Therapeutic Approaches Targeting Nucleolus in Cancer
Abstract
1. Introduction
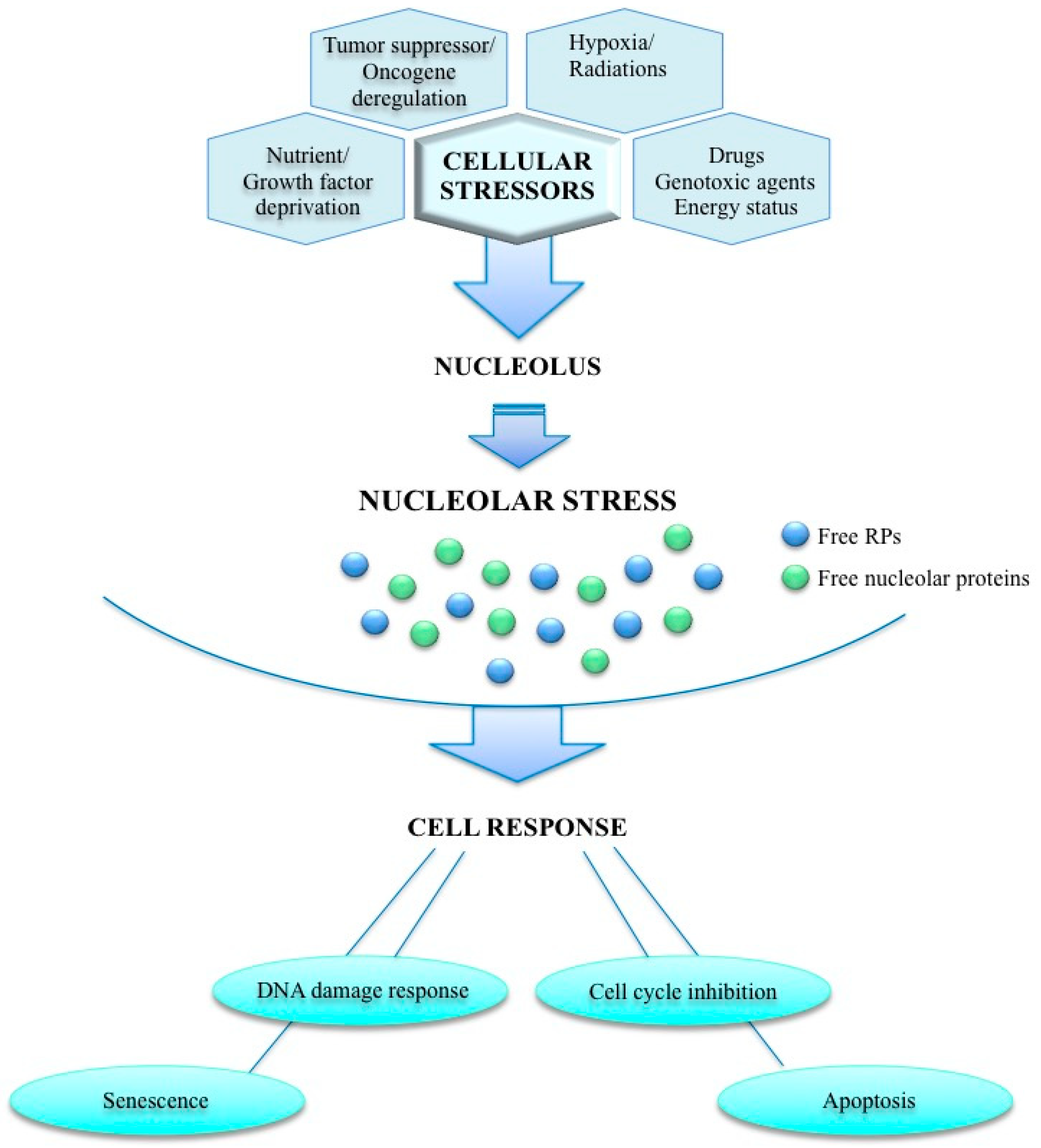
2. Nucleolus and Cancer
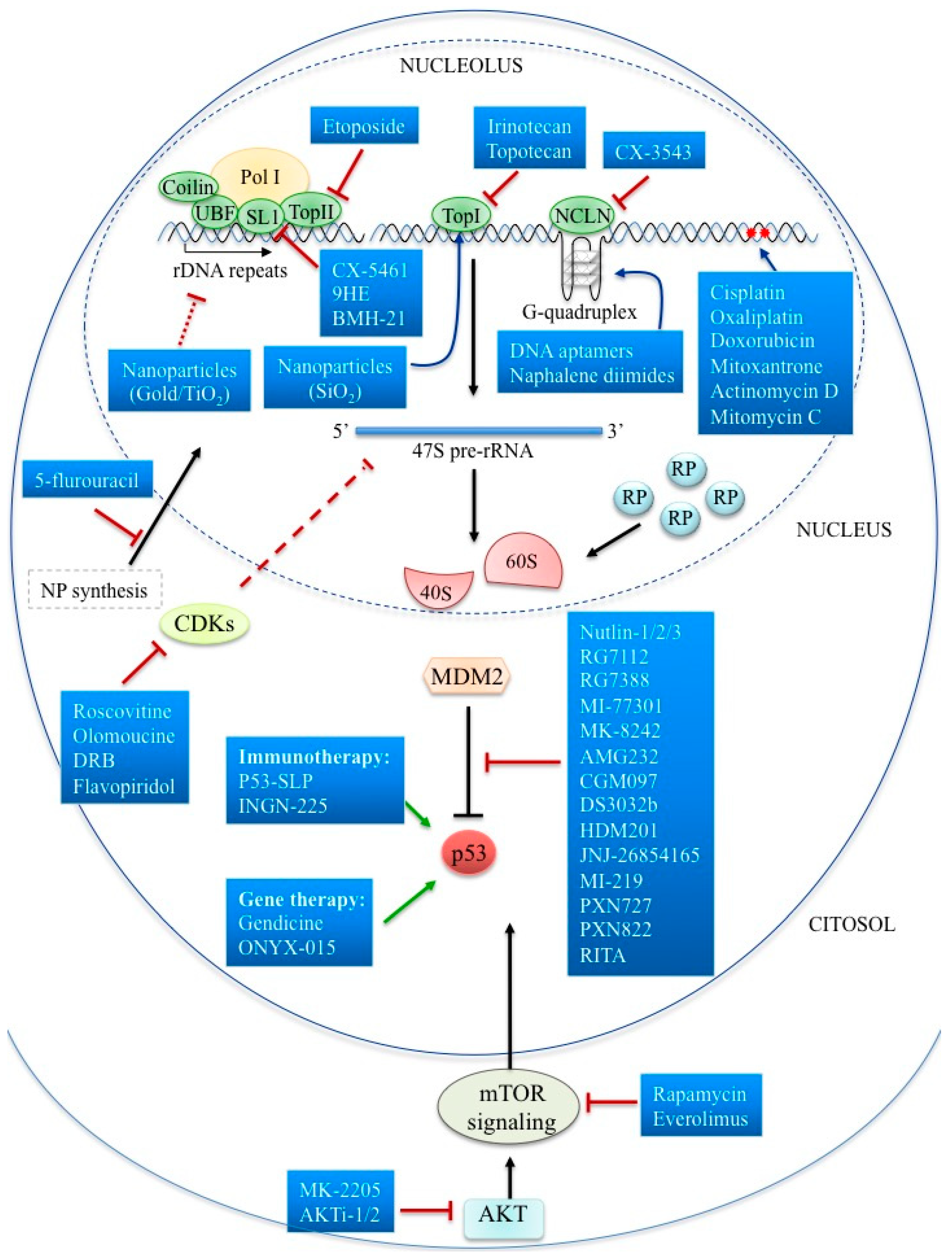
3. Targeting Nucleolus in Cancer
4. Targeting the Nucleolar Components
5. Targeting Nucleolus by Specific Inhibitors of Ribosome Biogenesis
6. Targeting Cell Signaling Pathways Functionally Regulating Nucleolus in Cancer
7. Targeting p53-dependent Nucleolar Stress Pathway
8. Concluding Remarks
Author Contributions
Funding
Conflicts of Interest
References
- Pederson, T. The nucleolus. Cold Spring Harb. Perspect. Biol. 2011, 3, a000638. [Google Scholar] [CrossRef] [PubMed]
- Tsekrekou, M.; Stratigi, K.; Chatzinikolaou, G. The Nucleolus: In Genome Maintenance and Repair. Int. J. Mol. Sci. 2017, 18, 1411. [Google Scholar] [CrossRef] [PubMed]
- Nazar, R.N. Ribosomal RNA processing and ribosome biogenesis in eukaryotes. IUBMB Life 2004, 56, 457–465. [Google Scholar] [CrossRef] [PubMed]
- Puvion-Dutilleul, F.; Puvion, E.; Bachellerie, J.P. Early stages of pre-rRNA formation within the nucleolar ultrastructure of mouse cells studied by in situ hybridization with a 5’ETS leader probe. Chromosoma 1997, 105, 496–505. [Google Scholar] [CrossRef] [PubMed]
- Russell, J.; Zomerdijk, J.C. The RNA polymerase I transcription machinery. Biochem. Soc. Symp. 2006, 73, 203–216. [Google Scholar] [CrossRef]
- Grummt, I. Life on a planet of its own: regulation of RNA polymerase I transcription in the nucleolus. Genes Dev. 2003, 17, 1691–1702. [Google Scholar] [CrossRef]
- Russo, G.; Ricciardelli, G.; Pietropaolo, C. Different domains cooperate to target the human ribosomal L7a protein to the nucleus and to the nucleoli. J. Biol. Chem. 1997, 272, 5229–5235. [Google Scholar] [CrossRef] [PubMed]
- Andersen, J.S.; Lam, Y.W.; Leung, A.K.; Ong, S.E.; Lyon, C.E.; Lamond, A.I.; Mann, M. Nucleolar proteome dynamics. Nature 2005, 433, 77–83. [Google Scholar] [CrossRef]
- Lindström, M.S.; Jurada, D.; Bursac, S.; Orsolic, I.; Bartek, J.; Volarevic, S. Nucleolus as an emerging hub in maintenance of genome stability and cancer pathogenesis. Oncogene 2018, 37, 2351–2366. [Google Scholar] [CrossRef]
- Schöfer, C.; Weipoltshammer, K. Nucleolus and chromatin. Histochem. Cell Biol. 2018, 150, 209–225. [Google Scholar] [CrossRef]
- Tsai, R.Y.; Pederson, T. Connecting the nucleolus to the cell cycle and human disease. FASEB J. 2014, 28, 3290–3296. [Google Scholar] [CrossRef] [PubMed]
- Dillinger, S.; Straub, T.; Németh, A. Nucleolus association of chromosomal domains is largely maintained in cellular senescence despite massive nuclear reorganisation. PLoS ONE 2017, 12, e0178821. [Google Scholar] [CrossRef]
- Russo, A.; Russo, G. Ribosomal Proteins Control or Bypass p53 during Nucleolar Stress. Int. J. Mol. Sci. 2017, 18, 140. [Google Scholar] [CrossRef]
- Golomb, L.; Volarevic, S.; Oren, M. p53 and ribosome biogenesis stress: the essentials. FEBS Lett. 2014, 588, 2571–2579. [Google Scholar] [CrossRef] [PubMed]
- Weeks, S.E.; Metge, B.J.; Samant, R.S. The nucleolus: A central response hub for the stressors that drive cancer progression. Cell Mol. Life Sci. 2019. [Google Scholar] [CrossRef] [PubMed]
- Iarovaia, O.V.; Minina, E.P.; Sheval, E.V.; Onichtchouk, D.; Dokudovskaya, S.; Razin, S.V.; Vassetzky, Y.S. Nucleolus: A Central Hub for Nuclear Functions. Trends Cell Biol. 2019, 29, 647–659. [Google Scholar] [CrossRef] [PubMed]
- Boisvert, F.M.; van Koningsbruggen, S.; Navascués, J.; Lamond, A.I. The multifunctional nucleolus. Nat. Rev. Mol. Cell Biol. 2007, 8, 574–585. [Google Scholar] [CrossRef] [PubMed]
- Woods, S.J.; Hannan, K.M.; Pearson, R.B.; Hannan, R.D. The nucleolus as a fundamental regulator of the p53 response and a new target for cancer therapy. Biochim. Biophys. Acta 2015, 1849, 821–829. [Google Scholar] [CrossRef]
- Derenzini, M.; Montanaro, L.; Treré, D. What the nucleolus says to a tumour pathologist. Histopathology 2009, 54, 753–762. [Google Scholar] [CrossRef]
- Catez, F.; Dalla Venezia, N.; Marcel, V.; Zorbas, C.; Lafontaine, D.L.J.; Diaz, J.J. Ribosome biogenesis: An emerging druggable pathway for cancer therapeutics. Biochem. Pharmacol. 2019, 159, 74–81. [Google Scholar] [CrossRef]
- Derenzini, M.; Trerè, D.; Pession, A.; Montanaro, L.; Sirri, V.; Ochs, R.L. Nucleolar function and size in cancer cells. Am. J. Pathol. 1998, 152, 1291–1297. [Google Scholar] [PubMed]
- Derenzini, M.; Nardi, F.; Farabegoli, F.; Ottinetti, A.; Roncaroli, F.; Bussolati, G. Distribution of silver-stained interphase nucleolar organizer regions as a parameter to distinguish neoplastic from nonneoplastic reactive cells in human effusions. Acta Cytol. 1989, 33, 491–498. [Google Scholar] [PubMed]
- Montanaro, L.; Treré, D.; Derenzini, M. Nucleolus, ribosomes, and cancer. Am. J. Pathol. 2008, 173, 301–310. [Google Scholar] [CrossRef] [PubMed]
- Ruggero, D. Revisiting the nucleolus: From marker to dynamic integrator of cancer signaling. Sci. Signal. 2012, 5, pe38. [Google Scholar] [CrossRef] [PubMed]
- van Sluis, M.; McStay, B. Ribosome biogenesis: Achilles heel of cancer? Genes Cancer 2014, 5, 152–153. [Google Scholar] [PubMed]
- Boussemart, L.; Malka-Mahieu, H.; Girault, I.; Allard, D.; Hemmingsson, O.; Tomasic, G.; Thomas, M.; Basmadjian, C.; Ribeiro, N.; Thuaud, F.; et al. eIF4F is a nexus of resistance to anti-BRAF and anti-MEK cancer therapies. Nature 2014, 513, 105–109. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, A.C.; Costa, M.; Zollo, O.; Davis, C.; Feldman, M.E.; Testa, J.R.; Meyuhas, O.; Shokat, K.M.; Ruggero, D. Genetic dissection of the oncogenic mTOR pathway reveals druggable addiction to translational control via 4EBP-eIF4E. Cancer Cell 2010, 17, 249–261. [Google Scholar] [CrossRef] [PubMed]
- Montanaro, L.; Treré, D.; Derenzini, M. Changes in ribosome biogenesis may induce cancer by down-regulating the cell tumor suppressor potential. Biochim. Biophys. Acta 2012, 1825, 101–110. [Google Scholar] [CrossRef]
- Bywater, M.J.; Pearson, R.B.; McArthur, G.A.; Hannan, R.D. Dysregulation of the basal RNA polymerase transcription apparatus in cancer. Nat. Rev. Cancer 2013, 13, 299–314. [Google Scholar] [CrossRef]
- Uemura, M.; Zheng, Q.; Koh, C.M.; Nelson, W.G.; Yegnasubramanian, S.; De Marzo, A.M. Overexpression of ribosomal RNA in prostate cancer is common but not linked to rDNA promoter hypomethylation. Oncogene 2012, 31, 1254–1263. [Google Scholar] [CrossRef]
- Williamson, D.; Lu, Y.J.; Fang, C.; Pritchard-Jones, K.; Shipley, J. Nascent pre-rRNA overexpression correlates with an adverse prognosis in alveolar rhabdomyosarcoma. Genes Chromosomes Cancer 2006, 45, 839–845. [Google Scholar] [CrossRef]
- Bywater, M.J.; Poortinga, G.; Sanij, E.; Hein, N.; Peck, A.; Cullinane, C.; Wall, M.; Cluse, L.; Drygin, D.; Anderes, K.; et al. Inhibition of RNA polymerase I as a therapeutic strategy to promote cancer-specific activation of p53. Cancer Cell 2012, 22, 51–65. [Google Scholar] [CrossRef] [PubMed]
- Drygin, D.; Rice, W.G.; Grummt, I. The RNA polymerase I transcription machinery: an emerging target for the treatment of cancer. Annu. Rev. Pharmacol. Toxicol. 2010, 50, 131–156. [Google Scholar] [CrossRef] [PubMed]
- Whittaker, S.; Martin, M.; Marais, R. All roads lead to the ribosome. Cancer Cell 2010, 18, 5–6. [Google Scholar] [CrossRef][Green Version]
- Tzoneva, G.; Perez-Garcia, A.; Carpenter, Z.; Khiabanian, H.; Tosello, V.; Allegretta, M.; Paietta, E.; Racevskis, J.; Rowe, J.M.; Tallman, M.S.; et al. Activating mutations in the NT5C2 nucleotidase gene drive chemotherapy resistance in relapsed ALL. Nat. Med. 2013, 19, 368–371. [Google Scholar] [CrossRef] [PubMed]
- De Keersmaecker, K.; Atak, Z.K.; Li, N.; Vicente, C.; Patchett, S.; Girardi, T.; Gianfelici, V.; Geerdens, E.; Clappier, E.; Porcu, M.; et al. Exome sequencing identifies mutation in CNOT3 and ribosomal genes RPL5 and RPL10 in T-cell acute lymphoblastic leukemia. Nat. Genet. 2013, 45, 186–190. [Google Scholar] [CrossRef]
- Hein, N.; Hannan, K.M.; George, A.J.; Sanij, E.; Hannan, R.D. The nucleolus: An emerging target for cancer therapy. Trends Mol. Med. 2013, 19, 643–654. [Google Scholar] [CrossRef]
- Peddibhotla, S.; Wei, Z.; Papineni, R.; Lam, M.H.; Rosen, J.M.; Zhang, P. The DNA damage effector Chk1 kinase regulates Cdc14B nucleolar shuttling during cell cycle progression. Cell Cycle 2011, 10, 671–679. [Google Scholar] [CrossRef]
- Sasaki, M.; Kawahara, K.; Nishio, M.; Mimori, K.; Kogo, R.; Hamada, K.; Itoh, B.; Wang, J.; Komatsu, Y.; Yang, Y.R.; et al. Regulation of the MDM2-P53 pathway and tumor growth by PICT1 via nucleolar RPL11. Nat. Med. 2011, 17, 944–951. [Google Scholar] [CrossRef]
- Andrique, L.; Fauvin, D.; El Maassarani, M.; Colasson, H.; Vannier, B.; Séité, P. ErbB3(80 kDa), a nuclear variant of the ErbB3 receptor, binds to the Cyclin D1 promoter to activate cell proliferation but is negatively controlled by p14ARF. Cell Signal. 2012, 24, 1074–1085. [Google Scholar] [CrossRef]
- Audas, T.E.; Jacob, M.D.; Lee, S. Immobilization of proteins in the nucleolus by ribosomal intergenic spacer noncoding RNA. Mol. Cell 2012, 45, 147–157. [Google Scholar] [CrossRef] [PubMed]
- Ebina, M.; Tsuruta, F.; Katoh, M.C.; Kigoshi, Y.; Someya, A.; Chiba, T. Myeloma overexpressed 2 (Myeov2) regulates L11 subnuclear localization through Nedd8 modification. PLoS ONE 2013, 8, e65285. [Google Scholar] [CrossRef] [PubMed]
- Qiu, W.; Wang, G.; Sun, X.; Ye, J.; Wei, F.; Shi, X.; Lv, G. The involvement of cell surface nucleolin in the initiation of CCR6 signaling in human hepatocellular carcinoma. Med. Oncol. 2015, 32, 75. [Google Scholar] [CrossRef] [PubMed]
- Daniely, Y.; Dimitrova, D.D.; Borowiec, J.A. Stress-dependent nucleolin mobilization mediated by p53-nucleolin complex formation. Mol. Cell Biol. 2002, 22, 6014–6022. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, J.; Fujimoto, H.; Sato, J.; Hayashi, I.; Burma, S.; Matsuura, S.; Chen, D.J.; Komatsu, K. Nucleolin participates in DNA double-strand break-induced damage response through MDC1-dependent pathway. PLoS ONE 2012, 7, e49245. [Google Scholar] [CrossRef] [PubMed]
- Ishimaru, D.; Zuraw, L.; Ramalingam, S.; Sengupta, T.K.; Bandyopadhyay, S.; Reuben, A.; Fernandes, D.J.; Spicer, E.K. Mechanism of regulation of bcl-2 mRNA by nucleolin and A+U-rich element-binding factor 1 (AUF1). J. Biol. Chem. 2010, 285, 27182–27191. [Google Scholar] [CrossRef] [PubMed]
- Lau, A.W.; Fukushima, H.; Wei, W. The Fbw7 and betaTRCP E3 ubiquitin ligases and their roles in tumorigenesis. Front. Biosci. (Landmark Ed.) 2012, 17, 2197–2212. [Google Scholar] [CrossRef]
- Boulon, S.; Westman, B.J.; Hutten, S.; Boisvert, F.M.; Lamond, A.I. The nucleolus under stress. Mol. Cell 2010, 40, 216–227. [Google Scholar] [CrossRef] [PubMed]
- Fumagalli, S.; Ivanenkov, V.V.; Teng, T.; Thomas, G. Suprainduction of p53 by disruption of 40S and 60S ribosome biogenesis leads to the activation of a novel G2/M checkpoint. Genes Dev. 2012, 26, 1028–1040. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Lu, H. Signaling to p53: ribosomal proteins find their way. Cancer Cell 2009, 16, 369–377. [Google Scholar] [CrossRef] [PubMed]
- Di Matteo, A.; Franceschini, M.; Chiarella, S.; Rocchio, S.; Travaglini-Allocatelli, C.; Federici, L. Molecules that target nucleophosmin for cancer treatment: an update. Oncotarget 2016, 7, 44821–44840. [Google Scholar] [CrossRef] [PubMed]
- Pfister, A.S.; Keil, M.; Kühl, M. The Wnt Target Protein Peter Pan Defines a Novel p53-independent Nucleolar Stress-Response Pathway. J. Biol. Chem. 2015, 290, 10905–10918. [Google Scholar] [CrossRef] [PubMed]
- Ayrault, O.; Andrique, L.; Fauvin, D.; Eymin, B.; Gazzeri, S.; Séité, P. Human tumor suppressor p14ARF negatively regulates rRNA transcription and inhibits UBF1 transcription factor phosphorylation. Oncogene 2006, 25, 7577–7586. [Google Scholar] [CrossRef] [PubMed]
- Gjerset, R.A.; Bandyopadhyay, K. Regulation of p14ARF through subnuclear compartmentalization. Cell Cycle 2006, 5, 686–690. [Google Scholar] [CrossRef] [PubMed]
- Russo, A.; Esposito, D.; Catillo, M.; Pietropaolo, C.; Crescenzi, E.; Russo, G. Human rpL3 induces G₁/S arrest or apoptosis by modulating p21 (waf1/cip1) levels in a p53-independent manner. Cell Cycle 2013, 12, 76–87. [Google Scholar] [CrossRef] [PubMed]
- Esposito, D.; Crescenzi, E.; Sagar, V.; Loreni, F.; Russo, A.; Russo, G. Human rpL3 plays a crucial role in cell response to nucleolar stress induced by 5-FU and L-OHP. Oncotarget 2014, 5, 11737–11751. [Google Scholar] [CrossRef] [PubMed]
- Russo, G.; Cuccurese, M.; Monti, G.; Russo, A.; Amoresano, A.; Pucci, P.; Pietropaolo, C. Ribosomal protein L7a binds RNA through two distinct RNA-binding domains. Biochem. J. 2005, 385, 289–299. [Google Scholar] [CrossRef]
- Russo, A.; Russo, G.; Cuccurese, M.; Garbi, C.; Pietropaolo, C. The 3’-untranslated region directs ribosomal protein-encoding mRNAs to specific cytoplasmic regions. Biochim. Biophys. Acta 2006, 1763, 833–843. [Google Scholar] [CrossRef]
- Russo, A.; Cirulli, C.; Amoresano, A.; Pucci, P.; Pietropaolo, C.; Russo, G. cis-acting sequences and trans-acting factors in the localization of mRNA for mitochondrial ribosomal proteins. Biochim. Biophys. Acta 2008, 1779, 820–829. [Google Scholar] [CrossRef]
- Cuccurese, M.; Russo, G.; Russo, A.; Pietropaolo, C. Alternative splicing and nonsense-mediated mRNA decay regulate mammalian ribosomal gene expression. Nucleic Acids Res. 2005, 33, 5965–5977. [Google Scholar] [CrossRef]
- Russo, A.; Siciliano, G.; Catillo, M.; Giangrande, C.; Amoresano, A.; Pucci, P.; Pietropaolo, C.; Russo, G. hnRNP H1 and intronic G runs in the splicing control of the human rpL3 gene. Biochim. Biophys. Acta 2010, 1799, 419–428. [Google Scholar] [CrossRef] [PubMed]
- Russo, A.; Catillo, M.; Esposito, D.; Briata, P.; Pietropaolo, C.; Russo, G. Autoregulatory circuit of human rpL3 expression requires hnRNP H1, NPM and KHSRP. Nucleic Acids Res. 2011, 39, 7576–7585. [Google Scholar] [CrossRef] [PubMed]
- d’Emmanuele di Villa Bianca, R.; Mitidieri, E.; Fusco, F.; Russo, A.; Pagliara, V.; Tramontano, T.; Donnarumma, E.; Mirone, V.; Cirino, G.; Russo, G.; et al. Urothelium muscarinic activation phosphorylates CBS(Ser227) via cGMP/PKG pathway causing human bladder relaxation through H2S production. Sci. Rep. 2016, 6, 31491. [Google Scholar] [CrossRef] [PubMed]
- Russo, A.; Saide, A.; Cagliani, R.; Cantile, M.; Botti, G.; Russo, G. rpL3 promotes the apoptosis of p53 mutated lung cancer cells by down-regulating CBS and NFκB upon 5-FU treatment. Sci. Rep. 2016, 6, 38369. [Google Scholar] [CrossRef] [PubMed]
- Pagliara, V.; Saide, A.; Mitidieri, E.; d’Emmanuele di Villa Bianca, R.; Sorrentino, R.; Russo, G.; Russo, A. 5-FU targets rpL3 to induce mitochondrial apoptosis via cystathionine-β-synthase in colon cancer cells lacking p53. Oncotarget 2016, 7, 50333–50348. [Google Scholar] [CrossRef] [PubMed]
- De Filippis, D.; Russo, A.; D’Amico, A.; Esposito, G.; Pietropaolo, C.; Concetta, P.; Cinelli, M.; Russo, G.; Iuvone, T. Cannabinoids reduce granuloma-associated angiogenesis in rats by controlling transcription and expression of mast cell protease-5. Br. J. Pharmacol. 2008, 154, 1672–1679. [Google Scholar] [CrossRef]
- De Filippis, D.; Russo, A.; De Stefano, D.; Cipriano, M.; Esposito, D.; Grassia, G.; Carnuccio, R.; Russo, G.; Iuvone, T. Palmitoylethanolamide inhibits rMCP-5 expression by regulating MITF activation in rat chronic granulomatous inflammation. Eur. J. Pharmacol. 2014, 725, 64–69. [Google Scholar] [CrossRef]
- Russo, A.; Pagliara, V.; Albano, F.; Esposito, D.; Sagar, V.; Loreni, F.; Irace, C.; Santamaria, R.; Russo, G. Regulatory role of rpL3 in cell response to nucleolar stress induced by Act D in tumor cells lacking functional p53. Cell Cycle 2016, 15, 41–51. [Google Scholar] [CrossRef]
- Russo, A.; Pellosi, D.S.; Pagliara, V.; Milone, M.R.; Pucci, B.; Caetano, W.; Hioka, N.; Budillon, A.; Ungaro, F.; Russo, G.; et al. Biotin-targeted Pluronic(®) P123/F127 mixed micelles delivering niclosamide: A repositioning strategy to treat drug-resistant lung cancer cells. Int. J. Pharm. 2016, 511, 127–139. [Google Scholar] [CrossRef]
- Russo, A.; Saide, A.; Smaldone, S.; Faraonio, R.; Russo, G. Role of uL3 in Multidrug Resistance in p53-Mutated Lung Cancer Cells. Int. J. Mol. Sci. 2017, 18, 547. [Google Scholar] [CrossRef]
- Russo, A.; Maiolino, S.; Pagliara, V.; Ungaro, F.; Tatangelo, F.; Leone, A.; Scalia, G.; Budillon, A.; Quaglia, F.; Russo, G. Enhancement of 5-FU sensitivity by the proapoptotic rpL3 gene in p53 null colon cancer cells through combined polymer nanoparticles. Oncotarget 2016, 7, 79670–79687. [Google Scholar] [CrossRef] [PubMed]
- Yoshikawa, M.; Fujii, Y.R. Human Ribosomal RNA-Derived Resident MicroRNAs as the Transmitter of Information upon the Cytoplasmic Cancer Stress. Biomed. Res. Int. 2016, 2016, 7562085. [Google Scholar] [CrossRef] [PubMed]
- Zheng, D.; Zhang, J.; Ni, J.; Luo, J.; Wang, J.; Tang, L.; Zhang, L.; Wang, L.; Xu, J.; Su, B.; et al. Small nucleolar RNA 78 promotes the tumorigenesis in non-small cell lung cancer. J. Exp. Clin. Cancer Res. 2015, 34, 49. [Google Scholar] [CrossRef] [PubMed]
- Carotenuto, P.; Fassan, M.; Pandolfo, R.; Lampis, A.; Vicentini, C.; Cascione, L.; Paulus-Hock, V.; Boulter, L.; Guest, R.; Quagliata, L.; et al. Wnt signalling modulates transcribed-ultraconserved regions in hepatobiliary cancers. Gut 2017, 66, 1268–1277. [Google Scholar] [CrossRef] [PubMed]
- Pirogov, S.A.; Gvozdev, V.A.; Klenov, M.S. Long Noncoding RNAs and Stress Response in the Nucleolus. Cells 2019, 8, 668. [Google Scholar] [CrossRef]
- Xu, B.; Li, H.; Perry, J.M.; Singh, V.P.; Unruh, J.; Yu, Z.; Zakari, M.; McDowell, W.; Li, L.; Gerton, J.L. Ribosomal DNA copy number loss and sequence variation in cancer. PLoS Genet. 2017, 13, e1006771. [Google Scholar] [CrossRef]
- Quin, J.E.; Devlin, J.R.; Cameron, D.; Hannan, K.M.; Pearson, R.B.; Hannan, R.D. Targeting the nucleolus for cancer intervention. Biochim. Biophys. Acta 2014, 1842, 802–816. [Google Scholar] [CrossRef]
- Cheung-Ong, K.; Giaever, G.; Nislow, C. DNA-damaging agents in cancer chemotherapy: Serendipity and chemical biology. Chem. Biol. 2013, 20, 648–659. [Google Scholar] [CrossRef]
- Burger, K.; Mühl, B.; Harasim, T.; Rohrmoser, M.; Malamoussi, A.; Orban, M.; Kellner, M.; Gruber-Eber, A.; Kremmer, E.; Hölzel, M.; et al. Chemotherapeutic drugs inhibit ribosome biogenesis at various levels. J. Biol. Chem. 2010, 285, 12416–12425. [Google Scholar] [CrossRef]
- Sun, X.X.; Dai, M.S.; Lu, H. 5-fluorouracil activation of p53 involves an MDM2-ribosomal protein interaction. J. Biol. Chem. 2007, 282, 8052–8059. [Google Scholar] [CrossRef]
- Burger, K.; Mühl, B.; Rohrmoser, M.; Coordes, B.; Heidemann, M.; Kellner, M.; Gruber-Eber, A.; Heissmeyer, V.; Strässer, K.; Eick, D. Cyclin-dependent kinase 9 links RNA polymerase II transcription to processing of ribosomal RNA. J. Biol. Chem. 2013, 288, 21173–21183. [Google Scholar] [CrossRef] [PubMed]
- Beljanski, M.; Crochet, S. The anticancer agent pb-100 concentrates in the nucleus and nucleoli of human glioblastoma cells but does not enter normal astrocytes. Int. J. Oncol. 1995, 7, 81–85. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; von Mikecz, A. Formation of nucleoplasmic protein aggregates impairs nuclear function in response to SiO2 nanoparticles. Exp. Cell Res. 2005, 305, 51–62. [Google Scholar] [CrossRef] [PubMed]
- Paunesku, T.; Vogt, S.; Lai, B.; Maser, J.; Stojićević, N.; Thurn, K.T.; Osipo, C.; Liu, H.; Legnini, D.; Wang, Z.; et al. Intracellular distribution of TiO2-DNA oligonucleotide nanoconjugates directed to nucleolus and mitochondria indicates sequence specificity. Nano Lett. 2007, 7, 596–601. [Google Scholar] [CrossRef] [PubMed]
- Kodiha, M.; Mahboubi, H.; Maysinger, D.; Stochaj, U. Gold Nanoparticles Impinge on Nucleoli and the Stress Response in MCF7 Breast Cancer Cells. Nanobiomedicine (Rij) 2016, 3, 3. [Google Scholar] [CrossRef] [PubMed]
- Esposito, V.; Russo, A.; Vellecco, V.; Bucci, M.; Russo, G.; Mayol, L.; Virgilio, A.; Galeone, A. Thrombin binding aptamer analogues containing inversion of polarity sites endowed with antiproliferative and anti-motility properties against Calu-6 cells. Biochim. Biophys. Acta Gen. Subj. 2018, 1862, 2645–2650. [Google Scholar] [CrossRef] [PubMed]
- Pirota, V.; Nadai, M.; Doria, F.; Richter, S.N. Naphthalene Diimides as Multimodal G-Quadruplex-Selective Ligands. Molecules 2019, 24, 426. [Google Scholar] [CrossRef]
- Drygin, D.; Siddiqui-Jain, A.; O’Brien, S.; Schwaebe, M.; Lin, A.; Bliesath, J.; Ho, C.B.; Proffitt, C.; Trent, K.; Whitten, J.P.; et al. Anticancer activity of CX-3543: A direct inhibitor of rRNA biogenesis. Cancer Res. 2009, 69, 7653–7661. [Google Scholar] [CrossRef]
- Drygin, D.; Lin, A.; Bliesath, J.; Ho, C.B.; O’Brien, S.E.; Proffitt, C.; Omori, M.; Haddach, M.; Schwaebe, M.K.; Siddiqui-Jain, A.; et al. Targeting RNA polymerase I with an oral small molecule CX-5461 inhibits ribosomal RNA synthesis and solid tumor growth. Cancer Res. 2011, 71, 1418–1430. [Google Scholar] [CrossRef]
- Andrews, W.J.; Panova, T.; Normand, C.; Gadal, O.; Tikhonova, I.G.; Panov, K.I. Old drug, new target: ellipticines selectively inhibit RNA polymerase I transcription. J. Biol. Chem. 2013, 288, 4567–4582. [Google Scholar] [CrossRef]
- Peltonen, K.; Colis, L.; Liu, H.; Jäämaa, S.; Zhang, Z.; Af Hällström, T.; Moore, H.M.; Sirajuddin, P.; Laiho, M. Small molecule BMH-compounds that inhibit RNA polymerase I and cause nucleolar stress. Mol. Cancer Ther. 2014, 13, 2537–2546. [Google Scholar] [CrossRef] [PubMed]
- Colis, L.; Peltonen, K.; Sirajuddin, P.; Liu, H.; Sanders, S.; Ernst, G.; Barrow, J.C.; Laiho, M. DNA intercalator BMH-21 inhibits RNA polymerase I independent of DNA damage response. Oncotarget 2014, 5, 4361–4369. [Google Scholar] [CrossRef] [PubMed]
- Peltonen, K.; Colis, L.; Liu, H.; Trivedi, R.; Moubarek, M.S.; Moore, H.M.; Bai, B.; Rudek, M.A.; Bieberich, C.J.; Laiho, M. A targeting modality for destruction of RNA polymerase I that possesses anticancer activity. Cancer Cell 2014, 25, 77–90. [Google Scholar] [CrossRef] [PubMed]
- Wei, T.; Najmi, S.M.; Liu, H.; Peltonen, K.; Kucerova, A.; Schneider, D.A.; Laiho, M. Small-Molecule Targeting of RNA Polymerase I Activates a Conserved Transcription Elongation Checkpoint. Cell Rep. 2018, 23, 404–414. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.; Xu, L.; Qi, L.; Tian, H.; Yi, D.; Yu, Y.; Liu, S.; Li, S.; Xu, Y.; Wang, C. BMH-21 inhibits viability and induces apoptosis by p53-dependent nucleolar stress responses in SKOV3 ovarian cancer cells. Oncol. Rep. 2017, 38, 859–865. [Google Scholar] [CrossRef] [PubMed]
- Wall, M.; Poortinga, G.; Stanley, K.L.; Lindemann, R.K.; Bots, M.; Chan, C.J.; Bywater, M.J.; Kinross, K.M.; Astle, M.V.; Waldeck, K.; et al. The mTORC1 inhibitor everolimus prevents and treats Eμ-Myc lymphoma by restoring oncogene-induced senescence. Cancer Discov. 2013, 3, 82–95. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.C.; Hannan, K.M.; Riddell, K.; Ng, P.Y.; Peck, A.; Lee, R.S.; Hung, S.; Astle, M.V.; Bywater, M.; Wall, M.; et al. AKT promotes rRNA synthesis and cooperates with c-MYC to stimulate ribosome biogenesis in cancer. Sci. Signal. 2011, 4, ra56. [Google Scholar] [CrossRef] [PubMed]
- Andreeff, M.; Kelly, K.R.; Yee, K.; Assouline, S.; Strair, R.; Popplewell, L.; Bowen, D.; Martinelli, G.; Drummond, M.W.; Vyas, P.; et al. Results of the Phase I Trial of RG7112, a Small-Molecule MDM2 Antagonist in Leukemia. Clin. Cancer Res. 2016, 22, 868–876. [Google Scholar] [CrossRef]
- Cheok, C.F.; Verma, C.S.; Baselga, J.; Lane, D.P. Translating p53 into the clinic. Nat. Rev. Clin. Oncol. 2011, 8, 25–37. [Google Scholar] [CrossRef]
- Ding, Q.; Zhang, Z.; Liu, J.J.; Jiang, N.; Zhang, J.; Ross, T.M.; Chu, X.J.; Bartkovitz, D.; Podlaski, F.; Janson, C.; et al. Discovery of RG7388, a potent and selective p53-MDM2 inhibitor in clinical development. J. Med. Chem. 2013, 56, 5979–5983. [Google Scholar] [CrossRef]
- Wang, S.; Zhao, Y.; Aguilar, A.; Bernard, D.; Yang, C.Y. Targeting the MDM2-p53 Protein-Protein Interaction for New Cancer Therapy: Progress and Challenges. Cold Spring Harb. Perspect. Med. 2017, 7, a026245. [Google Scholar] [CrossRef] [PubMed]
- Canon, J.; Osgood, T.; Olson, S.H.; Saiki, A.Y.; Robertson, R.; Yu, D.; Eksterowicz, J.; Ye, Q.; Jin, L.; Chen, A.; et al. The MDM2 Inhibitor AMG 232 Demonstrates Robust Antitumor Efficacy and Potentiates the Activity of p53-Inducing Cytotoxic Agents. Mol. Cancer Ther. 2015, 14, 649–658. [Google Scholar] [CrossRef] [PubMed]
- Holzer, P.; Masuya, K.; Furet, P.; Kallen, J.; Valat-Stachyra, T.; Ferretti, S.; Berghausen, J.; Bouisset-Leonard, M.; Buschmann, N.; Pissot-Soldermann, C.; et al. Discovery of a Dihydroisoquinolinone Derivative (NVP-CGM097): A Highly Potent and Selective MDM2 Inhibitor Undergoing Phase 1 Clinical Trials in p53wt Tumors. J. Med. Chem. 2015, 58, 6348–6358. [Google Scholar] [CrossRef] [PubMed]
- Tisato, V.; Voltan, R.; Gonelli, A.; Secchiero, P.; Zauli, G. MDM2/X inhibitors under clinical evaluation: perspectives for the management of hematological malignancies and pediatric cancer. J. Hematol. Oncol. 2017, 10, 133. [Google Scholar] [CrossRef] [PubMed]
- Ratnadiwakara, M.; Änkö, M. mRNA Stability Assay Using Transcription Inhibition by Actinomycin D in Mouse Pluripotent Stem Cells. Bio-protocol, 2018; 8, e133. [Google Scholar]
- Latonen, L.; Moore, H.M.; Bai, B.; Jäämaa, S.; Laiho, M. Proteasome inhibitors induce nucleolar aggregation of proteasome target proteins and polyadenylated RNA by altering ubiquitin availability. Oncogene 2011, 30, 790–805. [Google Scholar] [CrossRef] [PubMed]
- Navon, A.; Ciechanover, A. The 26 S proteasome: From basic mechanisms to drug targeting. J. Biol. Chem. 2009, 284, 33713–33718. [Google Scholar] [CrossRef]
- Galimberti, V.; Kinor, N.; Shav-Tal, Y.; Biggiogera, M.; Brüning, A. The stress-inducible transcription factor ATF4 accumulates at specific rRNA-processing nucleolar regions after proteasome inhibition. Eur. J. Cell Biol. 2016, 95, 389–400. [Google Scholar] [CrossRef]
- Maiolino, S.; Russo, A.; Pagliara, V.; Conte, C.; Ungaro, F.; Russo, G.; Quaglia, F. Biodegradable nanoparticles sequentially decorated with Polyethyleneimine and Hyaluronan for the targeted delivery of docetaxel to airway cancer cells. J. Nanobiotechnol. 2015, 13, 29. [Google Scholar] [CrossRef]
- d’Angelo, I.; Costabile, G.; Durantie, E.; Brocca, P.; Rondelli, V.; Russo, A.; Russo, G.; Miro, A.; Quaglia, F.; Petri-Fink, A.; et al. Hybrid Lipid/Polymer Nanoparticles for Pulmonary Delivery of siRNA: Development and Fate Upon In Vitro Deposition on the Human Epithelial Airway Barrier. J. Aerosol. Med. Pulm. Drug Deliv. 2018, 31, 170–181. [Google Scholar] [CrossRef]
- Esposito, V.; Russo, A.; Amato, T.; Varra, M.; Vellecco, V.; Bucci, M.; Russo, G.; Virgilio, A.; Galeone, A. Backbone modified TBA analogues endowed with antiproliferative activity. Biochim. Biophys. Acta Gen. Subj. 2017, 1861, 1213–1221. [Google Scholar] [CrossRef]
- Virgilio, A.; Russo, A.; Amato, T.; Russo, G.; Mayol, L.; Esposito, V.; Galeone, A. Monomolecular G-quadruplex structures with inversion of polarity sites: new topologies and potentiality. Nucleic Acids Res. 2017, 45, 8156–8166. [Google Scholar] [CrossRef] [PubMed]
- Esposito, V.; Russo, A.; Amato, T.; Vellecco, V.; Bucci, M.; Mayol, L.; Russo, G.; Virgilio, A.; Galeone, A. The “Janus face” of the thrombin binding aptamer: Investigating the anticoagulant and antiproliferative properties through straightforward chemical modifications. Bioorg Chem. 2018, 76, 202–209. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Di Antonio, M.; McKinney, S.; Mathew, V.; Ho, B.; O’Neil, N.J.; Santos, N.D.; Silvester, J.; Wei, V.; Garcia, J.; et al. CX-5461 is a DNA G-quadruplex stabilizer with selective lethality in BRCA1/2 deficient tumours. Nat. Commun. 2017, 8, 14432. [Google Scholar] [PubMed]
- Drygin, D.; O’Brien, S.E.; Hannan, R.D.; McArthur, G.A.; Von Hoff, D.D. Targeting the nucleolus for cancer-specific activation of p53. Drug Discov. Today 2014, 19, 259–265. [Google Scholar] [PubMed]
- Peltonen, K.; Colis, L.; Liu, H.; Jäämaa, S.; Moore, H.M.; Enbäck, J.; Laakkonen, P.; Vaahtokari, A.; Jones, R.J.; af Hällström, T.M.; et al. Identification of novel p53 pathway activating small-molecule compounds reveals unexpected similarities with known therapeutic agents. PLoS ONE 2010, 5, e12996. [Google Scholar] [CrossRef] [PubMed]
- Voit, R.; Hoffmann, M.; Grummt, I. Phosphorylation by G1-specific cdk-cyclin complexes activates the nucleolar transcription factor UBF. EMBO J. 1999, 18, 1891–1899. [Google Scholar] [CrossRef] [PubMed]
- Zhai, W.; Comai, L. Repression of RNA polymerase I transcription by the tumor suppressor p53. Mol. Cell Biol. 2000, 20, 5930–5938. [Google Scholar] [CrossRef] [PubMed]
- Kruhlak, M.; Crouch, E.E.; Orlov, M.; Montaño, C.; Gorski, S.A.; Nussenzweig, A.; Misteli, T.; Phair, R.D.; Casellas, R. The ATM repair pathway inhibits RNA polymerase I transcription in response to chromosome breaks. Nature 2007, 447, 730–734. [Google Scholar] [CrossRef]
- Ma, H.; Pederson, T. The nucleolus stress response is coupled to an ATR-Chk1-mediated G2 arrest. Mol. Biol. Cell 2013, 24, 1334–1342. [Google Scholar] [CrossRef]
- Calkins, A.S.; Iglehart, J.D.; Lazaro, J.B. DNA damage-induced inhibition of rRNA synthesis by DNA-PK and PARP-1. Nucleic Acids Res. 2013, 41, 7378–7386. [Google Scholar] [CrossRef]
- Bierhoff, H.; Dundr, M.; Michels, A.A.; Grummt, I. Phosphorylation by casein kinase 2 facilitates rRNA gene transcription by promoting dissociation of TIF-IA from elongating RNA polymerase I. Mol. Cell Biol. 2008, 28, 4988–4998. [Google Scholar] [CrossRef] [PubMed]
- Bakshi, R.; Zaidi, S.K.; Pande, S.; Hassan, M.Q.; Young, D.W.; Montecino, M.; Lian, J.B.; van Wijnen, A.J.; Stein, J.L.; Stein, G.S. The leukemogenic t(8;21) fusion protein AML1-ETO controls rRNA genes and associates with nucleolar-organizing regions at mitotic chromosomes. J. Cell Sci. 2008, 121, 3981–3990. [Google Scholar] [CrossRef] [PubMed]
- Anglin, I.; Passaniti, A. Runx protein signaling in human cancers. Cancer Treat. Res. 2004, 119, 189–215. [Google Scholar] [PubMed]
- Fukawa, T.; Ono, M.; Matsuo, T.; Uehara, H.; Miki, T.; Nakamura, Y.; Kanayama, H.O.; Katagiri, T. DDX31 regulates the p53-HDM2 pathway and rRNA gene transcription through its interaction with NPM1 in renal cell carcinomas. Cancer Res. 2012, 72, 5867–5877. [Google Scholar] [CrossRef] [PubMed]
- Delloye-Bourgeois, C.; Goldschneider, D.; Paradisi, A.; Therizols, G.; Belin, S.; Hacot, S.; Rosa-Calatrava, M.; Scoazec, J.Y.; Diaz, J.J.; Bernet, A.; et al. Nucleolar localization of a netrin-1 isoform enhances tumor cell proliferation. Sci. Signal. 2012, 5, ra57. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Forys, J.T.; Miceli, A.P.; Gwinn, A.S.; Weber, J.D. Identification of DHX33 as a mediator of rRNA synthesis and cell growth. Mol. Cell Biol. 2011, 31, 4676–4691. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Liang, P.; Geng, H.; Wang, Z.; Li, L.; Cheng, S.H.; Ying, J.; Su, X.; Ng, K.M.; Ng, M.H.; et al. A novel 19q13 nucleolar zinc finger protein suppresses tumor cell growth through inhibiting ribosome biogenesis and inducing apoptosis but is frequently silenced in multiple carcinomas. Mol. Cancer Res. 2012, 10, 925–936. [Google Scholar] [CrossRef]
- Hannan, K.M.; Brandenburger, Y.; Jenkins, A.; Sharkey, K.; Cavanaugh, A.; Rothblum, L.; Moss, T.; Poortinga, G.; McArthur, G.A.; Pearson, R.B.; et al. mTOR-dependent regulation of ribosomal gene transcription requires S6K1 and is mediated by phosphorylation of the carboxy-terminal activation domain of the nucleolar transcription factor UBF. Mol. Cell Biol. 2003, 23, 8862–8877. [Google Scholar] [CrossRef]
- Brown, C.J.; Cheok, C.F.; Verma, C.S.; Lane, D.P. Reactivation of p53: from peptides to small molecules. Trends Pharmacol. Sci. 2011, 32, 53–62. [Google Scholar] [CrossRef]
- Inoue, K.; Fry, E.A. Aberrant splicing of the DMP1-ARF-MDM2-p53 pathway in cancer. Int. J. Cancer 2016, 139, 33–41. [Google Scholar] [CrossRef]
- Patnaik, A.; Tolcher, A.; Beeram, M.; Nemunaitis, J.; Weiss, G.J.; Bhalla, K.; Agrawal, M.; Nichols, G.; Middleton, S.; Beryozkina, A.; et al. Clinical pharmacology characterization of RG7112, an MDM2 antagonist, in patients with advanced solid tumors. Cancer Chemother. Pharmacol. 2015, 76, 587–595. [Google Scholar] [CrossRef] [PubMed]
| Drug | Class of Compounds | Mechanism of Action | Cancer Type |
|---|---|---|---|
| Doxorubicin | Anthracycline | rDNA intercalating agent/ topoisomerase II inhibitors | Haematological cancers, bladder, breast, stomach, lung, ovarian and thyroid cancer, sarcoma [29] |
| Cisplatin, Oxaliplatin | Platinum compound | rDNA crosslinking agent | Sarcoma, lymphoma, carcinoma [37,78,79] |
| Actinomycin D | Antibiotic | DNA intercalating agent | Wilms’ tumour, sarcoma [37] |
| Mitomycin C | Antibiotic | rDNA alkylating/crosslinking agent | Stomach or pancreatic adenocarcinoma; anal, bladder, breast, cervical, colorectal, head, neck, non-small-cell lung cancer [29] |
| Irinotecan/Topotecan | Camptothecins | Topoisomerase I inhibition | Ovarian, lung, cervical cancer [79] |
| Etoposide | Epipodophyllotoxins | Topoisomerase II inhibition | Sarcoma, glioblastoma, lung, testicular, haematological cancers [79] |
| 5-flurouracil | Pyrimidine nucleotide analogue | Thymidylate synthase/rRNA/rDNA synthesis inhibitor | Colon, rectum, head, neck cancers [37,79,80] |
| Roscovitine/Olomoucine DRB | Cdk inhibitors | Disrupting nucleolar integrity | Adenocarcinoma, B-cell malignancies, breast cancer [81] |
| Flavopereirine (PB-100) | Alkaloid | Accumulating in the nucleoli | Glioblastoma [82] |
| Nanoparticles (SiO2) | Nanoparticles | Inducing nucleolar protein aggregates | ND [83] |
| Nanoparticles (TiO2) | Oligonucleotide-conjugated Nanoparticles | Depleting rDNA | ND [84] |
| Nanoparticles (Gold) | Nanoparticles | Interfering with the transcription of ribosomal DNA | Breast cancer [85] |
| DNA aptamers Naphthalene diimides | G-quadruplex interacting compounds | Binding to rDNA | Breast, lung cancer [86,87] |
| Drug | Class of Compounds | Mechanism of Action | Cancer Type |
|---|---|---|---|
| CX-3543 | Selective inhibitor of RNA Pol I | Targeting and disrupting nucleolin/rDNA G-quadruplex complexes | Carcinoid/neuroendocrine tumours [88] |
| CX-5461 9-Hydroxyellipticine (9HE) BMH-21 | Selective inhibitor of RNA Pol I | Inhibiting RNA Pol I activity | Haematological cancers [32,89], breast cancer [90] Pre-clinical models [91,92,93,94,95] |
| Rapamycin Everolimus | mTOR signalling inhibitor | Inhibiting mTOR signalling resulting in inhibition of ribosome biogenesis | Renal cell carcinoma, breast cancer and lymphoma [96] |
| AKTi-1/2 MK-2206 | AKT signalling inhibitor | Inhibiting AKT signalling resulting in suppression of rDNA gene transcription | Non–small cell lung cancer [97] |
| Nutlin-1/2/3, RG7112 RG7388 PXN727 PXN822 | Nutlins and derivatives | Targeting the MDM2-P53 interaction | Haematological tumours, solid tumours [98,99]; osteosarcoma, head and neck cancer [100]; pre-clinical models [99] |
| MI-77301 MI-219 | Spirooxindole-based compound | Targeting the MDM2-P53 interaction | Osteosarcoma, acute leukemia, prostate and colon cancer cells [101]; hematologic neoplasms and advanced solid tumors [99] |
| MK-8242 | antimetabolite analogue of cytidine | Targeting the MDM2-P53 interaction | Solid tumors and haematological cancers [101] |
| AMG 232 | Piperidinone derivative | Targeting the MDM2-P53 interaction | Breast cancer [102] |
| CGM097 | dihydroisoquinolinone derivative | Targeting the MDM2-P53 interaction | Hematological and pediatric cancers [103] |
| DS3032b HDM201 | Imidazopyrrolidinone derivative | Targeting the MDM2-P53 interaction | Haematological malignancies and advanced solid tumors [104] |
| JNJ-26854165 | Tryptamine derivative | Targeting the MDM2-P53 interaction | Solid tumours [99] |
| RITA | Thiophen derivative | Targeting the MDM2-P53 interaction | Pre-clinical models [99] |
| p53-SLP | P53-synthetic long peptide vaccine | Stimulating immunoresponse against P53 | Ovarian and colorectal cancer [99] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carotenuto, P.; Pecoraro, A.; Palma, G.; Russo, G.; Russo, A. Therapeutic Approaches Targeting Nucleolus in Cancer. Cells 2019, 8, 1090. https://doi.org/10.3390/cells8091090
Carotenuto P, Pecoraro A, Palma G, Russo G, Russo A. Therapeutic Approaches Targeting Nucleolus in Cancer. Cells. 2019; 8(9):1090. https://doi.org/10.3390/cells8091090
Chicago/Turabian StyleCarotenuto, Pietro, Annalisa Pecoraro, Gaetano Palma, Giulia Russo, and Annapina Russo. 2019. "Therapeutic Approaches Targeting Nucleolus in Cancer" Cells 8, no. 9: 1090. https://doi.org/10.3390/cells8091090
APA StyleCarotenuto, P., Pecoraro, A., Palma, G., Russo, G., & Russo, A. (2019). Therapeutic Approaches Targeting Nucleolus in Cancer. Cells, 8(9), 1090. https://doi.org/10.3390/cells8091090





