MiR-34b-5p Mediates the Proliferation and Differentiation of Myoblasts by Targeting IGFBP2
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals and Cells
2.2. Cell Culture and Transfection
2.3. Plasmid Construction
2.4. Dual-Luciferase Reporter Assay
2.5. RNA Isolation, Complementary DNA (cDNA) Synthesis, and Quantitative Real-Time PCR (qRT-PCR)
2.6. EdU (5-Ethynyl-2′-Deoxyuridine) Assay
2.7. Flow Cytometric Analysis
2.8. Western Blotting
2.9. Immunofluorescence
2.10. Statistical Analysis
3. Results
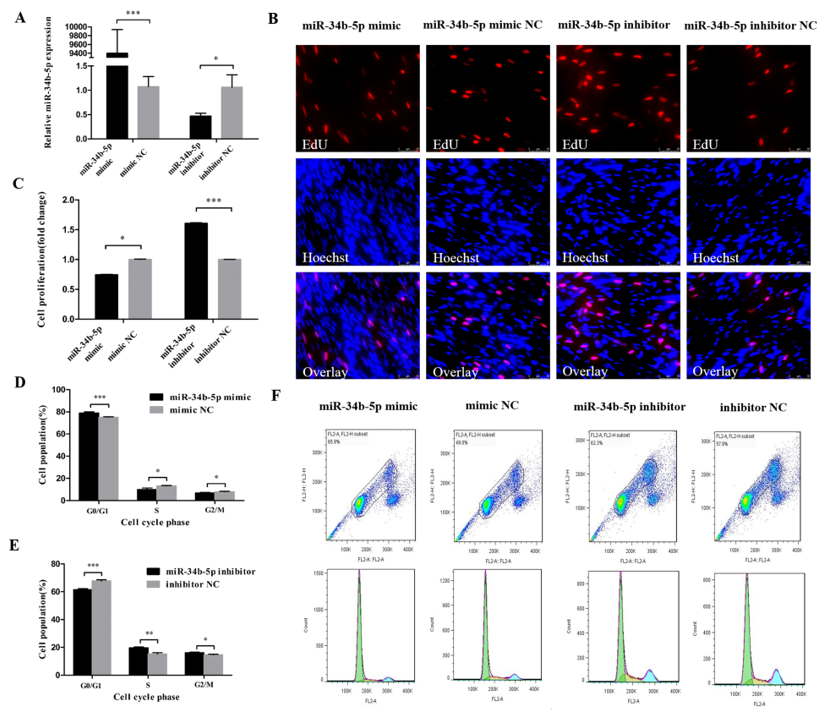
3.1. miR-34b-5p Represses the Cell Cycle Progression of Myoblasts
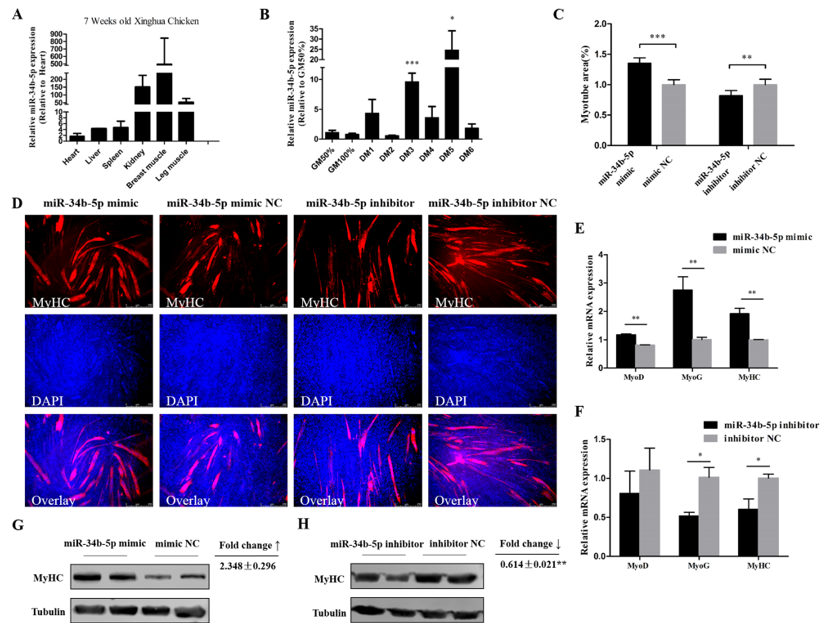
3.2. miR-34b-5p Promotes the Formation of Myotubes
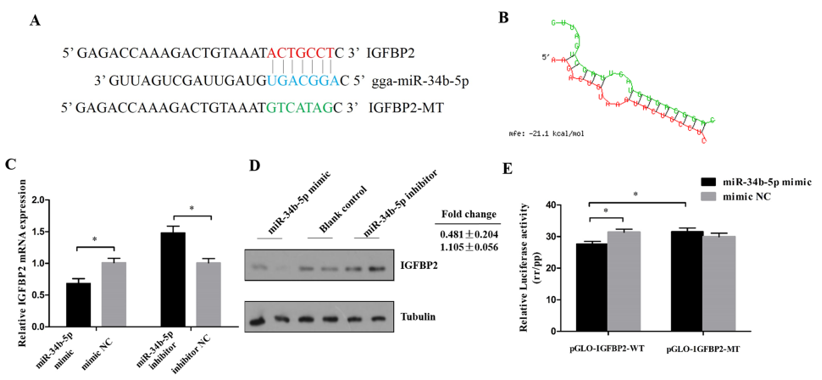
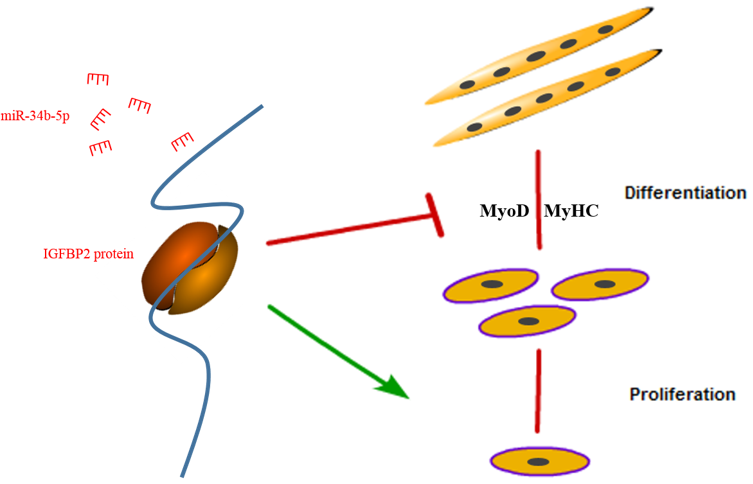
3.3. IGFBP2 is a Direct Target of miR-34b-5p
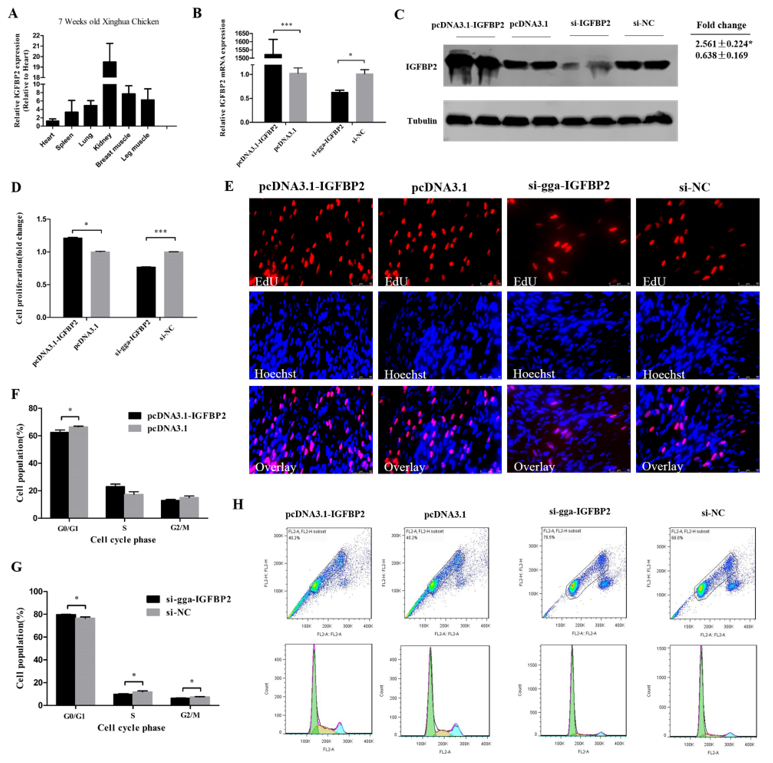
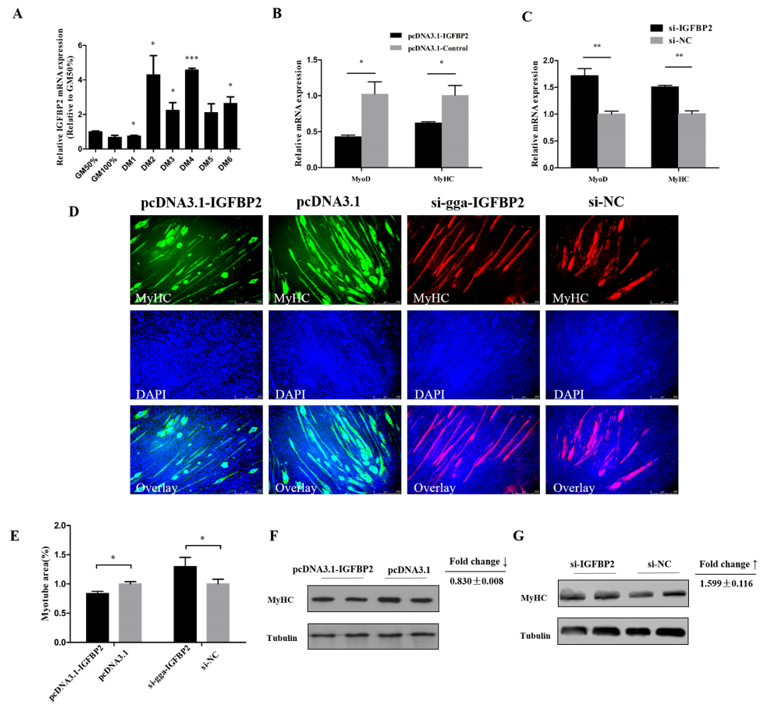
3.4. IGFBP2 Facilitates the Cell Cycle Progression of Myoblasts
3.5. IGFBP2 is an Inhibitor of Myotube Formation
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Saunders, M.A.; Liang, H.; Li, W. Human polymorphism at microRNAs and microRNA target sites. Proc. Natl. Acad. Sci. USA 2007, 104, 3300–3305. [Google Scholar] [CrossRef] [PubMed]
- Cullen, B.R. MicroRNAs as mediators of viral evasion of the immune system. Nat. Immunol. 2013, 14, 205–210. [Google Scholar] [CrossRef] [PubMed]
- Pandey, R.; Ahmed, R.P.H. MicroRNAs Inducing Proliferation of Quiescent Adult Cardiomyocytes. Cardiovasc. Regener. Med. 2015, 2, e519. [Google Scholar]
- Taft, R.J.; Pang, K.C.; Mercer, T.R.; Dinger, M.; Mattick, J.S. Non-coding RNAs: Regulators of disease. J. Pathol. 2010, 220, 126–139. [Google Scholar] [CrossRef]
- Yu, J.; Peng, J.; Luan, Z.; Zheng, F.; Su, W. MicroRNAs as a Novel Tool in the Diagnosis of Liver Lipid Dysregulation and Fatty Liver Disease. Molecules 2019, 24, 230. [Google Scholar] [CrossRef] [PubMed]
- Ouyang, H.; He, X.; Li, G.; Xu, H.; Jia, X.; Nie, Q.; Zhang, X. Deep Sequencing Analysis of miRNA Expression in Breast Muscle of Fast-Growing and Slow-Growing Broilers. Int. J. Mol. Sci. 2015, 16, 16242–16262. [Google Scholar] [CrossRef] [PubMed]
- Jebessa, E.; Ouyang, H.; Abdalla, B.A.; Li, Z.; Abdullahi, A.Y.; Liu, Q.; Nie, Q.; Zhang, X. Characterization of miRNA and their target gene during chicken embryo skeletal muscle development. Oncotarget 2018, 9, 17309–17324. [Google Scholar] [CrossRef] [PubMed]
- He, L.; He, X.; Lim, L.P.; De Stanchina, E.; Xuan, Z.; Liang, Y.; Xue, W.; Zender, L.; Magnus, J.; Ridzon, D.; et al. A microRNA component of the p53 tumour suppressor network. Nature 2007, 447, 1116–1130. [Google Scholar] [CrossRef] [PubMed]
- Bommer, G.T.; Gerin, I.; Feng, Y.; Kaczorowski, A.J.; Kuick, R.; Love, R.E.; Zhai, Y.; Giordano, T.J.; Qin, Z.S.; Moore, B.B.; et al. p53-mediated activation of miRNA34 candidate tumor-suppressor genes. Curr. Biol. 2007, 17, 1298–1307. [Google Scholar] [CrossRef] [PubMed]
- Hermeking, H. The miR-34 family in cancer and apoptosis. Cell Death Differ. 2010, 17, 193–199. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Luo, Q.; Xu, H.; Zheng, M.; Abdalla, B.A.; Feng, M.; Cai, B.; Zhang, X.; Nie, Q.; Zhang, X. MiR-34b-5p Suppresses Melanoma Differentiation-Associated Gene 5 (MDA5) Signaling Pathway to Promote Avian Leukosis Virus Subgroup J (ALV-J)-Infected Cells Proliferaction and ALV-J Replication. Front. Cell. Infect. Microbiol. 2017, 7. [Google Scholar] [CrossRef][Green Version]
- Huang, C.W.; Li, Y.H.; Hu, S.Y.; Chi, J.R.; Lin, G.H.; Lin, C.C.; Gong, H.Y.; Chen, J.Y.; Chen, R.H.; Chang, S.J.; et al. Differential expression patterns of growth-related microRNAs in the skeletal muscle of Nile tilapia (Oreochromis niloticus). J. Anim. Sci. 2012, 90, 4266–4279. [Google Scholar] [CrossRef]
- Bae, Y.; Yang, T.; Zeng, H.; Campeau, P.M.; Chen, Y.; Bertin, T.; Dawson, B.C.; Munivez, E.; Tao, J.; Lee, B.H. miRNA-34c regulates Notch signaling during bone development. Hum. Mol. Genet. 2012, 21, 2991–3000. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.; Shi, Y.; Zheng, L.; Zhou, B.; Inose, H.; Wang, J.; Guo, X.E.; Grosschedl, R.; Karsenty, G. miR-34s inhibit osteoblast proliferation and differentiation in the mouse by targeting SATB2. J. Cell Biol. 2012, 197, 509–521. [Google Scholar] [CrossRef]
- Wang, P.; Xu, J.; Hou, Z.; Wang, F.; Song, Y.; Wang, J.; Zhu, H.; Jin, H. miRNA-34a promotes proliferation of human pulmonary artery smooth muscle cells by targeting PDGFRA. Cell Prolif. 2016, 49, 484–493. [Google Scholar] [CrossRef]
- Chen, Q.; Yang, F.; Guo, M.; Wen, G.; Zhang, C.; Le Anh, L.; Zhu, J.; Xiao, Q.; Zhang, L. miRNA-34a reduces neointima formation through inhibiting smooth muscle cell proliferation and migration. J. Mol. Cell. Cardiol. 2015, 89, 75–86. [Google Scholar] [CrossRef]
- Choe, N.; Kwon, J.; Kim, Y.S.; Eom, G.H.; Ahn, Y.K.; Baik, Y.H.; Park, H.; Kook, H. The microRNA miR-34c inhibits vascular smooth muscle cell proliferation and neointimal hyperplasia by targeting stem cell factor. Cell Signal. 2015, 27, 1056–1065. [Google Scholar] [CrossRef]
- Hou, L.; Xu, J.; Li, H.; Ou, J.; Jiao, Y.; Hu, C.; Wang, C. MiR-34c represses muscle development by forming a regulatory loop with Notch1. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Baxter, R.C. Insulin-like growth factor (IGF)-binding proteins: Interactions with IGFs and intrinsic bioactivities. Am. J. Physiol. Endocrinol. Metab. 2000, 278, E967–E976. [Google Scholar] [CrossRef] [PubMed]
- Pickard, A.; McCance, D.J. IGF-binding protein 2-oncogene or tumor suppressor? Front. Endocrinol. 2015, 6. [Google Scholar] [CrossRef]
- Schoen, T.J.; Mazuruk, K.; Waldbillig, R.J.; Potts, J.; Beebe, D.C.; Chader, G.J.; Rodriguez, I.R. Cloning and characterization of a chick embryo cDNA and gene for IGF-binding protein-2. J. Mol. Endocrinol. 1995, 15, 49–59. [Google Scholar] [CrossRef] [PubMed]
- Gerrard, D.E.; Okamura, C.S.; Grant, A.L. Expression and location of IGF binding proteins-2, -4, and -5 in developing fetal tissues. J. Anim. Sci. 1999, 77, 1431–1441. [Google Scholar] [CrossRef][Green Version]
- Sironen, A.I.; Uimari, P.; Serenius, T.; Mote, B.; Rothschild, M.; Vilkki, J. Effect of polymorphisms in candidate genes on reproduction traits in Finnish pig populations. J. Anim. Sci. 2010, 88, 821–827. [Google Scholar] [CrossRef] [PubMed]
- Prasongsook, S.; Choi, I.; Bates, R.O.; Raney, N.E.; Ernst, C.W.; Tumwasorn, S. Association of Insulin-like growth factor binding protein 2 genotypes with growth, carcass and meat quality traits in pigs. J. Anim. Sci. Technol. 2015, 57, 31. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wang, Z.; Ouyang, H.; Chen, X.; Yu, J.; Abdallat, B.A.; Chen BNie, Q. Gga-miR-205a Affecting Myoblast Proliferation and Differentiation by Targeting CDH11. Front. Genet. 2018, 9. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Bass, J.J.; Wilkinson, D.J.; Rankin, D.; Phillips, B.E.; Szewczyk, N.J.; Smith KAtherton, P.J. An overview of technical considerations for Western blotting applications to physiological research. Scand. J. Med. Sci. Sports 2017, 27, 4–25. [Google Scholar] [CrossRef]
- Chen, E.H.; Grote, E.; Mohler, W.; Vignery, A. Cell-cell fusion. FEBS Lett. 2007, 581, 2181–2193. [Google Scholar] [CrossRef]
- Luo, W.; Nie, Q.; Zhang, X. MicroRNAs Involved in Skeletal Muscle Differentiation. J. Genet. Genom. 2013, 40, 107–116. [Google Scholar] [CrossRef]
- Rokavec, M.; Li, H.; Jiang, L.; Hermeking, H. The p53/miR-34 axis in development and disease. J. Mol. Cell Biol. 2014, 6, 214–230. [Google Scholar] [CrossRef]
- Tang, Z.; Qiu, H.; Luo, L.; Liu, N.; Zhong, J.; Kang, K.; Gou, D. miR-34b Modulates Skeletal Muscle Cell Proliferation and Differentiation. J. Cell. Biochem. 2017, 118, 4285–4295. [Google Scholar] [CrossRef]
- Venuti, J.M.; Morris, J.H.; Vivian, J.L.; Olson, E.N.; Klein, W.H. Myogenin is required for late but not early aspects of myogenesis during mouse development. J. Cell Biol. 1995, 128, 563–576. [Google Scholar] [CrossRef]
- Rudnicki, M.A.; Schnegelsberg, P.N.; Stead, R.H.; Braun, T.; Arnold, H.H.; Jaenisch, R. MyoD or Myf-5 is required for the formation of skeletal muscle. Cell 1993, 75, 1351–1359. [Google Scholar] [CrossRef]
- Zammit, P.S. Function of the myogenic regulatory factors Myf5, MyoD, Myogenin and MRF4 in skeletal muscle, satellite cells and regenerative myogenesis. Semin. Cell Dev. Biol. 2017, 72, 19–32. [Google Scholar] [CrossRef] [PubMed]
- Pette, D.; Staron, R.S. Myosin isoforms, muscle fiber types, and transitions. Microsc. Res. Tech. 2000, 50, 500–509. [Google Scholar] [CrossRef]
- Azar, W.J.; Zivkovic, S.; Werther, G.A.; Russo, V.C. IGFBP-2 nuclear translocation is mediated by a functional NLS sequence and is essential for its pro-tumorigenic actions in cancer cells. Oncogene 2014, 33, 578–588. [Google Scholar] [CrossRef]
- Lei, M.M.; Nie, Q.H.; Peng, X.; Zhang, D.X.; Zhang, X.Q. Single nucleotide polymorphisms of the chicken insulin-like factor binding protein 2 gene associated with chicken growth and carcass traits. Poultry Sci. 2005, 84, 1191–1198. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.H.; Li, H.; Zhang, H.; Wang, S.Z.; Wang, Q.G.; Wang, Y.X. Identification of a single nucleotide polymorphism of the insulin-like growth factor binding protein 2 gene and its association with growth and body composition traits in the chicken. J. Anim. Sci. 2006, 84, 2902–2906. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Luo, W.; Abdalla, B.A.; Ouyang, H.; Yu, J.; Hu, F.; Nie, Q.; Zhang, X. miRNA-223 upregulated by MYOD inhibits myoblast proliferation by repressing IGF2 and facilitates myoblast differentiation by inhibiting ZEB1. Cell Death Dis. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Luo, W.; Chen, J.; Li, L.; Ren, X.; Cheng, T.; Lu, S.; Lawal, R.A.; Nie, Q.; Zhang, X.; Hanotte, O. c-Myc inhibits myoblast differentiation and promotes myoblast proliferation and muscle fibre hypertrophy by regulating the expression of its target genes, miRNAs and lincRNAs. Cell Death Differ. 2018. [Google Scholar] [CrossRef] [PubMed]
| Fragment Name | Sequence (5′ to 3′) |
|---|---|
| miR-34b-5p mimic | CAGGCAGUGUAGUUAGCUGAUUG |
| miR-34b-5p inhibitor | CUUUCUGCUUUCUUCUCUGCCUG |
| si-gga-IGFBP2 | GGGAGTGTCTCTCTTTCTT |
| Primer Name | Primer Sequences (5′ to 3′) | Size (bp) |
|---|---|---|
| IGFBP2-WT | F:ACCATTTCCCTCTTCCTCC | 295 |
| R:ACCAAGCATTCAGCTCCAC | ||
| IGFBP2-MT | F:CGAGACCAAAGACTGTAAATTGTGAGTCTTGTGTCCTGCC | 2987 |
| R:GGCAGGACACAAGACTCACAATTTACAGTCTTTGGTCTCG | ||
| IGFBP2 | F:AGCGGCAGATGGGCAAAGT | 184 |
| R:GGGGATGTGGAGGGAGTAGAGG | ||
| β-actin | F:TCATTGTGCTAGGTGCCA | 160 |
| R:TCATTGTGCTAGGTGCCA | ||
| MyoD | F: GCTACTACACGGAATCACCAAAT | 200 |
| R: CTGGGCTCCACTGTCACTCA | ||
| MyoG | F: CGGAGGCTGAAGAAGGTGAA | 320 |
| R: CGGTCCTCTGCCTGGTCAT | ||
| MyHC | F: CTCCTCACGCTTTGGTAA | 213 |
| R: TGATAGTCGTATGGGTTGGT |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Z.; Zhang, X.; Li, Z.; Abdalla, B.A.; Chen, Y.; Nie, Q. MiR-34b-5p Mediates the Proliferation and Differentiation of Myoblasts by Targeting IGFBP2. Cells 2019, 8, 360. https://doi.org/10.3390/cells8040360
Wang Z, Zhang X, Li Z, Abdalla BA, Chen Y, Nie Q. MiR-34b-5p Mediates the Proliferation and Differentiation of Myoblasts by Targeting IGFBP2. Cells. 2019; 8(4):360. https://doi.org/10.3390/cells8040360
Chicago/Turabian StyleWang, Zhijun, Xiaocui Zhang, Zhenhui Li, Bahareldin Ali Abdalla, Yangfeng Chen, and Qinghua Nie. 2019. "MiR-34b-5p Mediates the Proliferation and Differentiation of Myoblasts by Targeting IGFBP2" Cells 8, no. 4: 360. https://doi.org/10.3390/cells8040360
APA StyleWang, Z., Zhang, X., Li, Z., Abdalla, B. A., Chen, Y., & Nie, Q. (2019). MiR-34b-5p Mediates the Proliferation and Differentiation of Myoblasts by Targeting IGFBP2. Cells, 8(4), 360. https://doi.org/10.3390/cells8040360







