Genome Editing in Plants: Exploration of Technological Advancements and Challenges
Abstract
1. Introduction
2. Cas Variants and Other Nucleases for Plant Genome Editing
3. Genes Targeted for Genome Editing in Plants
3.1. Evaluation of CRISPR/Cas Efficiency Using Easily Scorable Marker Genes
3.2. Translational Efforts by Targeting Genes Previously Annotated with RNAi
3.3. Genes for the Enhancement of Resistance against Biotic Stresses
3.4. Genes for the Enhancement of Abiotic Stress Tolerance in Plants
4. Editing Polyploidy Genomes—Challenges and Perspective
5. Multi-Targeting Genome Editing Approaches
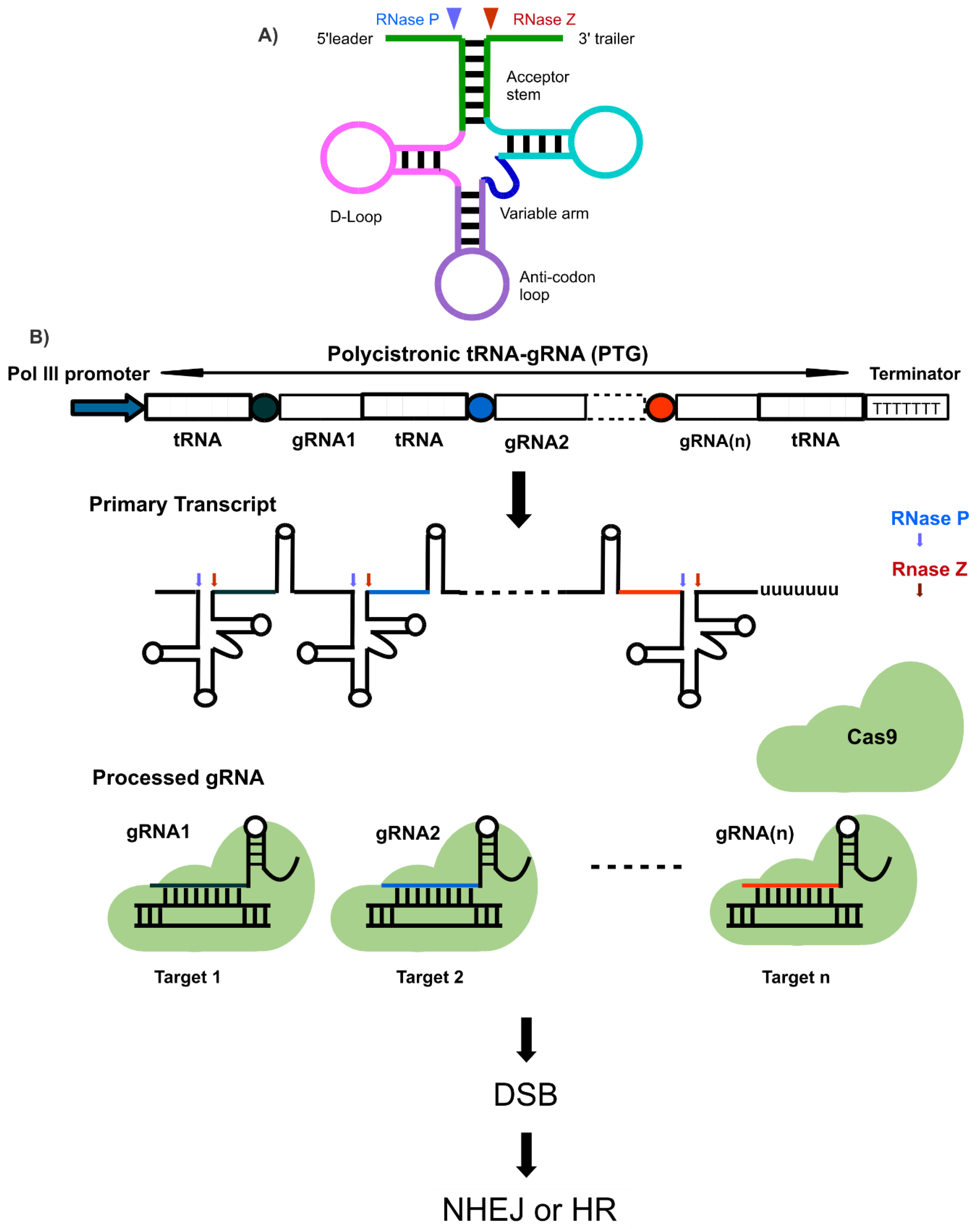
5.1. t-RNA Mediated Multi-Targeting Genome Editing
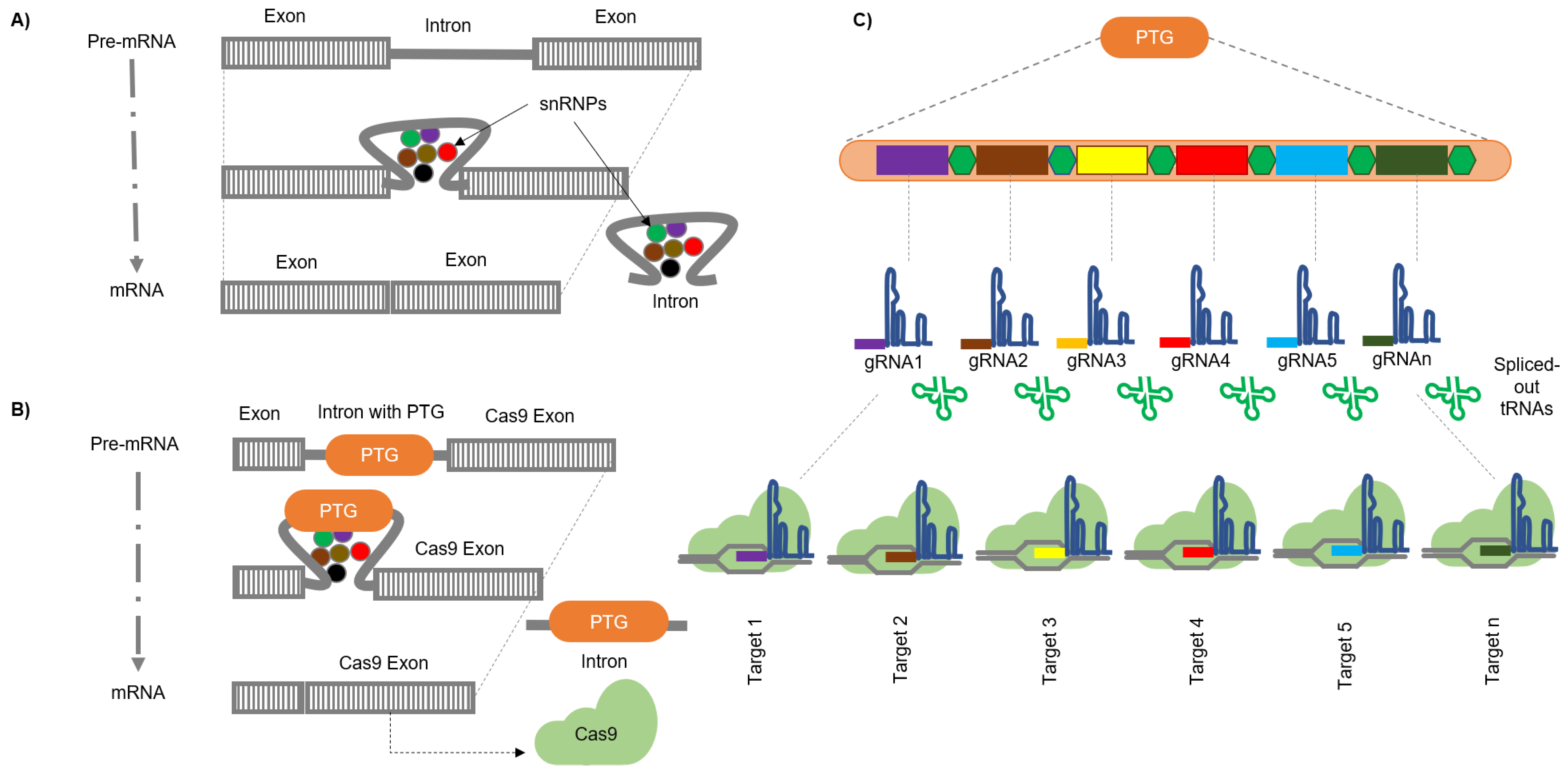
5.2. Engineering Introns to Express sgRNAs
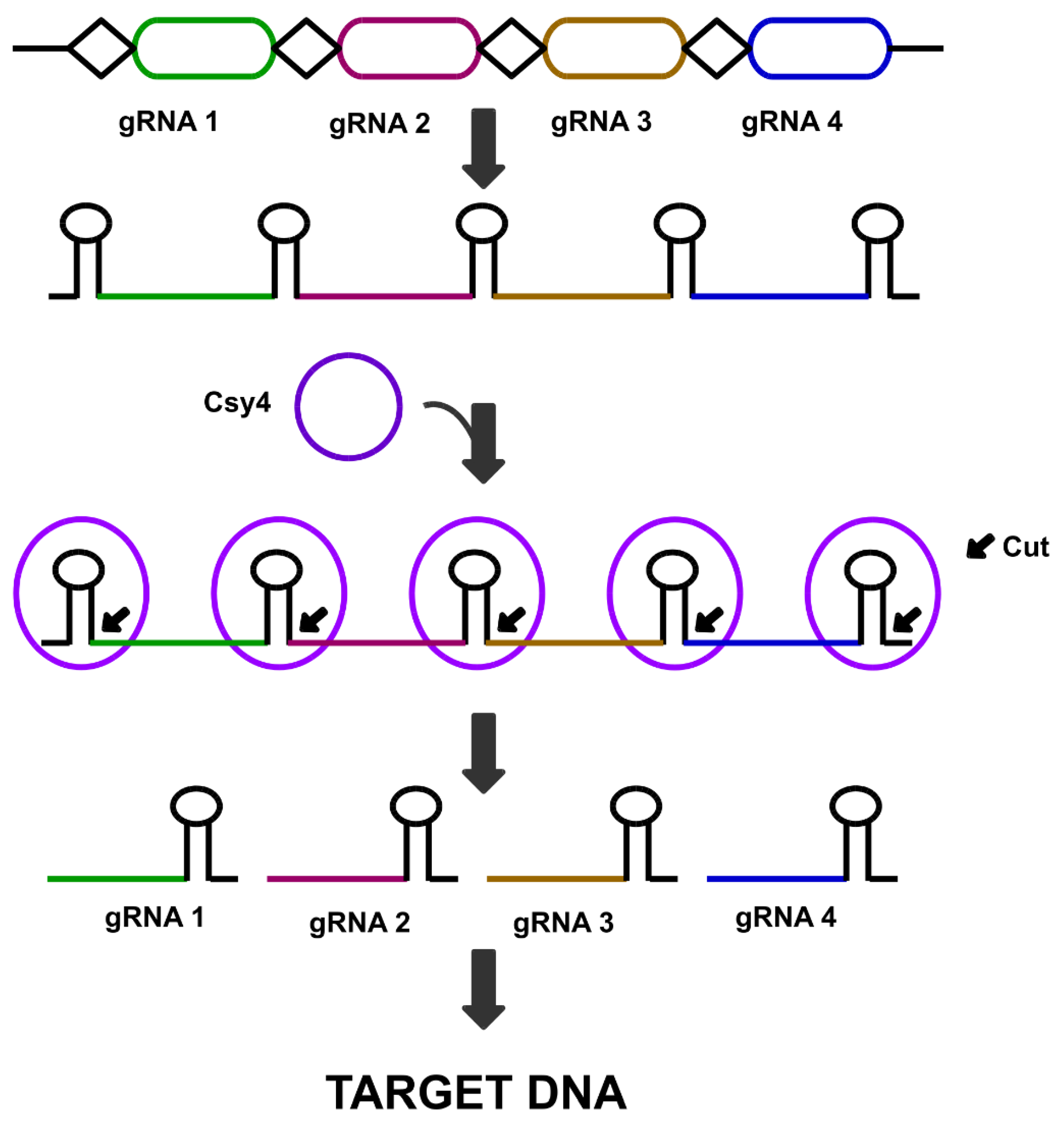
5.3. Csy4 Nuclease Mediated Multi-Targeting Genome Editing
5.4. Drosha-Based Multi-Targeting Genome Editing
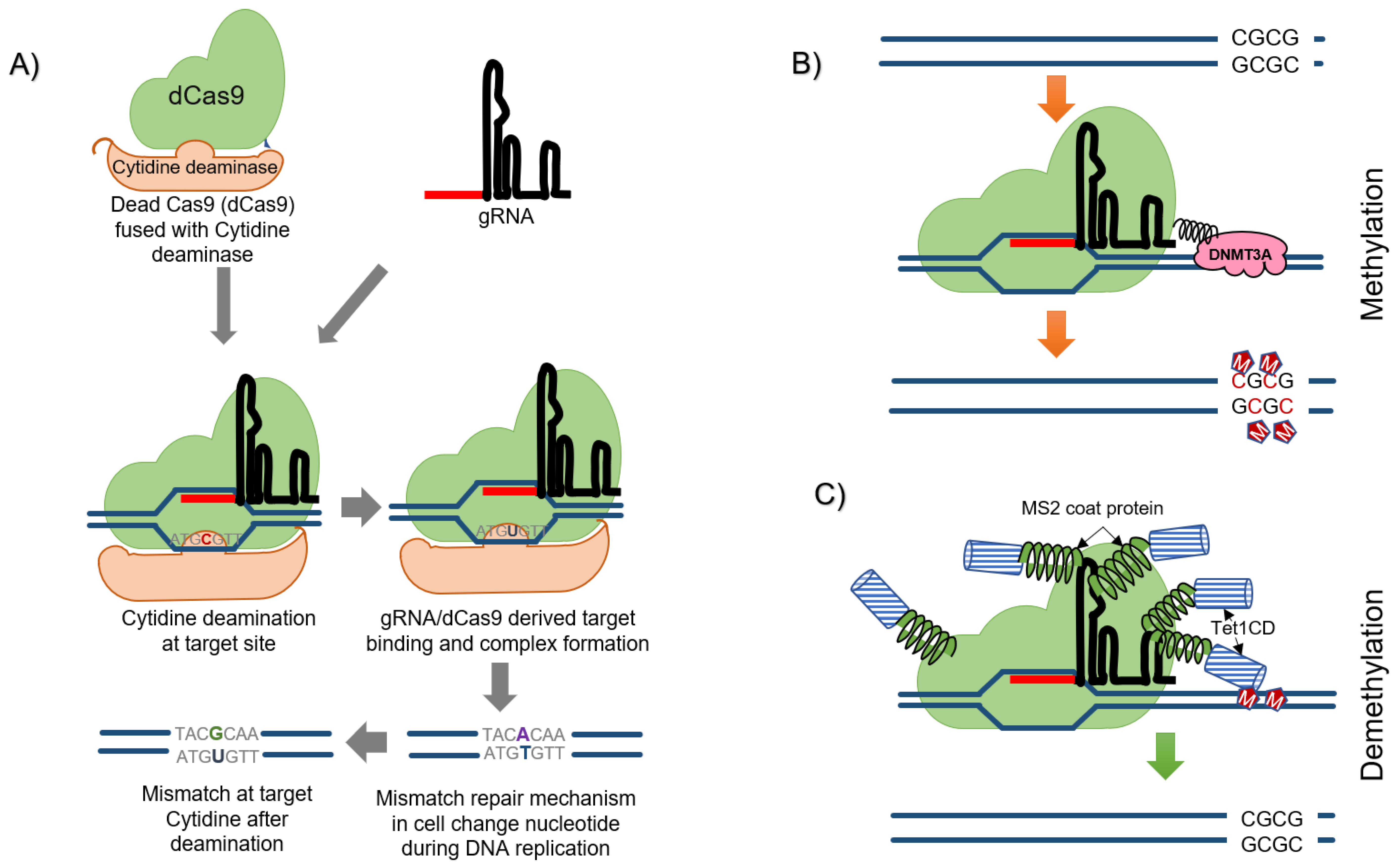
6. Precision Editing/Base-Editing Approach
7. CRISPR Mediated Manipulation of Gene Expression: Promoter Bashing
8. CRISPR Mediated Editing of Methylome
9. Concerns about Off-Targeting Activities of Cas Protein
10. Tools Available for Designing sgRNA and Detection of Potential Off-Target Sites
11. Challenges for Plants Difficult to Transform
Hand-In-Hand Improved Transformation Approaches
12. DNA-Free Reagent Delivery Methods
13. Germline-Specific Gene Expression
14. Future Prospects
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rudin, N.; Sugarman, E.; Haber, J.E. Genetic and physical analysis of double-strand break repair and recombination in Saccharomyces cerevisiae. Genetics 1989, 122, 519–534. [Google Scholar] [PubMed]
- Plessis, A.; Perrin, A.; Haber, J.; Dujon, B. Site-specific recombination determined by I-SceI, a mitochondrial group I intron-encoded endonuclease expressed in the yeast nucleus. Genetics 1992, 130, 451–460. [Google Scholar] [PubMed]
- Faruqi, A.F.; Egholm, M.; Glazer, P.M. Peptide nucleic acid-targeted mutagenesis of a chromosomal gene in mouse cells. Proc. Natl. Acad. Sci. USA 1998, 95, 1398–1403. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Zimmerly, S.; Perlman, P.S.; Lambowitz, A.M. Efficient integration of an intron RNA into double-stranded DNA by reverse splicing. Nature 1996, 381, 332. [Google Scholar] [CrossRef] [PubMed]
- Sullenger, B.A.; Cech, T.R. Ribozyme-mediated repair of defective mRNA by targeted trans-splicing. Nature 1994, 371, 619. [Google Scholar] [CrossRef]
- Chevalier, B.S.; Kortemme, T.; Chadsey, M.S.; Baker, D.; Monnat, R.J., Jr.; Stoddard, B.L. Design, activity, and structure of a highly specific artificial endonuclease. Mol. Cell 2002, 10, 895–905. [Google Scholar] [CrossRef]
- Jacquier, A.; Dujon, B. An intron-encoded protein is active in a gene conversion process that spreads an intron into a mitochondrial gene. Cell 1985, 41, 383–394. [Google Scholar] [CrossRef]
- Rouet, P.; Smih, F.; Jasin, M. Introduction of double-strand breaks into the genome of mouse cells by expression of a rare-cutting endonuclease. Mol. Cell. Biol. 1994, 14, 8096–8106. [Google Scholar] [CrossRef]
- Rai, K.M.; Ghose, K.; Rai, A.; Singh, H.; Srivastava, R.; Mendu, V. Genome Engineering Tools in Plant Synthetic Biology. In Current Developments in Biotechnology and Bioengineering; Elsevier: Amsterdam, The Netherlands, 2019; pp. 47–73. [Google Scholar]
- Bibikova, M.; Beumer, K.; Trautman, J.K.; Carroll, D. Enhancing gene targeting with designed zinc finger nucleases. Science 2003, 300, 764. [Google Scholar] [CrossRef]
- Bibikova, M.; Golic, M.; Golic, K.G.; Carroll, D. Targeted chromosomal cleavage and mutagenesis in Drosophila using zinc-finger nucleases. Genetics 2002, 161, 1169–1175. [Google Scholar]
- Kim, Y.-G.; Cha, J.; Chandrasegaran, S. Hybrid restriction enzymes: Zinc finger fusions to Fok I cleavage domain. Proc. Natl. Acad. Sci. USA 1996, 93, 1156–1160. [Google Scholar] [CrossRef] [PubMed]
- Lloyd, A.; Plaisier, C.L.; Carroll, D.; Drews, G.N. Targeted mutagenesis using zinc-finger nucleases in Arabidopsis. Proc. Natl. Acad. Sci. USA 2005, 102, 2232–2237. [Google Scholar] [CrossRef] [PubMed]
- Osakabe, K.; Osakabe, Y.; Toki, S. Site-directed mutagenesis in Arabidopsis using custom-designed zinc finger nucleases. Proc. Natl. Acad. Sci. USA 2010, 107, 12034–12039. [Google Scholar] [CrossRef] [PubMed]
- Townsend, J.A.; Wright, D.A.; Winfrey, R.J.; Fu, F.; Maeder, M.L.; Joung, J.K.; Voytas, D.F. High-frequency modification of plant genes using engineered zinc-finger nucleases. Nature 2009, 459, 442. [Google Scholar] [CrossRef] [PubMed]
- Wright, D.A.; Townsend, J.A.; Winfrey, R.J., Jr.; Irwin, P.A.; Rajagopal, J.; Lonosky, P.M.; Hall, B.D.; Jondle, M.D.; Voytas, D.F. High-frequency homologous recombination in plants mediated by zinc-finger nucleases. Plant J. 2005, 44, 693–705. [Google Scholar] [CrossRef]
- Petolino, J.F.; Worden, A.; Curlee, K.; Connell, J.; Moynahan, T.L.S.; Larsen, C.; Russell, S. Zinc finger nuclease-mediated transgene deletion. Plant Mol. Biol. 2010, 73, 617–628. [Google Scholar] [CrossRef]
- Shukla, V.K.; Doyon, Y.; Miller, J.C.; DeKelver, R.C.; Moehle, E.A.; Worden, S.E.; Mitchell, J.C.; Arnold, N.L.; Gopalan, S.; Meng, X. Precise genome modification in the crop species Zea mays using zinc-finger nucleases. Nature 2009, 459, 437. [Google Scholar] [CrossRef]
- Boch, J.; Scholze, H.; Schornack, S.; Landgraf, A.; Hahn, S.; Kay, S.; Lahaye, T.; Nickstadt, A.; Bonas, U. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 2009, 326, 1509–1512. [Google Scholar] [CrossRef]
- Christian, M.; Cermak, T.; Doyle, E.L.; Schmidt, C.; Zhang, F.; Hummel, A.; Bogdanove, A.J.; Voytas, D.F. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 2010, 186, 757–761. [Google Scholar] [CrossRef]
- Shan, Q.; Wang, Y.; Chen, K.; Liang, Z.; Li, J.; Zhang, Y.; Zhang, K.; Liu, J.; Voytas, D.F.; Zheng, X. Rapid and efficient gene modification in rice and Brachypodium using TALENs. Mol. Plant 2013, 6, 1365–1368. [Google Scholar] [CrossRef]
- Li, T.; Liu, B.; Spalding, M.H.; Weeks, D.P.; Yang, B. High-efficiency TALEN-based gene editing produces disease-resistant rice. Nat. Biotechnol. 2012, 30, 390. [Google Scholar] [CrossRef] [PubMed]
- Kopischke, S.; Schüßler, E.; Althoff, F.; Zachgo, S. TALEN-mediated genome-editing approaches in the liverwort Marchantia polymorpha yield high efficiencies for targeted mutagenesis. Plant methods 2017, 13, 20. [Google Scholar] [CrossRef]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef] [PubMed]
- Cong, L.; Ran, F.A.; Cox, D.; Lin, S.; Barretto, R.; Habib, N.; Hsu, P.D.; Wu, X.; Jiang, W.; Marraffini, L. Multiplex genome engineering using CRISPR/Cas systems. Science 2013, 339, 819–823. [Google Scholar] [CrossRef] [PubMed]
- Ishino, Y.; Shinagawa, H.; Makino, K.; Amemura, M.; Nakata, A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J. Bacteriol. 1987, 169, 5429–5433. [Google Scholar] [CrossRef]
- Bolotin, A.; Quinquis, B.; Sorokin, A.; Ehrlich, S.D. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 2005, 151, 2551–2561. [Google Scholar]
- Mojica, F.J.; García-Martínez, J.; Soria, E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J. Mol. Evol. 2005, 60, 174–182. [Google Scholar] [CrossRef]
- Barrangou, R.; Fremaux, C.; Deveau, H.; Richards, M.; Boyaval, P.; Moineau, S.; Romero, D.A.; Horvath, P. CRISPR provides acquired resistance against viruses in prokaryotes. Science 2007, 315, 1709–1712. [Google Scholar] [CrossRef]
- Cho, S.W.; Kim, S.; Kim, J.M.; Kim, J.-S. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat. Biotechnol. 2013, 31, 230. [Google Scholar] [CrossRef]
- Hwang, W.Y.; Fu, Y.; Reyon, D.; Maeder, M.L.; Tsai, S.Q.; Sander, J.D.; Peterson, R.T.; Yeh, J.J.; Joung, J.K. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat. Biotechnol. 2013, 31, 227. [Google Scholar] [CrossRef]
- Jinek, M.; East, A.; Cheng, A.; Lin, S.; Ma, E.; Doudna, J. RNA-programmed genome editing in human cells. elife 2013, 2, e00471. [Google Scholar] [CrossRef] [PubMed]
- Mali, P.; Yang, L.; Esvelt, K.M.; Aach, J.; Guell, M.; DiCarlo, J.E.; Norville, J.E.; Church, G.M. RNA-guided human genome engineering via Cas9. Science 2013, 339, 823–826. [Google Scholar] [CrossRef] [PubMed]
- Li, J.-F.; Norville, J.E.; Aach, J.; McCormack, M.; Zhang, D.; Bush, J.; Church, G.M.; Sheen, J. Multiplex and homologous recombination–mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat. Biotechnol. 2013, 31, 688. [Google Scholar] [CrossRef] [PubMed]
- Nekrasov, V.; Staskawicz, B.; Weigel, D.; Jones, J.D.; Kamoun, S. Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat. Biotechnol. 2013, 31, 691. [Google Scholar] [CrossRef]
- Shan, Q.; Wang, Y.; Li, J.; Zhang, Y.; Chen, K.; Liang, Z.; Zhang, K.; Liu, J.; Xi, J.J.; Qiu, J. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat. Biotechnol. 2013, 31, 686. [Google Scholar] [CrossRef]
- Deltcheva, E.; Chylinski, K.; Sharma, C.M.; Gonzales, K.; Chao, Y.; Pirzada, Z.A.; Eckert, M.R.; Vogel, J.; Charpentier, E. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 2011, 471, 602. [Google Scholar] [CrossRef]
- Rong, Y.S.; Golic, K.G. Gene targeting by homologous recombination in Drosophila. Science 2000, 288, 2013–2018. [Google Scholar] [CrossRef]
- Nishitani, C.; Hirai, N.; Komori, S.; Wada, M.; Okada, K.; Osakabe, K.; Yamamoto, T.; Osakabe, Y. Efficient genome editing in apple using a CRISPR/Cas9 system. Sci. Rep. 2016, 6, 31481. [Google Scholar] [CrossRef]
- Feng, Z.; Zhang, B.; Ding, W.; Liu, X.; Yang, D.-L.; Wei, P.; Cao, F.; Zhu, S.; Zhang, F.; Mao, Y. Efficient genome editing in plants using a CRISPR/Cas system. Cell Res. 2013, 23, 1229. [Google Scholar] [CrossRef]
- Kaur, N.; Alok, A.; Kaur, N.; Pandey, P.; Awasthi, P.; Tiwari, S. CRISPR/Cas9-mediated efficient editing in phytoene desaturase (PDS) demonstrates precise manipulation in banana cv. Rasthali Genome Funct. Integr. Genom. 2018, 18, 89–99. [Google Scholar] [CrossRef]
- Zheng, X.; Yang, S.; Zhang, D.; Zhong, Z.; Tang, X.; Deng, K.; Zhou, J.; Qi, Y.; Zhang, Y. Effective screen of CRISPR/Cas9-induced mutants in rice by single-strand conformation polymorphism. Plant Cell Rep. 2016, 35, 1545–1554. [Google Scholar] [CrossRef] [PubMed]
- Begemann, M.B.; Gray, B.N.; January, E.; Gordon, G.C.; He, Y.; Liu, H.; Wu, X.; Brutnell, T.P.; Mockler, T.C.; Oufattole, M. Precise insertion and guided editing of higher plant genomes using Cpf1 CRISPR nucleases. Sci. Rep. 2017, 7, 11606. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Li, J.; Zhang, J.; Du, W.; Fu, J.; Sutar, S.; Zhao, Y.; Xia, L. Synthesis-dependent repair of Cpf1-induced double-strand DNA breaks enables targeted gene replacement in rice. J. Exp. Bot. 2018, 69, 4715–4721. [Google Scholar] [CrossRef] [PubMed]
- Miki, D.; Zhang, W.; Zeng, W.; Feng, Z.; Zhu, J.-K. CRISPR/Cas9-mediated gene targeting in Arabidopsis using sequential transformation. Nat. Commun. 2018, 9, 1967. [Google Scholar] [CrossRef]
- Svitashev, S.; Young, J.; Schwartz, C.; Gao, H.; Falco, S.C.; Cigan, A. Targeted mutagenesis, precise gene editing and site-specific gene insertion in maize using Cas9 and guide RNA. Plant Physiol. 2015, 169, 931–945. [Google Scholar] [CrossRef]
- Xie, K.; Zhang, J.; Yang, Y. Genome-wide prediction of highly specific guide RNA spacers for CRISPR–Cas9-mediated genome editing in model plants and major crops. Mol. Plant 2014, 7, 923–926. [Google Scholar] [CrossRef]
- Zetsche, B.; Gootenberg, J.S.; Abudayyeh, O.O.; Slaymaker, I.M.; Makarova, K.S.; Essletzbichler, P.; Volz, S.E.; Joung, J.; Van Der Oost, J.; Regev, A. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 2015, 163, 759–771. [Google Scholar] [CrossRef]
- Stella, S.; Alcón, P.; Montoya, G.J.N. Structure of the Cpf1 endonuclease R-loop complex after target DNA cleavage. Nature 2017, 546, 559. [Google Scholar] [CrossRef]
- Bernabé-Orts, J.M.; Casas-Rodrigo, I.; Minguet, E.G.; Landolfi, V.; Garcia-Carpintero, V.; Gianoglio, S.; Vázquez-Vilar, M.; Granell, A.; Orzaez, D. Assessment of Cas12a-mediated gene editing efficiency in plants. Plant Biotechnol. J. 2019. [Google Scholar] [CrossRef]
- Wang, M.; Mao, Y.; Lu, Y.; Tao, X.; Zhu, J.-K. Multiplex gene editing in rice using the CRISPR-Cpf1 system. Mol. Plant 2017, 10, 1011–1013. [Google Scholar] [CrossRef]
- Tang, X.; Liu, G.; Zhou, J.; Ren, Q.; You, Q.; Tian, L.; Xin, X.; Zhong, Z.; Liu, B.; Zheng, X. A large-scale whole-genome sequencing analysis reveals highly specific genome editing by both Cas9 and Cpf1 (Cas12a) nucleases in rice. Genome Boil. 2018, 19, 84. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Lowder, L.G.; Zhang, T.; Malzahn, A.A.; Zheng, X.; Voytas, D.F.; Zhong, Z.; Chen, Y.; Ren, Q.; Li, Q. A CRISPR–Cpf1 system for efficient genome editing and transcriptional repression in plants. Nat. Plants 2017, 3, 17018. [Google Scholar] [CrossRef] [PubMed]
- Karvelis, T.; Bigelyte, G.; Young, J.K.; Hou, Z.; Zedaveinyte, R.; Pociute, K.; Silanskas, A.; Venclovas, C.; Siksnys, V. PAM recognition by miniature CRISPR-Cas14 triggers programmable double-stranded DNA cleavage. bioRxiv 2019. [Google Scholar] [CrossRef]
- Makarova, K.S.; Wolf, Y.I.; Alkhnbashi, O.S.; Costa, F.; Shah, S.A.; Saunders, S.J.; Barrangou, R.; Brouns, S.J.; Charpentier, E.; Haft, D. An updated evolutionary classification of CRISPR–Cas systems. Nat. Rev. Genet. 2015, 13, 722. [Google Scholar] [CrossRef] [PubMed]
- Gao, W.; Long, L.; Tian, X.; Xu, F.; Liu, J.; Singh, P.K.; Botella, J.R.; Song, C. Genome editing in cotton with the CRISPR/Cas9 system. Front. Plant Sci. 2017, 8, 1364. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wang, S.; Li, D.; Zhang, Q.; Li, L.; Zhong, C.; Liu, Y.; Huang, H. Optimized paired-sgRNA/Cas9 cloning and expression cassette triggers high-efficiency multiplex genome editing in kiwifruit. Plant Biotechnol. J. 2018, 16, 1424–1433. [Google Scholar] [CrossRef] [PubMed]
- Breitler, J.-C.; Dechamp, E.; Campa, C.; Rodrigues, L.A.Z.; Guyot, R.; Marraccini, P.; Etienne, H.; Tissue; Culture, O. CRISPR/Cas9-mediated efficient targeted mutagenesis has the potential to accelerate the domestication of Coffea canephora. Plant Celltissue Organ Cult. 2018, 134, 383–394. [Google Scholar] [CrossRef]
- Odipio, J.; Alicai, T.; Ingelbrecht, I.; Nusinow, D.A.; Bart, R.; Taylor, N. Efficient CRISPR/Cas9 genome editing of phytoene desaturase in cassava. Front. Plant Sci. 2017, 8, 1780. [Google Scholar] [CrossRef]
- Osakabe, Y.; Liang, Z.; Ren, C.; Nishitani, C.; Osakabe, K.; Wada, M.; Komori, S.; Malnoy, M.; Velasco, R.; Poli, M. CRISPR–Cas9-mediated genome editing in apple and grapevine. Nat. Protoc. 2018, 13, 2844. [Google Scholar] [CrossRef]
- Wang, P.; Zhang, J.; Sun, L.; Ma, Y.; Xu, J.; Liang, S.; Deng, J.; Tan, J.; Zhang, Q.; Tu, L. High efficient multisites genome editing in allotetraploid cotton (Gossypium hirsutum) using CRISPR/Cas9 system. Plant Biotechnol. J. 2018, 16, 137–150. [Google Scholar] [CrossRef]
- Estévez, J.M.; Cantero, A.; Romero, C.; Kawaide, H.; Jiménez, L.F.; Kuzuyama, T.; Seto, H.; Kamiya, Y.; León, P. Analysis of the expression of CLA1, a gene that encodes the 1-deoxyxylulose 5-phosphate synthase of the 2-C-methyl-D-erythritol-4-phosphate pathway in Arabidopsis. Plant Physiol. 2000, 124, 95–104. [Google Scholar] [CrossRef] [PubMed]
- Qin, G.; Gu, H.; Ma, L.; Peng, Y.; Deng, X.W.; Chen, Z.; Qu, L. Disruption of phytoene desaturase gene results in albino and dwarf phenotypes in Arabidopsis by impairing chlorophyll, carotenoid, and gibberellin biosynthesis. Cell Res. 2007, 17, 471. [Google Scholar] [CrossRef] [PubMed]
- Pessina, S.; Lenzi, L.; Perazzolli, M.; Campa, M.; Dalla Costa, L.; Urso, S.; Valè, G.; Salamini, F.; Velasco, R.; Malnoy, M. Knockdown of MLO genes reduces susceptibility to powdery mildew in grapevine. Hortic. Res. 2016, 3, 16016. [Google Scholar] [CrossRef] [PubMed]
- Soyk, S.; Müller, N.A.; Park, S.J.; Schmalenbach, I.; Jiang, K.; Hayama, R.; Zhang, L.; Van Eck, J.; Jiménez-Gómez, J.M.; Lippman, Z.B. Variation in the flowering gene SELF PRUNING 5G promotes day-neutrality and early yield in tomato. Nat. Genet. 2017, 49, 162. [Google Scholar] [CrossRef]
- Uluisik, S.; Chapman, N.H.; Smith, R.; Poole, M.; Adams, G.; Gillis, R.B.; Besong, T.M.; Sheldon, J.; Stiegelmeyer, S.; Perez, L. Genetic improvement of tomato by targeted control of fruit softening. Nat. Biotechnol. 2016, 34, 950. [Google Scholar] [CrossRef]
- Wang, D.; Samsulrizal, N.H.; Yan, C.; Allcock, N.S.; Craigon, J.; Blanco-Ulate, B.; Ortega-Salazar, I.; Marcus, S.E.; Bagheri, H.M.; Fons, L. Characterization of CRISPR Mutants Targeting Genes Modulating Pectin Degradation in Ripening Tomato. Plant Physiol. 2019, 179, 544–557. [Google Scholar]
- van Regenmortel, M.H.; Mahy, B.W. Desk Encyclopedia of Plant and Fungal Virology; Elsevier: San Diego, CA, USA, 2009. [Google Scholar]
- Zaidi, S.; Tashkandi, M.; Mansoor, S.; Mahfouz, M. Engineering plant immunity: Using CRISPR/Cas9 to generate virus resistance. Front. Plant Sci. 2016, 7, 1673. [Google Scholar] [CrossRef]
- Ali, Z.; Ali, S.; Tashkandi, M.; Zaidi, S.; Mahfouz, M. CRISPR/Cas9-mediated immunity to geminiviruses: Differential interference and evasion. Sci. Rep. 2016, 6, 26912. [Google Scholar] [CrossRef]
- Tashkandi, M.; Ali, Z.; Aljedaani, F.; Shami, A.; Mahfouz, M. Engineering resistance against Tomato yellow leaf curl virus via the CRISPR/Cas9 system in tomato. Plant Signal. Behav. 2018, 13, e1525996. [Google Scholar] [CrossRef]
- O’connell, M.R.; Oakes, B.L.; Sternberg, S.H.; East-Seletsky, A.; Kaplan, M.; Doudna, J. Programmable RNA recognition and cleavage by CRISPR/Cas9. Nature 2014, 516, 263. [Google Scholar] [CrossRef]
- Hale, C.R.; Zhao, P.; Olson, S.; Duff, M.O.; Graveley, B.R.; Wells, L.; Terns, R.M.; Terns, M. RNA-guided RNA cleavage by a CRISPR RNA-Cas protein complex. Cell 2009, 139, 945–956. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Zheng, Q.; Yi, X.; An, H.; Zhao, Y.; Ma, S.; Zhou, G. Establishing RNA virus resistance in plants by harnessing CRISPR immune system. Plant Biotechnol. J. 2018, 16, 1415–1423. [Google Scholar] [CrossRef] [PubMed]
- Pyott, D.E.; Sheehan, E.; Molnar, A. Engineering of CRISPR/Cas9-mediated potyvirus resistance in transgene-free Arabidopsis plants. Mol. Plant Pathol. 2016, 17, 1276–1288. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekaran, J.; Brumin, M.; Wolf, D.; Leibman, D.; Klap, C.; Pearlsman, M.; Sherman, A.; Arazi, T.; Gal-On, A. Development of broad virus resistance in non-transgenic cucumber using CRISPR/Cas9 technology. Mol. Plant Pathol. 2016, 17, 1140–1153. [Google Scholar] [CrossRef] [PubMed]
- Sanfaçon, H.J.V. Plant translation factors and virus resistance. Viruses 2015, 7, 3392–3419. [Google Scholar] [CrossRef] [PubMed]
- Borrelli, V.M.; Brambilla, V.; Rogowsky, P.; Marocco, A.; Lanubile, A. The enhancement of plant disease resistance using CRISPR/Cas9 technology. Front. plant Sci. 2018, 9, 1245. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Cheng, X.; Shan, Q.; Zhang, Y.; Liu, J.; Gao, C.; Qiu, J. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat. Biotechnol. 2014, 32, 947. [Google Scholar] [CrossRef]
- Nekrasov, V.; Wang, C.; Win, J.; Lanz, C.; Weigel, D.; Kamoun, S. Rapid generation of a transgene-free powdery mildew resistant tomato by genome deletion. Sci. Rep. 2017, 7, 482. [Google Scholar] [CrossRef]
- Malnoy, M.; Viola, R.; Jung, M.-H.; Koo, O.-J.; Kim, S.; Kim, J.-S.; Velasco, R.; Nagamangala Kanchiswamy, C. DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Front. Plant Sci. 2016, 7, 1904. [Google Scholar] [CrossRef]
- Wang, F.; Wang, C.; Liu, P.; Lei, C.; Hao, W.; Gao, Y.; Liu, Y.-G.; Zhao, K. Enhanced rice blast resistance by CRISPR/Cas9-targeted mutagenesis of the ERF transcription factor gene OsERF922. PLoS ONE 2016, 11, e0154027. [Google Scholar] [CrossRef]
- Bhat, J.A.; Ali, S.; Salgotra, R.K.; Mir, Z.A.; Dutta, S.; Jadon, V.; Tyagi, A.; Mushtaq, M.; Jain, N.; Singh, P. Genomic selection in the era of next generation sequencing for complex traits in plant breeding. Front. Genet. 2016, 7, 221. [Google Scholar] [CrossRef] [PubMed]
- Deshmukh, R.; Sonah, H.; Patil, G.; Chen, W.; Prince, S.; Mutava, R.; Vuong, T.; Valliyodan, B.; Nguyen, H. Integrating omic approaches for abiotic stress tolerance in soybean. Front. Plant Sci. 2014, 5, 244. [Google Scholar] [CrossRef] [PubMed]
- Kumawat, S.; Rana, N.; Bansal, R.; Vishwakarma, G.; Mehetre, S.T.; Das, B.K.; Kumar, M.; Kumar Yadav, S.; Sonah, H.; Sharma, T. Expanding Avenue of Fast Neutron Mediated Mutagenesis for Crop Improvement. Plants 2019, 8, 164. [Google Scholar] [CrossRef] [PubMed]
- Chaudhary, J.; Alisha, A.; Bhatt, V.; Chandanshive, S.; Kumar, N.; Mir, Z.; Kumar, A.; Yadav, S.K.; Shivaraj, S.; Sonah, H.J.P. Mutation Breeding in Tomato: Advances, Applicability and Challenges. Plants 2019, 8, 128. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Gao, H.; Wang, H.; Lafitte, H.R.; Archibald, R.L.; Yang, M.; Hakimi, S.M.; Mo, H.; Habben, J.E. ARGOS 8 variants generated by CRISPR-Cas9 improve maize grain yield under field drought stress conditions. Plant Biotechnol. J. 2017, 15, 207–216. [Google Scholar] [CrossRef] [PubMed]
- Osakabe, Y.; Osakabe, K. Genome Editing to Improve Abiotic Stress Responses in Plants. In Progress in Molecular Biology and Translational Science; Elsevier: Amsterdam, The Netherlands, 2017; Volume 149, pp. 99–109. [Google Scholar]
- Zhang, A.; Liu, Y.; Wang, F.; Li, T.; Chen, Z.; Kong, D.; Bi, J.; Zhang, F.; Luo, X.; Wang, J. Enhanced rice salinity tolerance via CRISPR/Cas9-targeted mutagenesis of the OsRR22 gene. Mol. Breed. 2019, 39, 47. [Google Scholar] [CrossRef]
- Bo, W.; Zhaohui, Z.; Huanhuan, Z.; Xia, W.; Binglin, L.; Lijia, Y.; Xiangyan, H.; Deshui, Y.; Xuelian, Z.; Chunguo, W. Targeted Mutagenesis of NAC Transcription Factor Gene, OsNAC041, Leading to Salt Sensitivity in Rice. Rice Sci. 2019, 26, 98–108. [Google Scholar] [CrossRef]
- Campa, C.C.; Weisbach, N.R.; Santinha, A.J.; Incarnato, D.; Platt, R. Multiplexed genome engineering by Cas12a and CRISPR arrays encoded on single transcripts. Nat. Methods 2019, 16, 887–893. [Google Scholar] [CrossRef]
- Zargar, S.M.; Nagar, P.; Deshmukh, R.; Nazir, M.; Wani, A.A.; Masoodi, K.Z.; Agrawal, G.K.; Rakwal, R. Aquaporins as potential drought tolerance inducing proteins: Towards instigating stress tolerance. J. Proteom. 2017, 169, 233–238. [Google Scholar] [CrossRef]
- Sonah, H.; Deshmukh, R.K.; Labbé, C.; Bélanger, R. Analysis of aquaporins in Brassicaceae species reveals high-level of conservation and dynamic role against biotic and abiotic stress in canola. Sci. Rep. 2017, 7, 2771. [Google Scholar] [CrossRef]
- Shivaraj, S.; Deshmukh, R.; Sonah, H.; Bélanger, R. Identification and characterization of aquaporin genes in Arachis duranensis and Arachis ipaensis genomes, the diploid progenitors of peanut. Bmc Genom. 2019, 20, 222. [Google Scholar] [CrossRef] [PubMed]
- Vishwakarma, K.; Mishra, M.; Patil, G.; Mulkey, S.; Ramawat, N.; Pratap Singh, V.; Deshmukh, R.; Kumar Tripathi, D.; Nguyen, H.T.; Sharma, S. Avenues of the membrane transport system in adaptation of plants to abiotic stresses. Crit. Rev. Biotechnol. 2019, 39, 861–883. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, P.; Tripathi, D.K.; Deshmukh, R.; Pratap Singh, V.; Corpas, F.J. Revisiting the Role of ROS and RNS in Plants under Changing Environment; Elsevier: Amsterdam, The Netherlands, 2019. [Google Scholar]
- Bhat, J.A.; Shivaraj, S.; Singh, P.; Navadagi, D.B.; Tripathi, D.K.; Dash, P.K.; Solanke, A.U.; Sonah, H.; Deshmukh, R.J.P. Role of silicon in mitigation of heavy metal stresses in crop plants. Plants 2019, 8, 71. [Google Scholar] [CrossRef] [PubMed]
- Mushtaq, M.; Bhat, J.A.; Mir, Z.A.; Sakina, A.; Ali, S.; Singh, A.K.; Tyagi, A.; Salgotra, R.K.; Dar, A.A.; Bhat, R. CRISPR/Cas approach: A new way of looking at plant-abiotic interactions. J. Plant Physiol. 2018, 224, 156–162. [Google Scholar] [CrossRef]
- Chen, X.; Lu, X.; Shu, N.; Wang, S.; Wang, J.; Wang, D.; Guo, L.; Ye, W. Targeted mutagenesis in cotton (Gossypium hirsutum L.) using the CRISPR/Cas9 system. Sci. Rep. 2017, 7, 44304. [Google Scholar] [CrossRef]
- Li, C.; Unver, T.; Zhang, B. A high-efficiency CRISPR/Cas9 system for targeted mutagenesis in cotton (Gossypium hirsutum L.). Sci. Rep. 2017, 7, 43902. [Google Scholar] [CrossRef]
- Janga, M.R.; Campbell, L.M.; Rathore, K.S. CRISPR/Cas9-mediated targeted mutagenesis in upland cotton (Gossypium hirsutum L.). Plant Mol. Biol. 2017, 94, 349–360. [Google Scholar] [CrossRef]
- Wang, Y.; Meng, Z.; Liang, C.; Meng, Z.; Wang, Y.; Sun, G.; Zhu, T.; Cai, Y.; Guo, S.; Zhang, R. Increased lateral root formation by CRISPR/Cas9-mediated editing of arginase genes in cotton. Sci. China Life Sci. 2017, 60, 524–527. [Google Scholar] [CrossRef]
- Jia, H.; Orbovic, V.; Jones, J.B.; Wang, N. Modification of the PthA4 effector binding elements in Type I Cs LOB 1 promoter using Cas9/sg RNA to produce transgenic Duncan grapefruit alleviating XccΔpthA4: dCs LOB 1.3 infection. Plant Biotechnol. J. 2016, 14, 1291–1301. [Google Scholar] [CrossRef]
- Jia, H.; Zhang, Y.; Orbović, V.; Xu, J.; White, F.F.; Jones, J.B.; Wang, N. Genome editing of the disease susceptibility gene Cs LOB 1 in citrus confers resistance to citrus canker. Plant Biotechnol. J. 2017, 15, 817–823. [Google Scholar] [CrossRef]
- Andersson, M.; Turesson, H.; Nicolia, A.; Fält, A.-S.; Samuelsson, M.; Hofvander, P. Efficient targeted multiallelic mutagenesis in tetraploid potato (Solanum tuberosum) by transient CRISPR-Cas9 expression in protoplasts. Plant Cell Rep. 2017, 36, 117–128. [Google Scholar] [CrossRef] [PubMed]
- Butler, N.M.; Atkins, P.A.; Voytas, D.F.; Douches, D. Generation and inheritance of targeted mutations in potato (Solanum tuberosum L.) using the CRISPR/Cas system. PLoS ONE 2015, 10, e0144591. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Zhang, S.; Wang, W.; Xiong, X.; Meng, F.; Cui, X. Efficient targeted mutagenesis in potato by the CRISPR/Cas9 system. Plant Cell Rep. 2015, 34, 1473–1476. [Google Scholar] [CrossRef] [PubMed]
- Butler, N.M.; Baltes, N.J.; Voytas, D.F.; Douches, D.S. Geminivirus-mediated genome editing in potato (Solanum tuberosum L.) using sequence-specific nucleases. Front. Plant Sci. 2016, 7, 1045. [Google Scholar] [CrossRef]
- Souza, G.M.; Berges, H.; Bocs, S.; Casu, R.; D’Hont, A.; Ferreira, J.E.; Henry, R.; Ming, R.; Potier, B.; Van Sluys, M. The sugarcane genome challenge: Strategies for sequencing a highly complex genome. Trop. Plant Biol. 2011, 4, 145–156. [Google Scholar] [CrossRef]
- Irvine, J.J.T.; Genetics, A. Saccharum species as horticultural classes. Theor. Appl. Genet. 1999, 98, 186–194. [Google Scholar] [CrossRef]
- Birch, R.G.; Bower, R.S.; Elliott, A. Highly efficient, 5′-sequence-specific transgene silencing in a complex polyploid. Trop. Plant Biol. 2010, 3, 88–97. [Google Scholar] [CrossRef]
- Mohan, C. Genome editing in sugarcane: Challenges ahead. Front. Plant Sci. 2016, 7, 1542. [Google Scholar] [CrossRef]
- Minkenberg, B.; Wheatley, M.; Yang, Y. CRISPR/Cas9-enabled multiplex genome editing and its application. In Progress in Molecular Biology and Translational Science; Elsevier: Amsterdam, The Netherlands, 2017; Volume 149, pp. 111–132. [Google Scholar]
- Xie, K.; Minkenberg, B.; Yang, Y. Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc. Natl. Acad. Sci. USA 2015, 112, 3570–3575. [Google Scholar] [CrossRef]
- Tsai, S.Q.; Wyvekens, N.; Khayter, C.; Foden, J.A.; Thapar, V.; Reyon, D.; Goodwin, M.J.; Aryee, M.J.; Joung, J. Dimeric CRISPR RNA-guided FokI nucleases for highly specific genome editing. Nat. Biotechnol. 2014, 32, 569. [Google Scholar] [CrossRef]
- Ran, F.A.; Hsu, P.D.; Lin, C.-Y.; Gootenberg, J.S.; Konermann, S.; Trevino, A.E.; Scott, D.A.; Inoue, A.; Matoba, S.; Zhang, Y.J.C. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 2013, 154, 1380–1389. [Google Scholar] [CrossRef] [PubMed]
- Guilinger, J.P.; Thompson, D.B.; Liu, D. Fusion of catalytically inactive Cas9 to FokI nuclease improves the specificity of genome modification. Nat. Biotechnol. 2014, 32, 577. [Google Scholar] [CrossRef] [PubMed]
- Gilbert, L.A.; Larson, M.H.; Morsut, L.; Liu, Z.; Brar, G.A.; Torres, S.E.; Stern-Ginossar, N.; Brandman, O.; Whitehead, E.H.; Doudna, J. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 2013, 154, 442–451. [Google Scholar] [CrossRef] [PubMed]
- Mali, P.; Aach, J.; Stranges, P.B.; Esvelt, K.M.; Moosburner, M.; Kosuri, S.; Yang, L.; Church, G. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat. Biotechnol. 2013, 31, 833. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, R.; Ueta, R.; Abe, C.; Osakabe, Y.; Osakabe, K. Efficient multiplex genome editing induces precise, and self-ligated type mutations in tomato plants. Front. Plant Sci. 2018, 9, 916. [Google Scholar] [CrossRef]
- Qi, W.; Zhu, T.; Tian, Z.; Li, C.; Zhang, W.; Song, R. High-efficiency CRISPR/Cas9 multiplex gene editing using the glycine tRNA-processing system-based strategy in maize. BMC Biotechnol. 2016, 16, 58. [Google Scholar] [CrossRef]
- Wang, W.; Akhunova, A.; Chao, S.; Akhunov, E.J.B. Optimizing multiplex CRISPR/Cas9-based genome editing for wheat. BioRxiv 2016. [Google Scholar] [CrossRef]
- Ding, D.; Chen, K.; Chen, Y.; Li, H.; Xie, K. Engineering introns to express RNA guides for Cas9-and Cpf1-mediated multiplex genome editing. Mol. Plant 2018, 11, 542–552. [Google Scholar] [CrossRef]
- Nissim, L.; Perli, S.D.; Fridkin, A.; Perez-Pinera, P.; Lu, T. Multiplexed and programmable regulation of gene networks with an integrated RNA and CRISPR/Cas toolkit in human cells. Mol. Cell 2014, 54, 698–710. [Google Scholar] [CrossRef]
- Ferreira, R.; Skrekas, C.; Nielsen, J.; David, F. Multiplexed CRISPR/Cas9 genome editing and gene regulation using Csy4 in Saccharomyces cerevisiae. ACS Synth. Biol. 2017, 7, 10–15. [Google Scholar] [CrossRef]
- Qi, L. RNA Processing enables predictable programming of gene expression. Nat. Biotechnol. 2012, 30, 1002. [Google Scholar] [CrossRef] [PubMed]
- Cermak, T.; Curtin, S.J.; Gil-Humanes, J.; Čegan, R.; Kono, T.J.; Konečná, E.; Belanto, J.J.; Starker, C.G.; Mathre, J.W.; Greenstein, R. A multi-purpose toolkit to enable advanced genome engineering in plants. Plant Cell 2017, 29, 1196–1217. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Chen, Y.-L.; Wang, D.-F.; Wang, Y.-L.; Zhang, T.-P.; Li, H.; Liang, F.; Zhao, Y.; Zhang, G. SgRNA Expression of CRIPSR-Cas9 System Based on MiRNA Polycistrons as a Versatile Tool to Manipulate Multiple and Tissue-Specific Genome Editing. Sci. Rep. 2017, 7, 5795. [Google Scholar] [CrossRef] [PubMed]
- Hess, G.T.; Tycko, J.; Yao, D.; Bassik, M. Methods and applications of CRISPR-mediated base editing in eukaryotic genomes. Mol. Cell 2017, 68, 26–43. [Google Scholar] [CrossRef]
- Komor, A.C.; Kim, Y.B.; Packer, M.S.; Zuris, J.A.; Liu, D. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 2016, 533, 420. [Google Scholar] [CrossRef]
- Hess, G.T.; Frésard, L.; Han, K.; Lee, C.H.; Li, A.; Cimprich, K.A.; Montgomery, S.B.; Bassik, M. Directed evolution using dCas9-targeted somatic hypermutation in mammalian cells. Nat. Methods 2016, 13, 1036. [Google Scholar] [CrossRef]
- Ma, Y.; Zhang, J.; Yin, W.; Zhang, Z.; Song, Y.; Chang, X. Targeted AID-mediated mutagenesis (TAM) enables efficient genomic diversification in mammalian cells. Nat. Methods 2016, 13, 1029. [Google Scholar] [CrossRef]
- Lu, Y.; Zhu, J.-K. Precise editing of a target base in the rice genome using a modified CRISPR/Cas9 system. Mol. Plant 2017, 10, 523–525. [Google Scholar] [CrossRef]
- Shimatani, Z.; Kashojiya, S.; Takayama, M.; Terada, R.; Arazoe, T.; Ishii, H.; Teramura, H.; Yamamoto, T.; Komatsu, H.; Miura, K. Targeted base editing in rice and tomato using a CRISPR-Cas9 cytidine deaminase fusion. Nat. Biotechnol. 2017, 35, 441. [Google Scholar] [CrossRef]
- Zong, Y.; Wang, Y.; Li, C.; Zhang, R.; Chen, K.; Ran, Y.; Qiu, J.-L.; Wang, D.; Gao, C. Precise base editing in rice, wheat and maize with a Cas9-cytidine deaminase fusion. Nat. Biotechnol. 2017, 35, 438. [Google Scholar] [CrossRef]
- Veillet, F.; Perrot, L.; Chauvin, L.; Kermarrec, M.-P.; Guyon-Debast, A.; Chauvin, J.-E.; Nogué, F.; Mazier, M. Transgene-free genome editing in tomato and potato plants using agrobacterium-mediated delivery of a CRISPR/Cas9 cytidine base editor. Int. J. Mol. Sci. 2019, 20, 402. [Google Scholar] [CrossRef] [PubMed]
- Qin, L.; Li, J.; Wang, Q.; Xu, Z.; Sun, L.; Alariqi, M.; Manghwar, H.; Wang, G.; Li, B.; Ding, X. High Efficient and Precise Base Editing of C• G to T• A in the Allotetraploid Cotton (Gossypium hirsutum) Genome Using a Modified CRISPR/Cas9 System. Plant Biotechnol. J. 2019. [Google Scholar] [CrossRef] [PubMed]
- Tieman, D.; Zhu, G.; Resende, M.F.; Lin, T.; Nguyen, C.; Bies, D.; Rambla, J.L.; Beltran, K.S.O.; Taylor, M.; Zhang, B.J.S. A chemical genetic roadmap to improved tomato flavor. Science 2017, 355, 391–394. [Google Scholar] [CrossRef] [PubMed]
- Mao, Y.; Zhang, H.; Xu, N.; Zhang, B.; Gou, F.; Zhu, J. Application of the CRISPR–Cas system for efficient genome engineering in plants. Mol. Plant 2013, 6, 2008. [Google Scholar] [CrossRef]
- Billon, P.; Bryant, E.E.; Joseph, S.A.; Nambiar, T.S.; Hayward, S.B.; Rothstein, R.; Ciccia, A. CRISPR-mediated base editing enables efficient disruption of eukaryotic genes through induction of STOP codons. Mol. Cell 2017, 67, 1068–1079.e1064. [Google Scholar] [CrossRef]
- Kuscu, C.; Parlak, M.; Tufan, T.; Yang, J.; Szlachta, K.; Wei, X.; Mammadov, R.; Adli, M. CRISPR-STOP: Gene silencing through base-editing-induced nonsense mutations. Nat. Methods 2017, 14, 710. [Google Scholar] [CrossRef]
- Cheng, A.W.; Wang, H.; Yang, H.; Shi, L.; Katz, Y.; Theunissen, T.W.; Rangarajan, S.; Shivalila, C.S.; Dadon, D.B.; Jaenisch, R. Multiplexed activation of endogenous genes by CRISPR-on, an RNA-guided transcriptional activator system. Cell Res. 2013, 23, 1163. [Google Scholar] [CrossRef]
- Maeder, M.L.; Linder, S.J.; Cascio, V.M.; Fu, Y.; Ho, Q.H.; Joung, J. CRISPR RNA–guided activation of endogenous human genes. Nat. Methods 2013, 10, 977. [Google Scholar] [CrossRef]
- Perez-Pinera, P.; Kocak, D.D.; Vockley, C.M.; Adler, A.F.; Kabadi, A.M.; Polstein, L.R.; Thakore, P.I.; Glass, K.A.; Ousterout, D.G.; Leong, K. RNA-guided gene activation by CRISPR-Cas9–based transcription factors. Nat. Methods 2013, 10, 973. [Google Scholar] [CrossRef]
- Piatek, A.; Ali, Z.; Baazim, H.; Li, L.; Abulfaraj, A.; Al-Shareef, S.; Aouida, M.; Mahfouz, M.M. RNA-guided transcriptional regulation in planta via synthetic dC as9-based transcription factors. Plant Biotechnol. J. 2015, 13, 578–589. [Google Scholar] [CrossRef]
- Lowder, L.G.; Paul, J.W.; Baltes, N.J.; Voytas, D.F.; Zhang, Y.; Zhang, D.; Tang, X.; Zheng, X.; Hsieh, T.-F.; Qi, Y. A CRISPR/Cas9 toolbox for multiplexed plant genome editing and transcriptional regulation. Plant Physiol. 2015, 169, 971–985. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Leal, D.; Lemmon, Z.H.; Man, J.; Bartlett, M.E.; Lippman, Z.B.J.C. Engineering quantitative trait variation for crop improvement by genome editing. Cell 2017, 171, 470–480.e478. [Google Scholar]
- Lister, R.; Pelizzola, M.; Dowen, R.H.; Hawkins, R.D.; Hon, G.; Tonti-Filippini, J.; Nery, J.R.; Lee, L.; Ye, Z.; Ngo, Q. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 2009, 462, 315. [Google Scholar] [CrossRef] [PubMed]
- Lister, R.; Mukamel, E.A.; Nery, J.R.; Urich, M.; Puddifoot, C.A.; Johnson, N.D.; Lucero, J.; Huang, Y.; Dwork, A.J.; Schultz, M. Global epigenomic reconfiguration during mammalian brain development. Science 2013, 341, 1237905. [Google Scholar] [CrossRef] [PubMed]
- Law, J.A.; Jacobsen, S. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 2010, 11, 204. [Google Scholar] [CrossRef]
- Liu, X.S.; Wu, H.; Ji, X.; Stelzer, Y.; Wu, X.; Czauderna, S.; Shu, J.; Dadon, D.; Young, R.A.; Jaenisch, R. Editing DNA methylation in the mammalian genome. Cell 2016, 167, 233–247.e217. [Google Scholar] [CrossRef]
- Papikian, A.; Liu, W.; Gallego-Bartolomé, J.; Jacobsen, S. Site-specific manipulation of Arabidopsis loci using CRISPR-Cas9 SunTag systems. Nat. Commun. 2019, 10, 729. [Google Scholar] [CrossRef]
- Cradick, T.J.; Fine, E.J.; Antico, C.J.; Bao, G. CRISPR/Cas9 systems targeting β-globin and CCR5 genes have substantial off-target activity. Nucleic Acids Res. 2013, 41, 9584–9592. [Google Scholar] [CrossRef]
- Fu, Y.; Foden, J.A.; Khayter, C.; Maeder, M.L.; Reyon, D.; Joung, J.K.; Sander, J. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. 2013, 31, 822. [Google Scholar] [CrossRef]
- Hsu, P.D.; Scott, D.A.; Weinstein, J.A.; Ran, F.A.; Konermann, S.; Agarwala, V.; Li, Y.; Fine, E.J.; Wu, X.; Shalem, O. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 2013, 31, 827. [Google Scholar] [CrossRef]
- Lin, Y.; Cradick, T.J.; Brown, M.T.; Deshmukh, H.; Ranjan, P.; Sarode, N.; Wile, B.M.; Vertino, P.M.; Stewart, F.J.; Bao, G. CRISPR/Cas9 systems have off-target activity with insertions or deletions between target DNA and guide RNA sequences. Nucleic Acids Res. 2014, 42, 7473–7485. [Google Scholar] [CrossRef] [PubMed]
- Pattanayak, V.; Lin, S.; Guilinger, J.P.; Ma, E.; Doudna, J.A.; Liu, D. High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nat. Biotechnol. 2013, 31, 839. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.W.; Kim, S.; Kim, Y.; Kweon, J.; Kim, H.S.; Bae, S.; Kim, J. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res. 2014, 24, 132–141. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Sander, J.D.; Reyon, D.; Cascio, V.M.; Joung, J. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 2014, 32, 279. [Google Scholar] [CrossRef]
- Ma, Y.; Zhang, L.; Huang, X. Genome modification by CRISPR/Cas9. Nat. Methods 2014, 281, 5186–5193. [Google Scholar] [CrossRef]
- Kuscu, C.; Arslan, S.; Singh, R.; Thorpe, J.; Adli, M. Genome-wide analysis reveals characteristics of off-target sites bound by the Cas9 endonuclease. Nat. Biotechnol. 2014, 32, 677. [Google Scholar] [CrossRef]
- Kim, D.; Bae, S.; Park, J.; Kim, E.; Kim, S.; Yu, H.R.; Hwang, J.; Kim, J.-I.; Kim, J. Digenome-seq: Genome-wide profiling of CRISPR-Cas9 off-target effects in human cells. Nat. Methods 2015, 12, 237. [Google Scholar] [CrossRef]
- Tsai, S.Q.; Zheng, Z.; Nguyen, N.T.; Liebers, M.; Topkar, V.V.; Thapar, V.; Wyvekens, N.; Khayter, C.; Iafrate, A.J.; Le, L.P. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 2015, 33, 187. [Google Scholar] [CrossRef]
- Kleinstiver, B.P.; Pattanayak, V.; Prew, M.S.; Tsai, S.Q.; Nguyen, N.T.; Zheng, Z.; Joung, J. High-fidelity CRISPR–Cas9 nucleases with no detectable genome-wide off-target effects. Nature 2016, 529, 490. [Google Scholar] [CrossRef]
- Sander, J.D.; Zaback, P.; Joung, J.K.; Voytas, D.F.; Dobbs, D. Zinc Finger Targeter (ZiFiT): An engineered zinc finger/target site design tool. Nucleic Acids Res. 2007, 35, W599–W605. [Google Scholar] [CrossRef]
- Naito, Y.; Hino, K.; Bono, H.; Ui-Tei, K.J.B. CRISPRdirect: Software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics 2014, 31, 1120–1123. [Google Scholar] [CrossRef] [PubMed]
- Heigwer, F.; Kerr, G.; Boutros, M. E-CRISP: Fast CRISPR target site identification. Nat. Methods 2014, 11, 122. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.J.; Holmes, B.R.; Aronin, N.; Brodsky, M. CRISPRseek: A bioconductor package to identify target-specific guide RNAs for CRISPR-Cas9 genome-editing systems. PLoS ONE 2014, 9, e108424. [Google Scholar] [CrossRef] [PubMed]
- Gratz, S.J.; Ukken, F.P.; Rubinstein, C.D.; Thiede, G.; Donohue, L.K.; Cummings, A.M.; O’Connor-Giles, K. Highly specific and efficient CRISPR/Cas9-catalyzed homology-directed repair in Drosophila. Genetics 2014, 196, 961–971. [Google Scholar] [CrossRef]
- Montague, T.G.; Cruz, J.M.; Gagnon, J.A.; Church, G.M.; Valen, E. CHOPCHOP: A CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res. 2014, 42, W401–W407. [Google Scholar] [CrossRef]
- Prykhozhij, S.V.; Rajan, V.; Gaston, D.; Berman, J. CRISPR multitargeter: A web tool to find common and unique CRISPR single guide RNA targets in a set of similar sequences. PLoS ONE 2015, 10, e0119372. [Google Scholar]
- Xie, S.; Shen, B.; Zhang, C.; Huang, X.; Zhang, Y. sgRNAcas9: A software package for designing CRISPR sgRNA and evaluating potential off-target cleavage sites. PLoS ONE 2014, 9, e100448. [Google Scholar] [CrossRef]
- Liu, H.; Ding, Y.; Zhou, Y.; Jin, W.; Xie, K.; Chen, L. CRISPR-P 2.0: An improved CRISPR-Cas9 tool for genome editing in plants. Mol. Plant 2017, 10, 530–532. [Google Scholar] [CrossRef]
- Upadhyay, S.K.; Sharma, S. SSFinder: High throughput CRISPR-Cas target sites prediction tool. BioMed Res. Int. 2014, 2014. [Google Scholar] [CrossRef]
- O’Brien, A.; Bailey, T. GT-Scan: Identifying unique genomic targets. Bioinformatics 2014, 30, 2673–2675. [Google Scholar] [CrossRef]
- Stemmer, M.; Thumberger, T.; del Sol Keyer, M.; Wittbrodt, J.; Mateo, J. CCTop: An intuitive, flexible and reliable CRISPR/Cas9 target prediction tool. PLoS ONE 2015, 10, e0124633. [Google Scholar] [CrossRef] [PubMed]
- Bae, S.; Park, J.; Kim, J. Cas-OFFinder: A fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 2014, 30, 1473–1475. [Google Scholar] [CrossRef] [PubMed]
- Oliveros, J.C.; Franch, M.; Tabas-Madrid, D.; San-León, D.; Montoliu, L.; Cubas, P.; Pazos, F. Breaking-Cas—Interactive design of guide RNAs for CRISPR-Cas experiments for ENSEMBL genomes. Nucleic Acids Res. 2016, 44, W267–W271. [Google Scholar] [CrossRef] [PubMed]
- Altpeter, F.; Springer, N.M.; Bartley, L.E.; Blechl, A.E.; Brutnell, T.P.; Citovsky, V.; Conrad, L.J.; Gelvin, S.B.; Jackson, D.P.; Kausch, A.P. Advancing Crop Transformation in the Era of Genome EditingOPEN. Plant Cell 2016, 28, 1510–1520. [Google Scholar]
- Baltes, N.J.; Gil-Humanes, J.; Cermak, T.; Atkins, P.A.; Voytas, D.F. DNA replicons for plant genome engineering. Plant Cell 2014, 26, 151–163. [Google Scholar] [CrossRef]
- Zuniga-Soto, E.; Mullins, E.; Dedicova, B. Ensifer-mediated transformation: An efficient non-Agrobacterium protocol for the genetic modification of rice. Springerplus 2015, 4, 600. [Google Scholar] [CrossRef]
- Rathore, D.S.; Doohan, F.; Mullins, E.T.; Culture, O. Capability of the plant-associated bacterium, Ensifer adhaerens strain OV14, to genetically transform its original host Brassica napus. Plant Celltissue Organ Cult. 2016, 127, 85–94. [Google Scholar] [CrossRef]
- Wendt, T.; Doohan, F.; Mullins, E. Production of Phytophthora infestans-resistant potato (Solanum tuberosum) utilising Ensifer adhaerens OV14. Transgenic Res. 2012, 21, 567–578. [Google Scholar] [CrossRef]
- Broothaerts, W.; Mitchell, H.J.; Weir, B.; Kaines, S.; Smith, L.M.; Yang, W.; Mayer, J.E.; Roa-Rodriguez, C.; Jefferson, R. Gene transfer to plants by diverse species of bacteria. Nature 2005, 433, 629. [Google Scholar] [CrossRef]
- Lacroix, B.; Citovsky, V. A functional bacterium-to-plant DNA transfer machinery of Rhizobium etli. PLoS Pathog. 2016, 12, e1005502. [Google Scholar] [CrossRef]
- Ali, Z.; Abul-Faraj, A.; Li, L.; Ghosh, N.; Piatek, M.; Mahjoub, A.; Aouida, M.; Piatek, A.; Baltes, N.J.; Voytas, D.F. Efficient virus-mediated genome editing in plants using the CRISPR/Cas9 system. Mol. Plant 2015, 8, 1288–1291. [Google Scholar] [CrossRef] [PubMed]
- Lowe, K.; Wu, E.; Wang, N.; Hoerster, G.; Hastings, C.; Cho, M.-J.; Scelonge, C.; Lenderts, B.; Chamberlin, M.; Cushatt, J. Morphogenic regulators Baby boom and Wuschel improve monocot transformation. Plant Cell 2016, 28, 1998–2015. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Meng, Z.; Wang, Y.; Chen, W.; Sun, C.; Cui, B.; Cui, J.; Yu, M.; Zeng, Z.; Guo, S. Pollen magnetofection for genetic modification with magnetic nanoparticles as gene carriers. Nat. Plants 2017, 3, 956. [Google Scholar] [CrossRef] [PubMed]
- Tsuboyama, S.; Nonaka, S.; Ezura, H.; Kodama, Y. Improved G-AgarTrap: A highly efficient transformation method for intact gemmalings of the liverwort Marchantia polymorpha. Sci. Rep. 2018, 8, 10800. [Google Scholar] [CrossRef] [PubMed]
- Torney, F.; Trewyn, B.G.; Lin, V.S.-Y.; Wang, K. Mesoporous silica nanoparticles deliver DNA and chemicals into plants. Nat. Nanotechnol. 2007, 2, 295. [Google Scholar] [CrossRef] [PubMed]
- Martin-Ortigosa, S.; Valenstein, J.S.; Lin, V.S.Y.; Trewyn, B.G.; Wang, K. Gold functionalized mesoporous silica nanoparticle mediated protein and DNA codelivery to plant cells via the biolistic method. Adv. Funct. Mater. 2012, 22, 3576–3582. [Google Scholar] [CrossRef]
- McKnight, T.E.; Melechko, A.V.; Hensley, D.K.; Mann, D.G.; Griffin, G.D.; Simpson, M. Tracking gene expression after DNA delivery using spatially indexed nanofiber arrays. Nano Lett. 2004, 4, 1213–1219. [Google Scholar] [CrossRef]
- Liu, J.; Wang, F.-H.; Wang, L.-L.; Xiao, S.-Y.; Tong, C.-Y.; Tang, D.-Y.; Liu, X. Preparation of fluorescence starch-nanoparticle and its application as plant transgenic vehicle. J. Cent. South Univ. Technol. 2008, 15, 768–773. [Google Scholar] [CrossRef]
- Gao, X.; Chen, J.; Dai, X.; Zhang, D.; Zhao, Y. An effective strategy for reliably isolating heritable and Cas9-free Arabidopsis mutants generated by CRISPR/Cas9-mediated genome editing. Plant physiology 2016, 171, 1794–1800. [Google Scholar] [CrossRef]
- Aliaga-Franco, N.; Cunjin, Z.; Presa, S.; Srivastava, A.K.; Granell, A.; Alabadi, D.; Sadanandom, A.; Blazquez, M.A.; Minguet, E.G. Identification of transgene-free CRISPR edited plants of rice, tomato and Arabidopsis by monitoring DsRED fluorescence in dry seeds. Front. Plant Sci. 2019, 10, 1150. [Google Scholar] [CrossRef]
- Okada, A.; Arndell, T.; Borisjuk, N.; Sharma, N.; Watson-Haigh, N.S.; Tucker, E.J.; Baumann, U.; Langridge, P.; Whitford, R. CRISPR/Cas9-mediated knockout of Ms1 enables the rapid generation of male-sterile hexaploid wheat lines for use in hybrid seed production. Plant Biotechnol. J. 2019, 17, 1905–1913. [Google Scholar] [CrossRef] [PubMed]
- Zuo, J.; Niu, Q.-W.; Ikeda, Y.; Chua, N. Marker-free transformation: Increasing transformation frequency by the use of regeneration-promoting genes. Curr. Opin. Biotechnol. 2002, 13, 173–180. [Google Scholar] [CrossRef]
- Liang, Z.; Chen, K.; Li, T.; Zhang, Y.; Wang, Y.; Zhao, Q.; Liu, J.; Zhang, H.; Liu, C.; Ran, Y. Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes. Nat. Commun. 2017, 8, 14261. [Google Scholar] [CrossRef] [PubMed]
- Subburaj, S.; Chung, S.J.; Lee, C.; Ryu, S.-M.; Kim, D.H.; Kim, J.-S.; Bae, S.; Lee, G.-J. Site-directed mutagenesis in Petunia× hybrida protoplast system using direct delivery of purified recombinant Cas9 ribonucleoproteins. Plant Cell Rep. 2016, 35, 1535–1544. [Google Scholar] [CrossRef]
- Woo, J.W.; Kim, J.; Kwon, S.I.; Corvalán, C.; Cho, S.W.; Kim, H.; Kim, S.-G.; Kim, S.-T.; Choe, S.; Kim, J.-S. DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat. Biotechnol. 2015, 33, 1162. [Google Scholar] [CrossRef] [PubMed]
- Svitashev, S.; Schwartz, C.; Lenderts, B.; Young, J.K.; Cigan, A.M. Genome editing in maize directed by CRISPR–Cas9 ribonucleoprotein complexes. Nat. Commun. 2016, 7, 13274. [Google Scholar] [CrossRef] [PubMed]
- Feng, Z.; Mao, Y.; Xu, N.; Zhang, B.; Wei, P.; Yang, D.-L.; Wang, Z.; Zhang, Z.; Zheng, R.; Yang, L. Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in Arabidopsis. Proc. Natl. Acad. Sci. USA 2014, 111, 4632–4637. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.-P.; Xing, H.-L.; Dong, L.; Zhang, H.-Y.; Han, C.-Y.; Wang, X.-C.; Chen, Q. Egg cell-specific promoter-controlled CRISPR/Cas9 efficiently generates homozygous mutants for multiple target genes in Arabidopsis in a single generation. Genome Biol. 2015, 16, 144. [Google Scholar] [CrossRef]
- Mao, Y.; Zhang, Z.; Feng, Z.; Wei, P.; Zhang, H.; Botella, J.R.; Zhu, J.K. Development of germ-line-specific CRISPR-Cas9 systems to improve the production of heritable gene modifications in Arabidopsis. Plant Biotechnol. J. 2016, 14, 519–532. [Google Scholar] [CrossRef]
- Xu, P.; Su, H.; Chen, W.; Lu, P. The application of a meiocyte-specific CRISPR/Cas9 (MSC) system and a suicide-MSC system in generating inheritable and stable mutations in Arabidopsis. Front. Plant Sci. 2018, 9, 1007. [Google Scholar] [CrossRef]
- Hyun, Y.; Kim, J.; Cho, S.W.; Choi, Y.; Kim, J.-S.; Coupland, G.J.P. Site-directed mutagenesis in Arabidopsis thaliana using dividing tissue-targeted RGEN of the CRISPR/Cas system to generate heritable null alleles. Planta 2015, 241, 271–284. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Wei, S.; Wu, Y.; Hu, R.; Li, H.; Yang, W.; Xie, Q. High-efficiency genome editing in Arabidopsis using YAO promoter-driven CRISPR/Cas9 system. Mol. Plant 2015, 8, 1820–1823. [Google Scholar] [CrossRef] [PubMed]

| Mechanism of Action. | Plant Species | Target | Protein | Type of Mutation | Promoter | Vector | Reference |
|---|---|---|---|---|---|---|---|
| Knockout | Apple | PDS | pcoCas9 fused to GFBSD2 | InDels | NA | pEgP226-2A-gfbsd2 | [39] |
| Knockout | Arabidopsis | BRI1, GAI, JAZ1 | hSpCas9 | InDels | NA | NA | [40] |
| Knockout | Banana | PDS | Cas9 | InDels resulting in early stop codon | NA | pRGEB31 | [41] |
| Knockout | Rice | OsROC5, OsDEP1 | Arabidopsis codon-optimized Cas9 | Small deletions of up to 10 base pairs. | NA | pZHY988 | [42] |
| Knock-in | Rice | OsPDS | SpCas9 | Successful insertion of Oligo with KpnI+EcoRI sites resulting in gene disruption. | ZmUbi | pEASY-Blunt vector | [36] |
| Knock-in | Rice | Chlorophyllide-a oxygenase gene of rice (CAO1) | Monocot optimized FnCpf1 | Heritable targeted insertion of repair template having hygromcin resistance. | ZmUbi | pUC19 backbone | [43] |
| Knock-in | Rice | ALS | pcoLbCpf1 | Targeted ALS gene replacement resulting in Herbicide resistant plants. | NA | pCXUN-LbCpf1 | [44] |
| Knock-in | Arabidopsis | GLABRA2 (GL2); ROS1; DME | hSpCas9 | GFP-DME; DME-GFP; ROS1-GFP; ROS1-luc fusions were generated | AtU6 | pCambia1300; pCambia3301 | [45] |
| Knock-in | Maize | ALS2; LIG | SpCas9 | Targeted insertion of MoPAT gene in LIG locus by upto 83% and mutation of ALS2 with by HDR with two oligos. | Ubi | pUC19; pSB11 | [46] |
| Tool | Specialization | Specific Feature | Restriction Site Compatibility | Off-Target Analysis | Website | Reference |
|---|---|---|---|---|---|---|
| ZiFiT | Animal |
| Absent | Present | http://zifit.partners.org/ZiFiT/ | [165] |
| CRISPR Direct | Animal and plant |
| Present | Present | http://crispr.dbcls.jp/ | [166] |
| E-CRISP website | Animal, bacteria, fungi, and plant |
| Absent | Present | http://www.e-crisp.org/E-CRISP/ | [167] |
| CRISPRSeek | Animal, fungi, bacteria, and plant included in Bio String based genome data packages. |
| Present | Present | https://bioconductor.org/packages/release/bioc/html/CRISPRseek.html | [168] |
| flyCRISPR Optimal Target Finder | Animal; invertebrates |
| Absent | Present. | http://targetfinder.flycrispr.neuro.brown.edu/ | [169] |
| CHOPCHOP | Animal and plant |
| Absent | Present | http://chopchop.cbu.uib.no/ | [170] |
| CRISPR-Multitarget | Animals and plants |
| Absent. | CRISPR-Multitargeter gives links to GT-scan and Cas-OFFinder to perform off target analysis. | http://multicrispr.net/ | [171] |
| sgRNAcas9 | All organisms |
| -NA- | Present | https://sourceforge.net/projects/sgrnacas9/;www.biootools.com | [172] |
| CRISPR-P | Plants |
| Present | Present | http://cbi.hzau.edu.cn/cgi-bin/CRISPR | [173] |
| SSFinder | -NA- |
| -NA- | -NA- | https://omictools.com/ssfinder-tool | [174] |
| GT-Scan | Animals and plants |
| Absent | Present | http://gt-scan.braembl.org.au/gt-scan/ | [175] |
| CRISPR gRNA Design Tool (DNA2.0 design tool) | Animals and plants (Arabidopsis) |
| Absent | Present | https://www.atum.bio/eCommerce/cas9/input | |
| CCTop | Animals and Plants |
| Absent | Present | https://crispr.cos.uni-heidelberg.de/ | [176] |
| Cas-OFFinder | Animals, plants and others (fungi, bacteria, virus) |
| -NA- | Present | http://www.rgenome.net/cas-offinder/ | [177] |
| Breaking-Cas | All eukaryotic genomes present in ENSEMBL. (protists, fungi, bacteria, plants and animals) |
| Absent | Present | http://bioinfogp.cnb.csic.es/tools/breakingcas | [178] |
| S.No. | Specificity | Promoter | Reference |
|---|---|---|---|
| 1. | Egg cell specific | EC1.2 promoter | [203] |
| 2. | Germ-line-specific | SPOROCYTELESS | [204] |
| 3. | Meiocyte-specific promoter | AtDMC1 | [205] |
| 4. | Pollen-specific promoter | Lat52 | [204] |
| 5. | Egg cell- and early embryo-specific promoter | DD45 | [204] |
| 6. | Dividing tissue specific | INCURVATA2 | [206] |
| 7. | Cell-division specific | YAO | [207] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vats, S.; Kumawat, S.; Kumar, V.; Patil, G.B.; Joshi, T.; Sonah, H.; Sharma, T.R.; Deshmukh, R. Genome Editing in Plants: Exploration of Technological Advancements and Challenges. Cells 2019, 8, 1386. https://doi.org/10.3390/cells8111386
Vats S, Kumawat S, Kumar V, Patil GB, Joshi T, Sonah H, Sharma TR, Deshmukh R. Genome Editing in Plants: Exploration of Technological Advancements and Challenges. Cells. 2019; 8(11):1386. https://doi.org/10.3390/cells8111386
Chicago/Turabian StyleVats, Sanskriti, Surbhi Kumawat, Virender Kumar, Gunvant B. Patil, Trupti Joshi, Humira Sonah, Tilak Raj Sharma, and Rupesh Deshmukh. 2019. "Genome Editing in Plants: Exploration of Technological Advancements and Challenges" Cells 8, no. 11: 1386. https://doi.org/10.3390/cells8111386
APA StyleVats, S., Kumawat, S., Kumar, V., Patil, G. B., Joshi, T., Sonah, H., Sharma, T. R., & Deshmukh, R. (2019). Genome Editing in Plants: Exploration of Technological Advancements and Challenges. Cells, 8(11), 1386. https://doi.org/10.3390/cells8111386







