1. Introduction
Small GTPases have long been identified to act as upstream regulators for the actin cytoskeleton [
1]. Rac1 promotes lamellipodia formation by activating the Arp2/3 complex and by inducing branched actin formation [
2], while Cdc42 is well known to induce filopodial protrusion and invadopodium formation by formin Dia2 and FMNL2 [
3,
4]. The cooperation of Rac1 and Cdc42 is essential in generating cell polarity during cell migration [
5]. RhoA interacts with downstream effectors to influence actin stability and to generate actomyosin contraction [
6]. The binding of mDia1 to active RhoA (GTP-RhoA) promotes linear actin formation [
7,
8]. A ring of RhoA has been observed to localize to the site of the nascent cytokinetic furrow in early cytokinesis and facilitates the formation of cytokinetic furrow [
9]. Active RhoA enhances contractile ring assembly through its downstream effectors including the Rho kinase (ROCK) [
10]. ROCK phosphorylates and activates LIM kinase, which in turn inhibits the actin-severing protein cofilin, resulting in stabilization of the actin network [
11].
RhoA is activated by guanine–nucleotide exchange factors (GEFs) and inactivated by GTPase activating proteins (GAPs) [
12]. p190RhoGAP is a major negative regulator of Rho GTPases and is expressed in cells as two isoforms that share a common GTP-binding domain and that controls RhoA activity: p190ARhoGAP (also known as ARHGAP35) and p190BRhoGAP (also known as ARHGAP5) [
13,
14]. p190ARhoGAP contains a protrusion localization sequence (PLS), which is necessary for p190ARhoGAP targeting to the leading edge [
15]. PLS deletion or mutation (S866F and delta 865–870) dramatically enhances p190ARhoGAP activity toward RhoA [
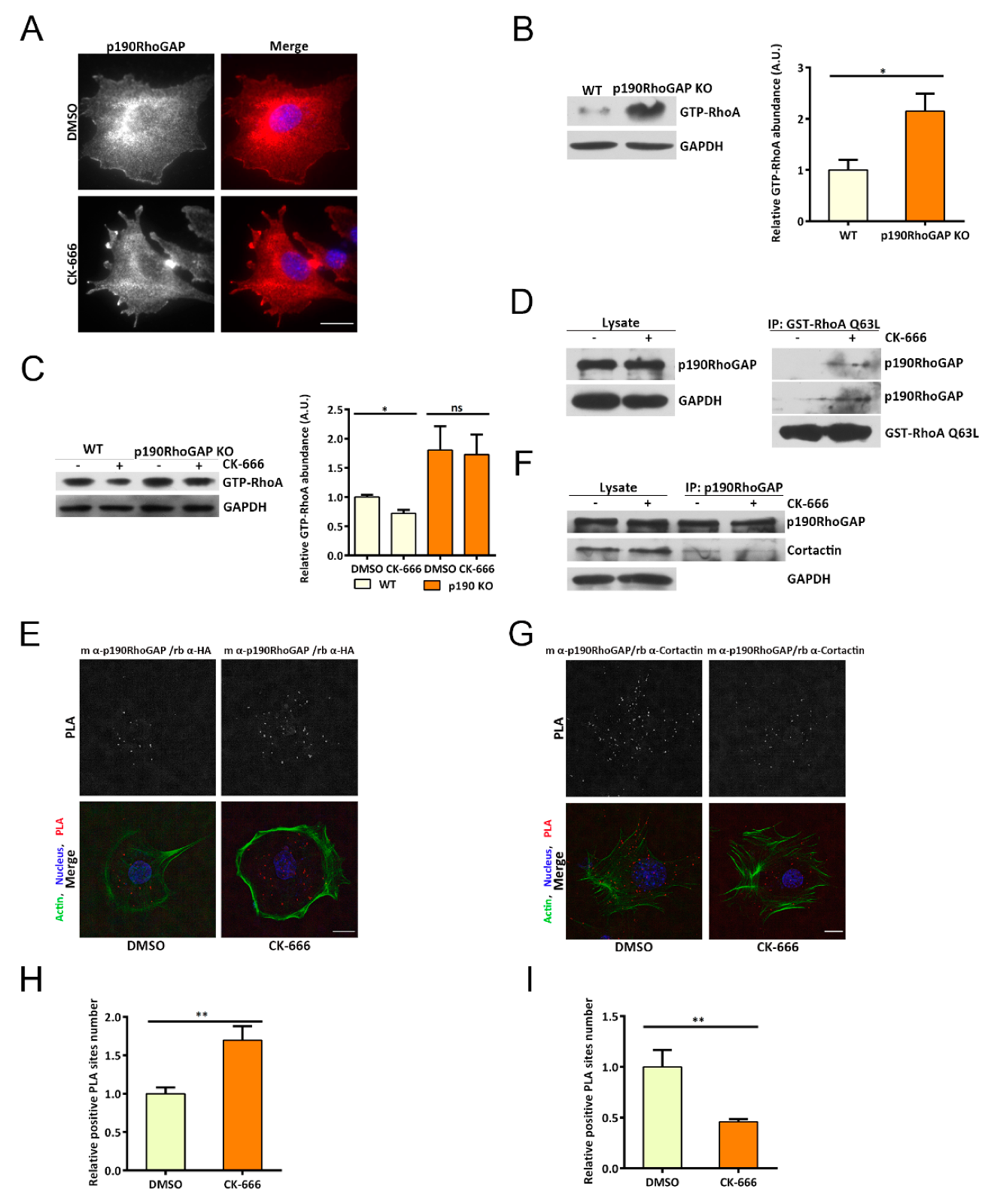
14]. The cortical actin-binding protein cortactin, which is a weak nucleator of the Arp2/3 complex, acts as a scaffold for p190ARhoGAP and is also required for leading edge localization of p190ARhoGAP [
15]. The binding of cortactin to PLS of p190ARhoGAP results in a closed and inactivated conformation of p190ARhoGAP, while its dissociation leads to opening of p190ARhoGAP and locally turns off RhoA [
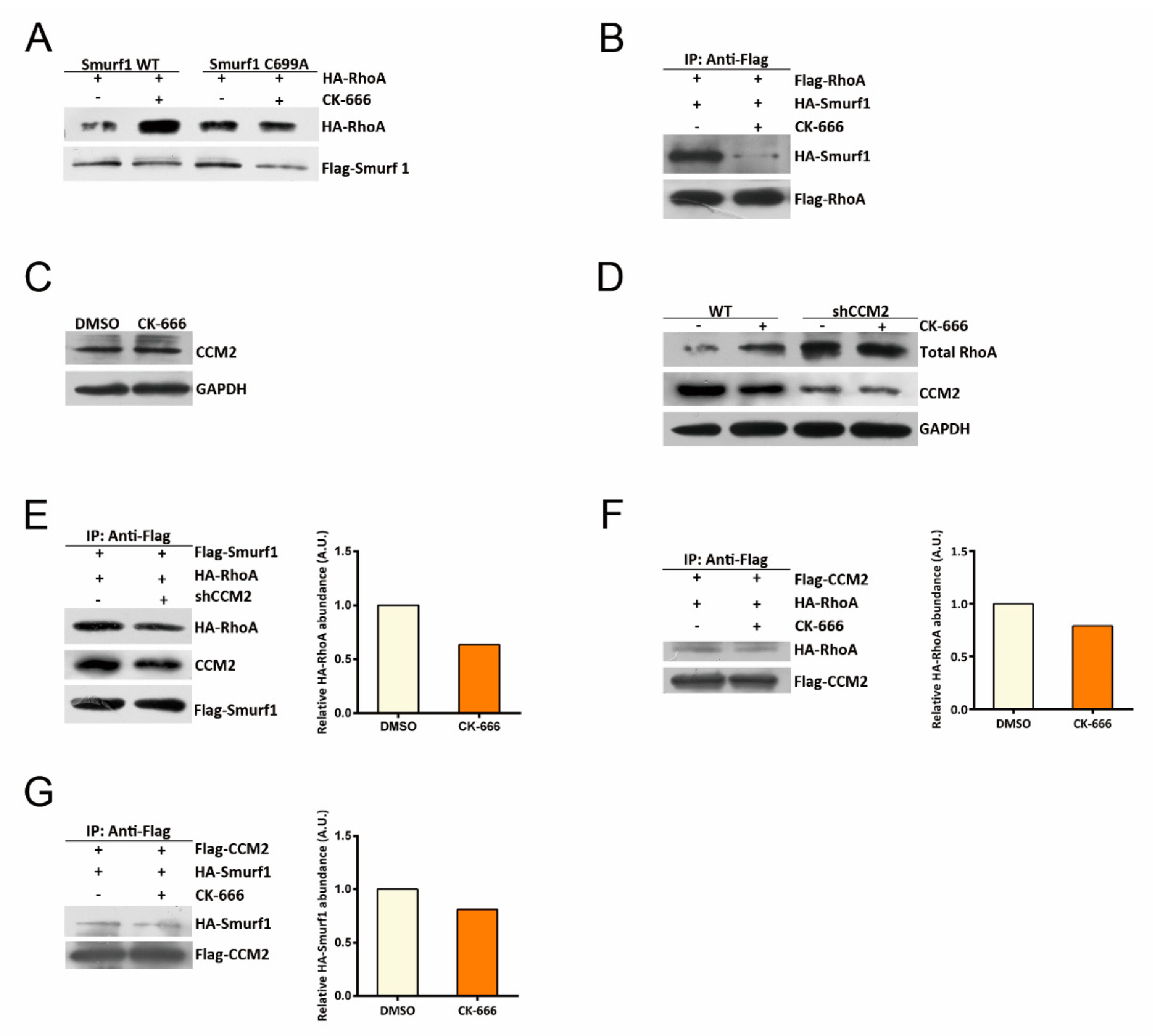
14]. Aside from the activity, protein abundance also affects cellular functions of small GTPases. The proteasomal degradation is the main pathway for the degradation of RhoA [
16]. Cullin3, SCF
FBXL19, and Smurf1 have been reported to be E3 ubiquitin ligases critical for proteasome-dependent RhoA degradation [
16,
17]. ERK2 is required for SCF
FBXL19-mediated RhoA ubiquitination [
16], while Cerebral Cavernous Malformation 2 (CCM2) has been demonstrated to act as an important scaffolding protein in facilitating Smurf1-mediated RhoA degradation [
18,
19].
Cross talks between different small GTPases has been observed. Rac1 has been reported to bind and activate p190RhoGAP, resulting in RhoA inhibition [
20]. Conversely, RhoA inhibits Rac1 activity by activating the Rac1 GAP FilGAP through ROCK [
21]. However, surprisingly scarce knowledge in the feedback regulation from the cytoskeleton to the small GTPases has been obtained [
22], which includes positive regulation of RacGEFs by F-actin through recruitment of PI3K to the plasma membrane [
23]. Precise spatiotemporal control of cell morphology and motility rely on feedback regulations, and such mechanisms involving downstream cytoskeleton in affecting upstream signaling factors have been largely understudied. Here, we report an intriguing finding that disruption of the Arp2/3-branched actin specifically reduces RhoA activity through altered cortactin/p190RhoGAP interaction but upregulates RhoA abundance by affecting CCM2/Smurf1. Moreover, we found that cytokinesis defects induced by Arp2/3 inhibition can be rescued by activating RhoA, demonstrating the physiological significance of this cytoskeleton–small GTPase signaling feedback loop. Our work sheds new lights on branched actin, highlighting its role in signal transduction aside from serving as critical cytoskeletal components to maintain cell morphology and motility.
2. Materials and Methods
2.1. Cell Lines and Cell Culture
HEK293T, Mouse embryonic fibroblasts (MEFs), and Hela cells were kept by our laboratory. All the cells were passed no more than 10 times and were mycoplasma free. Cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM) media supplemented with 10% fetal bovine serum (FBS) (PAN, Biotech, P30-3031, Aidenbach, Germany), 100 U/mL penicillin, and 100 µg/mL streptomycin at 37 °C with 5% CO2.
2.2. p190RhoGAP Knockout and CCM2 and ARHGAP29 Knockdown Cell-Line Generation
The following short hairpin RNA (shRNA) and primers were used to generate CCM2 knockdown HEK293T:
Human shRNA forward shRNA-1, 5′-GCCCAGGTCCTCTACTGTG-3′.
Human shRNA forward shRNA-2, 5′-GCTGAGCGACTATATTGAG-3′.
The following shRNA and primers were used to generate ARHGAP29 knockdown Hela:
Human shRNA forward shRNA-1, 5′-GCATCAGGTCAACTCTCTACT-3′.
Human shRNA forward shRNA-2, 5′-GGGCTCAAGTCCTTAAGTTCC-3′.
The following small guide RNA (sgRNA) and primers were used to generate p190RhoGAP knockout Hela:
Human sgRNA forward sgRNA-1, 5′-CACCGATTGAGTACATTGAAGCCAC-3′.
Human sgRNA forward sgRNA-2, 5′-CACCGTAACAACTGTCGTGACTCCA-3′.
Human sgRNA forward sgRNA-3, 5′-CTTCTTGACATTCTTTCTAGCAGTT-3′.
A single cell clone from lentivirus-infected pool cells was selected and verified by western blotting and DNA sequencing.
2.3. Western Blot
For western blotting, cells were washed with dulbecco’s phosphate buffered saline (DPBS) once and lysed in an appropriate volume of radio immunoprecipitation assay (RIPA) buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1% Triton X-100, 0.5% Na-deoxycholate, 0.1% SDS, 1 mM EDTA, and protease inhibitor cocktail) for 10 min on ice. Lysates were centrifuged at 12,000 rpm for 10 min, and the supernatants were collected. Five times SDS loading buffer was added to the supernatants and boiled for 10 min at 95 °C. Protein samples were run on 10% SDS PAGE acrylamide gels [H
2O, 30% acrylamide, 1.5 M Tris-HCl pH 8.8, 10% ammonium persulphate, and tetramethylethylenediamine (TEMED)] and transferred onto nitrocellulose membranes by wet electrophoretic transfer, followed by 10% nonfat-milk blocking at 4 °C overnight or at room temperature for 1 h. The first and second antibody incubations were at 4 °C overnight or at room temperature for 2 h. The following antibodies were used: anti-GAPDH (Abcam ab181602, 1:5000, Cambridge, MA, USA), anti-RhoA (Cytoskeleton ARH04, 1:500, Denver, CO, USA), anti-p190RhoGAP (BD Transduction Laboratories 610150, 1:1000, San Jose, CA, USA), anti-HA tag (M180-3, 1:1000), anti-Rac1 (Cytoskeleton ARC03, 1:500, Denver, CO, USA), anti-Cdc42 (Cytoskeleton ACD03, 1:500, Denver, CO, USA), anti-YAP (Cell Signaling Technology 14074s, 1:500, Danvers, MA, USA), anti-ARHGAP29 (Santa Cruz Biotechnology sc-377022, 1:1000), anti-CCM2 (Sigma SAB1400724, 1:500, Saint Louis, MO, USA), anti-pMLC (Cell Signaling Technology 3674s, 1:1000, Danvers, MA, USA), anti-lamin B (proteintech 66095-I-Ig, 1:2000, Rosemont, IL, USA), anti-HA tag (MBL 561, 1:1000, Woburn, MA, USA), anti-DDDDK-tag (MBL M185-3L, 1:1000, Woburn, MA, USA), anti-mouse (sc-2005, 1:5000), and anti-rabbit (sc-2004, 1:5000) Horseradish Peroxidase (HRP) -conjugated secondary antibodies from Santa Cruz. Immobilon Western chemilum HRP substrate (WBKLS0050, Millipore, Burlington, MA, USA) was used. Solution A and solution B were mixed at the ratio of 1:1; then, the mix was added onto nitrocellulose filter membrane. X-ray films were used to detect and record the band intensity. The fixed X-ray films were then scanned. The images were quantified by the Image J software (
https://imagej.nih.gov/ij/, image J→Analyze→Gels). The intensity was normalized to GAPDH.
2.4. Drug Treatment
The Arp2/3 complex inhibitor CK-666 (Sigma-Aldrich, 182515, Saint Louis, MO, USA) was used at the concentration of 100 μM for 5–6 h. SMIFH2 (Sigma-Aldrich, S4826, Saint Louis, MO, USA) was used to inhibit formin-induced linear actin at the concentration of 15 μM for 5–6 h. Rho activator II (Cytoskeleton CN03, Denver, CO, USA) was used at the concentration of 2 μg/mL or 4 μg/mL for 5 h. The proteasome inhibitor MG-132 (Millipore, 474790-20MG, Burlington, MA, USA) was used at the concentration of 20 μM. Cycloheximide (Sigma-Aldrich, C7698, Saint Louis, MO, USA) was used at the concentration of 10 μg/mL.
2.5. Active RhoA/Rac1/Cdc42 Pulldown Assay
Cells were lysed in RIPA buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1% Triton X-100, 0.5% Na-deoxycholate, 0.1% SDS, 1 mM EDTA, and protease inhibitor cocktail) for 10 min on ice. Lysates were centrifuged at 12,000 rpm for 10 min, and the supernatants were collected. Twenty μg Rhotekin-RBD Protein GST Beads (Cytoskeleton RT02, Denver, CO, USA) or PAK-GST Protein Beads (Cytoskeleton PAK02, Denver, CO, USA) were washed three times using RIPA buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1% Triton X-100, 0.5% Na-deoxycholate, 0.1% SDS, 1 mM EDTA, and protease inhibitor cocktail). Cell lysates (800 μg) were incubated with the beads at 4 °C for 1 h. After incubation, the beads were washed three times using wash buffer (25 mM Tris pH 7.5, 30 mM MgCl2, and 40 mM NaCl). Two times loading buffer (100 mM Tris-HCl, pH 6.8, 200 mM dithiothreitol (DTT), 4% SDS, 0.2% bromophenol blue, and 20% glycerol) were added to suspend the beads and boiled for 10 min at 95 °C. Active RhoA, active Rac1, or active Cdc42 was then detected by western blot by using anti-RhoA (Cytoskeleton ARH04, Denver, CO, USA), anti-Rac1 (Cytoskeleton ARC03, Denver, CO, USA), or anti-Cdc42 (Cytoskeleton ACD03, Denver, CO, USA).
2.6. Active p190RhoGAP Pulldown Assay
Fifty μL GST resins (Senhui Microsphere, SH-12-1605-13, Suzhou, China) were washed three times using binding buffer [10 mM N-2-hydroxyethylpiperazine-N-ethane-sulphonicacid (HEPES), pH 7.6, 60 mM NaCl, 1 mM MgCl2, 10 mM DTT, 10% glycerol, and 0.01% tween-20], 8 μM RhoA Q63L, and 50 μL GST resins in 200 μL binding buffer were incubated at 4 °C for 1 h. Cells were lysed in lysis buffer [150 mM NaCl, 20 mM HEPES, pH 7.6, 10 mM MgCl2, 1% Triton X-100, 1 mM Phenylmethylsulfonyl fluoride (PMSF), and 10 μg/mL aprotinin and leupeptin]. Cell lysates were incubated with 50 μg/mL glutathione Sepharose-bound GST-RhoAQ63L for 1 h at 4 °C. The beads were washed three times with lysis buffer and then boiled at 95 °C for 5 min. Anti-p190RhoGAP (BD Transduction Laboratories 610150, 1:1000, San Jose, CA, USA) was used to test active p190RhoGAP by western blot.
2.7. Proximity Ligation Assay (PLA)
To detect the interaction between p190RhoGAP with cortactin in vivo, the proximity ligation assay was performed with Duolink kits from Sigma-Aldrich (DUO92013). Cells were treated with CK-666 (Sigma-Aldrich, 182515, Saint Louis, MO, USA), followed by fixation with 4% paraformaldehyde at room temperature for 10 min. Then, the cells were permeabilized with 0.5% Triton X-100 in phosphate buffer saline (PBS) for 5 min, followed by 3 times wash with PBS. Commercial blocking solution (Sigma-Aldrich, DUO92013, Saint Louis, MO, USA) was added to the samples to incubate for 1 h at room temperature. Mouse anti-p190RhoGAP (1:25) was mixed with rabbit anti-Cortactin antibodies and diluted (1:200) in the antibody diluent. After removing the blocking solution, the diluted antibodies were incubated with cells for 1 h at room temperature followed by 3 × 5-min wash in PBS. The PLUS and MINUS PLA probes were mixed and diluted (1:5) in antibody diluent and incubated with samples for 1 h at 37 °C. Then, the samples were washed in 1× wash buffer A for 2 × 5 min. The ligase was diluted (1:40) in diluted ligation buffer (1:5 in H2O), and the mixture was incubated with samples for 1 h at 37 °C. After washing the samples in 1× wash buffer A for 2 × 2 min, the polymerase was diluted (1:80) in diluted amplification stock (1:5 in H2O), and the mix was incubated with samples for 100 min at 37 °C. The samples were then washed in 1× wash buffer B for 2 × 10 min followed by another wash in 0.01× wash buffer B for 1 min. Finally, the samples were mounted with Prolong Diamond Antifade with 4′,6-diamidino-2-phenylindole (DAPI) (Life Technologies, P36962, Carlsbad, CA, USA) for 30 min at room temperature.
2.8. Immunofluorescence and Imaging Analysis
Cells were plated on acid-washed coverslips overnight. Cells were then fixed with 4% paraformaldehyde (PFA) at room temperature for 15 min, permeabilized in 0.5% Triton X-100 in PBS for 10 min, washed with PBS once for 5 min, and blocked with 5% bovine serum albumin (BSA) for 1 h. For p-MLC (Cell Signaling Technology 3674s, 1:200, Danvers, MA, USA) and p190RhoGAP (1:100) staining, antibodies were diluted 1:200 in 1% BSA (Sigma, Saint Louis, MO, USA) and incubated for 1 h at room temperature. After 3 times washing with PBS, the coverslips were incubated with Alexa Fluor 488 secondary antibody (A-21202, Life Technologies, Carlsbad, CA, USA) and Acti-stain 488 phalloidin (PHDR1-A, Cytoskeleton, Denver, CO, USA) or Rhodamine Phalloidin (PHDR1, Cytoskeleton, Denver, CO, USA) for 0.5 h at room temperature. After another 3 times PBS washing, the coverslips were mounted with Prolong Diamond Antifade with DAPI. Images were captured using a spinning disk microscope (Dragonfly, Andor, Belfast, UK).
2.9. Co-IP Assay
HEK293T was plated on 10-cm dishes and transfected with different plasmids. Cells were treated with DMSO or CK-666 48 h before being washed with PBS and lysed with pierce lysis buffer (25 mM Tris-HCl pH 7.4, 150 mM NaCl, 1% NP-40, 1 mM EDTA, and 5% glycerol). Added into the cell lysate was 0.2–2 μg of anti-DDDDK-tag (MBL M185-3L, Woburn, MA, USA) antibody for 4 °C overnight incubation, and 20 μL protein A + protein G agarose (Beyotime, P2012, Shanghai, China) were added to the lysate and incubated for 3 h at 4 °C. Agarose beads were washed with pierce lysis buffer for 3 times and centrifuged at 2500 rpm for 5 min, and 30–40 μL 2× SDS-PAGE loading buffer was added into the beads and boiled for 5 min. Anti-HA tag (MBL 561, 1: 1000, Woburn, MA, USA) and anti-DDDDK-tag (MBL M185-3L, 1:1000, Woburn, MA, USA) were used for detection by western blot.
2.10. Live-Cell Imaging
RhoA biosensor live-cell imaging was acquired with an oil objective lens (NA 1.4, Olympus, Tokyo, Japan) on a total internal reflection fluorescent (TIRF) microscope (CellTIRF, Olympus, Tokyo, Japan). Cells were plated on glass-bottom cell-culture dish before imaging. For RhoA biosensor imaging, the interval was 10 min. Cells were maintained in DMEM media supplemented with 10% FBS (PAN, Biotech, Aidenbach, Germany), 100 U/mL penicillin, and 100 µg/mL streptomycin at 37 °C throughout the imaging process. Images were acquired for 6 h. For cytokinesis live-cell imaging, MEFs were plated onto 30-mm cell-culture dish (Corning, Corning, NY, USA), and images were acquired using a 10× dry objective lens (NA 0.3, Olympus, Tokyo, Japan) on a deconvolution microscope (CellTIRF). Single-plane multipoint acquisitions were captured every 5 min within a 37 °C chamber.
2.11. Cell Contractility Assay
Collagen was added into gel mix (10× DPBS, 0.23% 1N NaOH, and H2O) to generate the 2% collagen gel. MEFs were collected in a tube and centrifuged at 1000 rpm for 5 min at 4 °C. The cells were resuspended with 2% collagen gel at the density of 1 × 106/mL. The cell mix were seeded into a 48 well and incubated in a cell incubator for 30 min. Appropriate volume medium was added into the wells. The gels were isolated from the well and treated with DMSO/CK-666 for 5 h. The gels were imaged using a camera (Nikon, Tokyo, Japan), and the gel area was calculated by image J.
2.12. Stable Cell-Line Generation
Three plasmid-packing systems were used for lentivirus packing. The three plasmids are pLVX-EGFP-N1 inserted with the target genes, ps-PAX2, and pCMV-VSV-G. The packing cell line is HEK293T, and the lentivirus was harvested 48 h after plasmid transfection. Fresh lentivirus-containing media were used to infect Hela cells 3 times until we can see the expression of EGFP. Positive cells were selected by puromycin (Sigma-Aldrich, P8833, Saint Louis, MO, USA) at 2 μg/mL for 1 week.
2.13. Statistical Analysis
Statistical differences between two groups of data were analyzed with a two-tailed unpaired Student’s t test. All graphical data are represented as a mean with data bars representing SEM.
2.14. Polyacrylamide (PA) Gel Preparation and Functionalization
PA gel preparation and functionalization have been adapted from Knoll S.G. et al. [
24]. Specially, 40% acrylamide and 2% bis-acrylamide solutions were combined at 8.8% and 0.1%, respectively, to make a 13,800 Pa gel. To achieve a good coverage of fluorescent beads on the PA gel, 100-nm red (594/620) fluorescent beads (Invitrogen, Carlsbad, CA, USA) were mixed with the polyacrylamide gel solution at a volume ratio of 1:1000. Polymerization was initiated by adding ammonium persulfate (APS) and tetramethylethylenediamine (TEMED) at volume ratios of 1:10 and 1:20, respectively. Once initiated, a 15-μL gel solution was quickly pipetted between a confocal dish and a coverslip, forming a sandwich. The cover slip was treated with bind-silane (BioDee, Beijing, China) followed by treatment with 0.5% glutaraldehyde (Sangon Biotech, Beijing, China) solution. This results in the silanisation of the glass surface, which forms a covalent link with the polymerizing gel, assuring firm attachment of the underside of the gel to the coverslip. Once the sandwich was formed, the PA gel was allowed to polymerize for 15 min at room temperature. Once complete, the upper coverslip was peeled off the gel, leaving a thin layer of gel on the activated surface. The PA gel was then washed extensively in phosphate-buffered saline (PBS). The PA gel was functionalized with 2 mL soak solution (137 mM NaCl and 5% (
v/
v) glycerol) for 1 h at room temperature. After that, 2 mL of a mixed buffer of 1-ethyl-3-[3-dimethylaminopropyl] carbodiimide hydrochloride (EDC), N-hydroxysulfosuccinimide (NHS), and 2-(N-morpholino) ethanesulfonic acid (MES) (10% (
v/
v) glycerol and pH 4.5) in H
2O were added at concentrations of 15 mM, 25 mM, and 100 mM, respectively. The PA gel was incubated with the mixed buffer in the dark at room temperature for 30 min. Then, the PA gel was coated with fibronectin (50 μg/mL) in phosphate-buffered saline (PBS) for 35 min at room temperature. The PA gel was then washed with PBS three times and stored at 4 °C for up to 2 weeks.
2.15. Traction Force Microscopy (TFM) Assay and Data Analysis
TFM assay were conducted using a spinning disk confocal microscope (Dragonfly, Andor; Belfast, UK) equipped with a 63×/1.40 NA oil immersion objective lens. MEFs were incubated with CellTrackerTM Green (Invitrogen, 1:1000 dilution, 1 mM, Carlsbad, CA, USA) at 37 °C for 30 min the day before the experiment. Cells were seeded on fibronectin (FN)-coated polyacrylamide gel 8 h before the experiment. Live-cell imaging of MEFs treated with DMSO or CK-666 was taken for 15 min per cell, 1 min per frame. Both FluoSphereRed (561 nm) and green fluorescent protein (GFP) (488 nm) channels were imaged. Temperature control was maintained on the microscope stage using a live-cell chamber. After imaging, cells were perfused with 3 mL of 0.5% trypsin to release cells from the FN-coupled polyacrylamide (PA) substrate and an image of the unstrained substrate was taken in the 488 and 561 channels.
Once traction force maps and fluorescent images were acquired, average traction force magnitude exerted by the whole cell can be calculated. Firstly, cell contour was extracted by built-in morphological functions in MATLAB (R2017a). Then, cell contour was mapped onto traction force maps and set traction forces outside as zero. Finally, average traction force by the cell can be calculated by
where
indicates traction force magnitude per pixel and
represents the total area of cell contour. Stress ratio is analyzed by comparison of the before-to-after traction stresses for application of 0.5% trypsin.
4. Discussion
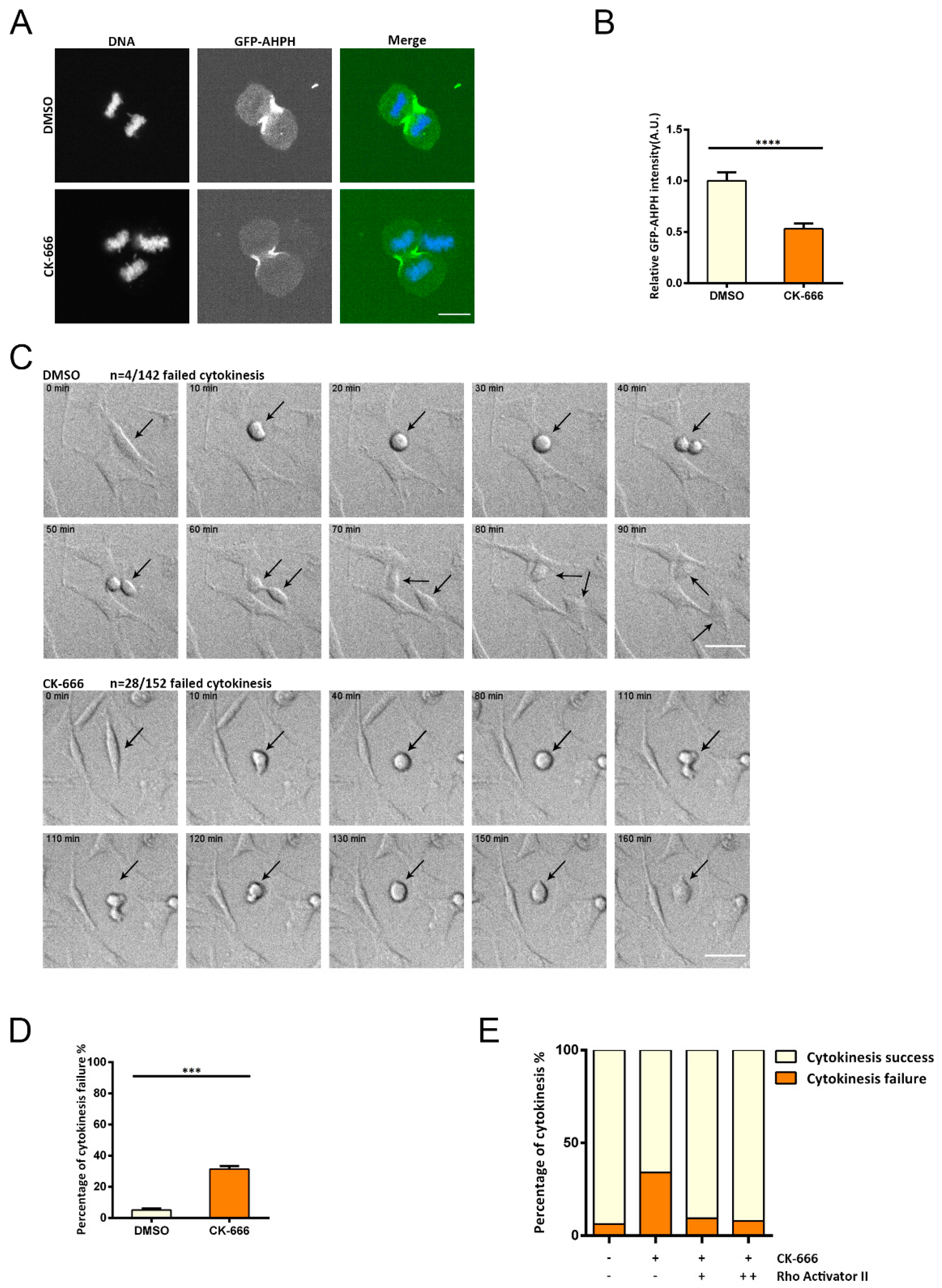
In this study, we investigated potential feedback regulation from the cytoskeleton to small GTPases. We identified reduced RhoA activity but increased RhoA abundance upon inhibiting the Arp2/3-branched actin and unveiled altered cortactin/p190RhoGAP and CCM2/Smurf1 interactions to underly these changes. Moreover, we found that Arp2/3 inhibition leads to cytokinesis defects and that this phenotype can be rescued by activating RhoA. Our work highlights the role of Arp2/3-branched actin mediating cellular signal transduction. Whether other forms of cytoskeletal components serve as signaling hubs aside from their fundamental role in maintaining cell morphology and motility remains to be explored.
Disruption of the Arp2/3-branched actin leads to reduced GTP-RhoA while increases RhoA abundance; as a net result, there would be increased GDP-RhoA residing in the cell. How the enrichment of GDP-RhoA would affect cellular functions remains an intriguing question to explore. It has been noted that the alteration of certain small GTPase may influence the homeostasis of the GTPases network through Rho-GDI [
32]. In our system, inhibition of the Arp2/3 complex by CK-666 did not change the protein abundance or activity level of either Rac1 or Cdc42. However, whether other small GTPase were indirectly affected is unknown and requires systematic studies to be unveiled.
It has been reported that Profilin-1 serves as a gatekeeper for actin assembly by Arp2/3-dependent and -independent pathways [
33,
34]. Upon depletion or inhibition of the Arp2/3 complex, the liner actin filaments nucleated by formins might enhance. In
C. elegans embryos, both Arp2/3 RNAi depletion or CK-666 treatment led to cytokinesis delays, which was proposed to be regulated by increases in the cortical and contractile ring levels of both F-actin and the diaphanous formin CYK-1 [
35]. In
C. elegans, inhibition of RGA-5—the homolog of p190RhoGAP—did not cause mitotic defects in
C. elegans [
36,
37], while p190RhoGAP is required for proper cytokinesis in Hela cells [
28]. This indicates that the function of RGA-5 in
C. elegans is not the same compared with p190RhoGAP in mammalian cells. Whether inhibition of the Arp2/3 complex in
C. elegans may cause similar RhoA activity changes though remains an intriguing question to be investigated.
Here, we reported the role of Arp2/3-branched actin in mediating cellular signal transduction. Many factors regulating actin dynamics and Arp2/3 activity have been reported to show altered expression in cancer [
38,
39,
40]. There may exist the possibility that changes in the actin cytoskeleton would affect RhoA abundance and/or activity. Since RhoA is critical for multiple cellular functions such as cell motility and division, whether this regulation existed in cancer would be highly informative in understanding the significance of cytoskeleton and signal transduction in human disease.