Emerging Roles of Long Non-Coding RNAs as Drivers of Brain Evolution
Abstract
1. Introduction
2. Main Text
2.1. Biogenesis and Functional Diversity of lncRNAs
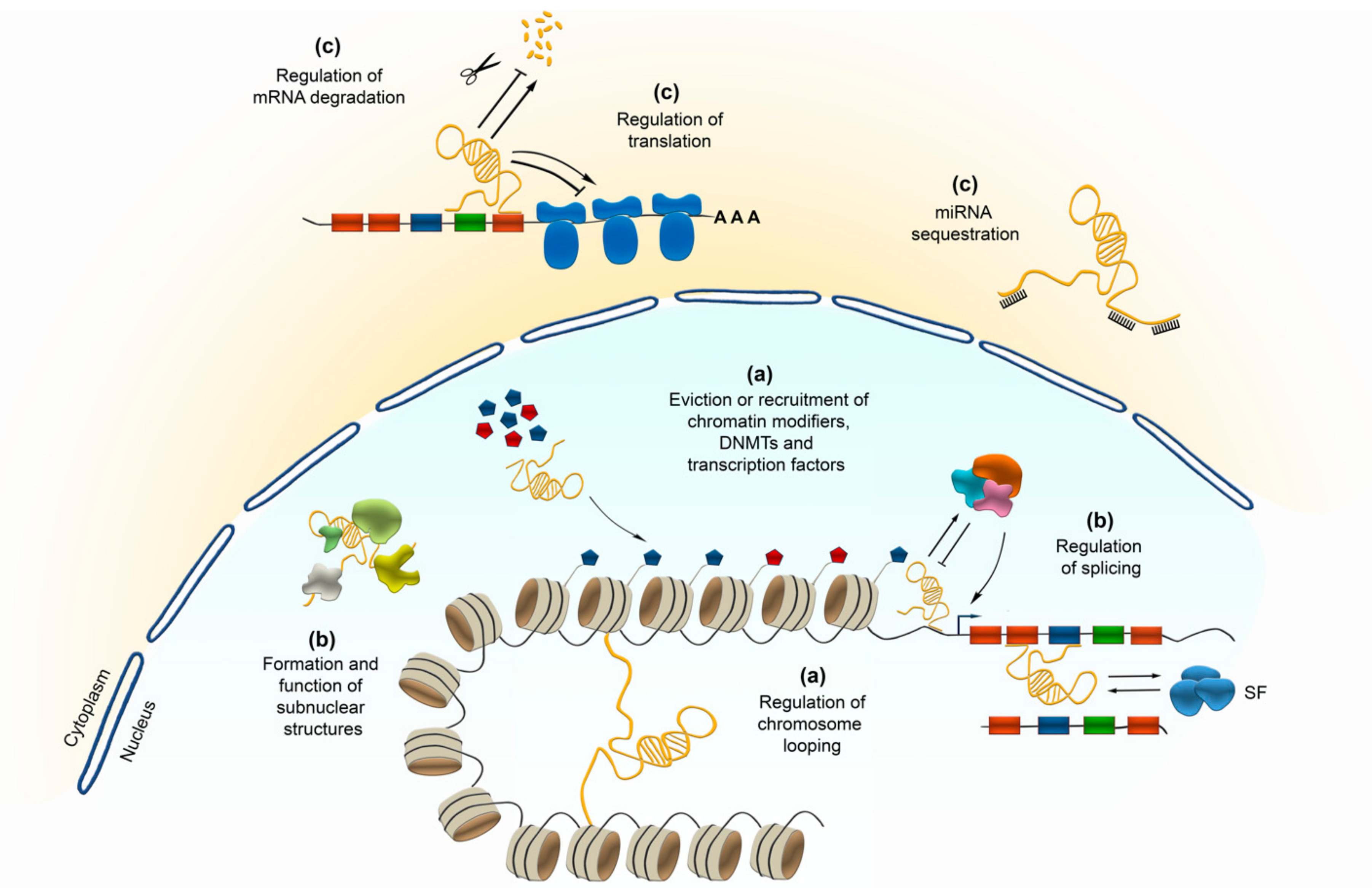
2.2. Transcriptional Control by lncRNAs
2.3. Implications of lncRNAs in Posttranscriptional and Translational Regulation
2.4. Indications for Potential Implications of lncRNAs in Human Brain Evolution
2.5. Evolutionary Innervations of the Human Brain
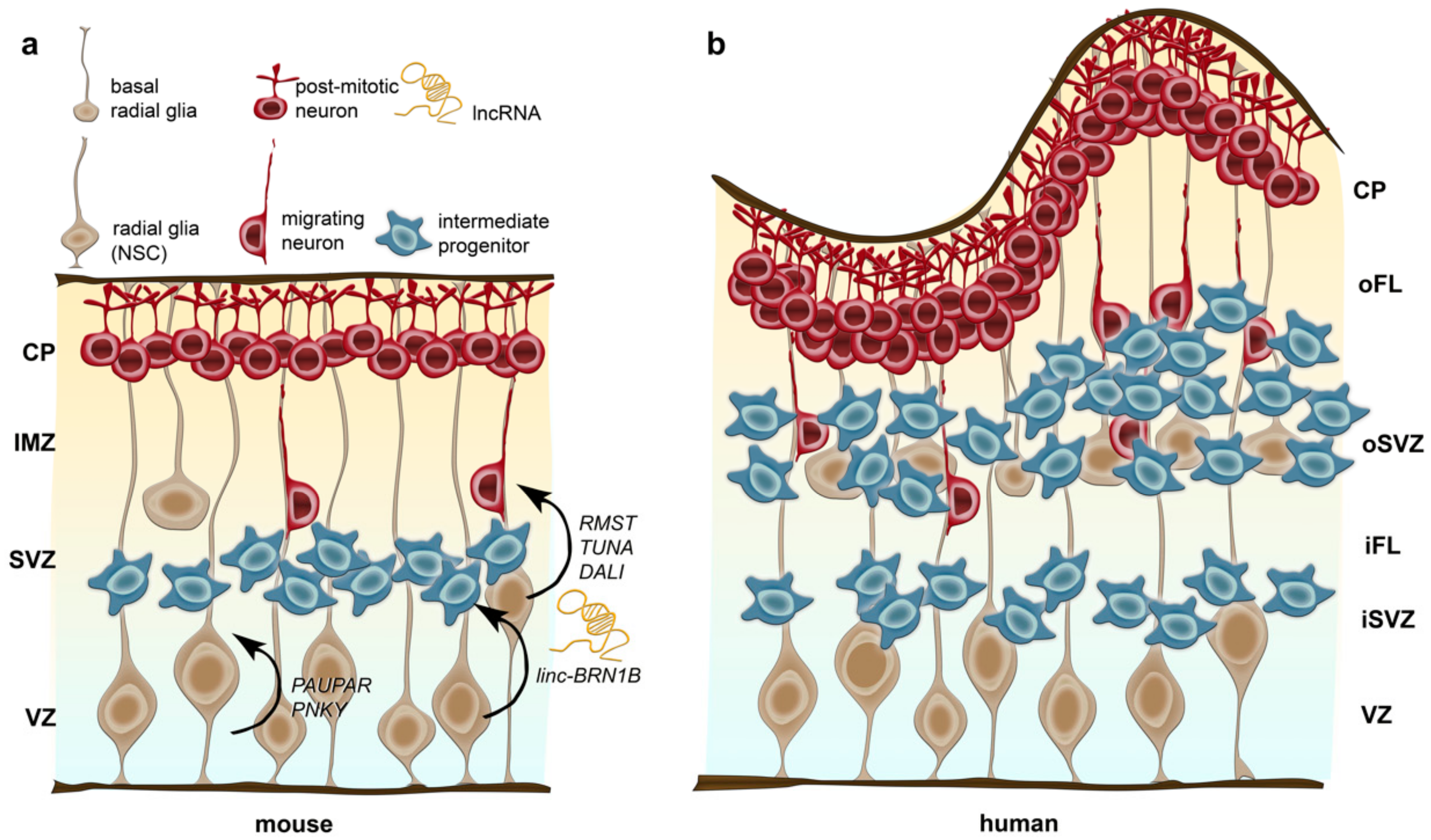
2.6. Hallmarks of Cortical Development in View of Potential Implications for Evolution
2.7. Implications of lncRNAs in Processes Potentially Relevant for Human Brain Evolution
2.7.1. Cis- and Trans-lncRNA Regulatory Control over Neuronal Differentiation
2.7.2. lncRNA-Mediated Regulation of Neurite Outgrowth and Synaptogenesis
2.7.3. lncRNA-Mediated Regulation of Synaptic Plasticity
3. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- The FANTOM Consortium; Carninci, P.; Kasukawa, T.; Katayama, S.; Gough, J.; Frith, M.; Maeda, N.; Oyama, R.; Ravasi, T.; Lenhard, B.; et al. The Transcriptional Landscape of the Mammalian Genome. Science 2005, 309, 1559–1563. [Google Scholar] [PubMed]
- The ENCODE Project Consortium. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 2007, 447, 799–816. [Google Scholar] [CrossRef] [PubMed]
- RIKEN Genome Exploration Research Group and Genome Science Group (Genome Network Project Core Group) and the FANTOM Consortium; Katayama, S.; Tomaru, Y.; Kasukawa, T.; Waki, K.; Nakanishi, M.; Nakamura, M.; Nishida, H.; Yap, C.C.; Suzuki, M.; et al. Antisense Transcription in the Mammalian Transcriptome. Science 2005, 309, 1564–1566. [Google Scholar] [PubMed]
- Costa, M.C.; Leitao, A.L.; Enguita, F.J. Biogenesis and Mechanism of Action of Small Non-Coding RNAs: Insights from the Point of View of Structural Biology. Int. J. Mol. Sci. 2012, 13, 10268–10295. [Google Scholar] [CrossRef] [PubMed]
- Qureshi, I.A.; Mehler, M.F. Emerging roles of non-coding RNAs in brain evolution, development, plasticity and disease. Nat. Rev. Neurosci. 2012, 13, 528–541. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.C.; Chang, H.Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 2011, 43, 904–914. [Google Scholar] [CrossRef]
- Derrien, T.; Johnson, R.; Bussotti, G.; Tanzer, A.; Djebali, S.; Tilgner, H.; Guernec, G.; Martín, D.; Merkel, A.; Knowles, D.G.; et al. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genome Res. 2012, 22, 1775–1789. [Google Scholar] [CrossRef]
- Harrow, J.; Frankish, A.; González, J.M.; Tapanari, E.; Diekhans, M.; Kokocinski, F.; Aken, B.L.; Barrell, D.; Zadissa, A.; Searle, S.; et al. GENCODE: The reference human genome annotation for The ENCODE Project. Genome Res. 2012, 22, 1760–1774. [Google Scholar] [CrossRef]
- Aprea, J.; Prenninger, S.; Dori, M.; Ghosh, T.; Monasor, L.S.; Wessendorf, E.; Zocher, S.; Massalini, S.; Alexopoulou, D.; Lesche, M.; et al. Transcriptome sequencing during mouse brain development identifies long non-coding RNAs functionally involved in neurogenic commitment. EMBO J. 2013, 32, 3145–3160. [Google Scholar] [CrossRef]
- Belgard, T.G.; Marques, A.C.; Oliver, P.L.; Abaan, H.O.; Sirey, T.M.; Hoerder-Suabedissen, A.; García-Moreno, F.; Molnár, Z.; Margulies, E.H.; Ponting, C.P. A Transcriptomic Atlas of Mouse Neocortical Layers. Neuron 2011, 71, 605–616. [Google Scholar] [CrossRef]
- Mercer, T.R.; Dinger, M.E.; Sunkin, S.M.; Mehler, M.F.; Mattick, J.S. Specific expression of long noncoding RNAs in the mouse brain. Proc. Natl. Acad. Sci. USA 2008, 105, 716–721. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.Q.; Habegger, L.; Noisa, P.; Szekely, A.; Qiu, C.; Hutchison, S.; Raha, D.; Egholm, M.; Lin, H.; Weissman, S.; et al. Dynamic transcriptomes during neural differentiation of human embryonic stem cells revealed by short, long, and paired-end sequencing. Proc. Natl. Acad. Sci. USA 2010, 107, 5254–5259. [Google Scholar] [CrossRef] [PubMed]
- Barry, G.; Briggs, J.A.; Vanichkina, D.P.; Poth, E.M.; Beveridge, N.J.; Ratnu, V.S.; Nayler, S.P.; Nones, K.; Hu, J.; Bredy, T.W.; et al. The long non-coding RNA Gomafu is acutely regulated in response to neuronal activation and involved in schizophrenia-associated alternative splicing. Mol. Psychiatry 2014, 19, 486. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.-K.; Hemberg, M.; Gray, J.M.; Costa, A.M.; Bear, D.M.; Wu, J.; Harmin, D.A.; Laptewicz, M.; Barbara-Haley, K.; Kuersten, S.; et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 2010, 465, 182–187. [Google Scholar] [CrossRef]
- Lipovich, L.; Dachet, F.; Cai, J.; Bagla, S.; Balan, K.; Jia, H.; Loeb, J.A. Activity-Dependent Human Brain Coding/Noncoding Gene Regulatory Networks. Genetics 2012, 192, 1133–1148. [Google Scholar] [CrossRef]
- Pereira Fernandes, D.; Bitar, M.; Jacobs, F.M.; Barry, G. Long non-coding RNAs in neuronal aging. Non-Coding RNA 2018, 4, 12. [Google Scholar] [CrossRef]
- Briggs, J.A.; Wolvetang, E.J.; Mattick, J.S.; Rinn, J.L.; Barry, G. Mechanisms of Long Non-coding RNAs in Mammalian Nervous System Development, Plasticity, Disease, and Evolution. Neuron 2015, 88, 861–877. [Google Scholar] [CrossRef]
- Neubauer, S.; Gunz, P. Endocasts and the evo-devo approach to study human brain evolution. In Digital Endocasts; Bruner, E., Ogihara, N., Tanabe, H., Eds.; Springer: Tokyo, Japan, 2018. [Google Scholar]
- Fatica, A.; Bozzoni, I. Long non-coding RNAs: New players in cell differentiation and development. Nat. Rev. Genet. 2014, 15, 7–21. [Google Scholar] [CrossRef]
- Core, L.J.; Waterfall, J.J.; Lis, J.T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 2008, 322, 1845–1848. [Google Scholar] [CrossRef]
- Kapranov, P.; Cheng, J.; Dike, S.; Nix, D.A.; Duttagupta, R.; Willingham, A.T.; Stadler, P.F.; Hertel, J.; Hackermüller, J.; Hofacker, I.L.; et al. RNA Maps Reveal New RNA Classes and a Possible Function for Pervasive Transcription. Science 2007, 316, 1484–1488. [Google Scholar] [CrossRef]
- Khalil, A.M.; Guttman, M.; Huarte, M.; Garber, M.; Raj, A.; Morales, D.R.; Thomas, K.; Presser, A.; Bernstein, B.E.; van Oudenaarden, A.; et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl. Acad. Sci. USA 2009, 106, 11667–11672. [Google Scholar] [CrossRef] [PubMed]
- Ørom, U.A.; Derrien, T.; Beringer, M.; Gumireddy, K.; Gardini, A.; Bussotti, G.; Lai, F.; Zytnicki, M.; Notredame, C.; Huang, Q.; et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 2010, 143, 46–58. [Google Scholar] [CrossRef] [PubMed]
- Seila, A.C.; Calabrese, J.M.; Levine, S.S.; Yeo, G.W.; Rahl, P.B.; Flynn, R.A.; Young, R.A.; Sharp, P.A. Divergent transcription from active promoters. Science 2008, 322, 1849–1851. [Google Scholar] [CrossRef]
- Mercer, T.R.; Wilhelm, D.; Dinger, M.E.; Soldà, G.; Korbie, D.J.; Glazov, E.A.; Truong, V.; Schwenke, M.; Simons, C.; Matthaei, K.I.; et al. Expression of distinct RNAs from 3′ untranslated regions. Nucleic Acids Res. 2010, 39, 2393–2403. [Google Scholar] [CrossRef] [PubMed]
- Hung, T.; Wang, Y.; Lin, M.F.; Koegel, A.K.; Kotake, Y.; Grant, G.D.; Horlings, H.M.; Shah, N.; Umbricht, C.; Wang, P.; et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat. Genet. 2011, 43, 621–629. [Google Scholar] [CrossRef] [PubMed]
- Azzalin, C.M.; Reichenbach, P.; Khoriauli, L.; Giulotto, E.; Lingner, J. Telomeric Repeat Containing RNA and RNA Surveillance Factors at Mammalian Chromosome Ends. Science 2007, 318, 798–801. [Google Scholar] [CrossRef] [PubMed]
- Wapinski, O.; Chang, H.Y. Long noncoding RNAs and human disease. Trends Cell Biol. 2011, 21, 354–361. [Google Scholar] [CrossRef]
- Yoon, J.H.; Abdelmohsen, K.; Gorospe, M. Posttranscriptional gene regulation by long noncoding RNA. J. Mol. Biol. 2013, 425, 3723–3730. [Google Scholar] [CrossRef]
- Lee, J.T. Epigenetic Regulation by Long Noncoding RNAs. Science 2012, 338, 1435–1439. [Google Scholar] [CrossRef]
- Batista, P.J.; Chang, H.Y. Long noncoding RNAs: Cellular address codes in development and disease. Cell 2013, 152, 1298–1307. [Google Scholar] [CrossRef]
- Marchese, F.P.; Raimondi, I.; Huarte, M. The multidimensional mechanisms of long noncoding RNA function. Genome Biol. 2017, 18, 206. [Google Scholar] [CrossRef] [PubMed]
- Cabianca, D.S.; Casà, V.; Bodega, B.; Xynos, A.; Ginelli, E.; Tanaka, Y.; Gabellini, D. A long ncRNA links copy number variation to a polycomb/trithorax epigenetic switch in FSHD muscular dystrophy. Cell 2012, 149, 819–831. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.C.; Yang, Y.W.; Liu, B.; Sanyal, A.; Corces-Zimmerman, R.; Chen, Y.; Lajoie, B.R.; Protacio, A.; Flynn, R.A.; Gupta, R.A.; et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 2011, 472, 120–124. [Google Scholar] [CrossRef] [PubMed]
- Margueron, R.; Reinberg, D. The Polycomb complex PRC2 and its mark in life. Nature 2011, 469, 343–349. [Google Scholar] [CrossRef]
- Rinn, J.L.; Kertesz, M.; Wang, J.K.; Squazzo, S.L.; Xu, X.; Brugmann, S.A.; Goodnough, L.H.; Helms, J.A.; Farnham, P.J.; Segal, E.; et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007, 129, 1311–1323. [Google Scholar] [CrossRef]
- Tsai, M.-C.; Manor, O.; Wan, Y.; Mosammaparast, N.; Wang, J.K.; Lan, F.; Shi, Y.; Segal, E.; Chang, H.Y. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010, 329, 689–693. [Google Scholar] [CrossRef]
- Davidovich, C.; Goodrich, K.J.; Gooding, A.R.; Cech, T.R. A dimeric state for PRC2. Nucleic Acids Res. 2014, 42, 9236–9248. [Google Scholar] [CrossRef]
- Pandey, R.R.; Mondal, T.; Mohammad, F.; Enroth, S.; Redrup, L.; Komorowski, J.; Nagano, T.; Mancini-DiNardo, D.; Kanduri, C. Kcnq1ot1 Antisense Noncoding RNA Mediates Lineage-Specific Transcriptional Silencing through Chromatin-Level Regulation. Mol. Cell 2008, 32, 232–246. [Google Scholar] [CrossRef]
- Fitzpatrick, G.V.; Soloway, P.D.; Higgins, M.J. Regional loss of imprinting and growth deficiency in mice with a targeted deletion of KvDMR1. Nat. Genet. 2002, 32, 426–431. [Google Scholar] [CrossRef]
- Mancini-DiNardo, D.; Steele, S.J.; Levorse, J.M.; Ingram, R.S.; Tilghman, S.M. Elongation of the Kcnq1ot1 transcript is required for genomic imprinting of neighboring genes. Genome Res. 2006, 20, 1268–1282. [Google Scholar] [CrossRef]
- Guil, S.; Soler, M.; Portela, A.; Carrère, J.; Fonalleras, E.; Gómez, A.; Villanueva, A.; Esteller, M.; Moruno, A.G. Intronic RNAs mediate EZH2 regulation of epigenetic targets. Nat. Struct. Mol. Biol. 2012, 19, 664–670. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Ohsumi, T.K.; Kung, J.T.; Ogawa, Y.; Grau, D.J.; Sarma, K.; Song, J.J.; Kingston, R.E.; Borowsky, M.; Lee, J.T. Genome-wide identification of polycomb-associated RNAs by RIP-seq. Mol. Cell 2010, 40, 939–953. [Google Scholar] [CrossRef] [PubMed]
- Kotake, Y.; Nakagawa, T.; Kitagawa, K.; Suzuki, S.; Liu, N.; Kitagawa, M.; Xiong, Y. Long non-coding RNA ANRIL is required for the PRC2 recruitment to and silencing of p15p15 INK4B tumor suppressor gene. Oncogene 2011, 30, 1956. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, S.; Bonasio, R.; Saldaña-Meyer, R.; Yoshida, T.; Son, J.; Nishino, K.; Umezawa, A.; Reinberg, D. Interactions between JARID2 and noncoding RNAs regulate PRC2 recruitment to chromatin. Mol. Cell 2014, 53, 290–300. [Google Scholar] [CrossRef] [PubMed]
- Guttman, M.; Donaghey, J.; Carey, B.W.; Garber, M.; Grenier, J.K.; Munson, G.; Young, G.; Lucas, A.B.; Ach, R.; Bruhn, L.; et al. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature 2011, 477, 295–300. [Google Scholar] [CrossRef] [PubMed]
- Jang, H.S.; Shin, W.J.; Lee, J.E.; Do, J.T. CpG and Non-CpG Methylation in Epigenetic Gene Regulation and Brain Function. Genes 2017, 8, 148. [Google Scholar] [CrossRef]
- Chalei, V.; Sansom, S.N.; Kong, L.; Lee, S.; Montiel, J.F.; Vance, K.W.; Ponting, C.P. The long non-coding RNA Dali is an epigenetic regulator of neural differentiation. Elife 2014, 3, 04530. [Google Scholar] [CrossRef]
- Ariel, F.; Jégu, T.; Latrasse, D.; Romero-Barrios, N.; Christ, A.; Benhamed, M.; Crespi, M. Noncoding Transcription by Alternative RNA Polymerases Dynamically Regulates an Auxin-Driven Chromatin Loop. Mol. Cell 2014, 55, 383–396. [Google Scholar] [CrossRef]
- Bardou, F.; Ariel, F.; Simpson, C.G.; Romero-Barrios, N.; Laporte, P.; Balzergue, S.; Brown, J.W.; Crespi, M. Long Noncoding RNA Modulates Alternative Splicing Regulators in Arabidopsis. Dev. Cell 2014, 30, 166–176. [Google Scholar] [CrossRef]
- Romero-Barrios, N.; Legascue, M.F.; Benhamed, M.; Ariel, F.; Crespi, M. Splicing regulation by long noncoding RNAs. Nucleic Acids Res. 2018, 46, 2169–2184. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, X.-O.; Chen, T.; Xiang, J.-F.; Yin, Q.-F.; Xing, Y.-H.; Zhu, S.; Yang, L.; Chen, L.-L. Circular Intronic Long Noncoding RNAs. Mol. Cell 2013, 51, 792–806. [Google Scholar] [CrossRef] [PubMed]
- Cao, W.; Jamison, S.F.; Garcia-Blanco, M.A. Both phosphorylation and dephosphorylation of ASF/SF2 are required for pre-mRNA splicing in vitro. RNA 1997, 3, 1456–1467. [Google Scholar] [PubMed]
- Manley, J.L.; Xiao, S.H. Phosphorylation of the ASF/SF2 RS domain affects both protein-protein and protein-RNA interactions and is necessary for splicing. Genes Dev. 1997, 11, 334–344. [Google Scholar]
- Xiao, S.-H.; Manley, J.L. Phosphorylation-dephosphorylation differentially affects activities of splicing factor ASF/SF2. EMBO J. 1998, 17, 6359–6367. [Google Scholar] [CrossRef] [PubMed]
- Cáceres, J.F.; Misteli, T.; Screaton, G.R.; Spector, D.L.; Krainer, A.R. Role of the Modular Domains of SR Proteins in Subnuclear Localization and Alternative Splicing Specificity. J. Cell Bilo. 1997, 138, 225–238. [Google Scholar] [CrossRef] [PubMed]
- Misteli, T.; Cáceres, J.F.; Clement, J.Q.; Krainer, A.R.; Wilkinson, M.F.; Spector, D.L. Serine Phosphorylation of SR Proteins Is Required for Their Recruitment to Sites of Transcription In Vivo. J. Cell Biol. 1998, 143, 297–307. [Google Scholar] [CrossRef] [PubMed]
- Cooper, D.R.; Carter, G.; Li, P.; Patel, R.; Watson, J.E.; Patel, N.A. Long Non-Coding RNA NEAT1 Associates with SRp40 to Temporally Regulate PPARγ2 Splicing during Adipogenesis in 3T3-L1 Cells. Genes 2014, 5, 1050–1063. [Google Scholar] [CrossRef]
- Malakar, P.; Shilo, A.; Mogilevsky, A.; Stein, I.; Pikarsky, E.; Nevo, Y.; Benyamini, H.; Elgavish, S.; Zong, X.; Prasanth, K.V.; et al. Long noncoding RNA MALAT1 promotes hepatocellular carcinoma development by SRSF1 upregulation and mTOR activation. Cancer Res. 2017, 77, 1155–1167. [Google Scholar] [CrossRef]
- Wang, X.; Sehgal, L.; Jain, N.; Khashab, T.; Mathur, R.; Samaniego, F. LncRNA MALAT1 promotes development of mantle cell lymphoma by associating with EZH2. J. Transl. Med. 2016, 14, 346. [Google Scholar] [CrossRef]
- Tripathi, V.; Ellis, J.D.; Shen, Z.; Song, D.Y.; Pan, Q.; Watt, A.T.; Freier, S.M.; Bennett, C.F.; Sharma, A.; Bubulya, P.A.; et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol. Cell 2010, 39, 925–938. [Google Scholar] [CrossRef]
- Huang, Y.; Yario, T.A.; Steitz, J.A. A molecular link between SR protein dephosphorylation and mRNA export. Proc. Natl. Acad. Sci. USA 2004, 101, 9666–9670. [Google Scholar] [CrossRef] [PubMed]
- Sanford, J.R.; Ellis, J.D.; Cazalla, D.; Cáceres, J.F. Reversible phosphorylation differentially affects nuclear and cytoplasmic functions of splicing factor 2/alternative splicing factor. Proc. Natl. Acad. Sci. USA 2005, 102, 15042–15047. [Google Scholar] [CrossRef] [PubMed]
- Ji, Q.; Zhang, L.; Liu, X.; Zhou, L.; Wang, W.; Han, Z.; Sui, H.; Tang, Y.; Wang, Y.; Liu, N.; et al. Long non-coding RNA MALAT1 promotes tumour growth and metastasis in colorectal cancer through binding to SFPQ and releasing oncogene PTBP2 from SFPQ/PTBP2 complex. Br. J. Cancer 2014, 111, 736–748. [Google Scholar] [CrossRef] [PubMed]
- Rapicavoli, N.A.; Blackshaw, S. New meaning in the message: Noncoding RNAs and their role in retinal development. Dev. Dyn. Off. Publ. Am. Assoc. Anat. 2009, 238, 2103–2114. [Google Scholar] [CrossRef] [PubMed]
- Rapicavoli, N.A.; Poth, E.M.; Blackshaw, S. The long noncoding RNA RNCR2 directs mouse retinal cell specification. BMC Dev. Biol. 2010, 10, 49. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.R.; Qureshi, I.A.; Gokhan, S.; Dinger, M.E.; Li, G.; Mattick, J.S.; Mehler, M.F. Long noncoding RNAs in neuronal-glial fate specification and oligodendrocyte lineage maturation. BMC Neurosci. 2010, 11, 14. [Google Scholar] [CrossRef]
- Sone, M.; Hayashi, T.; Tarui, H.; Agata, K.; Takeichi, M.; Nakagawa, S. The mRNA-like noncoding RNA Gomafu constitutes a novel nuclear domain in a subset of neurons. J. Cell Sci. 2007, 120, 2498–2506. [Google Scholar] [CrossRef]
- Tsuiji, H.; Yoshimoto, R.; Hasegawa, Y.; Furuno, M.; Yoshida, M.; Nakagawa, S. Competition between a noncoding exon and introns: Gomafu contains tandem UACUAAC repeats and associates with splicing factor-1. Genes Cells 2011, 16, 479–490. [Google Scholar] [CrossRef]
- Rashid, F.; Shah, A.; Shan, G. Long Non-coding RNAs in the Cytoplasm. Genom. Proteom. Bioinform. 2016, 14, 73–80. [Google Scholar] [CrossRef]
- Gong, C.; Maquat, L.E. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3’ UTRs via Alu elements. Nature 2011, 470, 284–288. [Google Scholar] [CrossRef]
- Liu, X.; Li, D.; Zhang, W.; Guo, M.; Zhan, Q. Long non-coding RNA gadd7 interacts with TDP-43 and regulates Cdk6 mRNA decay. EMBO J. 2012, 31, 4415–4427. [Google Scholar] [CrossRef] [PubMed]
- Faghihi, M.A.; Modarresi, F.; Khalil, A.M.; Wood, D.E.; Sahagan, B.G.; Morgan, T.E.; Finch, C.E.; Laurent III, G.S.; Kenny, P.J.; Wahlestedt, C. Expression of a noncoding RNA is elevated in Alzheimer’s disease and drives rapid feed-forward regulation of β-secretase. Nat. Med. 2008, 14, 723. [Google Scholar] [CrossRef] [PubMed]
- Kretz, M. TINCR, staufen1, and cellular differentiation. RNA Biol. 2013, 10, 1597–1601. [Google Scholar] [CrossRef] [PubMed]
- Job, C.; Eberwine, J. Localization and translation of mRNA in dentrites and axons. Nat. Rev. Neurosci. 2001, 2, 889–898. [Google Scholar] [CrossRef] [PubMed]
- Carrieri, C.; Cimatti, L.; Biagioli, M.; Beugnet, A.; Zucchelli, S.; Fedele, S.; Pesce, E.; Ferrer, I.; Collavin, L.; Santoro, C.; et al. Long non-coding antisense RNA controls Uchl1 translation through an embedded SINEB2 repeat. Nature 2012, 491, 454–457. [Google Scholar] [CrossRef] [PubMed]
- Yoon, J.-H.; Abdelmohsen, K.; Srikantan, S.; Yang, X.; Martindale, J.L.; De, S.; Huarte, M.; Zhan, M.; Becker, K.G.; Gorospe, M. LincRNA-p21 suppresses target mRNA translation. Mol. Cell 2012, 47, 648–655. [Google Scholar] [CrossRef]
- Salmena, L.; Poliseno, L.; Tay, Y.; Kats, L.; Pandolfi, P.P. A ceRNA hypothesis: The Rosetta Stone of a hidden RNA language? Cell 2011, 146, 353–358. [Google Scholar] [CrossRef]
- Chen, M.-T.; Lin, H.-S.; Shen, C.; Ma, Y.-N.; Wang, F.; Zhao, H.-L.; Yu, J.; Zhang, J.-W. PU.1-Regulated Long Noncoding RNA lnc-MC Controls Human Monocyte/Macrophage Differentiation through Interaction with MicroRNA 199a-5p. Mol. Cell. Biol. 2015, 35, 3212–3224. [Google Scholar] [CrossRef]
- Franco-Zorrilla, J.M.; Valli, A.; Todesco, M.; Mateos, I.; Puga, M.I.; Rubio-Somoza, I.; Leyva, A.; Weigel, D.; García, J.A.; Paz-Ares, J. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 2007, 39, 1033–1037. [Google Scholar] [CrossRef]
- Karreth, F.A.; Tay, Y.; Perna, D.; Ala, U.; Tan, S.M.; Rust, A.G.; DeNicola, G.; Webster, K.A.; Weiss, D.; Perez-Mancera, P.A.; et al. In Vivo Identification of Tumor- Suppressive PTEN ceRNAs in an Oncogenic BRAF-Induced Mouse Model of Melanoma. Cell 2011, 147, 382–395. [Google Scholar] [CrossRef]
- Peng, W.; Si, S.; Zhang, Q.; Li, C.; Zhao, F.; Wang, F.; Yu, J.; Ma, R. Long non-coding RNA MEG3 functions as a competing endogenous RNA to regulate gastric cancer progression. J. Exp. Clin. Cancer Res. 2015, 34, 79. [Google Scholar] [CrossRef] [PubMed]
- Tay, Y.; Kats, L.; Salmena, L.; Weiss, D.; Tan, S.M.; Ala, U.; Karreth, F.; Poliseno, L.; Provero, P.; Di Cunto, F.; et al. Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell 2011, 147, 344–357. [Google Scholar] [CrossRef] [PubMed]
- He, S.; Su, H.; Liu, C.; Skogerbø, G.; He, H.; He, D.; Zhu, X.; Liu, T.; Zhao, Y.; Chen, R. MicroRNA-encoding long non-coding RNAs. BMC Genom. 2008, 9, 236. [Google Scholar] [CrossRef] [PubMed]
- Keniry, A.; Oxley, D.; Monnier, P.; Kyba, M.; Dandolo, L.; Smits, G.; Reik, W. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and Igf1r. Nature 2012, 14, 659–695. [Google Scholar] [CrossRef]
- Dey, B.K.; Pfeifer, K.; Dutta, A. The H19 long noncoding RNA gives rise to microRNAs miR-675-3p and miR-675-5p to promote skeletal muscle differentiation and regeneration. Genes Dev. 2014, 28, 491–501. [Google Scholar] [CrossRef]
- Cai, X.; Cullen, B.R. The imprinted H19 noncoding RNA is a primary microRNA precursor. RNA 2007, 13, 313–316. [Google Scholar] [CrossRef]
- Nitsche, A.; Rose, D.; Fasold, M.; Reiche, K.; Stadler, P.F. Comparison of splice sites reveals that long noncoding RNAs are evolutionarily well conserved. RNA 2015, 21, 801–812. [Google Scholar] [CrossRef]
- Pang, K.C.; Frith, M.C.; Mattick, J.S. Rapid evolution of noncoding RNAs: Lack of conservation does not mean lack of function. Trends Genet. 2006, 22, 1–5. [Google Scholar] [CrossRef]
- Ulitsky, I.; Shkumatava, A.; Jan, C.H.; Sive, H.; Bartel, D.P. Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell 2011, 147, 1537–1550. [Google Scholar] [CrossRef]
- Tay, S.K.; Blythe, J.; Lipovich, L. Global discovery of primate-specific genes in the human genome. Proc. Natl. Acad. Sci. USA 2009, 106, 12019–12024. [Google Scholar] [CrossRef]
- Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature 2002, 420, 520. [Google Scholar] [CrossRef] [PubMed]
- Lindblad-Toh, K.; Garber, M.; Zuk, O.; Lin, M.F.; Parker, B.J.; Washietl, S.; Kheradpour, P.; Ernst, J.; Jordan, G.; Mauceli, E.; et al. A high-resolution map of human evolutionary constraint using 29 mammals. Nature 2011, 478, 476–482. [Google Scholar] [CrossRef] [PubMed]
- The Chimpanzee Sequencing and Analysis Consortium; Waterson, R.H.; Lander, E.S.; Wilson, R.K. Initial sequence of the chimpanzee genome and comparison with the human genome. Nature 2005, 437, 69. [Google Scholar]
- Grossman, S.R.; Andersen, K.G.; Shlyakhter, I.; Tabrizi, S.; Winnicki, S.; Yen, A.; Park, D.J.; Griesemer, D.; Karlsson, E.K.; Wong, S.H.; et al. Identifying recent adaptations in large-scale genomic data. Cell 2013, 152, 703–713. [Google Scholar] [CrossRef] [PubMed]
- Pollard, K.S.; Salama, S.R.; Lambert, N.; Lambot, M.-A.; Coppens, S.; Pedersen, J.S.; Katzman, S.; King, B.; Onodera, C.; Siepel, A.; et al. An RNA gene expressed during cortical development evolved rapidly in humans. Nature 2006, 443, 167–172. [Google Scholar] [CrossRef] [PubMed]
- Shi, P.; Bakewell, M.A.; Zhang, J. Did brain-specific genes evolve faster in humans than in chimpanzees? Trends Genet. 2006, 22, 608–613. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.Y.; Chien, H.C.; Osada, N.; Hashimoto, K.; Sugano, S.; Gojobori, T.; Chou, C.K.; Tsai, S.F.; Wu, C.I.; Shen, C.K.J. Rate of evolution in brain-expressed genes in humans and other primates. PLoS Biol. 2006, 5, e13. [Google Scholar] [CrossRef]
- Call, J.; Tomasello, M. Does the chimpanzee have a theory of mind? 30 years later. Trends Cogn. Sci. 2008, 12, 187–192. [Google Scholar] [CrossRef]
- Herrmann, E.; Call, J.; Hernàndez-Lloreda, M.V.; Hare, B.; Tomasello, M. Humans Have Evolved Specialized Skills of Social Cognition: The Cultural Intelligence Hypothesis. Science 2007, 317, 1360–1366. [Google Scholar] [CrossRef]
- Tomasello, M.; Vaish, A. Origins of human cooperation and morality. Annu. Rev. Psychol. 2013, 64, 231–255. [Google Scholar] [CrossRef]
- Tennie, C.; Call, J.; Tomasello, M. Ratcheting up the ratchet: On the evolution of cumulative culture. Trans. R. Soc. B Biol. Sci. 2009, 364, 2405–2415. [Google Scholar] [CrossRef] [PubMed]
- Fitch, W.T. The Evolution of Language; Cambridge University Press: Cambridge, UK, 2010. [Google Scholar]
- Hauser, M.D.; Chomsky, N.; Fitch, W.T. The faculty of language: What is it, who has it, and how did it evolve? Science 2002, 298, 1569–1579. [Google Scholar] [CrossRef] [PubMed]
- Herculano-Houzel, S.; Catania, K.; Manger, P.R.; Kaas, J.H. Mammalian brains are made of these: A dataset of the numbers and densities of neuronal and nonneuronal cells in the brain of glires, primates, scandentia, eulipotyphlans, afrotherians and artiodactyls, and their relationship with body mass. Brain Behav. Evol. 2015, 86, 145–163. [Google Scholar] [CrossRef] [PubMed]
- Kaas, J.H. The evolution of brains from early mammals to humans. Wiley Interdiscip. Rev. Cogn. Sci. 2013, 4, 33–45. [Google Scholar] [CrossRef] [PubMed]
- Molnár, Z.; Clowry, G.J.; Šestan, N.; Alzu’bi, A.; Bakken, T.; Hevner, R.F.; Hüppi, P.S.; Kostović, I.; Rakic, P.; Anton, E. New insights into the development of the human cerebral cortex. J. Anat. 2019, 235, 432–451. [Google Scholar] [CrossRef]
- Kaas, J.H. Why does the brain have so many visual areas? J. Cogn. Neurosci. 1989, 1, 121–135. [Google Scholar] [CrossRef]
- Molnár, Z.; Pollen, A. How unique is the human neocortex? Development 2014, 141, 11–16. [Google Scholar] [CrossRef]
- Uylings, H.B.M.; van Eden, C.G. Qualitative and quantitative comparison of the prefrontal cortex in rat and in primates, including humans. Prog. Brain Res. 1991, 85, 31–62. [Google Scholar]
- Hladnik, A.; Džaja, D.; Darmopil, S.; Jovanov-Milošević, N.; Petanjek, Z. Spatio-temporal extension in site of origin for cortical calretinin neurons in primates. Front. Neuroanat. 2014, 8, 50. [Google Scholar] [CrossRef]
- Teffer, K.; Semendeferi, K. Human prefrontal cortex: Evolution, development, and pathology. Prog. Brain Res. 2012, 195, 191–218. [Google Scholar]
- Friederici, A.D. The brain basis of language processing: From structure to function. Physiol. Rev. 2011, 91, 1357–1392. [Google Scholar] [CrossRef]
- O’Rahilly, R.; Müller, F. Significant features in the early prenatal development of the human brain. Ann. Anat. Anat. Anz. 2008, 190, 105–118. [Google Scholar]
- Roth, G.; Dicke, U. Evolution of the brain and intelligence in primates. Prog. Brain Res. 2012, 195, 413–430. [Google Scholar] [PubMed]
- Orban, G.A.; Claeys, K.; Nelissen, K.; Smans, R.; Sunaert, S.; Todd, J.T.; Wardak, C.; Durand, J.-B.; Vanduffel, W. Mapping the parietal cortex of human and non-human primates. Neuropsychologia 2006, 44, 2647–2667. [Google Scholar] [CrossRef] [PubMed]
- Stout, D.; Chaminade, T. The evolutionary neuroscience of tool making. Neuropsychologia 2007, 45, 1091–1100. [Google Scholar] [CrossRef] [PubMed]
- Verendeev, A.; Sherwood, C.C.; Hopkins, W.D. Organization and Evolution of the Neural Control of the Hand in Primates: Motor Systems, Sensory Feedback, and Laterality. In The Evolution of The Primate Hand; Kivell, T., Lemelin, P., Richmond, B., Schmitt, D., Eds.; Springer: New York, NY, USA, 2016. [Google Scholar]
- Burkhalter, A.H. Many specialists for suppressing cortical excitation. Front. Neurosci. 2008, 2, 26. [Google Scholar] [CrossRef]
- Forbes, C.E.; Grafman, J. The role of the human prefrontal cortex in social cognition and moral judgment. Annu. Rev. Neurosci. 2010, 33, 299–324. [Google Scholar] [CrossRef]
- Buzsáki, G.; Wang, X.-J. Mechanisms of gamma oscillations. Annu. Rev. Neurosci. 2012, 35, 203–225. [Google Scholar] [CrossRef]
- Whittington, M.A.; Cunningham, M.O.; LeBeau, F.E.; Racca, C.; Traub, R.D. Multiple origins of the cortical gamma rhythm. Dev. Neurobiol. 2011, 71, 92–106. [Google Scholar] [CrossRef]
- Franco, S.J.; Gil-Sanz, C.; Martinez-Garay, I.; Espinosa, A.; Harkins-Perry, S.R.; Ramos, C.; Muller, U. Fate-restricted neural progenitors in the mammalian cerebral cortex. Science 2012, 337, 746–749. [Google Scholar] [CrossRef]
- Rakic, P. Defects of neuronal migration and the pathogenesis of cortical malformations. Prog. Brain Res. 1988, 73, 15–37. [Google Scholar] [PubMed]
- Gorski, J.A.; Talley, T.; Qiu, M.; Puelles, L.; Rubenstein, J.L.; Jones, K.R. Cortical excitatory neurons and glia, but not GABAergic neurons, are produced in the Emx1-expressing lineage. J. Neurosci. 2002, 22, 6309–6314. [Google Scholar] [CrossRef] [PubMed]
- Rakic, P. The radial edifice of cortical architecture: From neuronal silhouettes to genetic engineering. Brain Res. Rev. 2007, 55, 204–219. [Google Scholar] [CrossRef] [PubMed]
- Rakic, P. Evolution of the neocortex: A perspective from developmental biology. Nat. Rev. Neurosci. 2009, 10, 724. [Google Scholar] [CrossRef] [PubMed]
- McConnell, S.K. Strategies for the generation of neuronal diversity in the developing central nervous system. J. Neurosci. Off. J. Soc. Neurosci. 1995, 15, 6987–6998. [Google Scholar] [CrossRef]
- Rakic, P. A small step for the cell, a giant leap for mankind: A hypothesis of neocortical expansion during evolution. Trends Neurosci. 1995, 18, 383–388. [Google Scholar] [CrossRef]
- Haubensak, W.; Attardo, A.; Denk, W.; Huttner, W.B. Neurons arise in the basal neuroepithelium of the early mammalian telencephalon: A major site of neurogenesis. Proc. Natl. Acad. Sci. USA 2004, 101, 3196–3201. [Google Scholar] [CrossRef]
- Noctor, S.C.; Martínez-Cerdeño, V.; Ivic, L.; Kriegstein, A.R. Cortical neurons arise in symmetric and asymmetric division zones and migrate through specific phases. Nat. Neurosci. 2004, 7, 136–144. [Google Scholar] [CrossRef]
- Taverna, E.; Götz, M.; Huttner, W.B. The cell biology of neurogenesis. Nat. Rev. Mol. Cell Biol. 2005, 6, 777–788. [Google Scholar]
- Cheung, A.F.; Pollen, A.A.; Tavare, A.; DeProto, J.; Molnár, Z. Comparative aspects of cortical neurogenesis in vertebrates. J. Anat. 2007, 211, 164–176. [Google Scholar] [CrossRef]
- Molnár, Z.; Métin, C.; Stoykova, A.; Tarabykin, V.; Price, D.J.; Francis, F.; Meyer, G.; Dehay, C.; Kennedy, H. Comparative aspects of cerebral cortical development. Eur. J. Neurosci. 2006, 23, 921–934. [Google Scholar] [CrossRef] [PubMed]
- Nonaka-Kinoshita, M.; Reillo, I.; Artegiani, B.; Martínez-Martínez, M.Á.; Nelson, M.; Borrell, V.; Calegari, F. Regulation of cerebral cortex size and folding by expansion of basal progenitors. EMBO J. 2013, 32, 1817–1828. [Google Scholar] [CrossRef] [PubMed]
- Stahl, R.; Walcher, T.; Romero, C.D.J.; Pilz, G.A.; Cappello, S.; Irmler, M.; Sanz-Aquela, J.M.; Beckers, J.; Blum, R.; Borrell, V. Trnp1 regulates expansion and folding of the mammalian cerebral cortex by control of radial glial fate. Cell 2013, 153, 535–549. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Hou, S.; Han, Y.-G. Hedgehog signaling promotes basal progenitor expansion and the growth and folding of the neocortex. Nat. Neurosci. 2016, 19, 888. [Google Scholar] [CrossRef] [PubMed]
- Wong, F.K.; Fei, J.-F.; Mora-Bermúdez, F.; Taverna, E.; Haffner, C.; Fu, J.; Anastassiadis, K.; Stewart, A.F.; Huttner, W.B. Sustained Pax6 expression generates primate-like basal radial glia in developing mouse neocortex. PLoS Biol. 2015, 13, e1002217. [Google Scholar] [CrossRef] [PubMed]
- Lewitus, E.; Kelava, I.; Kalinka, A.T.; Tomancak, P.; Huttner, W.B. An adaptive threshold in mammalian neocortical evolution. PLoS Biol. 2014, 12, e1002000. [Google Scholar] [CrossRef] [PubMed]
- Lui, J.H.; Hansen, D.V.; Kriegstein, A.R. Development and evolution of the human neocortex. Cell 2011, 146, 18–36. [Google Scholar] [CrossRef]
- Dehay, C.; Kennedy, H. Cell-cycle control and cortical development. Nat. Rev. Neurosci. 2007, 8, 438. [Google Scholar] [CrossRef]
- Smart, I.H.; Dehay, C.; Giroud, P.; Berland, M.; Kennedy, H. Unique morphological features of the proliferative zones and postmitotic compartments of the neural epithelium giving rise to striate and extrastriate cortex in the monkey. Cereb. Cortex 2002, 12, 37–53. [Google Scholar] [CrossRef]
- Dehay, C.; Kennedy, H.; Kosik, K.S. The outer subventricular zone and primate-specific cortical complexification. Neuron 2015, 85, 683–694. [Google Scholar] [CrossRef]
- Fietz, S.A.; Kelava, I.; Vogt, J.; Wilsch-Bräuninger, M.; Stenzel, D.; Fish, J.L.; Corbeil, D.; Riehn, A.; Distler, W.; Nitsch, R. OSVZ progenitors of human and ferret neocortex are epithelial-like and expand by integrin signaling. Nat. Neurosci. 2010, 13, 690. [Google Scholar] [CrossRef] [PubMed]
- Hansen, D.V.; Lui, J.H.; Parker, P.R.; Kriegstein, A.R. Neurogenic radial glia in the outer subventricular zone of human neocortex. Nature 2010, 464, 554. [Google Scholar] [CrossRef] [PubMed]
- Betizeau, M.; Cortay, V.; Patti, D.; Pfister, S.; Gautier, E.; Bellemin-Ménard, A.; Afanassieff, M.; Huissoud, C.; Douglas, R.J.; Kennedy, H. Precursor diversity and complexity of lineage relationships in the outer subventricular zone of the primate. Neuron 2013, 80, 442–457. [Google Scholar] [CrossRef] [PubMed]
- Lukaszewicz, A.; Savatier, P.; Cortay, V.; Giroud, P.; Huissoud, C.; Berland, M.; Kennedy, H.; Dehay, C. G1 phase regulation, area-specific cell cycle control, and cytoarchitectonics in the primate cortex. Neuron 2005, 47, 353–364. [Google Scholar] [CrossRef]
- Shitamukai, A.; Konno, D.; Matsuzaki, F. Oblique radial glial divisions in the developing mouse neocortex induce self-renewing progenitors outside the germinal zone that resemble primate outer subventricular zone progenitors. J. Neurosci. 2011, 31, 3683–3695. [Google Scholar] [CrossRef]
- Wang, X.; Tsai, J.-W.; LaMonica, B.; Kriegstein, A.R. A new subtype of progenitor cell in the mouse embryonic neocortex. Nat. Neurosci. 2011, 14, 555. [Google Scholar] [CrossRef]
- Reillo, I.; de Juan Romero, C.; García-Cabezas, M.Á.; Borrell, V. A role for intermediate radial glia in the tangential expansion of the mammalian cerebral cortex. Cereb. Cortex 2010, 21, 1674–1694. [Google Scholar] [CrossRef]
- Tuoc, T.C.; Boretius, S.; Sansom, S.N.; Pitulescu, M.-E.; Frahm, J.; Livesey, F.J.; Stoykova, A. Chromatin regulation by BAF170 controls cerebral cortical size and thickness. Dev. Cell 2013, 25, 256–269. [Google Scholar] [CrossRef]
- Kriegstein, A.; Noctor, S.; Martínez-Cerdeño, V. Patterns of neural stem and progenitor cell division may underlie evolutionary cortical expansion. Nat. Rev. Neurosci. 2006, 7, 883. [Google Scholar] [CrossRef]
- García-Moreno, F.; Vasistha, N.A.; Trevia, N.; Bourne, J.A.; Molnar, Z. Compartmentalization of cerebral cortical germinal zones in a lissencephalic primate and gyrencephalic rodent. Cereb. Cortex 2011, 22, 482–492. [Google Scholar] [CrossRef]
- LaMonica, B.E.; Lui, J.H.; Wang, X.; Kriegstein, A.R. OSVZ progenitors in the human cortex: An updated perspective on neurodevelopmental disease. Curr. Opin. Neurobiol. 2012, 22, 747–753. [Google Scholar] [CrossRef] [PubMed]
- Kalebic, N.; Gilardi, C.; Stepien, B.; Wilsch-Bräuninger, M.; Long, K.R.; Namba, T.; Florio, M.; Langen, B.; Lombardot, B.; Shevchenko, A. Neocortical expansion due to increased proliferation of basal progenitors is linked to changes in their morphology. Cell Stem Cell 2019, 24, 535–550. [Google Scholar] [CrossRef] [PubMed]
- Baala, L.; Briault, S.; Etchevers, H.C.; Laumonnier, F.; Natiq, A.; Amiel, J.; Boddaert, N.; Picard, C.; Sbiti, A.; Asermouh, A. Homozygous silencing of T-box transcription factor EOMES leads to microcephaly with polymicrogyria and corpus callosum agenesis. Nat. Genet. 2007, 39, 454. [Google Scholar] [CrossRef] [PubMed]
- Kowalczyk, T.; Pontious, A.; Englund, C.; Daza, R.A.; Bedogni, F.; Hodge, R.; Attardo, A.; Bell, C.; Huttner, W.B.; Hevner, R.F. Intermediate neuronal progenitors (basal progenitors) produce pyramidal–projection Neurons for all layers of cerebral cortex. Cereb. Cortex 2009, 19, 2439–2450. [Google Scholar] [CrossRef]
- Hatakeyama, J.; Wakamatsu, Y.; Nagafuchi, A.; Kageyama, R.; Shigemoto, R.; Shimamura, K. Cadherin-based adhesions in the apical endfoot are required for active Notch signaling to control neurogenesis in vertebrates. Development 2014, 141, 1671–1682. [Google Scholar] [CrossRef]
- Nelson, B.R.; Hodge, R.D.; Bedogni, F.; Hevner, R.F. Dynamic interactions between intermediate neurogenic progenitors and radial glia in embryonic mouse neocortex: Potential role in Dll1-Notch signaling. J. Neurosci. 2013, 33, 9122–9139. [Google Scholar] [CrossRef]
- Gerstmann, K.; Pensold, D.; Symmank, J.; Khundadze, M.; Hübner, C.A.; Bolz, J.; Zimmer, G. Thalamic afferents influence cortical progenitors via ephrin A5-EphA4 interactions. Development 2015, 142, 140–150. [Google Scholar] [CrossRef]
- Markov, N.T.; Ercsey-Ravasz, M.; Van Essen, D.C.; Knoblauch, K.; Toroczkai, Z.; Kennedy, H. Cortical high-density counterstream architectures. Science 2013, 342, 1238406. [Google Scholar] [CrossRef]
- Berezovskii, V.K.; Nassi, J.J.; Born, R.T. Segregation of feedforward and feedback projections in mouse visual cortex. J. Comp. Neurol. 2011, 519, 3672–3683. [Google Scholar] [CrossRef]
- Enard, W. The molecular basis of human brain evolution. Curr. Biol. 2016, 26, R1109–R1117. [Google Scholar] [CrossRef]
- Greig, L.C.; Woodworth, M.B.; Galazo, M.J.; Padmanabhan, H.; Macklis, J.D. Molecular logic of neocortical projection neuron specification, development and diversity. Nat. Rev. Neurosci. 2013, 14, 755. [Google Scholar] [CrossRef] [PubMed]
- Molyneaux, B.J.; Goff, L.A.; Brettler, A.C.; Chen, H.-H.; Brown, J.R.; Hrvatin, S.; Rinn, J.L.; Arlotta, P. DeCoN: Genome-wide analysis of in vivo transcriptional dynamics during pyramidal neuron fate selection in neocortex. Neuron 2015, 85, 275–288. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.; Wang, J.; Liu, Y.; Zhou, Y. Mechanisms of long non-coding RNAs in the assembly and plasticity of neural circuitry. Front. Neural Circuits 2017, 11, 76. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y. Cortical development and asymmetric cell divisions. Front. Biol. 2012, 7, 297–306. [Google Scholar] [CrossRef]
- Mohamed, J.S.; Gaughwin, P.M.; Lim, B.; Robson, P.; Lipovich, L. Conserved long noncoding RNAs transcriptionally regulated by Oct4 and Nanog modulate pluripotency in mouse embryonic stem cells. RNA 2010, 16, 324–337. [Google Scholar] [CrossRef] [PubMed]
- Ng, S.Y.; Johnson, R.; Stanton, L.W. Human long non-coding RNAs promote pluripotency and neuronal differentiation by association with chromatin modifiers and transcription factors. EMBO J. 2012, 31, 522–533. [Google Scholar] [CrossRef]
- Lin, N.; Chang, K.-Y.; Li, Z.; Gates, K.; Rana, Z.A.; Dang, J.; Zhang, D.; Han, T.; Yang, C.-S.; Cunningham, T.J. An evolutionarily conserved long noncoding RNA TUNA controls pluripotency and neural lineage commitment. Mol. Cell 2014, 53, 1005–1019. [Google Scholar] [CrossRef]
- Vance, K.W.; Sansom, S.N.; Lee, S.; Chalei, V.; Kong, L.; Cooper, S.E.; Oliver, P.L.; Ponting, C.P. The long non-coding RNA Paupar regulates the expression of both local and distal genes. EMBO J. 2014, 33, 296–311. [Google Scholar] [CrossRef]
- Götz, M.; Stoykova, A.; Gruss, P. Pax6 controls radial glia differentiation in the cerebral cortex. Neuron 1998, 21, 1031–1044. [Google Scholar] [CrossRef]
- Bond, A.M.; VanGompel, M.J.; Sametsky, E.A.; Clark, M.F.; Savage, J.C.; Disterhoft, J.F.; Kohtz, J.D. Balanced gene regulation by an embryonic brain ncRNA is critical for adult hippocampal GABA circuitry. Nat. Neurosci. 2009, 12, 1020. [Google Scholar] [CrossRef]
- Ramos, A.D.; Andersen, R.E.; Liu, S.J.; Nowakowski, T.J.; Hong, S.J.; Gertz, C.C.; Salinas, R.D.; Zarabi, H.; Kriegstein, A.R.; Lim, D.A. The long noncoding RNA Pnky regulates neuronal differentiation of embryonic and postnatal neural stem cells. Cell Stem Cell 2015, 16, 439–447. [Google Scholar] [CrossRef] [PubMed]
- Sauvageau, M.; Goff, L.A.; Lodato, S.; Bonev, B.; Groff, A.F.; Gerhardinger, C.; Sanchez-Gomez, D.B.; Hacisuleyman, E.; Li, E.; Spence, M. Multiple knockout mouse models reveal lincRNAs are required for life and brain development. Elife 2013, 2, e01749. [Google Scholar] [CrossRef] [PubMed]
- Mateos-Aparicio, P.; Rodríguez-Moreno, A. The impact of the study of brain plasticity. Front. Cell. Neurosci. 2019, 13, 66. [Google Scholar] [CrossRef] [PubMed]
- Modarresi, F.; Faghihi, M.A.; Lopez-Toledano, M.A.; Fatemi, R.P.; Magistri, M.; Brothers, S.P.; Van Der Brug, M.P.; Wahlestedt, C. Inhibition of natural antisense transcripts in vivo results in gene-specific transcriptional upregulation. Nat. Biotechnol. 2012, 30, 453. [Google Scholar] [CrossRef]
- Zhang, B.; Arun, G.; Mao, Y.S.; Lazar, Z.; Hung, G.; Bhattacharjee, G.; Xiao, X.; Booth, C.J.; Wu, J.; Zhang, C. The lncRNA Malat1 is dispensable for mouse development but its transcription plays a cis-regulatory role in the adult. Cell Rep. 2012, 2, 111–123. [Google Scholar] [CrossRef] [PubMed]
- Bernard, D.; Prasanth, K.V.; Tripathi, V.; Colasse, S.; Nakamura, T.; Xuan, Z.; Zhang, M.Q.; Sedel, F.; Jourdren, L.; Coulpier, F. A long nuclear-retained non-coding RNA regulates synaptogenesis by modulating gene expression. EMBO J. 2010, 29, 3082–3093. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Robles, A.; Sherwood, C.C. Human brain evolution. How the increase of brain plasticity made us a cultural species. Métode Sci. Stud. J. 2017, 35–43. [Google Scholar] [CrossRef]
- West, A.E.; Greenberg, M.E. Neuronal activity–regulated gene transcription in synapse development and cognitive function. Cold Spring Harb. Perspect. Biol. 2011, 3, a005744. [Google Scholar] [CrossRef]
- Kaikkonen, M.U.; Spann, N.J.; Heinz, S.; Romanoski, C.E.; Allison, K.A.; Stender, J.D.; Chun, H.B.; Tough, D.F.; Prinjha, R.K.; Benner, C. Remodeling of the enhancer landscape during macrophage activation is coupled to enhancer transcription. Mol. Cell 2013, 51, 310–325. [Google Scholar] [CrossRef]
- Lai, F.; Orom, U.A.; Cesaroni, M.; Beringer, M.; Taatjes, D.J.; Blobel, G.A.; Shiekhattar, R. Activating RNAs associate with Mediator to enhance chromatin architecture and transcription. Nature 2013, 494, 497. [Google Scholar] [CrossRef]
- Lam, M.T.; Cho, H.; Lesch, H.P.; Gosselin, D.; Heinz, S.; Tanaka-Oishi, Y.; Benner, C.; Kaikkonen, M.U.; Kim, A.S.; Kosaka, M. Rev-Erbs repress macrophage gene expression by inhibiting enhancer-directed transcription. Nature 2013, 498, 511. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Notani, D.; Ma, Q.; Tanasa, B.; Nunez, E.; Chen, A.Y.; Merkurjev, D.; Zhang, J.; Ohgi, K.; Song, X. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature 2013, 498, 516. [Google Scholar] [CrossRef] [PubMed]
- Melo, C.A.; Drost, J.; Wijchers, P.J.; van de Werken, H.; de Wit, E.; Vrielink, J.A.O.; Elkon, R.; Melo, S.A.; Léveillé, N.; Kalluri, R. eRNAs are required for p53-dependent enhancer activity and gene transcription. Mol. Cell 2013, 49, 524–535. [Google Scholar]
- Mousavi, K.; Zare, H.; Dell’Orso, S.; Grontved, L.; Gutierrez-Cruz, G.; Derfoul, A.; Hager, G.L.; Sartorelli, V. eRNAs promote transcription by establishing chromatin accessibility at defined genomic loci. Mol. Cell 2013, 51, 606–617. [Google Scholar] [CrossRef]
- Maag, J.L.; Panja, D.; Sporild, I.; Patil, S.; Kaczorowski, D.C.; Bramham, C.R.; Dinger, M.E.; Wibrand, K. Dynamic expression of long noncoding RNAs and repeat elements in synaptic plasticity. Front. Neurosci. 2015, 9, 351. [Google Scholar] [CrossRef]
- Costa-Mattioli, M.; Sossin, W.S.; Klann, E.; Sonenberg, N. Translational control of long-lasting synaptic plasticity and memory. Neuron 2009, 61, 10–26. [Google Scholar] [CrossRef]
- Muslimov, I.A.; Santi, E.; Homel, P.; Perini, S.; Higgins, D.; Tiedge, H. RNA transport in dendrites: A cis-acting targeting element is contained within neuronal BC1 RNA. J. Neurosci. 1997, 17, 4722–4733. [Google Scholar] [CrossRef]
- Wang, H.; Iacoangeli, A.; Popp, S.; Muslimov, I.A.; Imataka, H.; Sonenberg, N.; Lomakin, I.B.; Tiedge, H. Dendritic BC1 RNA: Functional role in regulation of translation initiation. J. Neurosci. 2002, 22, 10232–10241. [Google Scholar] [CrossRef]
- Zalfa, F.; Giorgi, M.; Primerano, B.; Moro, A.; Di Penta, A.; Reis, S.; Oostra, B.; Bagni, C. The fragile X syndrome protein FMRP associates with BC1 RNA and regulates the translation of specific mRNAs at synapses. Cell 2003, 112, 317–327. [Google Scholar] [CrossRef]
- Lewejohann, L.; Skryabin, B.; Sachser, N.; Prehn, C.; Heiduschka, P.; Thanos, S.; Jordan, U.; Dell’Omo, G.; Vyssotski, A.; Pleskacheva, M. Role of a neuronal small non-messenger RNA: Behavioural alterations in BC1 RNA-deleted mice. Behav. Brain Res. 2004, 154, 273–289. [Google Scholar] [CrossRef]
- Skryabin, B.V.; Sukonina, V.; Jordan, U.; Lewejohann, L.; Sachser, N.; Muslimov, I.; Tiedge, H.; Brosius, J. Neuronal untranslated BC1 RNA: Targeted gene elimination in mice. Mol. Cell. Biol. 2003, 23, 6435–6441. [Google Scholar] [CrossRef] [PubMed]
- Zhong, J.; Chuang, S.-C.; Bianchi, R.; Zhao, W.; Lee, H.; Fenton, A.A.; Wong, R.K.; Tiedge, H. BC1 regulation of metabotropic glutamate receptor-mediated neuronal excitability. J. Neurosci. 2009, 29, 9977–9986. [Google Scholar] [CrossRef] [PubMed]
- Muslimov, I.A.; Banker, G.; Brosius, J.; Tiedge, H. Activity-dependent regulation of dendritic BC1 RNA in hippocampal neurons in culture. J. Cell Biol. 1998, 141, 1601–1611. [Google Scholar] [CrossRef] [PubMed]
- Dinger, M.E.; Mercer, T.R.; Mattick, J.S. RNAs as extracellular signaling molecules. J. Mol. Endocrinol. 2008, 40, 151–159. [Google Scholar] [CrossRef] [PubMed]
- Frühbeis, C.; Fröhlich, D.; Krämer-Albers, E.-M. Emerging roles of exosomes in neuron–glia communication. Front. Physiol. 2012, 3, 119. [Google Scholar] [CrossRef] [PubMed]
- Van der Vos, K.E.; Balaj, L.; Skog, J.; Breakefield, X.O. Brain tumor microvesicles: Insights into intercellular communication in the nervous system. Cell. Mol. Neurobiol. 2011, 31, 949–959. [Google Scholar] [CrossRef]
- Choi, S.-W.; Kim, H.-W.; Nam, J.-W. The small peptide world in long noncoding RNAs. Brief. Bioinform. 2018, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Hart, R.P.; Goff, L.A. Long noncoding RNAs: Central to nervous system development. Int. J. Dev. Neurosci. 2016, 55, 109–116. [Google Scholar] [CrossRef]
- Théry, C. Exosomes: Secreted vesicles and intercellular communications. F1000 Biol. Rep. 2011, 3, 15. [Google Scholar] [CrossRef]

| Process | lncRNA | Biological Function/Phenotype | Molecular Function |
|---|---|---|---|
| Neural stem cell proliferation and differentiation | Pnky | Regulates neuronal differentiation of embryonic and adult NSPCs | Pnky together with PTBP1 regulate the expression and alternative splicing of an overlapping set of transcripts to promote neurogenesis |
| Paupar | Knockdown of Paupar induces neural differentiation of Neuro-2a neuroblastoma cells | Paupar regulates Pax6 expression locally in cis. Trans: Paupar also associates with PAX6 protein and localizes at promoters of Sox2, Nanog, and Hes1 | |
| Rmst | Promotes neuronal differentiation | Rmst interacts with SOX2 to regulate neurogenic genes including Ascl1 and Dlx1 in trans | |
| Tuna | Regulates pluripotency and neural differentiation of ESCs | Tuna forms a complex with three pluripotency related RNA-binding proteins, PTBP1, hnRNP-K, and NCL | |
| linc-Brn1b | controls differentiation of delaminating neural progenitor cells | Cis regulation of neighbouring BRN1 | |
| Gomafu | Controls retinal development; Dysregulated in schizophrenia | Gomafu regulates splicing of neuronal genes, including DISC1, ERRB4, and WNT7B, probably via association with splicing factors SF1, SRSF1, and QKI | |
| Dali | Depletion of Dali in Neuro-2a neuroblastoma cell inhibits its neuronal differentiation induced by retinoic acid | Cis: Dali maintains Brn1 expression. Trans: Dali interacts with the DNMT1 to regulates DNA methylation status of CpG island-associated promoters; interacts with BRN1 to regulate expression of neural differentiation genes | |
| Neurite outgrowth and synaptogenesis | Bdnf-AS | Depletion of Bdnf-AS promotes neuronal outgrowth and adult neurogenesis | repression of the BDNF growth factor gene through the recruitment of the PRC2 to the Bdnf locus |
| BC1/BC200 | Regulates synaptic excitability, turnover and plasticity | represses local translation in synapses by interaction with FMRP and translational machineries like eIF4a and poly(A)-binding protein | |
| Malat1 | Promotes dendrite maturation and synaptogenesis in cultured hippocampal neurons | Malat1 associates with SR family splicing factors to controls expression of synaptic molecules | |
| Interneurons | Evf2 | Ensures proper formation of GABA-dependent neuronal circuitry | Evf2 associates with DLX1/2 and MECP2 at the regulatory elements in the Dlx5/6 intergenic region to control Dlx5, Dlx6 and Gad1 expression |
© 2019 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zimmer-Bensch, G. Emerging Roles of Long Non-Coding RNAs as Drivers of Brain Evolution. Cells 2019, 8, 1399. https://doi.org/10.3390/cells8111399
Zimmer-Bensch G. Emerging Roles of Long Non-Coding RNAs as Drivers of Brain Evolution. Cells. 2019; 8(11):1399. https://doi.org/10.3390/cells8111399
Chicago/Turabian StyleZimmer-Bensch, Geraldine. 2019. "Emerging Roles of Long Non-Coding RNAs as Drivers of Brain Evolution" Cells 8, no. 11: 1399. https://doi.org/10.3390/cells8111399
APA StyleZimmer-Bensch, G. (2019). Emerging Roles of Long Non-Coding RNAs as Drivers of Brain Evolution. Cells, 8(11), 1399. https://doi.org/10.3390/cells8111399





