R.D. Co-correction of the ERCC1, ERCC4 and xeroderma pigmentosum group F DNA repair defects in vitro. EMBO J. 1993, 12, 3685–3692. [Google Scholar] [CrossRef]
- Van Vuuren, A.; Appeldoorn, E.; Odijk, H.; Yasui, A.; Jaspers, N.G.J.; Bootsma, D.; Hoeijmakers, J. Evidence for a repair enzyme complex involving ERCC1 and complementing activities of ERCC4, ERCC11 and xeroderma pigmentosum group F. EMBO J. 1993, 12, 3693–3701. [Google Scholar] [CrossRef] [PubMed]
- Perez-Oliva, A.B.; Lachaud, C.; Szyniarowski, P.; Muñoz, I.M.; Macartney, T.; Hickson, I.; Rouse, J.; Alessi, D.R. USP 45 deubiquitylase controls ERCC 1– XPF endonuclease-mediated DNA damage responses. EMBO J. 2014, 34, 326–343. [Google Scholar] [CrossRef]
- Schärer, O.D. XPG: Its products and biological roles. Single Mol. Single Cell Seq. 2008, 637, 83–92. [Google Scholar] [CrossRef]
- Kelman, Z. PCNA: Structure, functions and interactions. Oncogene 1997, 14, 629–640. [Google Scholar] [CrossRef]
- Han, C.; Wani, G.; Zhao, R.; Qian, J.; Sharma, N.; He, J.; Zhu, Q.; Wang, Q.-E.; A Wani, A. Cdt2-mediated XPG degradation promotes gap-filling DNA synthesis in nucleotide excision repair. Cell Cycle 2014, 14, 1103–1115. [Google Scholar] [CrossRef]
- Ogi, T.; Lehmann, A.R. The Y-family DNA polymerase κ (pol κ) functions in mammalian nucleotide-excision repair. Nature 2006, 8, 640–642. [Google Scholar] [CrossRef]
- Araújo, S.J.; Tirode, F.; Coin, F.; Pospiech, H.; Syväoja, J.E.; Stucki, M.; Hübscher, U.; Egly, J.-M.; Wood, R.D. Nucleotide excision repair of DNA with recombinant human proteins: Definition of the minimal set of factors, active forms of TFIIH, and modulation by CAK. Genome Res. 2000, 14, 349–359. [Google Scholar]
- Zhang, S.; Zhao, H.; Darzynkiewicz, Z.; Zhou, P.; Zhang, Z.; Lee, E.Y.C.; Lee, M.Y.W.T. A Novel Function of CRL4Cdt2. J. Boil. Chem. 2013, 288, 29550–29561. [Google Scholar] [CrossRef] [PubMed]
- Moser, J.; Kool, H.; Giakzidis, I.; Caldecott, K.; Mullenders, L.H.; Fousteri, M. Sealing of Chromosomal DNA Nicks during Nucleotide Excision Repair Requires XRCC1 and DNA Ligase IIIα in a Cell-Cycle-Specific Manner. Mol. Cell 2007, 27, 311–323. [Google Scholar] [CrossRef] [PubMed]
- Van Houten, B.; Kuper, J.; Kisker, C. Role of XPD in cellular functions: To TFIIH and beyond. DNA Repair 2016, 44, 136–142. [Google Scholar] [CrossRef] [PubMed]
- Missura, M.; Buterin, T.; Hindges, R.; Hübscher, U.; Kasparkova, J.; Brabec, V.; Naegeli, H. Double-check probing of DNA bending and unwinding by XPA–RPA: An architectural function in DNA repair. EMBO J. 2001, 20, 3554–3564. [Google Scholar] [CrossRef] [PubMed]
- Krasikova, Y.S.; Rechkunova, N.I.; Maltseva, E.A.; Petruseva, I.O.; Lavrik, O.I. Localization of xeroderma pigmentosum group A protein and replication protein A on damaged DNA in nucleotide excision repair. Nucleic Acids Res. 2010, 38, 8083–8094. [Google Scholar] [CrossRef]
- Gilljam, K.M.; Müller, R.; Liabakk, N.B.; Otterlei, M. Nucleotide Excision Repair Is Associated with the Replisome and Its Efficiency Depends on a Direct Interaction between XPA and PCNA. PLoS ONE 2012, 7, e49199. [Google Scholar] [CrossRef]
- Hutton, R.D.; Craggs, T.; White, M.F.; Penedo, J.C. PCNA and XPF cooperate to distort DNA substrates. Nucleic Acids Res. 2009, 38, 1664–1675. [Google Scholar] [CrossRef]
- Kemp, M.G.; Reardon, J.T.; Lindsey-Boltz, L.; Sancar, A. Mechanism of Release and Fate of Excised Oligonucleotides during Nucleotide Excision Repair*. J. Boil. Chem. 2012, 287, 22889–22899. [Google Scholar] [CrossRef] [PubMed]
- Rockx, D.A.P.; Mason, R.; Van Hoffen, A.; Barton, M.C.; Citterio, E.; Bregman, D.B.; Van Zeeland, A.A.; Vrieling, H.; Mullenders, L.H.F. UV-induced inhibition of transcription involves repression of transcription initiation and phosphorylation of RNA polymerase II. Proc. Natl. Acad. Sci. USA 2000, 97, 10503–10508. [Google Scholar] [CrossRef]
- Svejstrup, J.Q. Contending with transcriptional arrest during RNAPII transcript elongation. Trends Biochem. Sci. 2007, 32, 165–171. [Google Scholar] [CrossRef]
- Kamura, T.; Burian, D.; Yan, Q.; Schmidt, S.L.; Lane, W.S.; Querido, E.; Branton, P.E.; Shilatifard, A.; Conaway, R.C.; Conaway, J.W. MUF1, A Novel Elongin BC-interacting Leucine-rich Repeat Protein That Can Assemble with Cul5 and Rbx1 to Reconstitute a Ubiquitin Ligase. J. Boil. Chem. 2001, 276, 29748–29753. [Google Scholar] [CrossRef] [PubMed]
- Pankotai, T.; Bonhomme, C.; Chen, D.; Soutoglou, E. DNAPKcs-dependent arrest of RNA polymerase II transcription in the presence of DNA breaks. Nat. Struct. Mol. Boil. 2012, 19, 276–282. [Google Scholar] [CrossRef] [PubMed]
- Caron, P.; Pankotai, T.; Wiegant, W.W.; Tollenaere, M.A.; Furst, A.; Bonhomme, C.; Helfricht, A.; De Groot, A.; Pastink, A.; Vertegaal, A.C.; et al. WWP2 ubiquitylates RNA polymerase II for DNA-PK-dependent transcription arrest and repair at DNA breaks. Genome Res. 2019, 33, 684–704. [Google Scholar] [CrossRef]
- Borsos, B.N.; Huliák, I.; Majoros, H.; Ujfaludi, Z.; Gyenis, Á.; Pukler, P.; Boros, I.M.; Pankotai, T. Human p53 interacts with the elongating RNAPII complex and is required for the release of actinomycin D induced transcription blockage. Sci. Rep. 2017, 7, 40960. [Google Scholar] [CrossRef] [PubMed]
- Scharf, A.; Grozdanov, P.N.; Veith, R.; Kubitscheck, U.; Meier, U.T.; Von Mikecz, A. Distant positioning of proteasomal proteolysis relative to actively transcribed genes. Nucleic Acids Res. 2011, 39, 4612–4627. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gillette, T.G.; Gonzalez, F.; Delahodde, A.; Johnston, S.A.; Kodadek, T. Physical and functional association of RNA polymerase II and the proteasome. Proc. Natl. Acad. Sci. USA 2004, 101, 5904–5909. [Google Scholar] [CrossRef]
- Ferdous, A.; Gonzalez, F.; Sun, L.; Kodadek, T.; Johnston, S.A. The 19S Regulatory Particle of the Proteasome Is Required for Efficient Transcription Elongation by RNA Polymerase II. Mol. Cell 2001, 7, 981–991. [Google Scholar] [CrossRef]
- Majoros, H.; Ujfaludi, Z.; Borsos, B.N.; Hudacsek, V.V.; Nagy, Z.; Coin, F.; Buzas, K.; Kovács, I.; Bíró, T.; Boros, I.M.; et al. SerpinB2 is involved in cellular response upon UV irradiation. Sci. Rep. 2019, 9, 2753. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Borsos, B.N.; Majoros, H.; Pankotai, T. Emerging Roles of Post-Translational Modifications in Nucleotide Excision Repair. Cells 2020, 9, 1466. https://doi.org/10.3390/cells9061466
Borsos BN, Majoros H, Pankotai T. Emerging Roles of Post-Translational Modifications in Nucleotide Excision Repair. Cells. 2020; 9(6):1466. https://doi.org/10.3390/cells9061466
Chicago/Turabian StyleBorsos, Barbara N., Hajnalka Majoros, and Tibor Pankotai. 2020. "Emerging Roles of Post-Translational Modifications in Nucleotide Excision Repair" Cells 9, no. 6: 1466. https://doi.org/10.3390/cells9061466
APA StyleBorsos, B. N., Majoros, H., & Pankotai, T. (2020). Emerging Roles of Post-Translational Modifications in Nucleotide Excision Repair. Cells, 9(6), 1466. https://doi.org/10.3390/cells9061466





