Aberrant hiPSCs-Derived from Human Keratinocytes Differentiates into 3D Retinal Organoids that Acquire Mature Photoreceptors
Abstract
1. Introduction
2. Material and Methods
2.1. NHEK Primary Cell Line and Growth Conditions
2.2. Generation of Feeder-Free hiPSCs from Human Keratinocytes
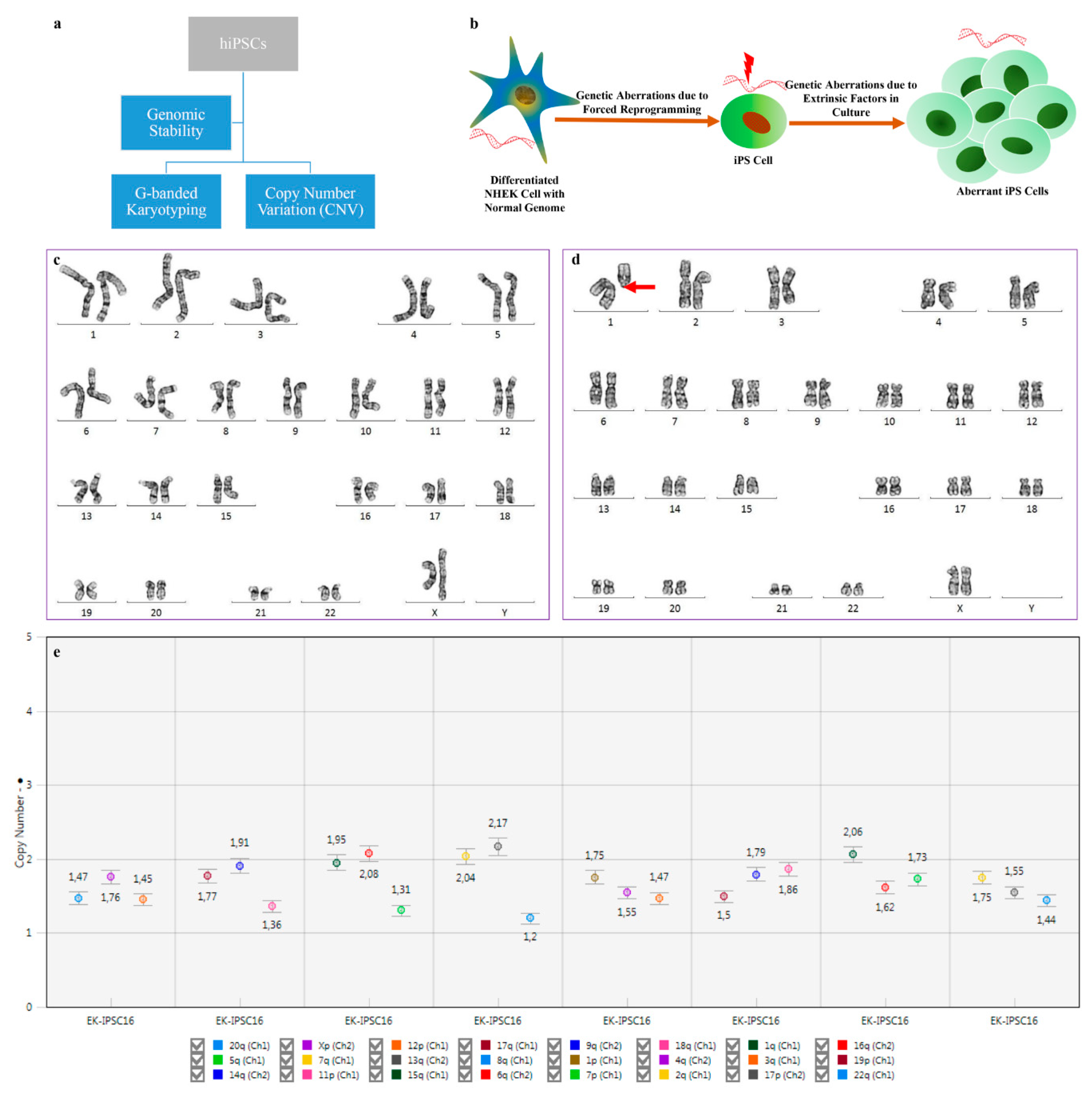
2.3. Genomic Stability of hiPSCs
2.3.1. G-Banded Karyotyping
2.3.2. Digital PCR: Copy Number Variation (CNV)
2.4. In Vitro Differentiation of hiPSCs into Embryoid Bodies (EBs)
2.5. In Vitro Differentiation of hiPSCs: Early Stages to 3-D Retinal Organoids
2.6. Immunostaining and Immunofluorescence Analysis
2.7. RNA Extraction and Reverse-Transcription PCR
2.8. TEM Ultrastructural Analysis
3. Results
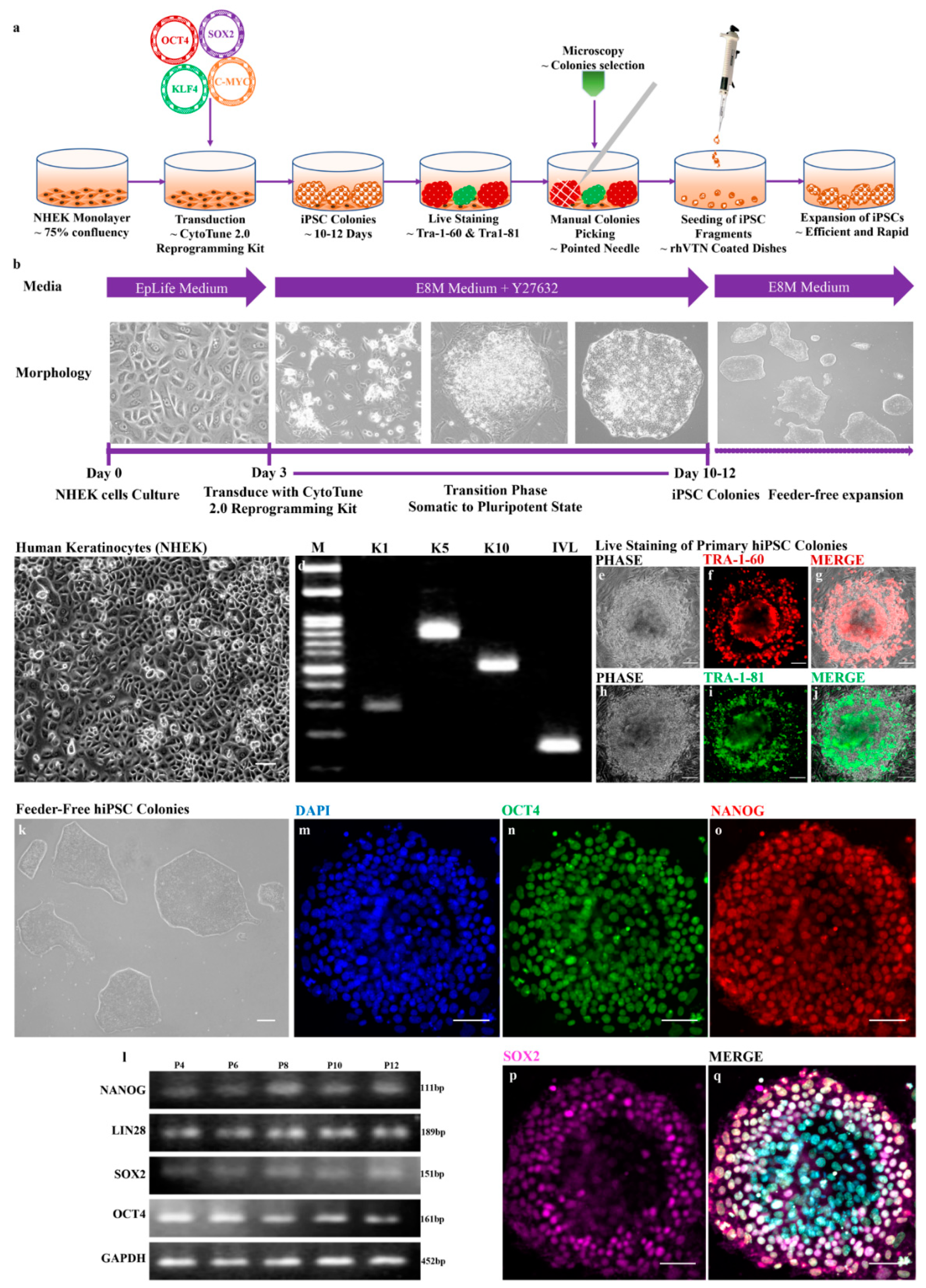
3.1. Rapid and Efficient Generation of Feeder-Free hiPSCs from Human Keratinocytes
3.2. Recurrent Chromosomal Abnormalities and Copy Number Variation in hiPSCs
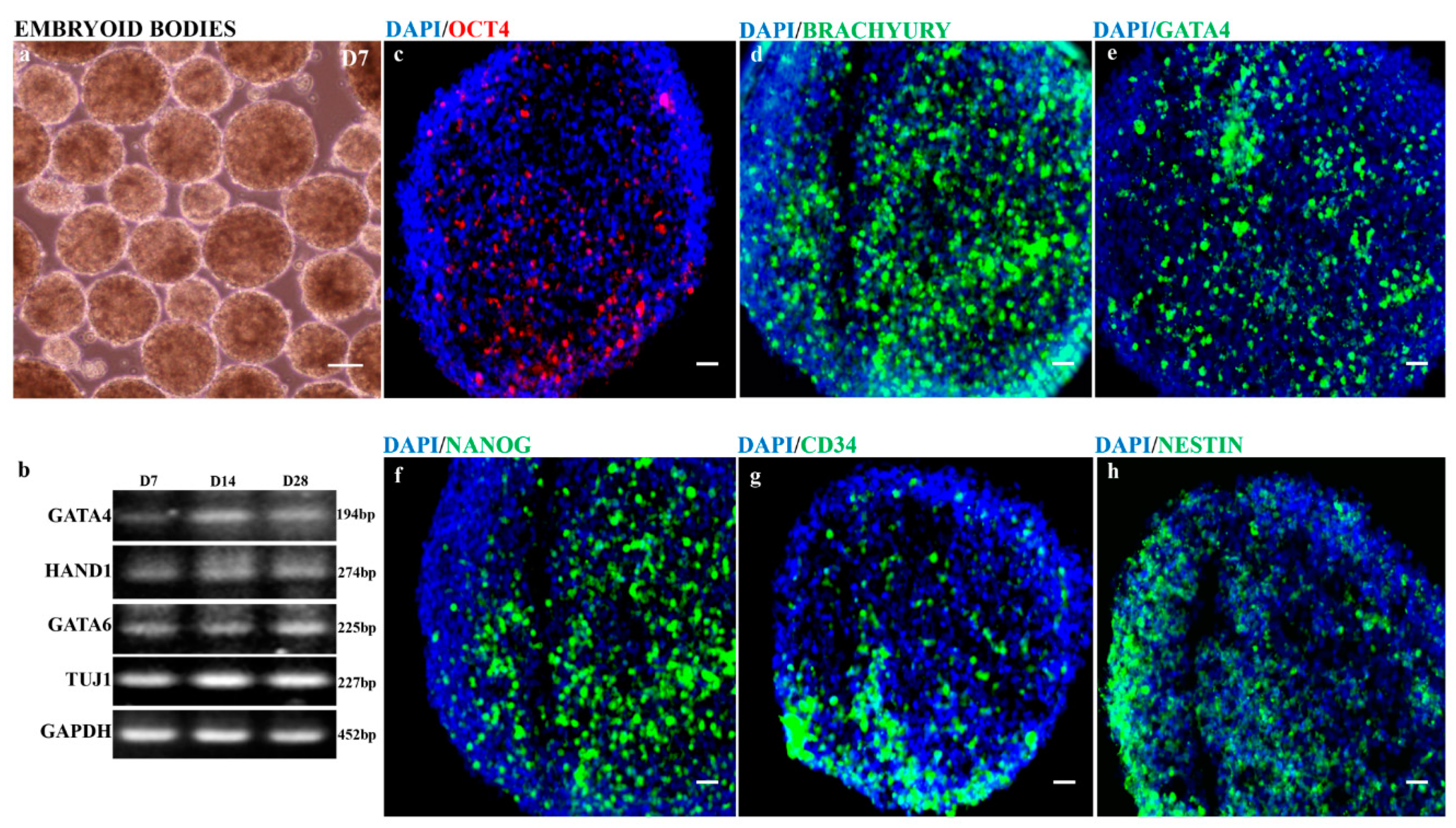
3.3. Self-Forming Embryoid Bodies (EBs) from Feeder-Free hiPSCs
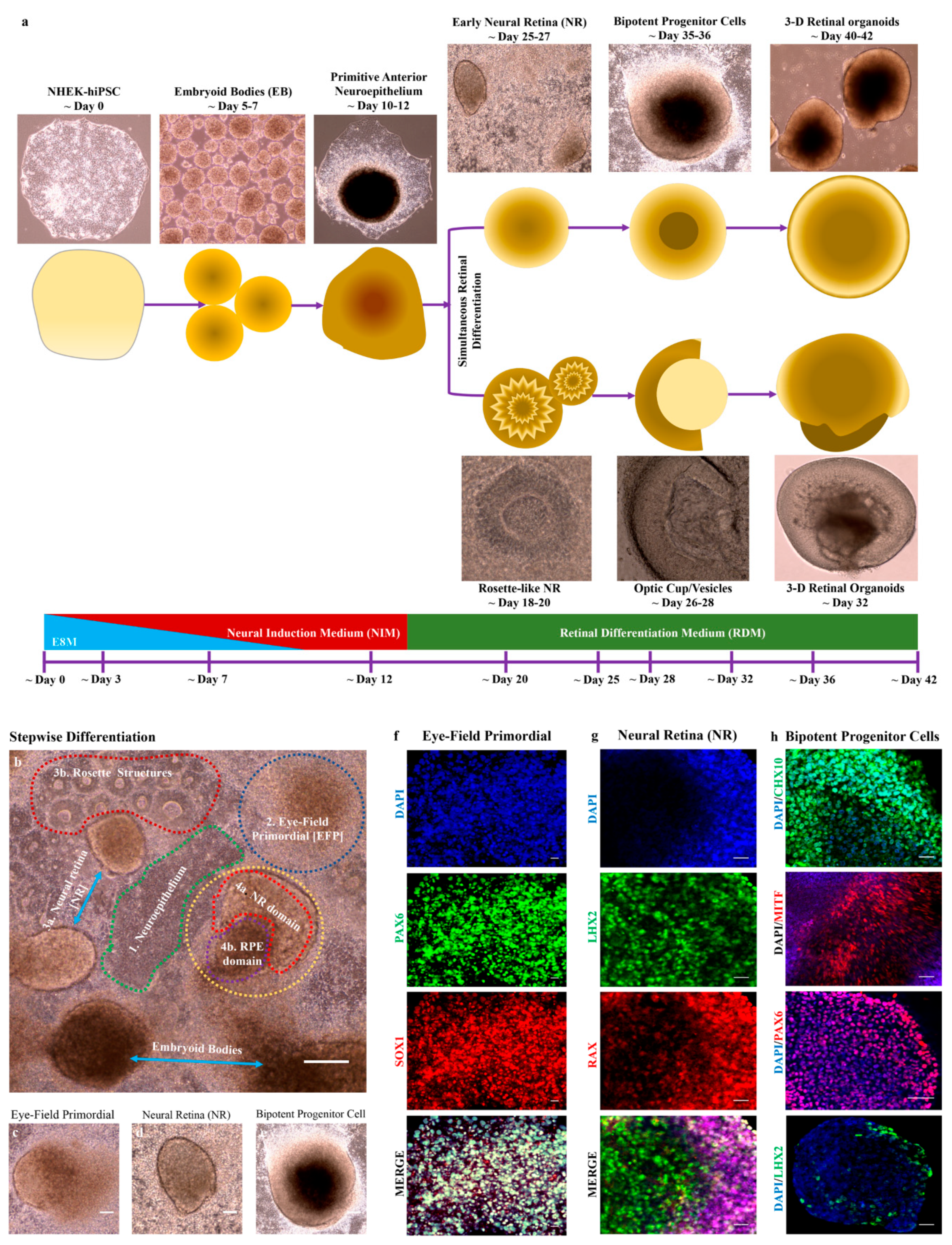
3.4. hiPSC-derived EBs Differentiated into Primitive Anterior Neuroepithelium and Early Neural Retinas
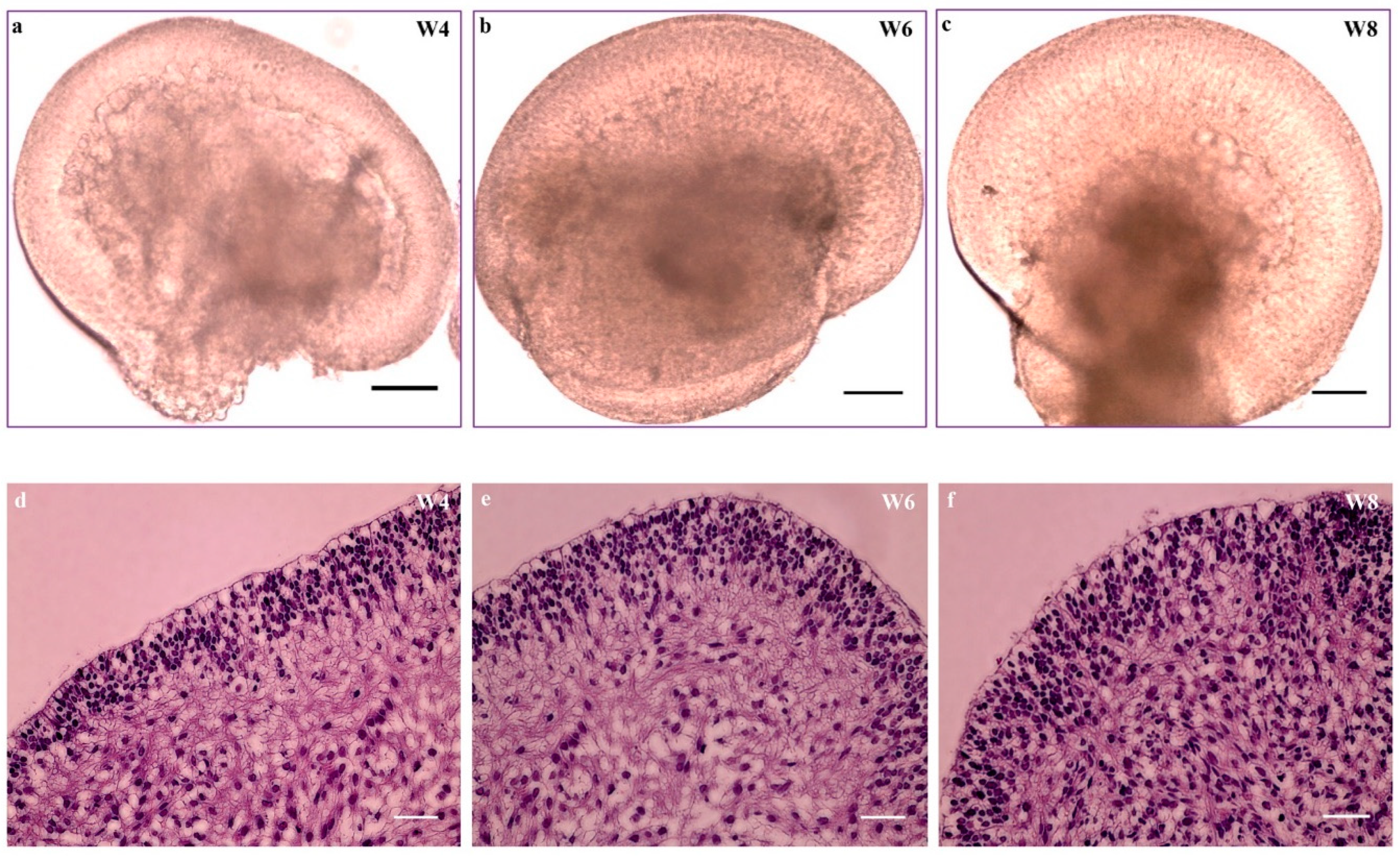
3.5. hiPSC-derived Retinal Organoids
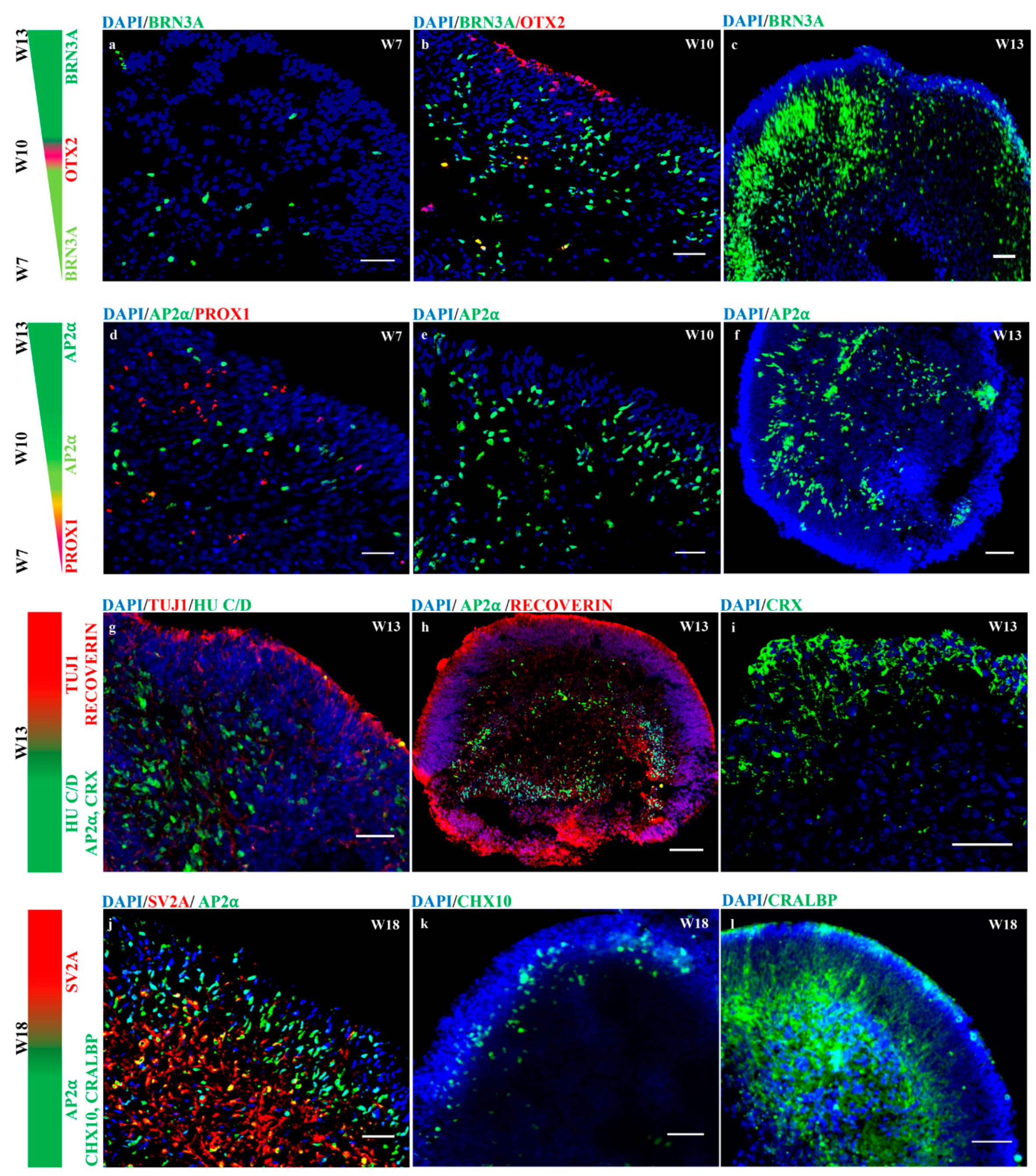
3.6. hiPSC-derived Organoids with Retinal Cell Specification and Laminated Structures
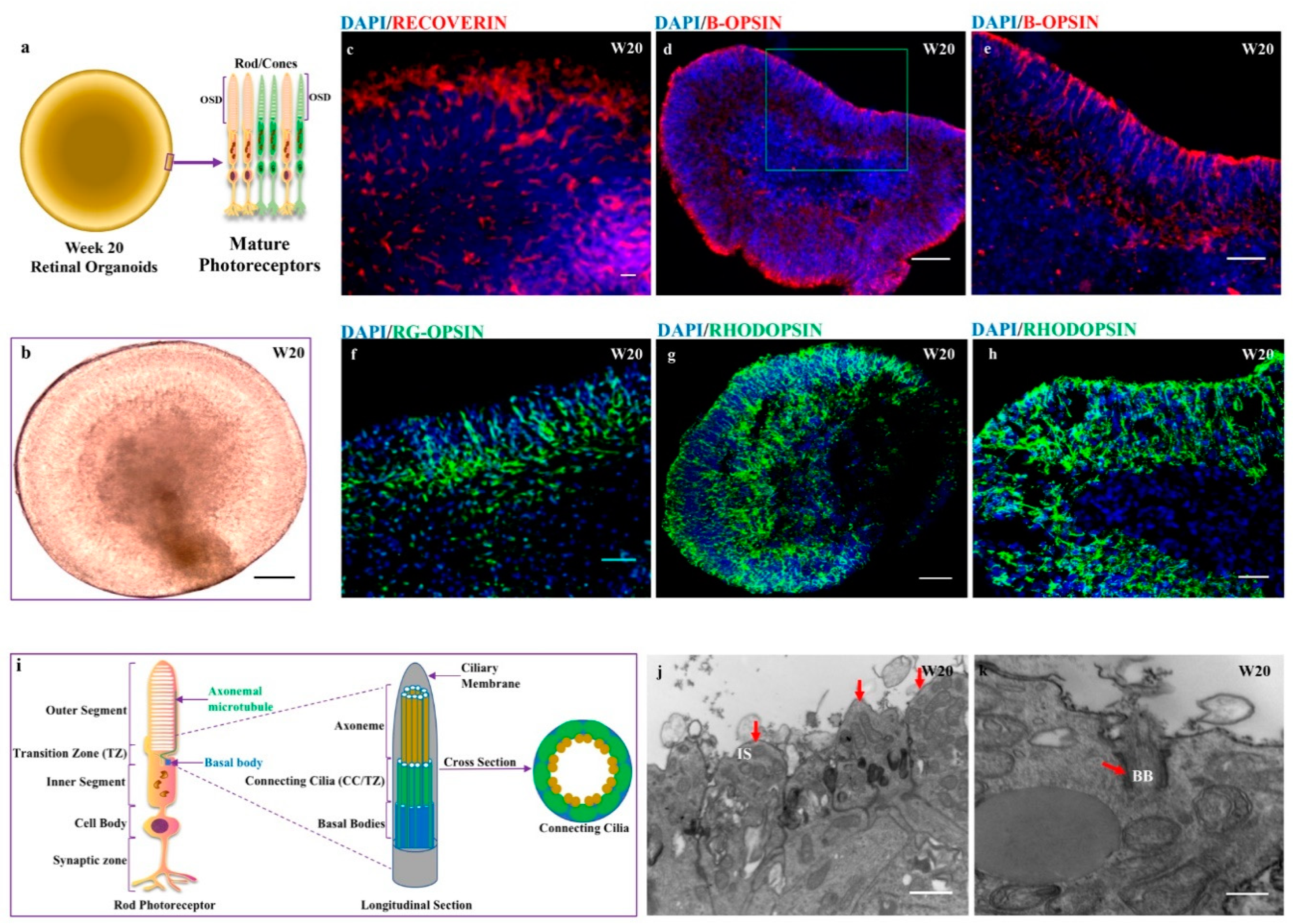
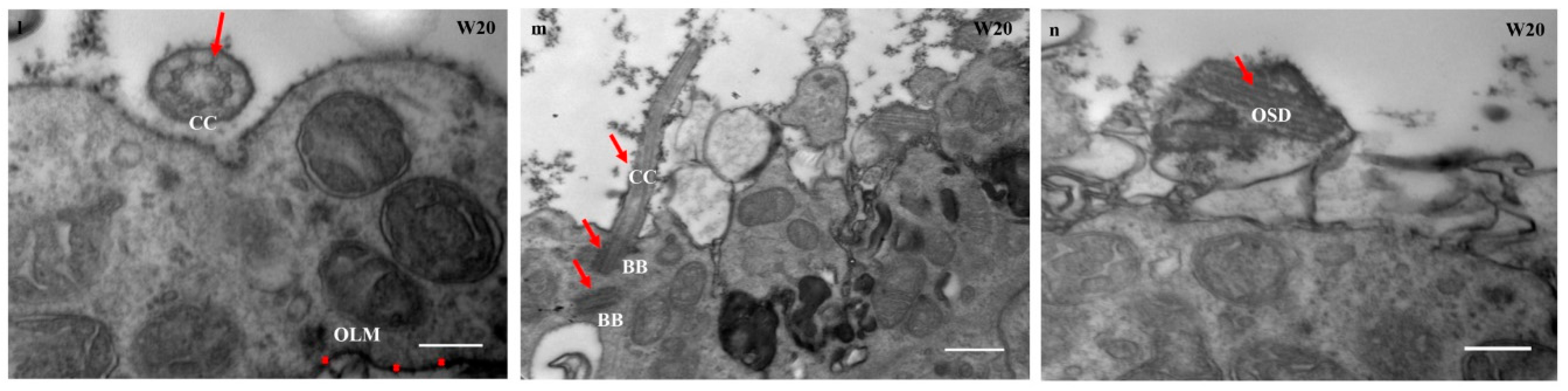
3.7. Acquisition of Mature Photoreceptors Expressing Highly Differentiated Rods and Cones Markers
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ho, B.X.; Pek, N.M.Q.; Soh, B.S. Disease modeling using 3D organoids derived from human induced pluripotent stem cells. Int. J. Mol. Sci. 2018, 19, 936. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.M.; Hochedlinger, K. Harnessing the potential of induced pluripotent stem cells for regenerative medicine. Nat. Cell Biol. 2011, 13, 497–505. [Google Scholar] [CrossRef] [PubMed]
- Orqueda, A.J.; Giménez, C.A.; Pereyra-Bonnet, F. IPSCs: A minireview from bench to bed, including organoids and the CRISPR system. Stem Cells Int. 2016, 2016. [Google Scholar] [CrossRef]
- Tabar, V.; Studer, L. Pluripotent stem cells in regenerative medicine: Challenges and recent progress. Nat. Rev. Genet. 2014, 15, 82–92. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Tanabe, K.; Ohnuki, M.; Narita, M.; Ichisaka, T.; Tomoda, K.; Yamanaka, S. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 2007, 131, 861–872. [Google Scholar] [CrossRef]
- Gagliardi, G.; Ben M’Barek, K.; Chaffiol, A.; Slembrouck-Brec, A.; Conart, J.B.; Nanteau, C.; Rabesandratana, O.; Sahel, J.A.; Duebel, J.; Orieux, G.; et al. Characterization and transplantation of CD73-positive photoreceptors isolated from human iPSC-derived retinal organoids. Stem Cell Rep. 2018, 11, 665–680. [Google Scholar] [CrossRef] [PubMed]
- Homma, K.; Okamoto, S.; Mandai, M.; Gotoh, N.; Rajasimha, H.K.; Chang, Y.S.; Chen, S.; Li, W.; Cogliati, T.; Swaroop, A.; et al. Developing rods transplanted into the degenerating retina of Crx-knockout mice exhibit neural activity similar to native photoreceptors. Stem Cells 2013, 31, 1149–1159. [Google Scholar] [CrossRef]
- Lamba, D.A.; McUsic, A.; Hirata, R.K.; Wang, P.R.; Russell, D.; Reh, T.A. Generation, purification and transplantation of photoreceptors derived from human induced pluripotent stem cells. PLoS ONE 2010, 5, e8763. [Google Scholar] [CrossRef]
- Tucker, B.A.; Park, I.H.; Qi, S.D.; Klassen, H.J.; Jiang, C.; Yao, J.; Redenti, S.; Daley, G.Q.; Young, M.J. Transplantation of adult mouse iPS cell-derived photoreceptor precursors restores retinal structure and function in degenerative mice. PLoS ONE 2011, 6, e18992. [Google Scholar] [CrossRef]
- Mandai, M.; Kurimoto, Y.; Takahashi, M. Autologous induced stem-cell-derived retinal cells for macular degeneration. N. Engl. J. Med. 2017, 377, 792–793. [Google Scholar] [CrossRef]
- Yoshihara, M.; Hayashizaki, Y.; Murakawa, Y. Genomic instability of iPSCs: Challenges towards their clinical applications. Stem Cell Rev. 2017, 13, 7–16. [Google Scholar] [CrossRef] [PubMed]
- Riggs, J.W.; Barrilleaux, B.L.; Varlakhanova, N.; Bush, K.M.; Chan, V.; Knoepfler, P.S. Induced pluripotency and oncogenic transformation are related processes. Stem Cells Dev. 2013, 22, 37–50. [Google Scholar] [CrossRef] [PubMed]
- Gore, A.; Li, Z.; Fung, H.L.; Young, J.E.; Agarwal, S.; Antosiewicz-Bourget, J.; Canto, I.; Giorgetti, A.; Israel, M.A.; Kiskinis, E.; et al. Somatic coding mutations in human induced pluripotent stem cells. Nature 2011, 471, 63–67. [Google Scholar] [CrossRef]
- Taapken, S.M.; Nisler, B.S.; Newton, M.A.; Sampsell-Barron, T.L.; Leonhard, K.A.; McIntire, E.M.; Montgomery, K.D. Karyotypic abnormalities in human induced pluripotent stem cells and embryonic stem cells. Nat. Biotechnol. 2011, 29, 313–314. [Google Scholar] [CrossRef]
- Martins-Taylor, K.; Nisler, B.S.; Taapken, S.M.; Compton, T.; Crandall, L.; Montgomery, K.D.; Lalande, M.; Xu, R.H. Recurrent copy number variations in human induced pluripotent stem cells. Nat. Biotechnol. 2011, 29, 488–491. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Kim, C.H.; Moon, J.I.; Chung, Y.G.; Chang, M.Y.; Han, B.S.; Ko, S.; Yang, E.; Cha, K.Y.; Lanza, R.; et al. Generation of human induced pluripotent stem cells by direct delivery of reprogramming proteins. Cell Stem Cell 2009, 4, 472–476. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Hu, K.; Smuga-Otto, K.; Tian, S.; Stewart, R.; Slukvin, I.I.; Thomson, J.A. Human induced pluripotent stem cells free of vector and transgene sequences. Science 2009, 324, 797–801. [Google Scholar] [CrossRef]
- Tanaka, T.; Yokoi, T.; Tamalu, F.; Watanabe, S.; Nishina, S.; Azuma, N. Generation of retinal ganglion cells with functional axons from human induced pluripotent stem cells. Sci. Rep. 2015, 5, 8344. [Google Scholar] [CrossRef]
- Zhu, J.; Reynolds, J.; Garcia, T.; Cifuentes, H.; Chew, S.; Zeng, X.; Lamba, D.A. Generation of transplantable retinal photoreceptors from a current good manufacturing practice-manufactured human induced pluripotent stem cell line. Stem Cells Transl. Med. 2018, 7, 210–219. [Google Scholar] [CrossRef]
- Meyer, J.S.; Howden, S.E.; Wallace, K.A.; Verhoeven, A.D.; Wright, L.S.; Capowski, E.E.; Pinilla, I.; Martin, J.M.; Tian, S.; Stewart, R.; et al. Optic vesicle-like structures derived from human pluripotent stem cells facilitate a customized approach to retinal disease treatment. Stem Cells 2011, 29, 1206–1218. [Google Scholar] [CrossRef]
- Phillips, M.J.; Wallace, K.A.; Dickerson, S.J.; Miller, M.J.; Verhoeven, A.D.; Martin, J.M.; Wright, L.S.; Shen, W.; Capowski, E.E.; Percin, E.F.; et al. Blood-derived human iPS cells generate optic vesicle-like structures with the capacity to form retinal laminae and develop synapses. Investig. Ophthalmol. Vis. Sci. 2012, 53, 2007–2019. [Google Scholar] [CrossRef] [PubMed]
- Reichman, S.; Terray, A.; Slembrouck, A.; Nanteau, C.; Orieux, G.; Habeler, W.; Nandrot, E.F.; Sahel, J.A.; Monville, C.; Goureau, O. From confluent human iPS cells to self-forming neural retina and retinal pigmented epithelium. Proc. Natl. Acad. Sci. USA 2014, 111, 8518–8523. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Xie, B.; He, L.; Zhou, T.; Gao, G.; Liu, S.; Pan, G.; Ge, J.; Peng, F.; Zhong, X. Generation of retinal organoids with mature rods and cones from urine-derived human induced pluripotent stem cells. Stem Cells Int. 2018, 2018. [Google Scholar] [CrossRef]
- Zhong, X.; Gutierrez, C.; Xue, T.; Hampton, C.; Vergara, M.N.; Cao, L.H.; Peters, A.; Park, T.S.; Zambidis, E.T.; Meyer, J.S.; et al. Generation of three-dimensional retinal tissue with functional photoreceptors from human iPSCs. Nat. Commun. 2014, 5, 4047. [Google Scholar] [CrossRef]
- Sridhar, A.; Steward, M.M.; Meyer, J.S. Nonxenogeneic growth and retinal differentiation of human induced pluripotent stem cells. Stem Cells Transl. Med. 2013, 2, 255–264. [Google Scholar] [CrossRef] [PubMed]
- Wahlin, K.J.; Maruotti, J.A.; Sripathi, S.R.; Ball, J.; Angueyra, J.M.; Kim, C.; Grebe, R.; Li, W.; Jones, B.W.; Zack, D.J. Photoreceptor outer segment-like structures in long-term 3D retinas from human pluripotent stem cells. Sci. Rep. 2017, 7, 766. [Google Scholar] [CrossRef] [PubMed]
- McLelland, B.T.; Lin, B.; Mathur, A.; Aramant, R.B.; Thomas, B.B.; Nistor, G.; Keirstead, H.S.; Seiler, M.J. Transplanted hESC-derived retina organoid sheets differentiate, integrate, and improve visual function in retinal degenerate rats. Investig. Ophthalmol. Vis. Sci. 2018, 59, 2586–2603. [Google Scholar] [CrossRef]
- Beers, J.; Gulbranson, D.R.; George, N.; Siniscalchi, L.I.; Jones, J.; Thomson, J.A.; Chen, G. Passaging and colony expansion of human pluripotent stem cells by enzyme-free dissociation in chemically defined culture conditions. Nat. Protoc. 2012, 7, 2029–2040. [Google Scholar] [CrossRef]
- Aasen, T.; Raya, A.; Barrero, M.J.; Garreta, E.; Consiglio, A.; Gonzalez, F.; Vassena, R.; Bilic, J.; Pekarik, V.; Tiscornia, G.; et al. Efficient and rapid generation of induced pluripotent stem cells from human keratinocytes. Nat. Biotechnol. 2008, 26, 1276–1284. [Google Scholar] [CrossRef]
- Mayshar, Y.; Ben-David, U.; Lavon, N.; Biancotti, J.C.; Yakir, B.; Clark, A.T.; Plath, K.; Lowry, W.E.; Benvenisty, N. Identification and classification of chromosomal aberrations in human induced pluripotent stem cells. Cell Stem Cell 2010, 7, 521–531. [Google Scholar] [CrossRef]
- Sheridan, S.D.; Surampudi, V.; Rao, R.R. Analysis of embryoid bodies derived from human induced pluripotent stem cells as a means to assess pluripotency. Stem Cells Int. 2012, 2012. [Google Scholar] [CrossRef]
- Marquardt, T.; Gruss, P. Generating neuronal diversity in the retina: One for nearly all. Trends Neurosci. 2002, 25, 32–38. [Google Scholar] [CrossRef]
- Bassett, E.A.; Wallace, V.A. Cell fate determination in the vertebrate retina. Trends Neurosci 2012, 35, 565–573. [Google Scholar] [CrossRef] [PubMed]
- Llonch, S.; Carido, M.; Ader, M. Organoid technology for retinal repair. Dev. Biol. 2018, 433, 132–143. [Google Scholar] [CrossRef] [PubMed]
- Osakada, F.; Jin, Z.B.; Hirami, Y.; Ikeda, H.; Danjyo, T.; Watanabe, K.; Sasai, Y.; Takahashi, M. In vitro differentiation of retinal cells from human pluripotent stem cells by small-molecule induction. J. Cell Sci. 2009, 122, 3169–3179. [Google Scholar] [CrossRef] [PubMed]
- Osakada, F.; Ikeda, H.; Mandai, M.; Wataya, T.; Watanabe, K.; Yoshimura, N.; Akaike, A.; Sasai, Y.; Takahashi, M. Toward the generation of rod and cone photoreceptors from mouse, monkey and human embryonic stem cells. Nat. Biotechnol. 2008, 26, 215–224. [Google Scholar] [CrossRef] [PubMed]
- Yamanaka, S. Patient-specific pluripotent stem cells become even more accessible. Cell Stem Cell 2010, 7, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Maherali, N.; Sridharan, R.; Xie, W.; Utikal, J.; Eminli, S.; Arnold, K.; Stadtfeld, M.; Yachechko, R.; Tchieu, J.; Jaenisch, R.; et al. Directly reprogrammed fibroblasts show global epigenetic remodeling and widespread tissue contribution. Cell Stem Cell 2007, 1, 55–70. [Google Scholar] [CrossRef] [PubMed]
- Wernig, M.; Meissner, A.; Foreman, R.; Brambrink, T.; Ku, M.; Hochedlinger, K.; Bernstein, B.E.; Jaenisch, R. In vitro reprogramming of fibroblasts into a pluripotent ES-cell-like state. Nature 2007, 448, 318–324. [Google Scholar] [CrossRef]
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef] [PubMed]
- Raab, S.; Klingenstein, M.; Liebau, S.; Linta, L. A comparative view on human somatic cell sources for iPSC generation. Stem Cells Int. 2014, 2014, 768391. [Google Scholar] [CrossRef] [PubMed]
- Wernig, M.; Lengner, C.J.; Hanna, J.; Lodato, M.A.; Steine, E.; Foreman, R.; Staerk, J.; Markoulaki, S.; Jaenisch, R. A drug-inducible transgenic system for direct reprogramming of multiple somatic cell types. Nat. Biotechnol. 2008, 26, 916–924. [Google Scholar] [CrossRef]
- Aoi, T.; Yae, K.; Nakagawa, M.; Ichisaka, T.; Okita, K.; Takahashi, K.; Chiba, T.; Yamanaka, S. Generation of pluripotent stem cells from adult mouse liver and stomach cells. Science 2008, 321, 699–702. [Google Scholar] [CrossRef] [PubMed]
- Linta, L.; Stockmann, M.; Kleinhans, K.N.; Bockers, A.; Storch, A.; Zaehres, H.; Lin, Q.; Barbi, G.; Bockers, T.M.; Kleger, A.; et al. Rat embryonic fibroblasts improve reprogramming of human keratinocytes into induced pluripotent stem cells. Stem Cells Dev. 2012, 21, 965–976. [Google Scholar] [CrossRef] [PubMed]
- Gandarillas, A.; Watt, F.M. c-Myc promotes differentiation of human epidermal stem cells. Genes Dev. 1997, 11, 2869–2882. [Google Scholar] [CrossRef] [PubMed]
- Segre, J.A.; Bauer, C.; Fuchs, E. Klf4 is a transcription factor required for establishing the barrier function of the skin. Nat. Genet. 1999, 22, 356–360. [Google Scholar] [CrossRef]
- Pettinato, G.; Vanden Berg-Foels, W.S.; Zhang, N.; Wen, X. ROCK inhibitor is not required for embryoid body formation from singularized human embryonic stem cells. PLoS ONE 2014, 9, e100742. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, K.; Ueno, M.; Kamiya, D.; Nishiyama, A.; Matsumura, M.; Wataya, T.; Takahashi, J.B.; Nishikawa, S.; Muguruma, K.; Sasai, Y. A ROCK inhibitor permits survival of dissociated human embryonic stem cells. Nat. Biotechnol. 2007, 25, 681–686. [Google Scholar] [CrossRef] [PubMed]
- Mathers, P.H.; Jamrich, M. Regulation of eye formation by the Rx and Pax6 homeobox genes. Cell. Mol. Life Sci. 2000, 57, 186–194. [Google Scholar] [CrossRef]
- Horsford, D.J.; Nguyen, M.T.; Sellar, G.C.; Kothary, R.; Arnheiter, H.; McInnes, R.R. Chx10 repression of Mitf is required for the maintenance of mammalian neuroretinal identity. Development 2005, 132, 177–187. [Google Scholar] [CrossRef]
- Rowan, S.; Chen, C.M.; Young, T.L.; Fisher, D.E.; Cepko, C.L. Transdifferentiation of the retina into pigmented cells in ocular retardation mice defines a new function of the homeodomain gene Chx10. Development 2004, 131, 5139–5152. [Google Scholar] [CrossRef] [PubMed]
- Meyer, J.S.; Shearer, R.L.; Capowski, E.E.; Wright, L.S.; Wallace, K.A.; McMillan, E.L.; Zhang, S.C.; Gamm, D.M. Modeling early retinal development with human embryonic and induced pluripotent stem cells. Proc. Natl. Acad. Sci. USA 2009, 106, 16698–16703. [Google Scholar] [CrossRef]
- Phillips, M.J.; Perez, E.T.; Martin, J.M.; Reshel, S.T.; Wallace, K.A.; Capowski, E.E.; Singh, R.; Wright, L.S.; Clark, E.M.; Barney, P.M.; et al. Modeling human retinal development with patient-specific induced pluripotent stem cells reveals multiple roles for visual system homeobox 2. Stem Cells 2014, 32, 1480–1492. [Google Scholar] [CrossRef] [PubMed]
- Eiraku, M.; Sasai, Y. Mouse embryonic stem cell culture for generation of three-dimensional retinal and cortical tissues. Nat. Protoc. 2011, 7, 69–79. [Google Scholar] [CrossRef]
- Nakano, T.; Ando, S.; Takata, N.; Kawada, M.; Muguruma, K.; Sekiguchi, K.; Saito, K.; Yonemura, S.; Eiraku, M.; Sasai, Y. Self-formation of optic cups and storable stratified neural retina from human ESCs. Cell Stem Cell 2012, 10, 771–785. [Google Scholar] [CrossRef] [PubMed]
- Eiraku, M.; Takata, N.; Ishibashi, H.; Kawada, M.; Sakakura, E.; Okuda, S.; Sekiguchi, K.; Adachi, T.; Sasai, Y. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 2011, 472, 51–56. [Google Scholar] [CrossRef]
- Volkner, M.; Zschatzsch, M.; Rostovskaya, M.; Overall, R.W.; Busskamp, V.; Anastassiadis, K.; Karl, M.O. Retinal organoids from pluripotent stem cells efficiently recapitulate retinogenesis. Stem Cell Rep. 2016, 6, 525–538. [Google Scholar] [CrossRef] [PubMed]
- Janssen, J.J.; Kuhlmann, E.D.; van Vugt, A.H.; Winkens, H.J.; Janssen, B.P.; Deutman, A.F.; Driessen, C.A. Retinoic acid delays transcription of human retinal pigment neuroepithelium marker genes in ARPE-19 cells. Neuroreport 2000, 11, 1571–1579. [Google Scholar] [CrossRef] [PubMed]

| Target | Brand-Catalogue Number | Dilution | Host |
|---|---|---|---|
| Primary Antibodies | |||
| Tra-1-60 | Invitrogen; 41–1000 | 1:100 | Mouse |
| Tra-1-81 | Invitrogen; 41–1100 | 1:100 | Mouse |
| OCT4 | Abcam; ab27985 | 1:200 | Goat |
| Alexa Fluor 647 anti-SOX2 | BioLegend; 656108 | 1:200 | Mouse |
| NANOG | Millipore; AB9220 | 1:200 | Rabbit |
| BRACHYURY | Abcam; AB20680 | 1:200 | Rabbit |
| CD34 | Abcam; ab81289 | 1:100 | Rabbit |
| GATA4 | Millipore; AB4132 | 1:200 | Rabbit |
| NESTIN | Sigma; N5413 | 1:100 | Rabbit |
| PAX6 | Millipore; AB2237 | 1:100 | Rabbit |
| SOX1 | Abcam; ab87775 | 1:500 | Rabbit |
| LHX2 | Santa Cruz Biotechnology; SC19344 | 1:50 | Goat |
| RX | Novus Biologicals; H00030062-M02 | 1:200 | Mouse |
| MITF | Abcam; ab80651 | 1:200 | Mouse |
| CHX10 | Novus Biologicals; NBP184476 | 1:1000 | Rabbit |
| BRN3A | Millipore; MAB1585 | 1:20 | Mouse |
| OTX2 | R&D; MAB1979 | 1:50 | Mouse |
| AP2α | Invitrogen; MA1872 | 1:50 | Mouse |
| PROX1 | Millipore; AB5475 | 1:2000 | Rabbit |
| TUJ1 | BioLegend; PRB-435P | 1:1000 | Rabbit |
| HU C/D | Invitrogen; A21271 | 1:100 | Mouse |
| CRX | Abnova; H00001406-M06 | 1:200 | Mouse |
| RECOVERIN | Millipore; AB5585 | 1:1000 | Rabbit |
| SV2A | Invitrogen, PA5-52476 | 1:50 | Rabbit |
| CRALBP | Abcam; ab15051 | 1:200 | Mouse |
| B-OPSIN | Millipore; AB5407 | 1:200 | Rabbit |
| RG-OPSIN | Millipore; AB5405 | 1:200 | Rabbit |
| RHODOPSIN | Abcam; ab5417 | 1:1000 | Mouse |
| Secondary Antibodies | |||
| Alexa Fluor 488 Anti-Goat | Invitrogen; A11055 | 1:500 | Donkey |
| Alexa Fluor 555 Anti-Goat | Invitrogen; A21432 | 1:500 | Donkey |
| Alexa Fluor 647 Anti-Goat | Invitrogen; A21447 | 1:500 | Donkey |
| Alexa Fluor 488 Anti-Mouse | Invitrogen; A21202 | 1:500 | Donkey |
| Alexa Fluor 555 Anti-Mouse | Invitrogen; A31570 | 1:500 | Donkey |
| Alexa Fluor 647 Anti-Mouse | Invitrogen; A31571 | 1:500 | Donkey |
| Alexa Fluor 488 Anti-Rabbit | Invitrogen; A21206 | 1:500 | Donkey |
| Alexa Fluor 555 Anti-Rabbit | Invitrogen; A31572 | 1:500 | Donkey |
| Alexa Fluor 647 Anti-Rabbit | Invitrogen; A31573 | 1:500 | Donkey |
| Genes | Forward Primers (5′-3′) | Reverse Primers (5′-3′) | Size (bp) |
|---|---|---|---|
| Human Keratinocytes-Specific Markers | |||
| IVL | CTGCCCACAAAGGGAGAAGT | AGCGGACCCGAAATAAGTGG | 166 |
| K1 | CCCTCCTGGTGGCATACAAG | GTTGGTCCACTCTCCTTCGG | 293 |
| K10 | CCCTGGGCTAAACAGCATCA | TCCAGAGCCCGAACTTTGTC | 525 |
| K5 | TCAACAAGCGTACCACTGCT | CTGCTACCTCCGGCAAGAC | 802 |
| hiPSC-Specific Markers | |||
| OCT4 | CAGTGCCCGAAACCCACAC | GGAGACCCAGCAGCCTCAAA | 161 |
| SOX2 | GGGAAATGGGAGGGGTGCAAAAGAGG | TTGCGTGAGTGTGGATGGGAT TGGTG | 151 |
| NANOG | CAGAAGGCCTCAGCACCTAC | ATTGTTCCAGGTCTGGTTGC | 111 |
| LIN28 | TGCACCAGAGTAAGCTGCAC | CTCCTTTTGATCTGCGCTTC | 189 |
| Embryoid Body-Specific Markers | |||
| GATA4 | TCCCTCTTCCCTCCTCAAAT | TCAGCGTGTAAAGGCATCTG | 194 |
| HAND1 | TGCCTGAGAAAGAGAACCAG | ATGGCAGGATGAACAAACAC | 274 |
| GATA6 | CCTCACTCCACTCGTGTCTGC | GTCCTGGCTTCTGGAAGTGG | 225 |
| TUJ1 | CAGAGCAAGAACAGCAGCTACTT | GTGAACTCCATCTCGTCCATGCCCTC | 227 |
| GAPDH | ACCACAGTCCATGCCATCAC | TCCACCACCCTGTTGCTGTA | 452 |
| Type of Abnormality | Cytogenetic Analysis | Metaphase |
|---|---|---|
| Normal | 46,XX | 6 |
| Deletion | 46,XX,del(1)(q11)[cp2] | 1 |
| Deletion | 46,XX,del(9)(q11) | 1 |
| Loss | 44,XX,-7,-11 | 1 |
| Loss | 45,XX,-4 | 1 |
| Loss | 45,XX,-14 | 1 |
| Derivative, Translocation and Loss | 46,XX,+der(9)t(9;?)(p13;?),-11 | 1 |
| Status: Abnormal | Karyotype: 46,XX,del(1)(q11)[cp2]∕46,XX [6] | Total: 12 |
| Category | Copy Number Values | Chromosomes |
|---|---|---|
| Loss | 1.3 | 8q |
| Trend to loss | 1.5 | 3q, 12p, 18q, 19p, 20q, 22q |
| Normal | 2.0 | 1p, 1q, 2q, 4q, 4q, 6q, 7p, 7q, 9q, 11p, 13q, 14q, 15q, 16q, 17p, 17q, |
| Trend to gain | 2.5 | - |
| Gain | 2.7 | - |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shrestha, R.; Wen, Y.-T.; Ding, D.-C.; Tsai, R.-K. Aberrant hiPSCs-Derived from Human Keratinocytes Differentiates into 3D Retinal Organoids that Acquire Mature Photoreceptors. Cells 2019, 8, 36. https://doi.org/10.3390/cells8010036
Shrestha R, Wen Y-T, Ding D-C, Tsai R-K. Aberrant hiPSCs-Derived from Human Keratinocytes Differentiates into 3D Retinal Organoids that Acquire Mature Photoreceptors. Cells. 2019; 8(1):36. https://doi.org/10.3390/cells8010036
Chicago/Turabian StyleShrestha, Rupendra, Yao-Tseng Wen, Dah-Ching Ding, and Rong-Kung Tsai. 2019. "Aberrant hiPSCs-Derived from Human Keratinocytes Differentiates into 3D Retinal Organoids that Acquire Mature Photoreceptors" Cells 8, no. 1: 36. https://doi.org/10.3390/cells8010036
APA StyleShrestha, R., Wen, Y.-T., Ding, D.-C., & Tsai, R.-K. (2019). Aberrant hiPSCs-Derived from Human Keratinocytes Differentiates into 3D Retinal Organoids that Acquire Mature Photoreceptors. Cells, 8(1), 36. https://doi.org/10.3390/cells8010036









